Abstract
The strictly regulated expression of most pleiotropic developmental control genes is critically dependent on the activity of long-range cis-regulatory elements. This was revealed by the identification of individuals with a genetic condition lacking coding-region mutations in the gene commonly associated with the disease but having a variety of nearby chromosomal abnormalities, collectively described as cis-ruption disease cases. The congenital eye malformation aniridia is caused by haploinsufficiency of the developmental regulator PAX6. We discovered a de novo point mutation in an ultraconserved cis-element located 150 kb downstream from PAX6 in an affected individual with intact coding region and chromosomal locus. The element SIMO acts as a strong enhancer in developing ocular structures. The mutation disrupts an autoregulatory PAX6 binding site, causing loss of enhancer activity, resulting in defective maintenance of PAX6 expression. These findings reveal a distinct regulatory mechanism for genetic disease by disruption of an autoregulatory feedback loop critical for maintenance of gene expression through development.
Main Text
Advances in genomic sequencing and epigenomics have firmly established that key developmental regulatory genes with pleiotropic functions (e.g., PAX6 [MIM 607108], SHH [MIM 600725], SOX9 [MIM 608160]) are surrounded by many, often highly conserved cis-regulatory elements, spread over hundreds of kilobases from the transcription start site.1,2 Most cis-elements function as transcriptional enhancers and are the primary determinants of precise tissue-specific gene regulation. Developments in human genetics have greatly contributed to recognition of the fact that the spatiotemporally and quantitatively appropriate activity of a gene requires not only the presence of intact coding sequence but also properly functioning regulatory control. However, in contrast to coding-region mutations, the identification of causal variants in regulatory regions and the validation of their role in disruption of appropriate gene expression remains a major problem. Nevertheless, because noncoding variation is now recognized as a major factor in both Mendelian and common genetic disease, the elucidation of the various control mechanisms whose disruption can have phenotypic consequences is essential for future diagnostic and therapeutic prospects.
Aniridia (MIM 106210) is a panocular disease characterized by a variable degree of iris hypoplasia, foveal hypoplasia, nystagmus, and ciliary body abnormalities. Affected individuals are often afflicted by progressive sight-threatening complications such as glaucoma, cataracts, and keratopathy resulting from corneal limbal stem cell deficiency.3 Haploinsufficiency of the developmental control gene PAX6 has been identified as the cause of classical aniridia, through loss-of-function point mutations or gene deletions in around 90% of patients (PAX6 mutation database) and abnormalities in the genomic region downstream of PAX6 in a further set of individuals.4–6 Transcriptional control of Pax6 expression depends on a large array of cis-elements.7–13 Analysis of people with aniridia carrying chromosomal abnormalities 3′ of the PAX6 transcription unit revealed the crucial role played by a cluster of distant, highly conserved control elements, the DRR, encoding a variety of eye and other tissue-specific activities (Figure 1A).13 The most distant breakpoints, in two familial cases with classical aniridia, were identified at 125 kb (SGL) and 150 kb (SIMO) downstream of the PAX6 P1 promoter site.4 Nevertheless, an additional group of aniridia-affected individuals remained for which the causative genetic defect remains unresolved. The well-established role of PAX6 coding mutations and chromosomal abnormalities in aniridia disease etiology presented the opportunity for exploration of additional mechanisms of cis-regulatory disruption in this group.
Figure 1.
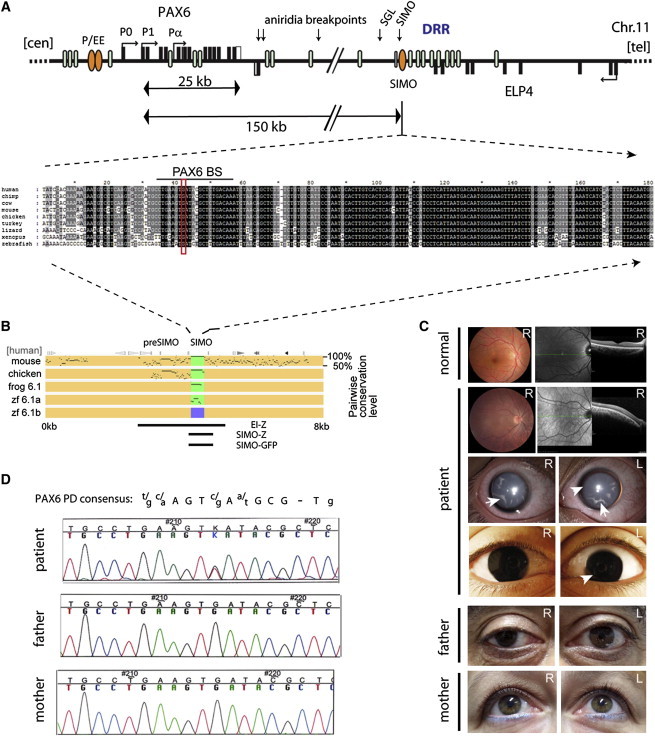
Aniridia Subject ID 1230P Carries a Mutation in a Remote, Ultraconserved PAX6 Regulatory Element
(A) Map of the PAX6 locus on human chromosome 11, displaying the exons of PAX6 (black rectangles, top strand) and adjacent ELP4 gene (black rectangles, bottom strand), whose introns contain long-range cis-elements (blue ellipses) for PAX6 (including the distal regulatory region [DRR]). The SIMO element (orange ellipse), located 150 kb downstream, is deeply conserved with strong sequence similarity across vertebrates. The alignment shows the centromeric part of the element. The mutation changes a 100% conserved residue in a highly conserved sequence block with strong similarity to a PAX6 binding site.
