Abstract
Bacteriorhodopsin is a light-driven proton pump in halobacteria that forms crystalline patches in the cell membrane. Isomerization of the bound retinal initiates a photocycle resulting in the extrusion of a proton. An electron crystallographic analysis of the N intermediate from the mutant F219L gives a three-dimensional view of the large conformational change that occurs on the cytoplasmic side after deprotonation of the retinal Schiff base. Helix F, together with helix E, tilts away from the center of the molecule, causing a shift of ∼3 Å at the EF loop. The top of helix G moves slightly toward the ground state location of helix F. These movements open a water-accessible channel in the protein, enabling the transfer of a proton from an aspartate residue to the Schiff base. The movement of helix F toward neighbors in the crystal lattice is so large that it would not allow all molecules to change conformation simultaneously, limiting the occupancy of this state in the membrane to 33%. This explains photocooperative phenomena in the purple membrane.
Keywords: bacteriorhodopsin/conformational change/electron crystallography/proton pump/trapped intermediates
Introduction
Bacteriorhodopsin (bR) is a small transmembrane protein in halobacteria, consisting of seven transmembrane helices (named A–G) and a retinal prosthetic group that is bound by a Schiff base linkage to the side chain of Lys216 on helix G. It acts as a light-driven proton pump. Capture of a photon initiates a cyclical series of changes (with intermediate states named J–O), resulting in the transport of a proton out of the cell. The initial light-driven event is a trans–cis isomerization of the retinal. In the transition from the L to the M intermediate, the Schiff base is deprotonated and the side chain of Asp85, located toward the extracellular side, is protonated. Later a proton is released to the medium, probably from a release complex including Glu194 and Glu204 (for recent reviews, see Lanyi, 1998; Haupts et al., 1999). The M to N transition is characterized by a reprotonation of the Schiff base, with Asp96 acting as the proton donor. Asp96 is located near the cytoplasmic surface, >10 Å from the Schiff base. No proton path between these two groups exists in the ground state, and therefore a conformational change has to take place to make Schiff base reprotonation possible. The amide bands in Fourier transform infra-red (FTIR) spectra differ between the M and the N intermediate, and this is thought to reflect this conformational change. The long-lived M intermediate in Asp96 mutants already has the N protein structure (Sasaki et al., 1992), and is therefore named MN.
Large-scale conformational changes can be studied by means of Fourier difference maps between the ground state and an intermediate using diffraction methods. In projection maps derived by electron diffraction of the early intermediates K (Bullough and Henderson, 1999) and L (Hendrickson et al., 1998) no changes are observed at 3.5 Å resolution. Consistent with this, a 2 Å resolution X-ray map of the low-temperature K intermediate shows very small changes in the side chains surrounding the Schiff base (Edman et al., 1999). Single membrane electron diffraction of the M and N intermediates trapped by various methods (Subramaniam et al., 1993, 1999; Han et al., 1994a; Vonck, 1996; Lindahl and Henderson, 1997; Hendrickson et al., 1998) as well as X-ray (Koch et al., 1991; Nakasako et al., 1991; Kamikubo et al., 1996, 1997; Sass et al., 1997) and neutron diffraction from stacks of membranes (Dencher et al., 1989; Hauss et al., 1994) all show a similar pattern of changes mainly involving the helices G, F and B. Electron crystallographic studies from slightly tilted samples indicate that the movements are restricted to the cytoplasmic side (Subramaniam et al., 1993; Vonck, 1996) and suggest that they involve a tilt of helix F, resulting in the opening of a channel from the cytoplasm to the Schiff base, as well as a change on helix G that may involve a local increase of order. However, the three-dimensional (3D) conformational change of the bR protein, which is crucial for understanding the molecular mechanism of proton pumping, has so far remained elusive.
Several models for the bR ground state have been published at increasing resolution, first by electron crystallography of the native two-dimensional (2D) crystals (Henderson et al., 1990; Grigorieff et al., 1996; Kimura et al., 1997; Mitsuoka et al., 1999) and then by X-ray crystallography of 3D crystals grown in cubic lipid phases (Pebay-Peyroula et al., 1997; Luecke et al., 1998, 1999b; Belrhali et al., 1999) and monoclinic crystals (Essen et al., 1998). The models are all very similar in the transmembrane part and only show small differences in the orientation of some side chains. In the highest resolution models, several water molecules can be identified that may have a role in proton transfer.
The early steps in the photocycle are becoming known at an atomic level. A 2 Å resolution X-ray difference map of the low-temperature K state (Edman et al., 1999) shows small changes in some side chains surrounding the Schiff base, notably Lys216, Asp85 and Tyr57. A model of the D96N MN intermediate (Luecke et al., 1999a), also at 2 Å resolution, has a 13-cis retinal, and reveals rearrangements of side chains and hydrogen bonds connecting them to each other and to water molecules, suggesting mechanisms for the pKa changes that induce proton transfers in the extracellular channel. However, the expected large-scale changes on helix F and G that should have occurred in the MN state were not visible in the map; instead, the top of helices E, F and G were so disordered that they could not be modeled. The cytoplasmic half of the protein shows higher temperature factors in all the ground state models, which indicates that this part of the structure is more flexible.
This paper presents the structure of the N intermediate from the mutant F219L at an in-plane resolution of 3.5 Å and a vertical resolution of ∼5.5 Å from electron diffraction studies. At this resolution, the main differences between the N map and the bR ground state map are tilts of the cytoplasmic part of helix F away from the center of the molecule and of the cytoplasmic part of helix G toward the ground state position of helix F. The occupancy of the N conformation is only ∼20–30%, probably because of steric restraints in the purple membrane. This has interesting implications for the photocycle.
Results
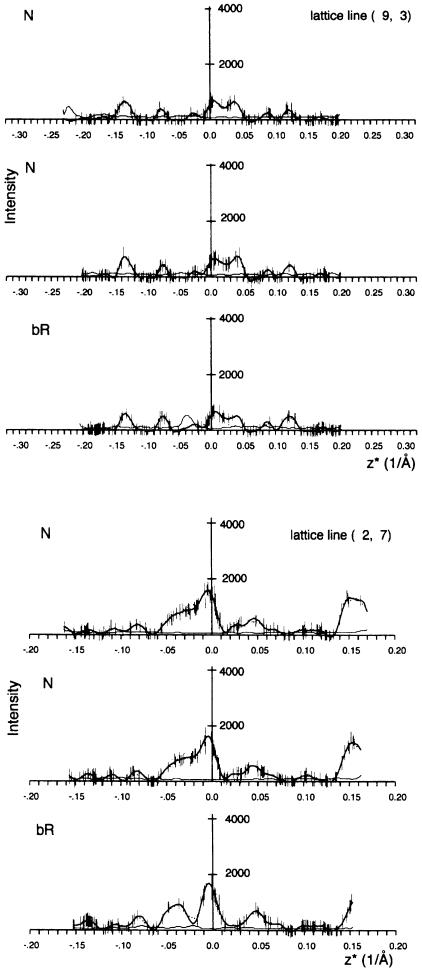
Electron diffraction data at 45° tilt were collected for the bR ground state and the N intermediate. For each data set, ∼120 patterns of flat, well diffracting crystals were selected, and the data to 3.5 Å resolution were merged together with patterns at 0° and 25° tilt (Table I). The data are almost complete to 3.5 Å resolution; the resolution is limited by the available image phases. The missing cone of data beyond 45° limits the vertical resolution to ∼5.5 Å. Examples of lattice lines are shown in Figure 1. To reduce noise, the data sets were split in two halves, which were merged independently, and two difference maps were calculated. The difference amplitudes from both difference maps were then averaged, which enhanced features present in both maps and reduced random noise.
