Abstract
Background
The capture and use of disease-related anatomic pathology data for both model organism phenotyping and human clinical practice requires a relatively simple nomenclature and coding system that can be integrated into data collection platforms (such as computerized medical record-keeping systems) to enable the pathologist to rapidly screen and accurately record observations. The MPATH ontology was originally constructed in 2,000 by a committee of pathologists for the annotation of rodent histopathology images, but is now widely used for coding and analysis of disease and phenotype data for rodents, humans and zebrafish.
Construction and content
MPATH is divided into two main branches describing pathological processes and structures based on traditional histopathological principles. It does not aim to include definitive diagnoses, which would generally be regarded as disease concepts. It contains 888 core pathology terms in an almost exclusively is_a hierarchy nine layers deep. Currently, 86% of the terms have textual definitions and contain relationships as well as logical axioms to other ontologies such the Gene Ontology.
Application and utility
MPATH was originally devised for the annotation of histopathological images from mice but is now being used much more widely in the recording of diagnostic and phenotypic data from both mice and humans, and in the construction of logical definitions for phenotype and disease ontologies. We discuss the use of MPATH to generate cross-products with qualifiers derived from a subset of the Phenotype and Trait Ontology (PATO) and its application to large-scale high-throughput phenotyping studies. MPATH provides a largely species-agnostic ontology for the descriptions of anatomic pathology, which can be applied to most amniotes and is now finding extensive use in species other than mice. It enables investigators to interrogate large datasets at a variety of depths, use semantic analysis to identify the relations between diseases in different species and integrate pathology data with other data types, such as pharmacogenomics.
Keywords: Pathology, Ontology, Disease, Mouse, Phenotype
Background
Since the late eighteenth century when achromatic lenses and reliable histological stains began to be available, investigators of anatomic pathology, and particularly in the mid -nineteenth century the innovators of cellular pathology such as Rudolf Virchow, developed and applied terminologies to describe their observations [1,2]. These depended on the “school” to which the pathologists belonged, but more importantly on the etiologic or mechanistic paradigm in which they were working [3]. One of the great achievements of the nineteenth century was the recognition of the universality of pathological processes and entities and their occurrence in multiple species as recognisable manifestations of the same underlying processes [4]. It was, nevertheless, a century before broadly accepted and rationally structured pathology terminologies were developed (e.g. [5]). The development of pathology terminologies has to an extent occurred independently of disease terminologies and nosologies, partly as a result of the much longer history of classifying diseases, and partly due to the inherited preconceptions of the nature of disease in clinical medicine.
The distinction between pathological and clinical descriptions of disease, disorders and predispositions is still not satisfactorily resolved. However, in recent years there have been attempts to rationalise the definitions of these concepts [6] and their relation to each other as part of a broadly applicable model of disease other than an unstructured collection of manifestations or phenotypes which are found in that class of individuals forming the basis of a diagnosis. Issues about severity, time course, organ involvement etc. are beginning to be addressed, but it is remarkable that even treating diseases as a “bag“ of phenotypes has been shown to provide a powerful approach in establishing the relationships between diseases, and the presence of related diseases in different organisms [7-10]. What has recently been identified as important, nevertheless, is that the tissue-specific resolution of the recording of lesions, and the ability to record the pattern of disease within an individual, has proved vital for GWAS mapping of predisposing genetic variants in inbred strains of mice allowing each class of lesion to be analysed in isolation [11,12].
The discipline of pathology may be broken down into clinical and anatomic pathology, the former is concerned with clinical chemistry, hematology, clinical microbiology and emerging sub-specialities such as molecular diagnostics and proteomics. The latter, which forms the domain of MPATH, deals with the histological, histochemical or immunohistochemical observations of alterations in tissue composition or architecture. Both branches of the medical specialty, which are increasingly merging, may be viewed as aspects of phenotyping, and both provide subtypes of the clinical signs associated with ongoing disease processes, the results of developmental abnormalities, or the historical presence of disease.
Anatomic pathology nomenclature and its applications
The universality of the repertoire of responses to underlying genetic or extrinsic insults means that gross and histopathologically-defined phenotypes are some of the most useful phenotypes for relating diseases between different species, and constitute some of the most species-agnostic phenotype descriptors. This makes a pathologic term-based ontology a crucial tool in experimental and clinical phenotype data capture [13].
