The endoplasmic reticulum (ER) is the major source of molecules that directly interact with the cell’s external environment. This occurs through the secretory pathway, which starts from the ER and regulates macromolecular traffic to the cell surface and inner hydrolytic compartments. It is thus no wonder that the ER functions as a major, very efficient station that integrates signaling from the environment and takes decisions on appropriate responses. The most promiscuous ER signaling mechanism is the unfolded protein response (UPR) (1). A major UPR sensing system appears to be conserved in all eukaryotes and monitors the workload of the ER in the folding and assembly of newly synthesized secretory proteins. This workload can be increased when environmental conditions negatively affect protein folding or when developmental response programs require higher synthesis of secretory proteins. In these situations, ER resident transmembrane sensors start a cascade of various events that eventually alters the expression of many genes, mainly stimulating the synthesis of ER folding helpers and at the same time repressing general synthesis of passenger secretory proteins (1). When the response is not sufficient, unresolved UPR induces programmed cell death (PCD). In PNAS, two studies (2, 3) now provide important progress in characterizing the activity of upstream and downstream players of plant UPR and their relationships with developmental programs as well as PCD.
Inositol requiring enzyme1 (IRE1), the only UPR sensor identified in budding yeast, is a transmembrane ER protein present also in plants and metazoans. Its luminal domain interacts with the major ER chaperone immunoglobulin binding protein (BiP) and thus senses the amount of resting BiP (4). When ER workload increases, BiP is displaced, IRE1 oligomerizes, authophosphorylates, and activates its ribonuclease cytosolic domain, starting a series of events that is not exactly the same in all eukaryotes but always leads to major gene expression alterations. Additional, kingdom-specific UPR transmembrane sensors exist, probably reflecting the diverse strategies available to unicellular eukaryotes, metazoans or plants: PKR-like ER kinase (PERK) and the activating transcription factor 6 (ATF6) in metazoans and the basic leucine zipper (bZIP) transcription factors bZIP28 and bZIP17 in plants (1). All of them are BiP interactors. The major substrate of plant IRE1 is the transcription factor bZIP60. This is normally an integral ER transmembrane protein, but IRE1 ribonuclease activity removes an intron, causing a frameshift leading to the synthesis of an alternative, soluble bZIP60 form that can thus migrate into the nucleus (5, 6). BiP release from bZIP28 and bZIP17 instead induces their traffic from the ER to the Golgi apparatus, followed by removal of the transmembrane domains by Golgi proteases (7–9). This mechanism is similar to the one that activates ATF6, whereas metazoan IRE1 acts slightly differently from the plant homolog: its substrate mRNA encodes a soluble but inactive transcription factor activated upon IRE1-mediated intron removal (1).
There are two IRE1 genes in Arabidopsis: IRE1a and IRE1b. Single KOs do not have growth defects, but the double ire1a ire1b mutant is defective in root growth also in the absence of imposed UPR stress, indicating that the sensors play a role during normal growth of a highly secreting tissue (10). This double mutant is also more sensitive to tunicamycin, an inhibitor of protein N-glycosylation and the most widely used UPR inducer (10). Deng et al. (2) stacked KO mutations of IRE1 and other known Arabidopsis UPR sensors and analyzed seedling survival, as well as root elongation in the absence of stress or on UPR (in this case induced by the reducing agent DTT). The triple mutant bzip28, ire1a ire1b is lethal. The phenotype of bzip28 ire1b is similar to that of ire1a ire1b, indicating that a fully active IRE1 system is needed in the absence of the second branch of UPR sensing. Surprisingly, bzip28 bzip60 has no growth defect, suggesting that IRE1 genes play a role in growth independently of their action on bZIP60 (2). Conversely, on DTT treatment, bzip28 bzip60 has much more severe root growth defects than bzip28 ire1b, confirming, as expected, an important involvement of bZIP60 in UPR stress response. The bZIP60-independent function of IRE1 requires both its kinase and ribonuclease activities, as indicated by expressing in the ire1a ire1b background ire1b mutated constructs impaired in either activity (2), but the direct molecular targets of this function are as yet unknown.