(B) Visualization of evolutionary sequence conservation by percentage identity plot (PIP). Green boxes highlight presence of the element and purple indicates its absence in the zebrafish pax6.1b locus. Fragments characterized in the EI-Z,13 SIMO-LacZ (SIMO-Z), and SIMO-GFP transgenic reporters are indicated.
(C) Eye phenotype of the affected individual and his unaffected parents. Slit lamp and close-up photographs of the eyes of subject ID 1230P show complete absence of an iris (arrowheads) and presence of lens cataracts (arrows) in both eyes. Ocular fundus photography and horizontal optical coherence tomography (OCT) in comparison with a normal eye reveal absence of a foveal depression in the subject.
(D) Sequence traces from affected individual and parents show the de novo heterozygous G/T transition in the element. A consensus PAX6 paired domain binding sequence is shown in alignment above the sequence traces.
To investigate whether mutations in individual regulatory elements could be a further cause of aniridia, we screened a selection of eye-related cis-regulatory elements (Table S1) in a panel of 60 subjects without PAX6 exonic mutations or large-scale chromosomal abnormalities, as assessed by exon sequencing, array-CGH (Roche Nimblegen 12X135k whole-genome array), or MLPA testing (SALSA MLPA P219-B2 reagent kit, MRC-Holland). Informed consent was obtained in keeping with the guidelines approved by the UK Multiregional Ethics Committee (reference: 06/MRE00/76) and the study protocol was in keeping with the tenets of the Declaration of Helsinki. We identified one individual with a single de novo nucleotide variant (chr11: 31,685,945G>T [UCSC Genome Browser human reference sequence hg19]) in an ultraconserved sequence, SIMO (uc.325),14 located 150 kb downstream of PAX6 (Figures 1A and 1B, and Figure S1 available online). Individual ID 1230P (normal karyotype: 46,XY) was diagnosed with bilateral iris hypoplasia at the age of two. Aniridia was confirmed at age 17 (Figure 1C) and detailed ophthalmologic evaluation was performed (Supplemental Data). The G/T variant affects a nucleotide residue conserved more than 350 million years across the spectrum of bony vertebrates (Figures 1A and 1D). In silico assessment of the variant base pair and flanking sequences revealed it as a potential binding site for the PAX6 paired domain.15,16 Based on the crystal structure of the human PAX6 paired domain-DNA complex,17 the G to T transversion would be predicted to affect binding by changing a nucleotide that is critical for paired domain linker-DNA contact, suggesting that interference with autoregulation of PAX6 could be the cause of disease in this individual.
To investigate the mechanism that leads from a single base pair substitution to a debilitating disease, we first performed a detailed characterization of the 800 bp SIMO cis-element in mouse and zebrafish reporter transgenics. The experiments were approved by the University of Edinburgh ethical committee (TR-20-10) and performed under UK Home Office license number PPL 60/3785. We observed LacZ reporter activity in early surface ectoderm, lens, diencephalon, and rhombencephalon of transgenic mouse embryos from E9.5 (Figures 2A, 2B, and S2B–S2G) and in the developing retina from E13.5 onward (Figures 2C, S2H, and S2I). Eye expression continues through later development and into adulthood (Figures 2C, S2L, and S2M), where it is seen in the lens epithelium, retina, and iris. To assess functional conservation of the element, we made zebrafish GFP reporter transgenics with the mouse and zebrafish elements. Both elements produced similar patterns, being inactive at 24 hpf, and driving expression in lens, diencephalon, and hindbrain subsequently (Figures 2D–2F and S2N–S2T). To assess whether disruption of the putative PAX6 binding site (BS) would affect enhancer activity, we tested two mutant versions of SIMO in mouse and zebrafish transgenic assays: the exact subject ID 1230P mutation (SIMO-T) and a multinucleotide disruption of the putative PAX6 BS (SIMO(Pax6mut)). Transgenic fish for both mutated versions (Figures 2G–2I) lacked GFP signal in the lens but retained expression in other parts of the embryos. In mouse transgenics, compared with the wild-type element, LacZ expression in both mutant versions was lost in the lens but remained in hindbrain and diencephalon (Figures 2J–2O). Later-stage expression in the retina is also lost in transgenic mouse embryos with the mutant elements (Figure 2L). We conclude that the mutation found in patient ID 1230P is sufficient to abolish activity of the SIMO enhancer in developing ocular tissues.
Figure 2.