Table I. Electron diffraction data.
| bR | N | |
|---|---|---|
| No. of patterns at 45° | 137 | 114 |
| No. of patterns at 25° | 46 | 37 |
| No. of patterns at 0° | 22 | 12 |
| Rmergea | 11.7 | 13.6 |
| Rsymb | 8.8 | 8.3 |
aRmerge = Σ|(Iobserved – Icurve|/ΣIcurve
bRsym is a measure of the difference between Friedel mates: Rsym = Σ|(I1 – I2)|/Σ(I1 + I2) (Baldwin and Henderson, 1984).
Fig. 1. Measured diffraction intensities and fitted curves for two representative lattice lines for two halves of the N data and one half of the bR data for comparison. Thick lines, fitted curves; thin lines, fitting error. Error bars, differences between Friedel mates. The (2,7) lattice line is at 6.6 Å resolution for z* = 0 and maximum at 4.4 Å for z* = 0.17 Å–1 (limited by the tilt angle). The (9,3) lattice line is at 5.0 Å resolution for z* = 0 and maximum at 3.5 Å for z* = 0.20 Å–1.
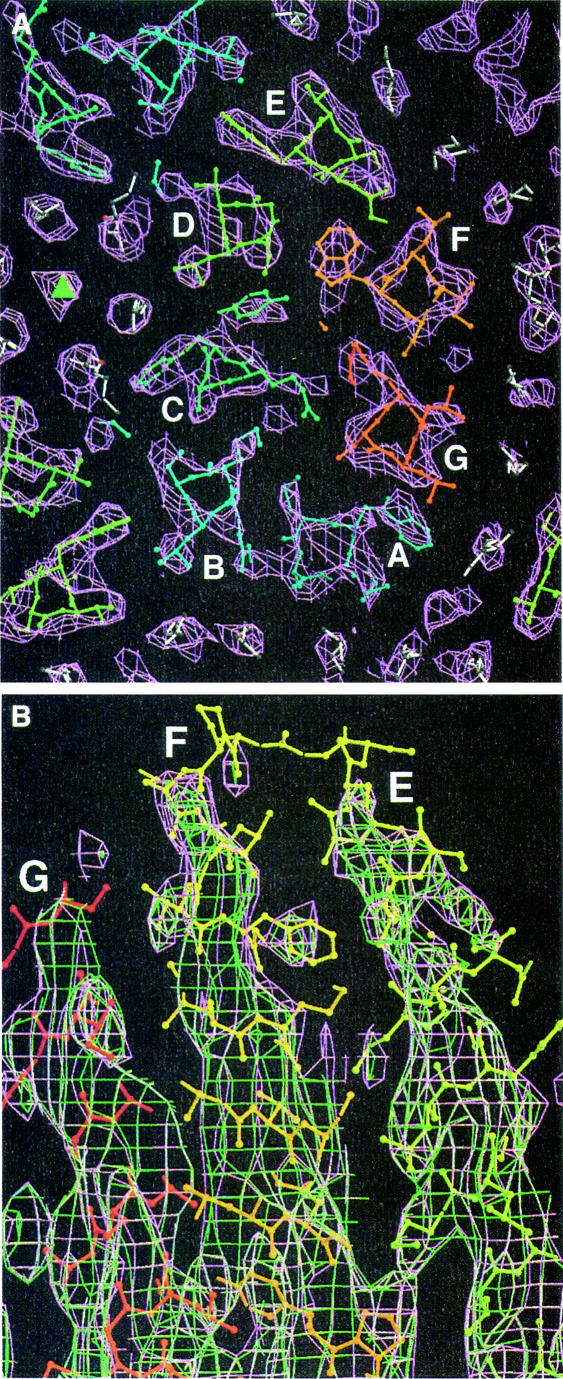
An electron density map was calculated for the ground state. As shown in Figure 2A, it fits the high-resolution X-ray models very well; especially on the extracellular side, even the lipid molecules from the model fit in electron density. A map from the N data looks very similar to the bR map. Only on the top of helices F and G is a slightly lower density seen in the N state, with no new density appearing (Figure 2B). This indicates that the N data consist of a mixture of the two configurations, with an occupancy of the ground state configuration of >50%.
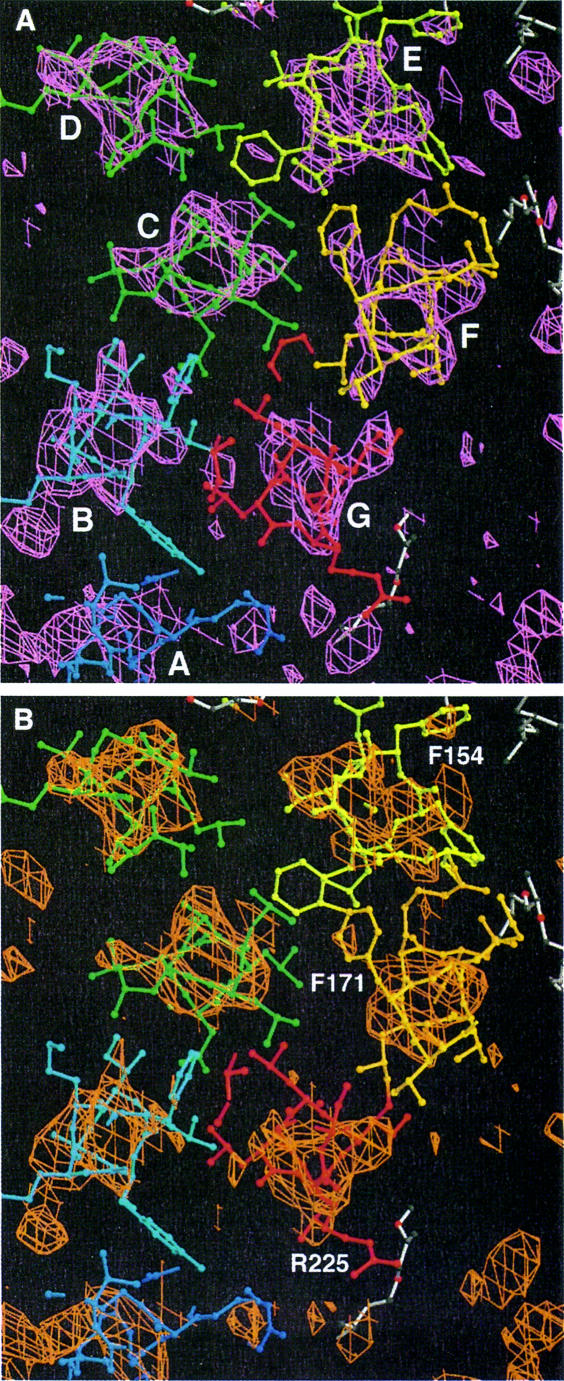
Fig. 2. Electron density maps with model 1qhj superimposed. White, lipids; from blue to red, protein. (A) A 5 Å slice of the bR density map on the extracellular side (purple). The center of the trimer is indicated with a triangle. (B) Side view of the bR (purple) and N (green) density map. Figures 2–6 were made with Bobscript (Esnouf, 1997, 1999), an extension of MOLSCRIPT (Kraulis, 1991), and Raster3D (Merritt and Bacon, 1997).