The development of systematic human pathologic nomenclatures has been driven by the efforts of the American College of Pathologists, initially with the development of the pathology specific nomenclature (SNOP) over 40 years ago [14] to the current SNOMED –CT with cross references to UMLS, the NCI thesaurus and other terminologies. The ICD [15], now in its 11th revision and the associated ICD-O v-3 for cancer, also contains descriptions of many pathological lesions.
The other driver for pathologic terminology standardisation has been coding of lesions from toxicopathology. The American Society of Toxicopathology (STP) working with Registry of Industrial Toxicology Animal-data (RITA) database group in Europe has produced several internationally accepted nomenclature systems, particularly focusing on proliferative lesions. Recently, the STP has undertaken a major harmonization exercise for rodent pathology – the INHAND (International Harmonization of Nomenclature and Diagnostic Criteria for Lesions in Rats and Mice) initiative [16]. So far this group has reported on the hepaticobiliary, respiratory, nervous and urinary systems [17-20]. For some time the National Cancer Institute’s Mouse Models of Human Cancer consortium (MMHCC) has been examining the classification of tumours in genetically engineered mice. MMHCC has produced a consensus base terminology for neoplasias of the major organ systems that have been presented in a series of papers over the last decade [21].
Despite the huge value of these resources, none is currently constructed as an ontology with meaningful axioms to support inference and automated reasoning, and to that end we developed MPATH to describe lesions that arise in laboratory mice.
Construction and content
The MPATH ontology was constructed ab initio by a group of clinical and veterinary pathologists in 2,000 and has since been revised and augmented by an evolving group of US and European pathologists on a regular basis. It is clear from more than a decade of experience that expert input and manual curation are essential to generate an accurate and functional resource. One strategy for building the ontology has been to integrate it into large-scale phenotyping and diagnostic programs so that the pathologists use it on a daily basis and have fields to add missing terms or synonyms that they are more familiar with thereby constantly increasing its coverage and utilitarian value.
MPATH contains 888 classes in an almost exclusively is_a hierarchy eight layers deep. Part_of relations are used only in the case of, for example, adenomatous polyp (MPATH:490) and intraductal papilloma (MPATH:285) where pathologists distinguish between a macroscopic lesion (“–osis”) and individual lesions which make them up; both need to be described. The top level distinction in MPATH is between pathophysiology (pathological processes) and anatomic pathology (pathological entities). One issue met frequently in development was the normal practice of referring to observation of a physical lesion by using the process term; for example “necrosis” or “sclerosis”. Thus the noun describing the real-world entity observed is homonymous with the inferred process. This problem has been addressed through the textual and logical definitions of terms but is a recurrent source of confusion in formal treatments of pathology nomenclature. Pathologists using MPATH almost exclusively use the anatomic pathology segment of the ontology with the exception of describing inflammation or other general processes where the process is described using qualifiers such as “acute” which are logically process-specific, consistent with the use of Phenotype And Trait Ontology (PATO) [22] qualifiers (see below). The upper levels of MPATH’s anatomic pathology branch include six broad domains familiar to all traditions of pathology training and comprehensively covering all known lesions; cell and tissue damage, circulatory disorder, developmental and structural abnormality, growth and differentiation defect, healing and repair structure, and neoplasm, and are as orthogonal as is feasible given the complexities of pathobiology (see Figure 1A). The upper levels of MPATH’s pathophysiology branch denote pathological processes that underlie lesions and include six broad domains; cell and tissue damage process (see Figure 1C), defective growth and differentiation process, developmental process abnormalities, healing and repair process, immunopathological process, and neoplasia. All pathological processes and entities can be placed within these upper level domains, which will be familiar to all pathologists and are common to all amniotes.
Figure 1.
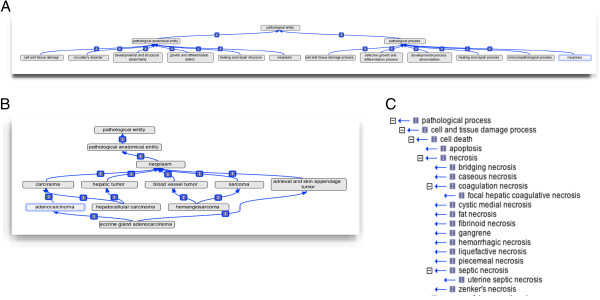
Structure of MPATH. A; overall structure of the top level classes in MPATH showing division into pathological process and physical entity for the major divisions. B; polyhierarchy showing multiple parentage for some anatomically predicated tumor classes and their anatomical and morphological classification. C; cell and tissue damage process segment of MPATH showing main pathological process classes in this branch.