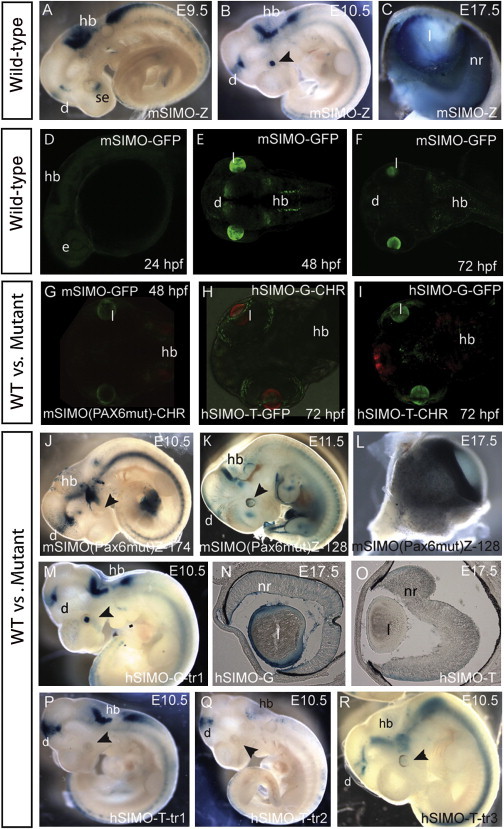
Characterization of SIMO Wild-Type and Mutant Enhancer Activity in Zebrafish and Mouse Reporter Transgenics
Wild-type and mutant versions of the SIMO element were cloned into appropriate transgenic reporter vectors by the method described in Ravi et al.35 The SIMO G>T mutation was PCR amplified from the subject’s DNA, and the artificial multinucleotide PAX6 BS mutation was made with Quickchange site-directed mutagenesis. Primers used are shown in Table S4.
(A and B) Lateral views of transgenic mouse embryos with the wild-type element show expression in surface ectoderm at E9.5 (se) and lens at E10.5 dpc and in hindbrain (hb) and diencephalon (d).
(C) At later developmental stages, X-gal staining is found in lens epithelium and neuroretina.
(D–F) Comparable expression is seen in stable transgenic zebrafish with mouse SIMO element (mSIMO).
(D) At 24 hr postfertilization (hpf), no GFP signal is yet detected.
(E) By 48 hpf, strong enhancer activity is seen in the lens and more variably in diencephalon and hindbrain.
(F) Expression is maintained at 72 hpf.
(G–I) Double fluorescent reporter transgenic zebrafish demonstrate the loss of lens activity for the mutant SIMO elements.
(G) Wild-type mouse SIMO linked to GFP (mSIMO-GFP) in green and the multinucleotide SIMO mutant linked to mCherry (mSIMO(Pax6)-CHR) in red.
(H) Loss of enhancer activity in the eye by the patient mutation is demonstrated in comparative analysis of wild-type human hSIMO-G-GFP versus the mutant hSIMO-T-Cherry.
(I) The same result is seen when the reporters are swapped.
(J–R) Absence of eye expression in SIMO mutant transgenic embryos.
(J and K) Two independent transgenic embryos for the multinucleotide SIMO mutant (mSIMO(Pax6)-Z) show absence of lens expression.
(L) Staining is absent in SIMO mutant eyes at E17.5.
(M) The human SIMO-G wild-type element is expressed in lens, diencephalon, and hindbrain.
(N and O) Sections through E17.5 hSIMO eyes show staining in lens epithelium, neuroretina, and ciliary margin (arrowhead) in hSIMO-G (N) but not in hSIMO-T (O).
(P–R) Three independent transgenic embryos for hSIMO-T at E10.5 demonstrate loss of expression in the lens while di- and rhombencephalon expression remains.
We next investigated whether enhancer activity of SIMO depends on the presence of Pax6. We first performed histone H3 chromatin immunoprecipitations to ascertain the active enhancer status of the endogenous SIMO fragment in chromatin prepared from mouse E14.5 eyes. qPCR revealed strong enrichment for monomethylated H3K4 (ab8895, Abcam) and acetylated H3K27 (ab4729, Abcam) at the SIMO fragment compared to negative (featureless intergenic fragments near mouse Wt1 and Rcn1) control sequences from the locus (Figure 3A). ChIP with Pax6 antibody (DSHB) revealed clear enrichment at the SIMO element and the positive control fragment (Pax6 P1 promoter) and a lack of enrichment at the negative controls (Figure 3B). A DNA affinity-capture assay via biotinylated double-stranded oligonucleotides as probe also efficiently pulled down Pax6 protein from a nuclear extract of E14.5 eyes when the wild-type SIMO-G PAX6 binding sequence was used, but the efficiency of the Pax6 pulldown was strongly reduced with the SIMO-T mutant version of the probe (Figure 3C). This was confirmed by an EMSA competition experiment where the G>T change strongly reduced the ability to compete for binding of purified PAX6 paired domain (Figure S3). These results show that SIMO is an active enhancer in E14.5 eyes and that Pax6 efficiently binds to a wild-type SIMO-G but not a mutant SIMO-T fragment, albeit with lower affinity than the optimal PAX6 consensus site.15
Figure 3.