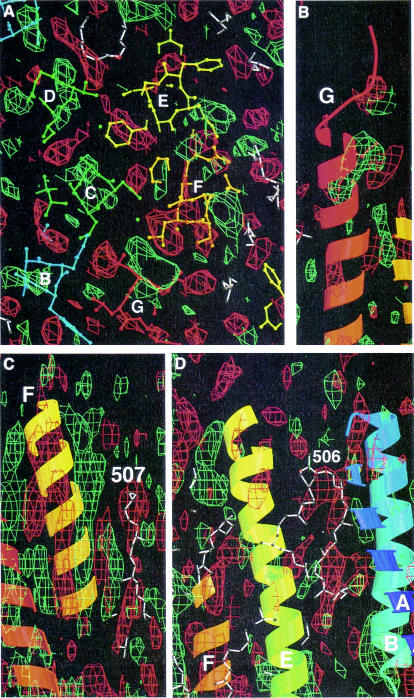
A 3D Fourier difference map reveals the changes between bR and N. As in the projection maps, the largest differences are on the helices F and G (Figure 3). The strongest difference peak is on the ground state position of helix G above the Schiff base, moving out toward helix F around residue 222. There is no associated negative density of similar magnitude, but some side chains on the opposite side (away from helix F) sit in negative density. This suggests a movement of the top of helix G toward the F helix.
Fig. 3. N–bR difference map. Positive peaks are green, negative peaks are red. Model 1qhj is superimposed with colors as in Figure 1. (A) A 5 Å slice on the cytoplasmic side. (B) Side view of the cytoplasmic part of helix G. The positive peak toward helix F can be seen. (C) Side view of helix F and lipid 507. Both sit in negative density and helix F moves toward the lipid. (D) Lipid 506 between helices E and F from one monomer and helices A and B from its neighbor in the trimer is associated with negative density.
Figure 3C illustrates the movement of helix F. The difference peaks on this helix consist of a positive and a negative component throughout, with the negative peak on the ground state position of the helix and the positive peak at an increasing distance when moving toward the cytoplasmic surface, a clear indication of a helix tilt in the N intermediate. Also on the top of helix E, positive density in the direction of helix F is visible. The side chain of Leu152 sits in strong negative density.
Many of the helices are found in positive density in the N–bR map, indicating an increase of order in the N state. The effect is strongest on helix G, as discussed before, but it is also present on most of helix C (residues 80–100) and parts of helices A (17–24) and B (49–60), as well as short stretches on the extracellular side of helices E (130–138) and F (190–196). The more highly ordered regions on helices A, B, C and G are located adjacent to each other in the region around the Schiff base and Asp85, and the better ordered region on helix F includes Glu194. These residues have all been identified as elements of the proton release pathway. In projection maps, the positive peak on helix B is the strongest, because this helix is nearly perpendicular to the membrane so that the positive density is not smeared out.
The prominent negative peak near helix B that is seen on most projection difference maps is also visible in the 3D map (Figure 3D). It coincides with part of a lipid molecule between molecules of a trimer in the cytoplasmic leaflet [lipid 612 in 1c3w (Luecke et al., 1999a); 506 in 1qhj (Belrhali et al., 1999)]. In model 1c3w, this lipid is the one with the lowest temperature factors and it is likely to play a role in the stability of the trimer. The negative peak indicates it is becoming disordered in the N intermediate.
Some other lipid molecules, all in the cytoplasmic leaflet, are also partly in negative density. Most prominent of these is a long stretch of lipid 609 of 1c3w (507 in qhj), which is located adjacent to the F helix and will obviously have to move when this helix tilts (Figure 3C).
Using the ground state model 1qhj (Belrhali et al., 1999) as a starting point, the new positions of the helices in the N state were determined by refitting parts of the structure to the difference densities. To account for the fact that the movements could be less than the resolution limit of 3.5 Å and thus the minimum separation between peak pairs, care was taken to make the distance between the center of the positive density peak and the N position the same as that between the center of the negative peak and the ground state position. For helix G, the part above Lys216 was tilted as a rigid body to bring the main chain of 222–223 and the Leu223 side chain into the strong positive density feature. This also put Arg227 into positive density and took the main chain of 224–225 out of negative density. The maximum displacement was ∼2 Å at Ile222 and the tilt angle ∼7°. Because of the difference densities seen on the E helix adjacent to F, the F and E helices and the EF loop were refitted together to the densities. Assuming an F tilt with a hinge at Pro186 resulted in changes that were too large at the lower part of helix F. Instead, residues 152–176 were moved as a rigid body and 176–186 separately. This resulted in the most satisfactory fit, with an outward movement of the EF loop of ∼3 Å and a tilt of helix F of ∼5°.