Relationship to upper level and other ontologies
MPATH is largely congruent with the upper level Ontology for General Medical Science (OGMS) [6], founded on the Basic Formal Ontology (BFO). Pathological bodily process (OGMS:0000061) and pathological anatomical structure (OGMS:0000077) are broadly mappable to the upper levels of MPATH; MPATH:603 (pathological anatomical entity) and MPATH: 596 (pathological process) respectively. However, more detailed mapping is difficult. For example the MPATH experts view congenital malformations as pathological anatomical structures, whereas OGMS views them as distinct, and similarly MPATH views inflammation as a pathological process whereas OGMS does not include this as a pathological bodily process. Until such discrepancies are resolved, integration of MPATH into the OGMS framework will be problematical.
From the point of view of application, the most important mappings for MPATH are to the Human Phenotype Ontology (125), the Mammalian Phenotype ontology (111), the Disease Ontology (231), SNOMED-CT (867) and the NCIt (566), reflecting the emphasis on the domain of anatomic pathology rather than disease.
Definitions and axiomatic relationships
Currently, 86% of classes have textual definitions. Each class is in the mouse pathology namespace and is uniquely identified by a URI of the form: http://purl.obolibrary.org/OBO/MPATH_n. The main ontology is available in both the OBO Flatfile Format and the Web Ontology Language (OWL). MPATH is housed in a subversion repository and is made available via OBO registry, Bioportal (http://purl.bioontology.org/ontology/MPATH) and on the project’s website http://mpath.googlecode.com/. MPATH contains relationships and other logical axioms to other ontologies such the Gene Ontology (GO) [23], Cell Type ontology (CL) [24] and the Phenotype And Trait Ontology (PATO) [22]. For example, the MPATH term transitional cell metaplasia (MPATH:172) represents a metaplastic response of the transitional epithelium, for example in the bladder to give squamous metaplasia and glandular metaplasia. To allow computational access to these relations, we use the derives-from relation and relate metaplasia (MPATH:549) (an MPATH term that denotes an abnormal transformation of a differentiated adult cell or tissue of one kind into a differentiated tissue of another kind) with the CL term transitional epithelial cell (CL:0000244).
Application and utility
A post-composition strategy for pathology coding
Traditionally pathologists have relied on a narrative form of recording their definitive diagnoses, making use of morphologic, etiologic, and disease-based terms that collectively provide a diagnosis useful for clinical patient management. This is particularly important for non–neoplastic lesions where it can be complex to capture important subtleties of distribution, severity, microscopic sub-type and anatomical location for example. Whilst this is the gold standard, it is not possible to compute on data recorded in this way and it is very difficult to tabulate and quantitatively analyse the collected information. There are strong arguments, mainly from experience in toxicologic pathology, that a descriptive (anatomic) rather than diagnostic coding is the most objective and useful way to code pathology-based observations. This is particularly relevant to examination of mutant mice where traditional etiologic or summative diagnostic terms are simply not available because of the novelty of the lesion or its presentation. This is particularly the case where mice are manipulated to model human conditions that have not been previously seen, for example lung or mammary tumours [11,25,26] which have not previously been reported to occur spontaneously in mice. In many cases, a disease diagnosis implies a particular pathogenesis or etiology based on the spontaneous disease, which is not appropriate for the disease caused by genetic and sometimes both genetic and external challenge combined. This latter issue is of particular concern to practicing pathologists and in the development of MPATH we have been urged to include some diagnostic terms as well as descriptive anatomic ones.
Many tissue responses are common to multiple anatomical sites and as far as possible the verbosity (ontology “bloat”) of specifying a particular response in multiple tissues has been avoided, with the additional topographical or anatomical information for description coming from an anatomy ontology, generally the MA [27] or EMAP ontologies [28] for the mouse, however, there is often an intrinsic anatomical element embedded in the term or traditional pathology includes information about the cell type or tissue of origin. This is most frequent with the neoplasias and we felt that such terms were best included in their familiar form. Figure 1B shows how anatomically predicated classes such as hepatocellular carcinoma have multiple parents, providing relations in this case to both carcinoma and hepatic tumor superclasses. Most observations made by pathologists using MPATH are, nevertheless, cross-products using a combination of an MPATH term and an anatomical (MA) or cell type (CL) [24,29] component. This strategy provides all of the necessary coverage.