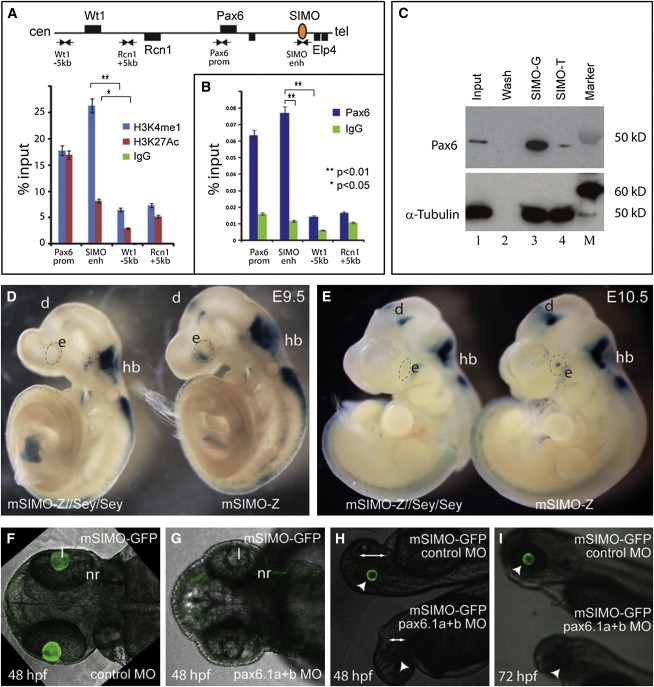
Pax6 Binds to the SIMO Enhancer and Is Essential for Its Function
(A) Chromatin immunoprecipitation on chromatin, prepared from approximately 200 dissected mouse E14.5 embryonic eyes as described,36 shows that the SIMO element is enriched for enhancer-associated histone modifications H3K4me1 (ab8895, Abcam) and H3K27Ac (ab4729, Abcam; Table S3). The Pax6 P1 promoter is also positive for these marks, whereas featureless control fragments located near the adjacent Wt1 and Rcn1 genes show minimal enrichment.
(B) ChIP with Pax6 antibody (DSHB) shows clear enrichment at the SIMO enhancer and Pax6 P1 promoter but not at the control regions. qPCR primer sets used are shown in Table S4.
Relative enrichments are shown as mean percentage of input ± SEM (standard error of the mean); ∗p < 0.05, ∗∗p < 0.01 (Student’s t test).
(C) Double-stranded DNA affinity capture assay from nuclear extract prepared from approximately 100 E14.5 mouse embryonic eyes via NE-PER Nuclear and Cytoplasmic Extraction Reagents (Thermo Scientific, catalog number 78833), performed as described in Jeong et al.28 Pax6 is efficiently pulled down with a probe representing the wild-type SIMO-G sequence (Table S4). Binding to the mutant SIMO-T fragment is greatly reduced. Immunoblot via a mix of monoclonal anti-Pax6 antibodies (Table S3). Lane 1: precleared supernatant (Input); lane 2: bead wash fraction after extract binding; lane 3: final bead fraction via the wild-type SIMO-G probe; lane 4: final bead fraction for the mutant SIMO-T probe; lane M: marker.
(D and E) Absence of functional Pax6 protein in vivo disrupts SIMO enhancer activity in the surface ectoderm and primordial lens of homozygous smalleye (Sey) embryos at E9.5 (D) and E10.5 (E).
(F, H, and I [bottom embryo]) Injection of morpholinos (MO; Gene Tools) against zebrafish pax6.1a (5′-AGTTCCAACAGCCTTTGTATCCTCG-3′) and pax6.1b (5′-GCCTGAGCCCTTCCGAGCAAATCAG-3′) in stable SIMO-GFP transgenic fish embryos results in loss of reporter expression specifically in the lens, with variable phenotypic deformity of the embryos, notably a reduced eye size (arrows).
(G, H, and I [top embryo]) Injection with the Gene Tools standard negative control morpholino: 5′-CCTCTTACCTCAGTTACAATTTATA-3′ has no effect. White arrowhead indicates the eye.
Abbreviations are as follows: d, diencephalon; e, eye; hb, hindbrain; l, lens; nr, neuroretina.
To determine how absence of Pax6 would affect functional activity of the SIMO enhancer in vivo, we crossed a SIMO-LacZ reporter allele onto the mouse mutant smalleye (Pax6 null).18 smalleye homozygous embryos carrying the SIMO-LacZ transgene showed a distinct lack of X-gal staining in the surface ectoderm and lens primordium at E9.5 (Figure 3D) and E10.5 (Figure 3E), in comparison with wild-type littermates, indicating that Pax6 is required for activation of the SIMO element. SIMO-driven reporter expression in the lens was also abolished in a zebrafish mSIMO-GFP transgenic line upon coinjection of morpholinos against zebrafish pax6.1a and pax6.1b. Reporter activity was unaffected in fish injected with a control morpholino (Figures 3F–3I). Immunoblotting of morpholino-injected embryos confirmed effective depletion of pax6 (Figure S3).