Several abiotic stresses and pathogen attack are UPR inducers in plants (1), which cannot change their physical location when dangers approach and often rapidly respond with increased synthesis of specific soluble or membrane proteins of the secretory pathway. The transcriptome changes induced by different stresses are of course not totally overlapping, but analysis of genes similarly induced by classic UPR inducers and common environmental stresses can give clues on signaling integration. A modest overlap in transcriptional response occurs in soybean on osmotic or tunicamycin stress (11). The few overlapping genes were considered candidate regulatory components that may integrate the osmotic stress and ER stress signaling pathways. Among them, two development and cell death domain–containing proteins [N-rich proteins (NRPs)] and the NRP-induced NAC transcription factor GmNAC81 (formerly GmNAC6) were identified (12, 13). Further work indicated that that NRP expression is positively regulated by the GmERD15 (glycine max early responsive to dehydration 15) transcription factor (14). In search for GmNAC81 interactors, Mendes et al. (3) now identify GmNAC30 as a partner. The authors show that GmNAC 81 and GmNAC30 interact in the nucleus of plant cells; bind to a common, specific DNA consensus sequence; enhance each other’s synthesis; and have a synergetic effect in inducing the synthesis of numerous hydrolases, including four of the five vacuolar processing enzymes (VPEs) of soybean (3). VPEs have caspase1-like activity and are major executors of both pathogen-induced and developmental PCD (15, 16). In mammals, UPR-induced apoptosis is mediated by the activation of IRE1 and PERK and thus relies on BiP-mediated sensing mechanisms (17). Consistently, a predicted and largely observed phenomenon is that BiP or IRE1 overexpression attenuates UPR, including apoptosis (18–20). However, the tunicamycin induction of NRPs in soybean is not relieved by BiP overexpression (12), and the classic folding helpers (including BiP) induced by tunicamycin are not induced by osmotic stress (3). This indicates that Mendes et al. have now identified the downstream executors of an important PCD mechanism that is induced by UPR but is dependent from upstream signaling performed by as yet unknown sensors.
Besides activating its major target bZIP60, the endonuclease activity of IRE1 also participates in attenuating secretory protein synthesis by selectively degrading ER-localized mRNAs, a process termed regulated IRE1-dependent decay (RIDD) (1). The UPR PERK sensor of metazoans is both implicated in attenuating protein synthesis (by inhibiting the activity of initiation factor eIF2α) and in apoptosis, but it seems to be absent in plants (1). It has thus been proposed that RIDD may be more important in UPR-induced cell death in plants than in other eukaryotes (1, 21). However, tunicamycin-induced PCD is enhanced in ire1a ire1b Arabidopsis with respect to WT (21), thus pointing to the existence of UPR-dependent IRE1-independent PCD signaling events in line with the results of Mendes et al. (3).
The signaling networks revealed by Mendes et al. and Deng et al. underline the intricate networks of UPR, plant stress response, and developmental programs but are still lacking the identification of important players (question marks in Fig. 1). A first catalog of Arabidopsis RIDD targets has been produced (21). Comparisons of this list with those of genes affected by different abiotic and biotic stresses, as well as more detailed analysis on how the plants mutated in known UPR players cope with stress and developmental programs, will help in future research. It should also be underlined that general protein folding inhibitors may negatively affect the activity of many soluble and membrane secretory proteins; thus, it is not always easy to separate the effects due to UPR activation from those that may directly result from the misfolding of proteins involved in signal transduction events. More refined UPR activation can be obtained by overexpressing an individual mutated protein with folding defects (22). These tools may be very valuable in defining the different pathways.