In addition to the core terms in MPATH, it is important to describe organ-specific topography, distribution, microscopic character, duration/chronicity and severity. These are not included in MPATH but drawn from other ontologies. These qualifiers or modifiers are generally applicable across a wide range of organs and lesions and so need to be coded separately to the core terms to allow post-composition as required. The pattern we have adopted is very close to that recommended by the INHAND proposals and also includes “compound” terms which lie beneath a definitive diagnosis or disease level of description, but bundle defined sets of descriptive terms, for example “nephropathy”, “alopecia”, “glomerulonephritis” which are in common use and well understood. These qualifiers have been incorporated into PATO, and some examples are given in Table 1. The strategy for composing pathology descriptions using the combination of MPATH, MA and PATO is summarised in Figure 2.
Table 1.
Examples of pathology term qualifiers now incorporated into PATO
| Qualifier | PATO | Class name | Definition |
|---|---|---|---|
| Severity |
0000461 |
Normal |
No lesions |
| |
0000394 |
Mild |
Lesion dependent; often size, number and characteristics. |
| |
0000395 |
Moderate |
|
| |
0000465 |
Marked |
|
| |
0000396 |
Severe |
|
| Duration |
0002387 |
Per-acute |
Extremely acute and aggressive |
| |
0000389 |
Acute |
Beginning abruptly with marked intensity |
| |
0002091 |
Subacute |
Between acute and chronic |
| |
0001863 |
Chronic |
Slow progress and long continuance |
| |
0002387 |
Chronic-active |
Coexistence of chronic process and superimposed acute process |
| Distribution |
0000627 |
Focal |
Single well delineated lesion |
| |
0002388 |
Focally extensive |
Single lesion with expansion into surrounding tissue |
| |
0001791 |
Multifocal |
Multiple lesions |
| |
0002389 |
Multifocal to coalescing |
Multiple lesions some interconnecting with each other |
| |
0000330 |
Random |
No appreciable pattern |
| |
0001566 |
Diffuse |
Not circumscribed or limited |
| |
0000635 |
Generalized |
Affecting all regions without specificity of distribution |
| |
0000634 |
Unilateral |
Confined to one side only |
| |
0000618 |
Bilateral |
Involving both sides |
| 0002389 | Segmental | Relating to a segment |
Figure 2.
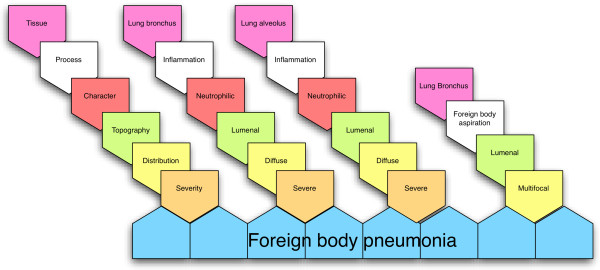
Post-composition coding strategy. Elements of the compound description are specified by the string of linked pentagons on the left hand side of the figure and specific examples given for three observations which taken together are indicative of foreign body pneumonia. Tissue terms are taken from the appropriate anatomy ontology, eg. MA, Process from MPATH, character, topology, distribution and severity from PATO. The combination of observations defines the disease foreign body pneumonia.
Implementation of MPATH coding strategy
The strategy adopted was originally designed to describe histopathology images for the Pathbase mouse pathology database [30], but lends itself readily to a wide range of coding applications. The MPATH strategy has been adopted by two major high-throughput studies. A combination of MPATH and PATO is being used for the capture of pathology data from the genome-wide mutant mouse phenotyping project, KOMP2 run as part of the International Mouse Phenotyping Consortium [31], where the MPATH approach is being used in the primary phenotyping pipeline by the Toronto Centre for Phenogenomics and other centres carrying out histopathology. MPATH has also been adopted for the MoDIS database [32] to capture and analyse pathology data from a massive aging study which has systematically phenotyped 31 of the most important inbred mouse strains. Complete necropsies of mice were carried out at 12 and 20 months of age (cross-sectional study) and moribund mice in the life span (longitudinal study). Nearly 2,000 mice were necropsied, generating more than 50,000 slides [33]. Lesion incidence and severity data for all organs is now being applied in highly successful GWAS studies of age-associated disease [33].
MPATH has proved to be additionally useful in dealing with the recoding of multi-species legacy data from non-standard nomenclatures, permitting integration of otherwise siloed data. Examples are the European Radiological archive (ERA) database where 6,700 human diagnoses were recoded from ICD-8 and the Klinischer Diagnosenschleussel [34] to MPATH/FMA [35], and with the Northwestern University Janus radiobiology database (http://janus.northwestern.edu/janus2/), who have coded 50,000 individual mouse records to MPATH to link the two datasets. Recently the ontology has been applied to zebrafish phenotype data in the Zfin database [36] indicating a useful application of MPATH to non-mammalian species which could be developed further.