Having demonstrated the dependence of SIMO enhancer activity on Pax6 and its disruption by the subject’s mutation, we wished to investigate whether SIMO-driven enhancer activity is essential for the expression of PAX6. In contrast to aniridia cases where chromosomal abnormalities had removed large collections of cis-regulatory elements from the locus,4–6,13 no other small deletions or regulatory mutations had been identified. This raised the question of whether disruption of a single cis-element, in the presence of multiple enhancers with overlapping tissue specificity, could be sufficient to cause a disease phenotype. To determine the consequence of SIMO element loss from its wider genomic context, we employed a BAC transgenic approach. Because the size of the known human PAX6 regulatory domain exceeds normal BAC content, we used a BAC containing the more compact X. tropicalis pax6 locus including all currently known lens and retinal enhancers and inserted a GFP reporter into the pax6 ATG start site by recombineering.19,20 Tol2 sites flanking the insert were inserted into the BAC vector to allow Tol2-mediated transgenesis.21 The resulting wild-type reporter BAC (BAC-XtPax6-GFP-Wt) was further modified by deletion of the SIMO element (BAC-XtPax6-GFP-ΔSIMO) (Figures 4A and S4). Zebrafish transgenic lines were established for both BAC versions. The wild-type reporter BAC recapitulates the combined pax6 expression patterns of the partially overlapping expression domains of the two zebrafish pax6.1 genes (Figures 4B–4F).22,23 The same pattern is seen in transgenic fish for the SIMO-deleted BAC at 24 hpf (Figures 4G and 4H). However, by 48 hpf, GFP expression is no longer detected in the lens and remains conspicuously absent subsequently (Figures 4I–4K). Thus, deletion of the SIMO element from the wider pax6 locus results in complete abolition of Pax6-GFP expression in the lens after 24 hpf, despite the continued presence of other defined lens enhancers on the BAC, demonstrating that activity of the remote SIMO enhancer element is critically required for continued tissue-specific expression from the Pax6 promoters. Requirement of PAX6 itself for the functional activity of the element suggested that the SIMO enhancer is not essential in the initial activating phase of PAX6 gene expression but has a role in maintenance of its transcription through a positive-feedback loop (Figure 4L). The finding that deletion of SIMO from the BAC transgene has no apparent effect on reporter expression in early embryos, but causes complete loss of lens expression at subsequent stages, supports this model. The model also highlights the notion that equivalent effects on PAX6 lens expression can be produced by deletion of a remote regulatory element, absence of a crucial trans-acting factor, or the mutation of a single nucleotide in its binding site (Figure 4L).
Figure 4.
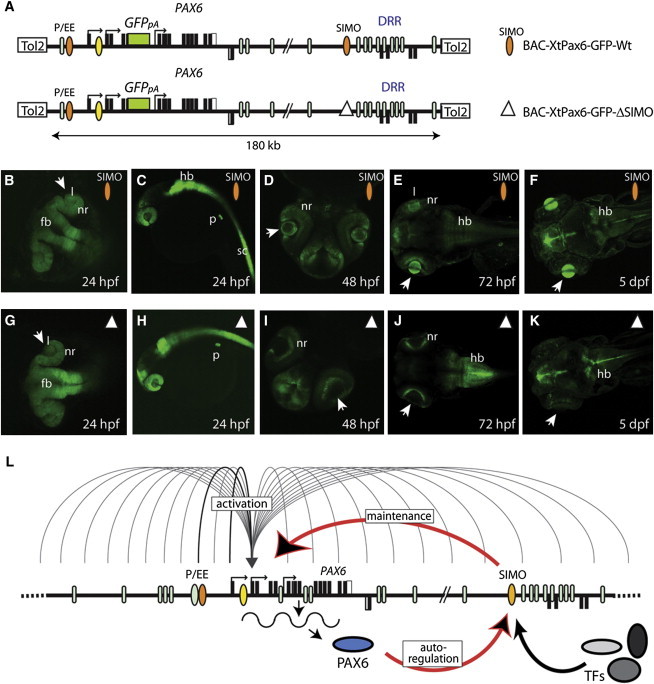
The SIMO Enhancer Is Essential for Expression of Pax6 in Its Wider Genomic Context
(A) Schematic representation of the modified BAC constructs containing the X. tropicalis Pax6 genomic locus, differing by the presence (orange ellipse) or absence (open triangle) of the SIMO element. Modified BACs are based on X. tropicalis BAC CH216-109E08 (obtained from CHORI BACPAC), transformed into the recombineering permissive bacterial strain SW102. A gfp3 reporter cassette was inserted into the pax6 exon 4 translational start site by GalK positive/negative selection.19 Tol2 recombinase arms were then placed in the BAC vector by the iTol2-ampicillin selection cassette.37 A 1.1 kb fragment covering the genomic coordinates from scaffold_399:567,254–568,353 (JGI 4.1/xenTro2), containing the X. tropicalis conserved SIMO sequence, was removed from the SIMO deleted BAC via rpsL/kanamycin selection recombineering.38 Primer sets used are shown in Table S4. The multitude of cis-elements present in the Pax6 locus is represented by blue ellipses, including the lens-specific ectodermal enhancer10 (EE, orange).
(B–K) Temporal series of stable transgenic zebrafish carrying wild-type (B–F) or SIMO-deleted (G–K) reporter BAC transgenes. GFP signal recapitulates the combined pax6 expression pattern, including forebrain (fb), hindbrain (hb), spinal cord (sc), neuroretina (nr), lens (l), and pancreas (p). Wild-type and SIMO-deleted reporter patterns are identical at 24 hpf and lens expression is seen in both (B, C, G, H). By 48 hpf, GFP signal is absent from the lenses of SIMO-deleted BAC transgenic embryos, although being maintained in wild-type BAC transgenics (D, I). GFP patterns remain similar in wild-type and SIMO-deleted BAC transgenic fish at 72 hpf and 5 days with the exception of lens expression (E, F, J, K).