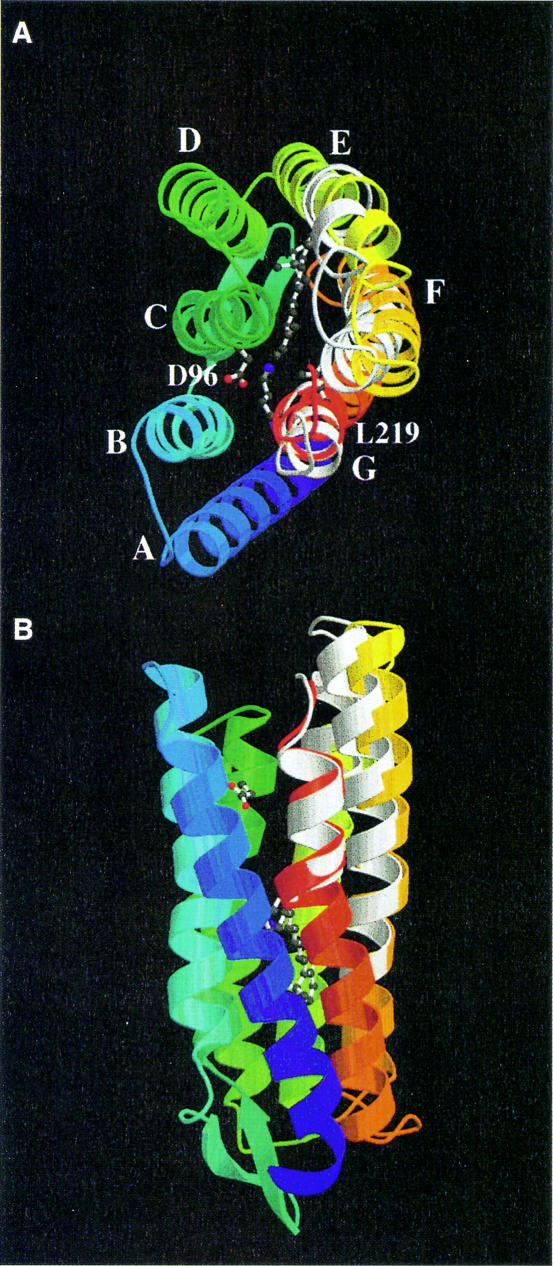
Many difference peaks are visible in the lower part of the structure, but the changes here are on the level of single side chains and thus much smaller than on the cytoplasmic half. These small changes could not be modeled at the present resolution. However, a positive/negative pair near Lys216 and the retinal was consistent with the change observed in the D96N M intermediate (Luecke et al., 1999a) and the cis-retinal structure and the changed G helix from 214 to 217 from model 1c8s were substituted. The model of the N intermediate is shown in Figure 4.
Fig. 4. The N model, colored from blue to red, with the ground state model 1qhj shown in white where the two differ. (A) Top view; (B) side view. Some key residues and the retinal are shown in ball-and-stick representation.
Although the N intermediate as trapped here consists of a mixture of the N and bR conformations in an unknown ratio, the structure in the N state can be calculated by subtracting the appropriate amount of bR from the amplitudes. If x and y are the proportion of N and bR in the N data set, respectively, so ‘N’ = xN + ybR, the bR data set can be represented as xbR + ybR. Subtracting ybR from ‘N’ will give the N intermediate only (xN + ybR – ybR = xN), although with a lower amplitude and consequently more noise. Subtracting <y will result in a map still similar to the ground state (movements too small), whereas subtracting too much will give a result similar to the N–bR difference map (movements too large). Maps were calculated with different amounts of bR subtracted and maps at values of 0.7–0.8 fitted the N structure (Figure 4) best, as shown in Figure 5B. From this it can be concluded that probably ∼20–30% of the molecules are in the N state conformation.
Fig. 5. A 10 Å thick slice of the bR map (purple) with model 1qhj (A) and the N–0.7bR (or 3.33N–2.33bR) map (orange) superimposed on the N model (B). Helices in (A) and residues for orientation in (B) are indicated.
Discussion
The structure of the N intermediate
The difference map of the F219L N intermediate presented here shows unambiguously that the E, F and G helices experience a significant tilt in this part of the photocycle. The tilts seem large enough to allow several water molecules to enter the channel. A role for water in the second half of the photocycle is well established, as dehydration inhibits M decay (Cao et al., 1991; Perkins et al., 1993), and increased hydration is an obvious means to decrease the pKa of Asp96 (Cao et al., 1993). The order of events in the second half of the photocycle can be summarized as follows.
After the proton transfer from the Schiff base to Asp85, the alternative (tilted) helix conformation has become energetically favorable and the helices tilt. The movement of helix G away from helix C makes room for water to hydrate Asp96, which in this environment has a lower pKa and consequently deprotonates. The tilt of helices E and F away from the center of the molecule opens a further water-accessible channel, probably extending to the Schiff base. The amount of water entering the channel cannot be determined yet from this analysis. It could be very small, as increased hydration in the M state of D96N could not be measured by neutron diffraction (Weik et al., 1998). The proton released by Asp96 lowers the pH in this channel to below the pKa of the Schiff base, and the Schiff base reprotonates. In the wild-type protein, the proton transfer will take place as soon as the water channel has formed, so that the MN intermediate does not accumulate, but the N intermediate is formed immediately. In an Asp96 mutant, the M state with a deprotonated Schiff base persists until a proton enters the channel on the cytoplasmic side. The decay of N occurs after Asp96 is reprotonated from the cytoplasm and is pH dependent (Otto et al., 1989; Ames and Mathies, 1990). Cis–trans isomerization and reversal of some of the protein structural changes also occur in this step (Rothschild, 1992); the fact that the N to O transition is pH dependent shows that the Asp96 reprotonation is the rate limiting step.
What factors influence the equilibrium between the closed and open state? Studies on wild-type and mutant bR under different conditions give clues on this. The physiologically important formation of the tilted helix conformation during the lifetime of M in the wild type is clearly induced by the deprotonated Schiff base and its altered hydrogen bonding pattern (Luecke et al., 1999a), as was suggested earlier (Kataoka et al., 1994). The D85N mutant has an open state in the dark at high pH when the Schiff base is deprotonated (Kataoka et al., 1994), which shows it is the deprotonation rather than the 13-cis retinal that causes the conformational change. The initial inducement for the tilt may be a broken hydrogen bond from Lys216 to Gly220 (Luecke et al., 1999a), which makes helix G tilt toward helix F, which moves away in consequence. Alternatively, the effect may be via residues on helix F that interact with retinal (Trp182 and Trp189) or via Tyr185 on helix F by its interaction with Asp212 (Rothschild et al., 1990), with the movement of helix G as a consequence of the change in helix F. As soon as the channel is open, proton transfer from Asp96 via the newly formed water channel to the Schiff base occurs. The N state does not decay until Asp96 is reprotonated (Otto et al., 1989; Ames and Mathies, 1990), indicating that the presence of an anionic Asp96 with a hydration shell stabilizes the open state of the channel. Mutants like F219L and F171C, where a large side chain lining the cytoplasmic channel is replaced by a smaller one, have a long-lived N intermediate (Brown et al., 1994) and some open channels are present in the ground state (Subramaniam et al., 1999). This is probably caused by the presence of water when the channel is closed, making the closed state less favorable than in the wild type where there is tight hydrophobic packing throughout. This effect is even stronger in the triple mutant D96G/F219L/F171C, where the maximum number of channels is already open in the dark and no light-induced structural change is seen (Subramaniam et al., 1999).