MPATH as a core ontology for PATO-based logical definitions
The PATO framework was built with the intention of providing an integration platform for phenotype data between species and between data types [22]. According to the PATO framework, phenotype data can be described by utilising species-specific ontologies (such as the various anatomy ontologies) or species-agnostic ontologies such as GO with the various qualities provided by the PATO ontology in order to describe affected entities in a phenotype manifestation. PATO can be used for annotation either directly in a so-called post-composed (post-coordinated) manner or for providing logical definitions (equivalence axioms) to ontologies containing a set of precomposed (pre-coordinated) phenotype terms [22,37-39]. For further discussion see [40].
Rather than using a pre-composed phenotype ontology such as MP [29] or HPO [41], phenotypes may be described using the Entity–Quality (EQ) formalism. In the EQ method, a phenotype is characterized by an affected Entity and a Quality (from PATO) that specifies how the entity is affected. The affected entity can either be a biological function or process such as specified in GO, or an anatomical entity. The phylogenetic conservation, at least within the amniotes, of most histopathologic lesions or processes makes MPATH an important core ontology in writing logical definitions and we have used it extensively in defining classes in the major pre-composed phenotype ontologies and MPATH is an important component ontology of our recently developed semantic approaches to comparative phenomics – PhenomeNET and Mousefinder [8,9].
Composition of logical definitions is a time-consuming task for which there are currently several approaches to automation using class label segmentation, entity recognition and lexical matching to core ontologies. This approach can be useful for suggesting definitions where the class label is a composite of for example, anatomy and process (MA + GO). Automated decomposition of unilexical terms such as are found in the neoplasias is much more difficult though approaches with text mining definitions from other ontologies such as NCIt for lexically matching labels may be useful to expert curators in establishing more simple definitions for these classes.
Conclusions
Whilst MPATH was originally designed to support rodent, and particularly mouse, pathology the extensive overlap with human pathology means that most of the terms may be used in a human context and linked to the foundational model of anatomy (FMA) [42] as the anatomy ontology. Extending MPATH to become a mammalian pathology ontology encompassing human pathology is a major undertaking, but we have established that the current structure and upper level classes would readily support the inclusion of human terminology. Initially we will import terms for neoplasias from the CINEAS codes (Central Information System for Hereditary Diseases and Synonyms; http://www.cineas.org/; Prof Rolf Sijmons, pers, comm). SNOMED-CT, UMLS and ICD-O v3 will be mined for terms not currently in MPATH which relate to anatomic pathology. Terms already covered by existing ontologies such as Disease Ontology (DO) [43] may be referenced using MIREOT [44]. DO classifies diseases largely by anatomical site and not by disease process or class, and overlaps only slightly with MPATH as it is concerned with summative diagnostic entities for the main part. For example there is no “inflammation” superclass in DO for the tissue specific inflammatory conditions described.
Use of MPATH to construct logical definitions for DO classes would potentially add a further dimension to the richness and applicability of DO.
The power of the description of pathological lesions to discriminate between diseases and therefore between models of human disease is substantial. We recently estimated the information content (IC) of pre-composed MP ontology terms used to code phenotypes in the EUMODIC mouse phenotyping pipeline [45], which included or excluded anatomic pathology descriptions, using their logical definitions. Pathology-related phenotypes were shown to have a significantly greater discriminatory power than other in vivo assays, strongly supporting the use of these assays in the development of mouse models of human diseases [13].
Further development and application of MPATH will inevitably depend on community engagement and we encourage anyone with an interest to provide feedback.
Abbreviations
CINEAS: Central Information System for Hereditary Diseases and Synonyms; CL: Cell ontology; DO: Disease ontology; EMAP: EMouse atlas project; ERA: European radiobiology archive; EUMODIC: European mouse disease clinic; FMA: Foundational model of anatomy; GWAS: Genome-wide association study; HPO: Human phenotype ontology; ICD: International classification of disease; ICD-O: International classification of disease – oncology; IMPC: International Mouse Phenotyping Consortium; INHAND: International Harmonization of Nomenclature and Diagnostic Criteria for Lesions in Rats and Mice; KOMP2: Knockout mouse project 2; MA: Mouse anatomy ontology; MIREOT: Minimum information to reference an external ontology term; MMHCC: Mouse Models of Human Cancer consortium; MPATH: Mouse pathology ontology; MP: Mouse phenotype ontology; NCIt: National Cancer Institute Thesaurus; OBO: Open biological ontology; PATO: Phenotype and trait ontology; RITA: Registry of Industrial Toxicology Animal-data; SNOMED-CT: Systematized nomenclature of medicine clinical terms; STP: Society for toxicopathology; UMLS: Unified medical language system.