(L) Model for the role of the autoregulatory PAX6 binding site in the SIMO enhancer, maintaining a positive feedback loop required for PAX6 expression during eye development. Undefined enhancer(s) in the locus, possibly including EE, initiate PAX6 transcription in early lens/surface ectoderm. Expression of PAX6 allows activation of the SIMO enhancer which, alone or in combination, maintains expression of PAX6 at subsequent stages of eye development.
We show that the SIMO enhancer is crucial for expression of PAX6 in developing ocular tissues. Disruption of the PAX6 binding site within SIMO abolishes both lens and late retinal expression in mouse transgenics, showing that PAX6 binding is a critical event for both these functions of the element. As a consequence it is presently unclear whether the developmental defects leading to the subject’s aniridia phenotype are due to misregulation of PAX6 in the lens, retina, or both. Based on observations from morula aggregation-derived Pax6+/+<>Pax6+/− chimeric eyes in which the iris was not affected, even in chimeras with a very high percentage of Pax6+/− cells in the iris, it was surmised that iris hypoplasia in aniridia patients is a secondary effect of defective lens development.24 However, heterozygous conditional inactivation of Pax6 in either lens or retina separately showed no obvious iris abnormality upon lens-specific inactivation of one Pax6 allele, but iris hypoplasia was seen after heterozygous Pax6 deletion in the retina.25 The latter observations suggest disruption of the retinal part of the SIMO expression as the more likely causative event, but considering the intertwining of lens and retinal development, further dissection of the SIMO element is needed to separate the subdomains of SIMO-driven expression. It is likely that activation of the SIMO enhancer in lens, retina, diencephalon, and hindbrain depends on the presence (and levels) of different combinations of common and unique TFs in those tissues, binding to the appropriate subsets of recognition sites within SIMO. Indeed, although the Pax6 BS mutation affects lens and retinal expression, reporter activity in hindbrain and diencephalon appears largely unaffected. Similarly, mutation of binding sites for other TFs may selectively affect SIMO activity in a different subset of tissues.
Genetic disease can be caused by mutation or deletion of the transcribed part of a gene or by the disruption of its appropriately regulated gene expression. A variety of mechanisms interfering with proper expression control and leading to congenital malformations have been observed.5 In the large majority of such cases, a substantial insult to the integrity of the cis-regulatory landscape surrounding the disease-associated gene has occurred. At present only a handful of cases involving single-nucleotide changes in a remote regulatory element have been described, representing a variety of subtly differing mechanisms:26–30 creation of a cryptic promoter diverting normal regulatory interaction in the α-globin locus in thalassemia (MIM 604131);26 multiple independent mutations in the ZRS, a long-range enhancer for SHH, create an ectopic gain-of-function effect in the limb bud causing limb deformities;27 mutation of a BS for the transcriptional activator SIX3 (MIM 603714) in the SHH SBE2 forebrain enhancer leads to holoprosencephaly (MIM 142945),28 whereas a mutation in the BS for the YY1 repressor (MIM 600013) leads to overexpression of HCFC1 (MIM 300019) in intellectual disability (MIM 309541);29 and a mutation improving the binding affinity of a MSX1 (MIM 142983) BS in the remote HCNE-F2 enhancer located 1.3 Mb upstream of SOX9 is implicated in Pierre Robin Sequence (MIM 261800).30 The mutation in the aniridia case described here adds a further mechanism to this list: disruption of a positive feedback loop by the mutation of an autoregulatory binding site. It further demonstrates that even a single point mutation in a remote enhancer located 150 kb from its target gene can have severe consequences, highlighting the sensitivity of embryonic development to disturbances at specific positions in our genome. This stands in stark contrast to reports that one can delete megabase segments of the genome31 or remove complete ultraconserved enhancers32 without any apparent effect, which had suggested widespread redundancy among enhancers for critical developmental regulatory genes.33,34 These contrasting observations underline the fact that our understanding of the cis-regulatory mechanisms governing the expression of our genome is far from complete. Our work highlights the fact that, despite the current bias toward mutation discovery in exonic regions of the genome, detailed study of the role of noncoding parts of the genome can make a valuable contribution to our understanding of the mechanistic variety of the causes of genetic disorders.
Acknowledgments
We are grateful to the affected individual and parents for their cooperation with the study. We thank Nick Hastie, David Fitzpatrick, Bob Hill, Jacky Guy, and Nick Gilbert for comments on the manuscript; Kathy Williamson for the original aniridia mutation screening of the PAX6 exonic regions; Philippe Gautier and Hajime Ogino for bio-informatics support; and Emma Murdoch, Jacek Mendrychowski, and Harris Morrison for technical assistance. The iTol2AMP plasmid used in BAC recombineering was kindly provided by K. Kawakami. S.B. was supported by National Institutes of Health grant 1RO1 EY01 8000-02. H.B. was supported by EU collaborative project NeuroXsys Health-F4-2009-223262. Work at the IGMM Human Genetics Unit was supported by the Medical Research Council UK.
Contributor Information
Veronica van Heyningen, Email: veronica.vanheyningen@igmm.ed.ac.uk.
Dirk A. Kleinjan, Email: dirk-jan.kleinjan@igmm.ed.ac.uk.