The occupancy of the open state
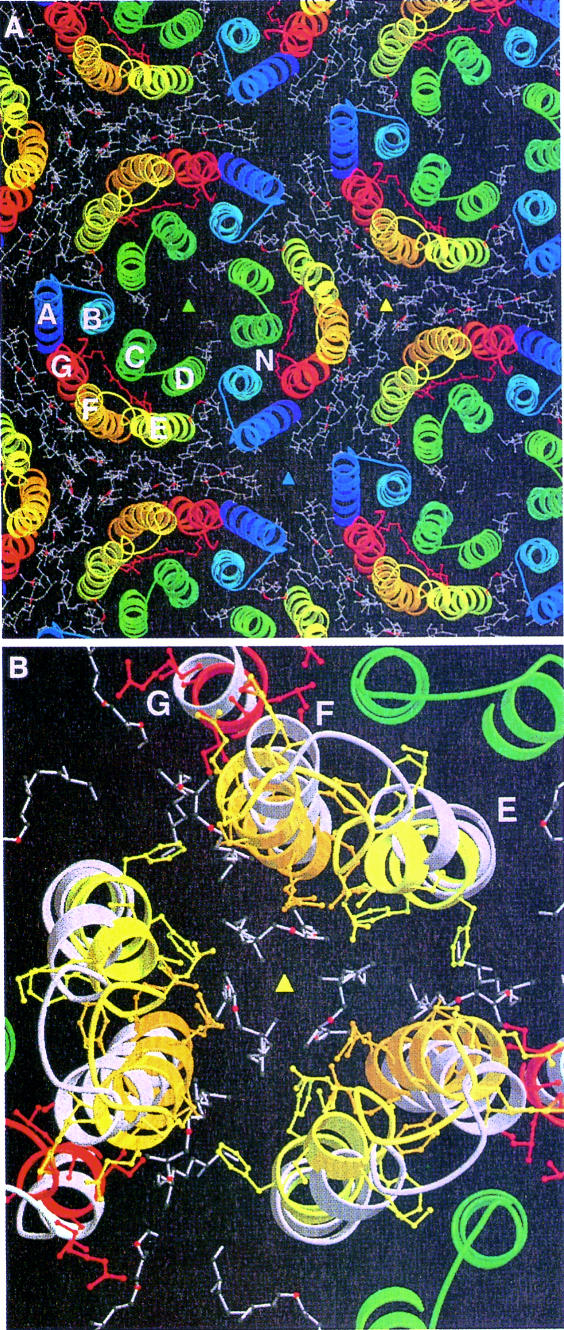
The N structure shown here with a clearly defined new position for helices F and G would seem to contradict the X-ray map of the D96N MN intermediate (Luecke et al., 1999a), where the upper parts of these helices were disordered, even though the crystals appeared to be 100% M. However, a model of the purple membrane in the N state (Figure 6) can explain this discrepancy. The position of the F helix in the purple membrane is such that it tilts toward a 3-fold axis between three monomers from different trimers. The extent of the tilt could be so large that it would not be possible for all three symmetry-related molecules surrounding the 3-fold axis to tilt at the same time because of steric clashes. It seems likely therefore that the maximum occupancy of the tilted helix conformation is 1/3 and molecules already in the M state can only proceed to MN or N when both neighbors (on different trimers) are still or again in the ground state conformation. The resulting structural heterogeneity (broken symmetry) would cause the lipids in the cytoplasmic leaflet to become disordered, rather than moving to a new position, as shown by the fact that in the difference map only negative peaks are associated with the lipid molecules. This model implies that at high occupancies of N, M would accumulate along with N. Detection of this by FTIR spectroscopy is complicated by the fact that the half life of N is limited (a few seconds at –3°C; Vonck, 1996) and M would only be present as long as >1/3 of the molecules are excited. In the spectroscopic studies on F219L (Brown et al., 1994; Vonck, 1996; Yamazaki et al., 1998), some M has only been detected by Yamazaki et al. (1998).
Fig. 6. A model of the purple membrane in the ground state (1qhj) with the molecule in the center (labeled N) replaced by the N model. Lipids are as in the ground state. Colored triangles indicate 3-fold axes. Helices E and F move toward the yellow 3-fold axis. (A) Overview; (B) close-up of the 3-fold axis with 1qhj white, the N model colored with ball-and-stick side chains. The lipid area becomes much smaller and there would not be space for all three molecules to move at the same time.
In the 3D crystals in the M state (Luecke et al., 1999a) that were spectroscopically 100% M, 1/3 of the molecules will have been in the MN state and 2/3 in earlier M according to this model, explaining the disorder that was seen in the tops of helices E, F and G. The fact that the retinal and the residues in the proton exit channel could be modeled without problems shows that no significant changes occur in this region between M and MN. The limited occupancy of the tilted-helix conformation means that it will not be possible to study the MN or N intermediate in a pure state in crystals where the purple membrane packing is retained. The monoclinic crystals used by Essen et al. (1998) are an interesting alternative, because in this crystal packing the three molecules within a trimer are in different environments. The monomers designated ‘B’ and ‘C’ from adjacent, oppositely oriented trimers pack tightly against each other along helices F and G in direct protein–protein contact without intervening lipids. These helices show very low temperature factors, unlike their counterparts in the purple membrane-like crystals, and it is unlikely that they can tilt at all. The monomer ‘A’, however, faces another across a considerable gap of 19 Å and shows the usual high temperature factors on the cytoplasmic side, so it should be able to undergo the conformational changes to form MN or N without interaction with a neighbor.
Photocooperativity
If this model of the interaction between the molecules in the purple membrane is correct, it can explain the so-called cooperative effect in the photocycle that has been an object of much debate because of the need to incorporate it in plausible photocycle models. The effect is seen as an increase in the proportion of a slow-decaying M species at increasing light intensities at the expense of a fast-decaying species. It has been explained variously as resulting from heterogeneity in the photoexcited molecules or from cooperativity within a trimer or within the purple membrane (Hendler et al., 1994; Shrager et al., 1995; Váró et al., 1996; Tokaji, 1998). The same effect is seen when a second light flash is given within a few milliseconds after the first (Tokaji and Dancsházy, 1991). The steric clash between adjacent F helices would explain this effect quite simply. The first of three molecules around an F helix 3-fold axis to reach the M state can decay freely to N with the fast rate constant. At low light intensity, where less than one in three molecules is hit, this situation will be true for most excited molecules. However, the second and third molecules cannot convert to MN until the first one has decayed to O and the F helix is back in the ground state position. This gives rise to the slow decay component, which is related to the N decay rate. In general, cooperativity will thus be seen in conditions where the N decay is slower than the M decay. The fact that at low pH no cooperativity occurs (Radionov and Kaulen, 1995) can be explained by the faster Asp96 reprotonation and thus increased N decay. Also, no cooperativity is observed in D96N (Radionov and Kaulen, 1995), this time because M decay is much slower than N decay. In fact, the seeming lack of cooperativity in this case is artificial because the fast-decaying molecules actually decay not to N but to MN, which cannot be distinguished from the earlier M spectroscopically. When M decay is accelerated by addition of azide, which acts as an external proton donor, cooperativity is restored (Radionov and Kaulen, 1995). M decay becomes faster than N decay and the different rate constants for the first and second excited molecules around the F helix 3-fold axis can now be detected.