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
All of the authors have made major contributions to the writing and coding of the ontology and its definitions over many years. The paper was written by PNS, JPS, and GVG. All authors read and approved the final manuscript.
Contributor Information
Paul N Schofield, Email: ps@mole.bio.cam.ac.uk.
John P Sundberg, Email: john.sundberg@jax.org.
Beth A Sundberg, Email: beth.sundberg@jax.org.
Colin McKerlie, Email: colin.mckerlie@phenogenomics.ca.
Georgios V Gkoutos, Email: geg18@aber.ac.uk.
Acknowledgements
The authors would like to thank those who have contributed to the development and application of MPATH over the years. This work was funded by the European Commission. Contract QLRI-1999-00320, the Ellison Medical Foundation, National Institutes of Health AG25707, for the Shock Aging Center, CA89713, and AR056635 and AR063781 to JPS, and HG004838-04 to PNS.
References
- Virchow R. Cellular pathology as based upon physiological and pathological histology. 2. New York: R M DeWitt; 1860. [DOI] [PubMed] [Google Scholar]
- Plaut A. Rudolf Virchow and today's physicians and scientists. Bull Hist Med. 1953;27:236–251. [PubMed] [Google Scholar]
- Hajdu SI. Inventors of new laboratory and pathology terms. Ann Clin Lab Sci. 2007;37:192–194. [PubMed] [Google Scholar]
- Sundberg JP, Schofield PN. One medicine, one pathology, and the one health concept. J Am Vet Med Assoc. 2009;234:1530–1531. doi: 10.2460/javma.234.12.1530. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taylor CR, Hartsock RJ. Classifications of lymphoma; reflections of time and technology. Virchows Archiv. 2011;458:637–648. doi: 10.1007/s00428-011-1083-0. [DOI] [PubMed] [Google Scholar]
- Scheuermann RH, Ceusters W, Smith B. Toward an ontological treatment of disease and diagnosis. Summit Transl bioinformatics. 2009;2009:116–120. [PMC free article] [PubMed] [Google Scholar]
- Washington NL, Haendel MA, Mungall CJ, Ashburner M, Westerfield M, Lewis SE. Linking human diseases to animal models using ontology-based phenotype annotation. PLoS Biol. 2009;7:e1000247. doi: 10.1371/journal.pbio.1000247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen CK, Mungall CJ, Gkoutos GV, Doelken SC, Kohler S, Ruef BJ, Smith C, Westerfield M, Robinson PN, Lewis SE. et al. MouseFinder: Candidate disease genes from mouse phenotype data. Hum Mutat. 2012;33:858–866. doi: 10.1002/humu.22051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoehndorf R, Schofield PN, Gkoutos GV. PhenomeNET: a whole-phenome approach to disease gene discovery. Nucleic Acids Res. 2011;39:e119. doi: 10.1093/nar/gkr538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oellrich A, Hoehndorf R, Gkoutos GV, Rebholz-Schuhmann D. Improving disease gene prioritization by comparing the semantic similarity of phenotypes in mice with those of human diseases. PloS One. 2012;7:e38937. doi: 10.1371/journal.pone.0038937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berndt A, Cario CL, Silva KA, Kennedy VE, Harrison DE, Paigen B, Sundberg JP. Identification of Fat4 and Tsc22d1 as novel candidate genes for spontaneous pulmonary adenomas. Cancer Res. 2011;71:5779–5791. doi: 10.1158/0008-5472.CAN-11-1418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Q, Berndt A, Guo H, Sundberg JP, Uitto J. A Novel Animal Model for Pseudoxanthoma Elasticum: The KK/HlJ Mouse. Am J Pathol. 2012;181(4):1190–1196. doi: 10.1016/j.ajpath.2012.06.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schofield PN, Vogel P, Gkoutos GV, Sundberg JP. Exploring the elephant: histopathology in high-throughput phenotyping of mutant mice. Dis Model Mech. 2011;5:19–25. doi: 10.1242/dmm.008334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cornet R, De Keizer N. Forty years of SNOMED: a literature review. BMC Med Inform Decis Mak. 2008;8(Suppl 1):2. doi: 10.1186/1472-6947-8-S1-S2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- World Health Organisation. International Statistical Classification of Diseases and Health Related Problems (The) ICD-10. Geneva: WHO; 2008. [Google Scholar]