Supplemental Data
Web Resources
The URLs for data presented herein are as follows:
Clustal Omega, http://www.ebi.ac.uk/Tools/msa/clustalo/
Online Mendelian Inheritance in Man (OMIM), http://www.omim.org/
PAX6 mutation database, http://lsdb.hgu.mrc.ac.uk/home.php?select_db=PAX6
Pipmaker, http://pipmaker.bx.psu.edu/pipmaker/
UCSC Genome Browser, http://genome.ucsc.edu
References
- 1.Woolfe A., Goodson M., Goode D.K., Snell P., McEwen G.K., Vavouri T., Smith S.F., North P., Callaway H., Kelly K. Highly conserved non-coding sequences are associated with vertebrate development. PLoS Biol. 2005;3:e7. doi: 10.1371/journal.pbio.0030007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Kleinjan D.J., Coutinho P. Cis-ruption mechanisms: disruption of cis-regulatory control as a cause of human genetic disease. Brief. Funct. Genomics Proteomics. 2009;8:317–332. doi: 10.1093/bfgp/elp022. [DOI] [PubMed] [Google Scholar]
- 3.Hingorani M., Hanson I., van Heyningen V. Aniridia. Eur. J. Hum. Genet. 2012;20:1011–1017. doi: 10.1038/ejhg.2012.100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Fantes J., Redeker B., Breen M., Boyle S., Brown J., Fletcher J., Jones S., Bickmore W., Fukushima Y., Mannens M. Aniridia-associated cytogenetic rearrangements suggest that a position effect may cause the mutant phenotype. Hum. Mol. Genet. 1995;4:415–422. doi: 10.1093/hmg/4.3.415. [DOI] [PubMed] [Google Scholar]
- 5.Lauderdale J.D., Wilensky J.S., Oliver E.R., Walton D.S., Glaser T. 3′ deletions cause aniridia by preventing PAX6 gene expression. Proc. Natl. Acad. Sci. USA. 2000;97:13755–13759. doi: 10.1073/pnas.240398797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Robinson D.O., Howarth R.J., Williamson K.A., van Heyningen V., Beal S.J., Crolla J.A. Genetic analysis of chromosome 11p13 and the PAX6 gene in a series of 125 cases referred with aniridia. Am. J. Med. Genet. A. 2008;146A:558–569. doi: 10.1002/ajmg.a.32209. [DOI] [PubMed] [Google Scholar]
- 7.Plaza S., Dozier C., Langlois M.C., Saule S. Identification and characterization of a neuroretina-specific enhancer element in the quail Pax-6 (Pax-QNR) gene. Mol. Cell. Biol. 1995;15:892–903. doi: 10.1128/mcb.15.2.892. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Kammandel B., Chowdhury K., Stoykova A., Aparicio S., Brenner S., Gruss P. Distinct cis-essential modules direct the time-space pattern of the Pax6 gene activity. Dev. Biol. 1999;205:79–97. doi: 10.1006/dbio.1998.9128. [DOI] [PubMed] [Google Scholar]
- 9.Griffin C., Kleinjan D.A., Doe B., van Heyningen V. New 3′ elements control Pax6 expression in the developing pretectum, neural retina and olfactory region. Mech. Dev. 2002;112:89–100. doi: 10.1016/s0925-4773(01)00646-3. [DOI] [PubMed] [Google Scholar]
- 10.Williams S.C., Altmann C.R., Chow R.L., Hemmati-Brivanlou A., Lang R.A. A highly conserved lens transcriptional control element from the Pax-6 gene. Mech. Dev. 1998;73:225–229. doi: 10.1016/s0925-4773(98)00057-4. [DOI] [PubMed] [Google Scholar]
- 11.Xu P.X., Zhang X., Heaney S., Yoon A., Michelson A.M., Maas R.L. Regulation of Pax6 expression is conserved between mice and flies. Development. 1999;126:383–395. doi: 10.1242/dev.126.2.383. [DOI] [PubMed] [Google Scholar]
- 12.Kleinjan D.A., Seawright A., Mella S., Carr C.B., Tyas D.A., Simpson T.I., Mason J.O., Price D.J., van Heyningen V. Long-range downstream enhancers are essential for Pax6 expression. Dev. Biol. 2006;299:563–581. doi: 10.1016/j.ydbio.2006.08.060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Kleinjan D.A., Seawright A., Schedl A., Quinlan R.A., Danes S., van Heyningen V. Aniridia-associated translocations, DNase hypersensitivity, sequence comparison and transgenic analysis redefine the functional domain of PAX6. Hum. Mol. Genet. 2001;10:2049–2059. doi: 10.1093/hmg/10.19.2049. [DOI] [PubMed] [Google Scholar]
- 14.Bejerano G., Pheasant M., Makunin I., Stephen S., Kent W.J., Mattick J.S., Haussler D. Ultraconserved elements in the human genome. Science. 2004;304:1321–1325. doi: 10.1126/science.1098119. [DOI] [PubMed] [Google Scholar]
- 15.Epstein J., Cai J., Glaser T., Jepeal L., Maas R. Identification of a Pax paired domain recognition sequence and evidence for DNA-dependent conformational changes. J. Biol. Chem. 1994;269:8355–8361. [PubMed] [Google Scholar]