Accumulation of an M-like intermediate ‘NM’ (with a different decay rate than the normal M) upon illumination of a sample containing a large amount of N has also been explained as resulting from a photoreaction of N (Kouyama et al., 1988; Kouyama and Nasuda-Kouyama, 1989; Fukuda and Kouyama, 1992). This M population can also be explained as resulting from illumination of bR, not N, while its different decay rate, not its formation, is a consequence of the presence of neighboring molecules in the N state.
This explanation of photocooperativity implies that it occurs between different trimers, not within a trimer as assumed in some models (Korenstein et al., 1979; Tokaji, 1998). That this is indeed the case is shown by the observation that mild Triton X-100 treatment abolished cooperativity before the trimers were disrupted (Mukhopadhyay et al., 1994).
Even at very low light intensities, the slow decay component is never totally absent (Fukuda and Kouyama, 1992; Hendler et al., 1994; Tokaji, 1998). The easiest explanation is that the ground state is heterogeneous and contains some molecules with a tilted-helix conformation. The existence of N- or O-like molecules has been shown experimentally at high pH and temperature (Fukuda and Kouyama, 1992) and is correlated with a larger amplitude of the slow decay component of M. Difference maps between mutants and wild type in the dark showed that the N-like conformation is present to some extent in the ground state of mutants with a long-lived N intermediate (Subramaniam et al., 1999). This includes F219L, the protein that was used in this study. The amount of N already present is difficult to quantify, but from the difference maps it appears to be comparable to the amount of subsequent light-induced change. This effect can explain the unexpectedly low proportion of N formed in the bR to N transition. The maximum number of molecules in the N conformation is 33%; those that are already in this conformation in the dark get subtracted in the difference map, so the amount of N produced was probably close to the maximum obtainable in this mutant.
Materials and methods
The N intermediate and bR ground state were trapped as described previously (Vonck, 1996): purple membranes of the F219L mutant (Brown et al., 1994) were fused to make larger crystals (Baldwin and Henderson, 1984). Glucose-embedded electron microscopy samples were made by the back-injection method (Glaeser and Downing, 1990) on molybdenum grids (Pacific Grid Tech, Palo Alto, CA). The use of molybdenum grids was essential to obtain flat crystals (Booy and Pawley, 1993; Han et al., 1994b). The grids were hydrated at 80% relative humidity (Perkins et al., 1993) before plunging into liquid nitrogen in the dark. For the N intermediate, the grid was warmed to 270 K, illuminated for 30 s with green light and rapidly cooled with liquid nitrogen again. To trap the bR ground state, the grid was kept in the dark.
Electron diffraction data of the N intermediate and bR ground state were recorded at 45° tilt in a Gatan cold state at liquid nitrogen temperatures at 400 or 120 kV on a Gatan or TVIPS 1K × 1K CCD camera. The data was processed using the MRC suite of programs (Baldwin and Henderson, 1984; Ceska and Henderson, 1990). Only patterns that diffracted to at least 3.5 Å and had an Rsym of <12% were selected for merging. N–bR difference Fourier maps were calculated using image phases (Grigorieff et al., 1996). Fitting of the model was done using the program O (Jones and Kjeldgaard, 1993).
Acknowledgments
Acknowledgements
I am grateful to Robert Glaeser and Werner Kühlbrandt for their support and discussions in different stages of this work. I thank Janos Lanyi for the gift of the F219L strain, Felicia Hendrickson for isolating the purple membrane and Deryck Mills for technical assistance. I thank Robert Glaeser, Felicia Hendrickson, Werner Kühlbrandt and Cécile Breyton for comments on the manuscript. This work was supported by DFG Sonderforschungsbereich 472.
References
- Ames J.B. and Mathies,R.A. (1990) The role of back-reactions and proton uptake during the N to O transition in bacteriorhodopsin’s photocycle: a kinetic resonance Raman study. Biochemistry, 29, 7181–7190. [DOI] [PubMed] [Google Scholar]
- Baldwin J.M. and Henderson,R. (1984) Measurement and evaluation of electron diffraction patterns from two-dimensional crystals. Ultramicroscopy, 14, 319–336. [Google Scholar]
- Belrhali H., Nollert,P., Royant,A., Menzel,C., Rosenbusch,J.P., Landau,E.M. and Pebay-Peyroula,E. (1999) Protein, lipid and water organization in bacteriorhodopsin crystals: a molecular view of the purple membrane at 1.9 Å resolution. Structure, 7, 909–917. [DOI] [PubMed] [Google Scholar]
- Booy F.P. and Pawley,J.B. (1993) Cryo-crinkling: what happens to carbon films on copper grids at low temperature. Ultramicroscopy, 48, 273–280. [DOI] [PubMed] [Google Scholar]
- Brown L.S., Yamazaki,Y., Maeda,A., Sun,L., Needleman,R. and Lanyi,J.K. (1994) The proton transfers in the cytoplasmic domain of bacteriorhodopsin are facilitated by a cluster of interacting residues. J. Mol. Biol., 239, 401–414. [DOI] [PubMed] [Google Scholar]
- Bullough P.A. and Henderson,R. (1999) The projection structure of the low temperature K intermediate of the bacteriorhodopsin photocycle determined by electron diffraction. J. Mol. Biol., 286, 1663–1671. [DOI] [PubMed] [Google Scholar]
- Cao Y., Váró,G., Chang,M., Ni,B., Needleman,R. and Lanyi,J.K. (1991) Water is required for proton transfer from aspartate-96 to the bacteriorhodopsin Schiff base. Biochemistry, 30, 10972–10979. [DOI] [PubMed] [Google Scholar]
- Cao Y., Váró,G., Klinger,A.L., Czajkowsky,D.M., Braiman,M.S., Needleman,R. and Lanyi,J.K. (1993) Proton transfer from Asp-96 to the bacteriorhodopsin Schiff base is caused by a decrease of the pKa of Asp-96 which follows a protein backbone conformational change. Biochemistry, 32, 1981–1990. [DOI] [PubMed] [Google Scholar]