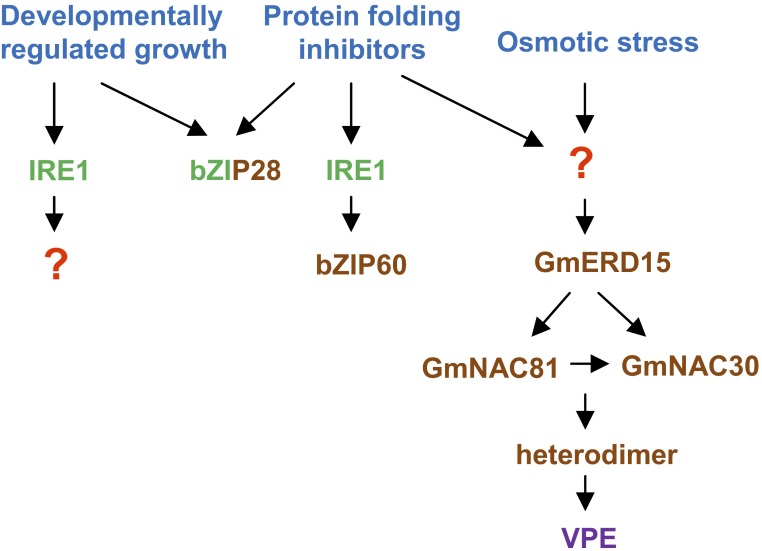
Fig. 1.
Signaling relationships between UPR, osmotic stress-induced PCD, and developmentally programmed root elongation. The inducing conditions (blue), upstream sensors (green), and induced transcription factors (brown) are shown. bZIP28 is both an upstream sensor and a transcription factor. The red question marks indicate as yet unknown players. Transcription of VPE genes (as well other genes not indicated in the figure) is the final outcome of the signaling pathway at right. For simplicity, the final transcriptional changes of IRE1- and bZIP28-mediated classic plant UPR are not shown.
Footnotes
References
- 1.Hollien J. Evolution of the unfolded protein response. Biochim Biophys Acta. 2013;1833(11):2458–2463. doi: 10.1016/j.bbamcr.2013.01.016. [DOI] [PubMed] [Google Scholar]
- 2.Deng Y, Srivastava R, Howell SH. Protein kinase and ribonuclease domains of IRE1 confer stress tolerance, vegetative growth, and reproductive development in Arabidopsis. Proc Natl Acad Sci USA. 2013;110:19633–19638. doi: 10.1073/pnas.1314749110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Mendes GC, et al. GmNAC30 and GmNAC81 integrate the endoplasmic reticulum stress- and osmotic stress-induced cell death responses through a vacuolar processing enzyme. Proc Natl Acad Sci USA. 2013;110:19627–19632. doi: 10.1073/pnas.1311729110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Carrara M, Prischi F, Ali MMU. UPR Signal activation by luminal sensor domains. Int J Mol Sci. 2013;14(3):6454–6466. doi: 10.3390/ijms14036454. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Deng Y, et al. Heat induces the splicing by IRE1 of a mRNA encoding a transcription factor involved in the unfolded protein response in Arabidopsis. Proc Natl Acad Sci USA. 2011;108(17):7247–7252. doi: 10.1073/pnas.1102117108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Nagashima Y, et al. Arabidopsis IRE1 catalyses unconventional splicing of bZIP60 mRNA to produce the active transcription factor. Sci Rep. 2011;1:29. doi: 10.1038/srep00029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Tajima H, Iwata Y, Iwano M, Takayama S, Koizumi N. Identification of an Arabidopsis transmembrane bZIP transcription factor involved in the endoplasmic reticulum stress response. Biochem Biophys Res Commun. 2008;374(2):242–247. doi: 10.1016/j.bbrc.2008.07.021. [DOI] [PubMed] [Google Scholar]
- 8.Che P, et al. Signaling from the endoplasmic reticulum activates brassinosteroid signaling and promotes acclimation to stress in Arabidopsis. Sci Signal. 2010;3(141):ra69. doi: 10.1126/scisignal.2001140. [DOI] [PubMed] [Google Scholar]
- 9.Liu JX, Srivastava R, Che P, Howell SH. An endoplasmic reticulum stress response in Arabidopsis is mediated by proteolytic processing and nuclear relocation of a membrane-associated transcription factor, bZIP28. Plant Cell. 2007;19(12):4111–4119. doi: 10.1105/tpc.106.050021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Chen Y, Brandizzi F. AtIRE1A/AtIRE1B and AGB1 independently control two essential unfolded protein response pathways in Arabidopsis. Plant J. 2012;69(2):266–277. doi: 10.1111/j.1365-313X.2011.04788.x. [DOI] [PubMed] [Google Scholar]