- Mann PC, Vahle J, Keenan CM, Baker JF, Bradley AE, Goodman DG, Harada T, Herbert R, Kaufmann W, Kellner R. et al. International harmonization of toxicologic pathology nomenclature: an overview and review of basic principles. Toxicol Pathol. 2012;40:7S–13S. doi: 10.1177/0192623312438738. [DOI] [PubMed] [Google Scholar]
- Kaufmann W, Bolon B, Bradley A, Butt M, Czasch S, Garman RH, George C, Groters S, Krinke G, Little P. et al. Proliferative and nonproliferative lesions of the rat and mouse central and peripheral nervous systems. Toxicol Pathol. 2011;40:87S–157S. doi: 10.1177/0192623312439125. [DOI] [PubMed] [Google Scholar]
- Frazier KS, Seely JC, Hard GC, Betton G, Burnett R, Nakatsuji S, Nishikawa A, Durchfeld-Meyer B, Bube A. Proliferative and nonproliferative lesions of the rat and mouse urinary system. Toxicol Pathol. 2011;40:14S–86S. doi: 10.1177/0192623312438736. [DOI] [PubMed] [Google Scholar]
- Thoolen B, Maronpot RR, Harada T, Nyska A, Rousseaux C, Nolte T, Malarkey DE, Kaufmann W, Kuttler K, Deschl U. et al. Proliferative and nonproliferative lesions of the rat and mouse hepatobiliary system. Toxicol Pathol. 2010;38:5S–81S. doi: 10.1177/0192623310386499. [DOI] [PubMed] [Google Scholar]
- Renne R, Brix A, Harkema J, Herbert R, Kittel B, Lewis D, March T, Nagano K, Pino M, Rittinghausen S. et al. Proliferative and nonproliferative lesions of the rat and mouse respiratory tract. Toxicol Pathol. 2009;37:5S–73S. doi: 10.1177/0192623309353423. [DOI] [PubMed] [Google Scholar]
- Marks C. Mouse Models of Human Cancers Consortium (MMHCC) from the NCI. Dis Model Mech. 2009;2:111. doi: 10.1242/dmm.002725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gkoutos GV, Green ECJ, Mallon A-M, Hancock JM, Davidson D. Building mouse phenotype ontologies. Pac Symp Biocomput. 2004;9:178–189. doi: 10.1142/9789812704856_0018. [DOI] [PubMed] [Google Scholar]
- Gene Ontology Consortium. The Gene Ontology in 2010: extensions and refinements. Nucleic Acids Res. 2010;38:331–335. doi: 10.1093/nar/gkp1018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bard J, Rhee SY, Ashburner M. An ontology for cell types. Genome Biol. 2005;6:R21. doi: 10.1186/gb-2005-6-2-r21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meuwissen R, Berns A. Mouse models for human lung cancer. Genes Dev. 2005;19:643–664. doi: 10.1101/gad.1284505. [DOI] [PubMed] [Google Scholar]
- Derksen PW, Liu X, Saridin F, van der Gulden H, Zevenhoven J, Evers B, Van Beijnum JR, Griffioen AW, Vink J, Krimpenfort P. et al. Somatic inactivation of E-cadherin and p53 in mice leads to metastatic lobular mammary carcinoma through induction of anoikis resistance and angiogenesis. Cancer Cell. 2006;10:437–449. doi: 10.1016/j.ccr.2006.09.013. [DOI] [PubMed] [Google Scholar]
- Hayamizu TF, Mangan M, Corradi JP, Kadin JA, Ringwald M. The adult mouse anatomical dictionary: a tool for annotating and integrating data. Genome Biol. 2005;6:R29. doi: 10.1186/gb-2005-6-3-r29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Richardson L, Venkataraman S, Stevenson P, Yang Y, Burton N, Rao J, Fisher M, Baldock RA, Davidson DR, Christiansen JH. EMAGE mouse embryo spatial gene expression database: 2010 update. Nucleic Acids Res. 2010;38:D703–709. doi: 10.1093/nar/gkp763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meehan TF, Masci AM, Abdulla A, Cowell LG, Blake JA, Mungall CJ, Diehl AD. Logical development of the cell ontology. BMC Bioinform. 2011;12:6. doi: 10.1186/1471-2105-12-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schofield PN, Gruenberger M, Sundberg JP. Pathbase and the MPATH Ontology: Community Resources for Mouse Histopathology. Vet Pathol. 