- Ceska T.A. and Henderson,R. (1990) Analysis of high-resolution electron diffraction patterns from purple membrane labelled with heavy-atoms. J. Mol. Biol., 213, 539–560. [DOI] [PubMed] [Google Scholar]
- Dencher N.A., Dresselhaus,D., Zaccai,G. and Büldt,G. (1989) Structural changes in bacteriorhodopsin during proton translocation revealed by neutron diffraction. Proc. Natl Acad. Sci. USA, 86, 7876–7879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Edman K., Nollert,P., Royant,A., Belrhali,H., Pebay-Peyroula,E., Hajdu,J., Neutze,R. and Landau,E.M. (1999) High-resolution X-ray structure of an early intermediate in the bacteriorhodopsin photocycle. Nature, 401, 822–826. [DOI] [PubMed] [Google Scholar]
- Esnouf R.M. (1997) An extensively modified version of MolScript that includes greatly enhanced coloring capabilities. J. Mol. Graph. Model., 15, 132–134. [DOI] [PubMed] [Google Scholar]
- Esnouf R.M. (1999) Further additions to MolScript version 1.4, including reading and contouring of electron-density maps. Acta Crystallogr. D, 55, 938–940. [DOI] [PubMed] [Google Scholar]
- Essen L.O., Siegert,R., Lehmann,W.D. and Oesterhelt,D. (1998) Lipid patches in membrane protein oligomers—crystal structure of the bacteriorhodopsin–lipid complex. Proc. Natl Acad. Sci. USA, 95, 11673–11678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fukuda K. and Kouyama,T. (1992) Photoreaction of bacteriorhodopsin at high pH: origins of the slow decay component of M. Biochemistry, 31, 11740–11747. [DOI] [PubMed] [Google Scholar]
- Glaeser R.M. and Downing,K.H. (1990) The ‘specimen flatness’ problem in high-resolution electron crystallography of biological macromolecules. In Peachy,L.D. and Williams,D.B. (eds), XIIth International Congress for Electron Microscopy. San Francisco Press, Inc., Seattle, WA, Vol. 1, pp. 98–99. [Google Scholar]
- Grigorieff N., Ceska,T.A., Downing,K.H., Baldwin,J.M. and Henderson,R. (1996) Electron-crystallographic refinement of the structure of bacteriorhodopsin. J. Mol. Biol., 259, 393–421. [DOI] [PubMed] [Google Scholar]
- Han B.-G., Vonck,J. and Glaeser,R.M. (1994a) The bacteriorhodopsin photocycle: direct structural study of two substates of the M-intermediate. Biophys. J., 67, 1179–1186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han B.-G., Wolf,S.G., Vonck,J. and Glaeser,R.M. (1994b) Specimen flatness in electron crystallography of glucose-embedded biological materials is significantly affected by the choice of carbon evaporation stock. Ultramicroscopy, 55, 1–5. [DOI] [PubMed] [Google Scholar]
- Haupts U., Tittor,J. and Oesterhelt,D. (1999) Closing in on bacteriorhodopsin: progress in understanding the photocycle. Annu. Rev. Biophys. Biomol. Struct., 28, 367–399. [DOI] [PubMed] [Google Scholar]
- Hauss T., Büldt,G., Heyn,M.P. and Dencher,N.A. (1994) Light-induced isomerization causes an increase in the chromophore tilt in the M intermediate of bacteriorhodopsin: a neutron diffraction study. Proc. Natl Acad. Sci. USA, 91, 11854–11858. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henderson R., Baldwin,J.M., Ceska,T.A., Zemlin,F., Beckmann,E. and Downing,K.H. (1990) A model for the structure of bacteriorhodopsin based on high resolution electron cryomicroscopy. J. Mol. Biol., 213, 899–929. [DOI] [PubMed] [Google Scholar]
- Hendler R.W., Dancsházy,Z., Bose,S., Shrager,R.I. and Tokaji,Z. (1994) Influence of excitation energy on the bacteriorhodopsin photocycle. Biochemistry, 33, 4604–4610. [DOI] [PubMed] [Google Scholar]
- Hendrickson F.M., Burkard,F. and Glaeser,R.M. (1998) Structural characterization of the L-to-M transition of the bacteriorhodopsin photocycle. Biophys. J., 75, 1446–1454. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones T.A. and Kjeldgaard,M. (1993) O—The Manual. Uppsala University, Uppsala, Sweden. [Google Scholar]
- Kamikubo H., Kataoka,M., Váró,G., Oka,T., Tokunaga,F., Needleman,R. and Lanyi,J.K. (1996) Structure of the N intermediate of bacteriorhodopsin revealed by X-ray diffraction. Proc. Natl Acad. Sci. USA, 93, 1386–1390. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kamikubo H., Oka,T., Imamoto,Y., Tokunaga,F., Lanyi,J.K. and Kataoka,M. (1997) The last phase of the reprotonation switch in bacteriorhodopsin: the transition between the M-type and the N-type protein conformation depends on hydration. Biochemistry, 36, 12282–12287. [DOI] [PubMed] [Google Scholar]
- Kataoka M., Kamikubo,H., Tokunaga,F., Brown,L.S., Yamazaki,Y., Maeda,A., Sheves,M., Needleman,R. and Lanyi,J.K. (1994) Energy coupling in an ion pump. The reprotonation switch of bacteriorhodopsin. J. Mol. Biol., 243, 621–638. [DOI] [PubMed] [Google Scholar]
- Kimura Y., Vassylyev,D.G., Miyazawa,A., Kidera,A., Matsushima,M., Mitsuoka,K., Murata,K., Hirai,T. and Fujiyoshi,Y. (1997) Surface of bacteriorhodopsin revealed by high-resolution electron crystallography. Nature, 389, 206–211. [DOI] [PubMed] [Google Scholar]
- Koch M.H.J., Dencher,N.A., Oesterhelt,D., Plöhn,H.-J., Rapp,G. and Büldt,G. (1991) Time-resolved X-ray diffraction study of structural changes associated with the photocycle of bacteriorhodopsin. EMBO J., 10, 521–526. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Korenstein R., Hess,B. and Markus,M. (1979) Cooperativity in the photocycle of purple membrane of Halobacterium halobium with a mechanism of free energy transduction. FEBS Lett., 102, 155–161. [Google Scholar]
- Kouyama T. and Nasuda-Kouyama,A. (1989) Turnover rate of the proton pumping cycle of bacteriorhodopsin: pH and light-intensity dependences. Biochemistry, 28, 5963–5970. [Google Scholar]
- Kouyama T., Nasuda-Kouyama,A., Ikegami,A., Mathew,M.K. and Stoeckenius,W. (1988) Bacteriorhodopsin photoreaction: identification of a long-lived N (P, R350) at high pH and its M-like photoproduct. Biochemistry, 27, 5855–5863. [DOI] [PubMed] [Google Scholar]
- Kraulis P.J. (1991) MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr., 24, 946–950. [Google Scholar]