- 11.Irsigler AS, et al. Expression profiling on soybean leaves reveals integration of ER- and osmotic-stress pathways. BMC Genomics. 2007;8:431. doi: 10.1186/1471-2164-8-431. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Costa MDL, et al. A new branch of endoplasmic reticulum stress signaling and the osmotic signal converge on plant-specific asparagine-rich proteins to promote cell death. J Biol Chem. 2008;283(29):20209–20219. doi: 10.1074/jbc.M802654200. [DOI] [PubMed] [Google Scholar]
- 13.Faria JA, et al. The NAC domain-containing protein, GmNAC6, is a downstream component of the ER stress- and osmotic stress-induced NRP-mediated cell-death signaling pathway. BMC Plant Biol. 2011;11:129. doi: 10.1186/1471-2229-11-129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Alves MS, et al. A novel transcription factor, ERD15 (Early Responsive to Dehydration 15), connects endoplasmic reticulum stress with an osmotic stress-induced cell death signal. J Biol Chem. 2011;286(22):20020–20030. doi: 10.1074/jbc.M111.233494. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Hatsugai N, et al. A plant vacuolar protease, VPE, mediates virus-induced hypersensitive cell death. Science. 2004;305(5685):855–858. doi: 10.1126/science.1099859. [DOI] [PubMed] [Google Scholar]
- 16.Hara-Nishimura I, Hatsugai N. The role of vacuole in plant cell death. Cell Death Differ. 2011;18(8):1298–1304. doi: 10.1038/cdd.2011.70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Tabas I, Ron D. Integrating the mechanisms of apoptosis induced by endoplasmic reticulum stress. Nat Cell Biol. 2011;13(3):184–190. doi: 10.1038/ncb0311-184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Gu X, et al. Bip overexpression, but not CHOP inhibition, attenuates fatty-acid-induced endoplasmic reticulum stress and apoptosis in HepG2 liver cells. Life Sci. 2010;87(23-26):724–732. doi: 10.1016/j.lfs.2010.10.012. [DOI] [PubMed] [Google Scholar]
- 19.Tirasophon W, Welihinda AA, Kaufman RJ. A stress response pathway from the endoplasmic reticulum to the nucleus requires a novel bifunctional protein kinase/endoribonuclease (Ire1p) in mammalian cells. Genes Dev. 1998;12(12):1812–1824. doi: 10.1101/gad.12.12.1812. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Wang XZ, et al. Cloning of mammalian Ire1 reveals diversity in the ER stress responses. EMBO J. 1998;17(19):5708–5717. doi: 10.1093/emboj/17.19.5708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Mishiba K, et al. Defects in IRE1 enhance cell death and fail to degrade mRNAs encoding secretory pathway proteins in the Arabidopsis unfolded protein response. Proc Natl Acad Sci USA. 2013;110(14):5713–5718. doi: 10.1073/pnas.1219047110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Oono Y, et al. Analysis of ER stress in developing rice endosperm accumulating beta-amyloid peptide. Plant Biotechnol J. 2010;8(6):691–718. doi: 10.1111/j.1467-7652.2010.00502.x. [DOI] [PubMed] [Google Scholar]