2010;47:1016–1020. doi: 10.1177/0300985810374845. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown SD, Moore MW. Towards an encyclopaedia of mammalian gene function: the International Mouse Phenotyping Consortium. Dis Model Mech. 2012;5:289–292. doi: 10.1242/dmm.009878. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sundberg JP, Sundberg BA, Schofield P. Integrating mouse anatomy and pathology ontologies into a phenotyping database: tools for data capture and training. Mamm Genome. 2008;19:413–419. doi: 10.1007/s00335-008-9123-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sundberg J, Berndt A, Sundberg B, Silva KA, Kennedy V, Bronson R, Yuan R, Paigen B, Harrison D, Schofield PN. The mouse as a model for understanding chronic diseases of aging: the histopathologic basis of aging in inbred mice. Pathobiol Aging Age-related Dis. 2011;1:71719. doi: 10.3402/pba.v1i0.7179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Immich H. Klinischer Diagnosenschlüssel. Stuttgart: Schattauer; 1968. [Google Scholar]
- Tapio S, Schofield PN, Adelmann C, Atkinson MJ, Bard JL, Bijwaard H, Birschwilks M, Dubus P, Fiette L, Gerber G. et al. Progress in updating the European Radiobiology Archives. Int J Radiat Biol. 2008;84:930–936. doi: 10.1080/09553000802460214. [DOI] [PubMed] [Google Scholar]
- Howe DG, Bradford YM, Conlin T, Eagle AE, Fashena D, Frazer K, Knight J, Mani P, Martin R, Moxon SA. et al. ZFIN, the Zebrafish Model Organism Database: increased support for mutants and transgenics. Nucleic Acids Res. 2013;41:D854–860. doi: 10.1093/nar/gks938. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gkoutos GV, Mungall C, Dolken S, Ashburner M, Lewis S, Hancock J, Schofield P, Kohler S, Robinson PN. Entity/quality-based logical definitions for the human skeletal phenome using PATO. Conf Proc IEEE Eng Med Biol Soc. 2009;1:7069–7072. doi: 10.1109/IEMBS.2009.5333362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mungall CJ, Gkoutos GV, Smith CL, Haendel MA, Lewis SE, Ashburner M. Integrating phenotype ontologies across multiple species. Genome Biol. 2009;11:R2. doi: 10.1186/gb-2010-11-1-r2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gkoutos GV, Mungall C, Dolken S, Ashburner M, Lewis S, Hancock J, Schofield P, Kohler S, Robinson PN. Entity/quality-based logical definitions for the human skeletal phenome using PATO. Conf Proc IEEE Eng Med Biol Soc. 2009;2009:7069–7072. doi: 10.1109/IEMBS.2009.5333362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schofield PN, Sundberg JP, Hoehndorf R, Gkoutos GV. New approaches to the representation and analysis of phenotype knowledge in human diseases and their animal models. Brief Funct Genomics. 2012;10:258–265. doi: 10.1093/bfgp/elr031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robinson PN, Mundlos S. The human phenotype ontology. Clin Genet. 2010;77:525–534. doi: 10.1111/j.1399-0004.2010.01436.x. [DOI] [PubMed] [Google Scholar]
- Rosse C, Mejino JL Jr. A reference ontology for biomedical informatics: the Foundational Model of Anatomy. J Biomed Inform. 2003;36:478–500. doi: 10.1016/j.jbi.2003.11.007. [DOI] [PubMed] [Google Scholar]
- Schriml LM, Arze C, Nadendla S, Chang YW, Mazaitis M, Felix V, Feng G, Kibbe WA. Disease Ontology: a backbone for disease semantic integration. Nucleic Acids Res. 2012;40:D940–946. doi: 10.1093/nar/gkr972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Courtot M, Frank G, Allyson LL, James M, Daniel S, Ryan RB, Alan R. MIREOT: The minimum information to reference an external ontology term. Appl Ontol. 2011;6:23–33. [Google Scholar]
- Morgan H, Beck T, Blake A, Gates H, Adams N, Debouzy G, Leblanc S, Lengger C, Maier H, Melvin D. et al. EuroPhenome: a repository for high-throughput mouse phenotyping data. Nucleic Acids Res. 2012;38:D577–585. doi: 10.1093/nar/gkp1007. [DOI] [PMC free article] [PubMed] [Google Scholar]