- Lanyi J.K. (1998) Understanding structure and function in the light-driven proton pump bacteriorhodopsin. J. Struct. Biol., 124, 164–178. [DOI] [PubMed] [Google Scholar]
- Lindahl M. and Henderson,R. (1997) Structure of the bacteriorhodopsin D85N/D96N double mutant showing substantial structural changes and a highly twinned, disordered lattice. Ultramicroscopy, 70, 95–106. [Google Scholar]
- Luecke H., Richter,H.-T. and Lanyi,J.K. (1998) Proton transfer pathways in bacteriorhodopsin at 2.3 Å resolution. Science, 280, 1934–1937. [DOI] [PubMed] [Google Scholar]
- Luecke H., Schobert,B., Richter,H.T., Cartailler,J.P. and Lanyi,J.K. (1999a) Structural changes in bacteriorhodopsin during ion transport at 2 Å resolution. Science, 286, 255–260. [DOI] [PubMed] [Google Scholar]
- Luecke H., Schobert,B., Richter,H.T., Cartailler,J.P. and Lanyi,J.K. (1999b) Structure of bacteriorhodopsin at 1.55 Å resolution. J. Mol. Biol., 291, 899–911. [DOI] [PubMed] [Google Scholar]
- Merritt E.A. and Bacon,D.J. (1997) Raster3D: photorealistic molecular graphics. Methods Enzymol., 277, 505–524. [DOI] [PubMed] [Google Scholar]
- Mitsuoka K., Hirai,T., Murata,K., Miyazawa,A., Kidera,A., Kimura,Y. and Fujiyoshi,Y. (1999) The structure of bacteriorhodopsin at 3.0 Å resolution based on electron crystallography: implication of the charge distribution. J. Mol. Biol., 286, 861–882. [DOI] [PubMed] [Google Scholar]
- Mukhopadhyay A.K., Bose,S. and Hendler,R.W. (1994) Membrane-mediated control of the bacteriorhodopsin photocycle. Biochemistry, 33, 10889–10895. [DOI] [PubMed] [Google Scholar]
- Nakasako M., Kataoka,M., Amemiya,Y. and Tokunaga,F. (1991) Crystallographic characterization by X-ray diffraction of the M-intermediate from the photocycle of bacteriorhodopsin at room temperature. FEBS Lett., 292, 73–75. [DOI] [PubMed] [Google Scholar]
- Otto H., Marti,T., Holz,M., Mogi,T., Lindau,M., Khorana,H.G. and Heyn,M.P. (1989) Aspartic acid-96 is the internal proton donor in the reprotonation of the Schiff base of bacteriorhodopsin. Proc. Natl Acad. Sci. USA, 86, 9228–9232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pebay-Peyroula E., Rummel,G., Rosenbusch,J.P. and Landau,E.M. (1997) X-ray structure of bacteriorhodopsin at 2.5 Å from microcrystals grown in lipidic cubic phases. Science, 277, 1676–1681. [DOI] [PubMed] [Google Scholar]
- Perkins G.A., Burkard,F., Liu,E. and Glaeser,R.M. (1993) Glucose alone does not completely hydrate bacteriorhodopsin in glucose-embedded purple membrane. J. Microsc., 169, 61–65. [DOI] [PubMed] [Google Scholar]
- Radionov A.N. and Kaulen,A.D. (1995) Cooperative phenomena in the photocycle of D96N mutant bacteriorhodopsin. FEBS Lett., 377, 330–332. [DOI] [PubMed] [Google Scholar]
- Rothschild K.J. (1992) FTIR difference spectroscopy of bacterio rhodopsin: toward a molecular model. J. Bioenerg. Biomembr., 24, 147–167. [DOI] [PubMed] [Google Scholar]
- Rothschild K.J., Braiman,M.S., He,Y.-W., Marti,T. and Khorana,H.G. (1990) Vibrational spectroscopy of bacteriorhodopsin mutants. Evidence for the interaction of aspartic acid 212 with tyrosine 185 and possible role in the proton pump mechanism. J. Biol. Chem., 265, 16985–16991. [PubMed] [Google Scholar]
- Sasaki J., Shichida,Y., Lanyi,J.K. and Maeda,A. (1992) Protein changes associated with reprotonation of the Schiff base in the photocycle of Asp96Asn bacteriorhodopsin. The MN intermediate with unprotonated Schiff base but N-like protein structure. J. Biol. Chem., 267, 20782–20786. [PubMed] [Google Scholar]
- Sass H.J., Schachowa,I.W., Rapp,G., Koch,M.H.J., Oesterhelt,D., Dencher,N.A. and Büldt,G. (1997) The tertiary structural changes in bacteriorhodopsin occur between M states: X-ray diffraction and Fourier transform infrared spectroscopy. EMBO J., 16, 1484–1491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shrager R.I., Hendler,R.W. and Bose,S. (1995) The ability of actinic light to modify the bacteriorhodopsin photocycle. Heterogeneity or photocooperativity? Eur. J. Biochem., 229, 589–595. [DOI] [PubMed] [Google Scholar]
- Subramaniam S., Gerstein,M., Oesterhelt,D. and Henderson,R. (1993) Electron diffraction analysis of structural changes in the photocycle of bacteriorhodopsin. EMBO J., 8, 1–12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Subramaniam S., Lindahl,M., Bullough,P., Faruqi,A.R., Tittor,J., Oesterhelt,D., Brown,L., Lanyi,J. and Henderson,R. (1999) Protein conformational changes in the bacteriorhodopsin photocycle. J. Mol. Biol., 287, 145–161. [DOI] [PubMed] [Google Scholar]
- Tokaji Z. (1998) Quantitative model for the cooperative interaction of the bacteriorhodopsin molecules in purple membrane. FEBS Lett., 423, 343–346. [DOI] [PubMed] [Google Scholar]
- Tokaji Z. and Dancsházy,Z. (1991) Light-induced, long-lived perturbation of the photocycle of bacteriorhodopsin. FEBS Lett., 281, 170–172. [DOI] [PubMed] [Google Scholar]
- Váró G., Needleman,R. and Lanyi,J.K. (1996) Protein structural change at the cytoplasmic surface as the cause of cooperativity in the bacteriorhodopsin photocycle. Biophys. J., 70, 461–467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vonck J. (1996) A three-dimensional difference map of the N intermediate in the bacteriorhodopsin photocycle: part of the F helix tilts in the M to N transition. Biochemistry, 35, 5870–5878. [DOI] [PubMed] [Google Scholar]
- Weik M., Zaccai,G., Dencher,N.A., Oesterhelt,D. and Hauss,T. (1998) Structure and hydration of the M-state of the bacteriorhodopsin mutant D96N studied by neutron diffraction. J. Mol. Biol., 275, 625–634. [DOI] [PubMed] [Google Scholar]
- Yamazaki Y., Kandori,H., Needleman,R., Lanyi,J.K. and Maeda,A. (1998) Interaction of the protonated Schiff base with the peptide backbone of valine 49 and the intervening water molecule in the N photointermediate of bacteriorhodopsin. Biochemistry, 37, 1559–1564. [DOI] [PubMed] [Google Scholar]