- 16.Coutinho P., Pavlou S., Bhatia S., Chalmers K.J., Kleinjan D.A., van Heyningen V. Discovery and assessment of conserved Pax6 target genes and enhancers. Genome Res. 2011;21:1349–1359. doi: 10.1101/gr.124115.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Xu H.E., Rould M.A., Xu W., Epstein J.A., Maas R.L., Pabo C.O. Crystal structure of the human Pax6 paired domain-DNA complex reveals specific roles for the linker region and carboxy-terminal subdomain in DNA binding. Genes Dev. 1999;13:1263–1275. doi: 10.1101/gad.13.10.1263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Hill R.E., Favor J., Hogan B.L., Ton C.C., Saunders G.F., Hanson I.M., Prosser J., Jordan T., Hastie N.D., van Heyningen V. Mouse small eye results from mutations in a paired-like homeobox-containing gene. Nature. 1991;354:522–525. doi: 10.1038/354522a0. [DOI] [PubMed] [Google Scholar]
- 19.Warming S., Costantino N., Court D.L., Jenkins N.A., Copeland N.G. Simple and highly efficient BAC recombineering using galK selection. Nucleic Acids Res. 2005;33:e36. doi: 10.1093/nar/gni035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Fish M.B., Nakayama T., Grainger R.M. Simple, fast, tissue-specific bacterial artificial chromosome transgenesis in Xenopus. Genesis. 2012;50:307–315. doi: 10.1002/dvg.20819. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Suster M.L., Abe G., Schouw A., Kawakami K. Transposon-mediated BAC transgenesis in zebrafish. Nat. Protoc. 2011;6:1998–2021. doi: 10.1038/nprot.2011.416. [DOI] [PubMed] [Google Scholar]
- 22.Nornes S., Clarkson M., Mikkola I., Pedersen M., Bardsley A., Martinez J.P., Krauss S., Johansen T. Zebrafish contains two pax6 genes involved in eye development. Mech. Dev. 1998;77:185–196. doi: 10.1016/s0925-4773(98)00156-7. [DOI] [PubMed] [Google Scholar]
- 23.Kleinjan D.A., Bancewicz R.M., Gautier P., Dahm R., Schonthaler H.B., Damante G., Seawright A., Hever A.M., Yeyati P.L., van Heyningen V., Coutinho P. Subfunctionalization of duplicated zebrafish pax6 genes by cis-regulatory divergence. PLoS Genet. 2008;4:e29. doi: 10.1371/journal.pgen.0040029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Collinson J.M., Quinn J.C., Buchanan M.A., Kaufman M.H., Wedden S.E., West J.D., Hill R.E. Primary defects in the lens underlie complex anterior segment abnormalities of the Pax6 heterozygous eye. Proc. Natl. Acad. Sci. USA. 2001;98:9688–9693. doi: 10.1073/pnas.161144098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Davis-Silberman N., Kalich T., Oron-Karni V., Marquardt T., Kroeber M., Tamm E.R., Ashery-Padan R. Genetic dissection of Pax6 dosage requirements in the developing mouse eye. Hum. Mol. Genet. 2005;14:2265–2276. doi: 10.1093/hmg/ddi231. [DOI] [PubMed] [Google Scholar]
- 26.De Gobbi M., Viprakasit V., Hughes J.R., Fisher C., Buckle V.J., Ayyub H., Gibbons R.J., Vernimmen D., Yoshinaga Y., de Jong P. A regulatory SNP causes a human genetic disease by creating a new transcriptional promoter. Science. 2006;312:1215–1217. doi: 10.1126/science.1126431. [DOI] [PubMed] [Google Scholar]
- 27.Lettice L.A., Hill A.E., Devenney P.S., Hill R.E. Point mutations in a distant sonic hedgehog cis-regulator generate a variable regulatory output responsible for preaxial polydactyly. Hum. Mol. Genet. 2008;17:978–985. doi: 10.1093/hmg/ddm370. [DOI] [PubMed] [Google Scholar]
- 28.Jeong Y., Leskow F.C., El-Jaick K., Roessler E., Muenke M., Yocum A., Dubourg C., Li X., Geng X., Oliver G., Epstein D.J. Regulation of a remote Shh forebrain enhancer by the Six3 homeoprotein. Nat. Genet. 2008;40:1348–1353. doi: 10.1038/ng.230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Huang L., Jolly L.A., Willis-Owen S., Gardner A., Kumar R., Douglas E., Shoubridge C., Wieczorek D., Tzschach A., Cohen M. A noncoding, regulatory mutation implicates HCFC1 in nonsyndromic intellectual disability. Am. J. Hum. Genet. 2012;91:694–702. doi: 10.1016/j.ajhg.2012.08.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Benko S., Fantes J.A., Amiel J., Kleinjan D.J., Thomas S., Ramsay J., Jamshidi N., Essafi A., Heaney S., Gordon C.T. Highly conserved non-coding elements on either side of SOX9 associated with Pierre Robin sequence. Nat. Genet. 2009;41:359–364. doi: 10.1038/ng.329. [DOI] [PubMed] [Google Scholar]
- 31.Nóbrega M.A., Zhu Y., Plajzer-Frick I., Afzal V., Rubin E.M. Megabase deletions of gene deserts result in viable mice. Nature. 2004;431:988–993. doi: 10.1038/nature03022. [DOI] [PubMed] [Google Scholar]
- 32.Ahituv N., Zhu Y., Visel A., Holt A., Afzal V., Pennacchio L.A., Rubin E.M. Deletion of ultraconserved elements yields viable mice. PLoS Biol. 2007;5:e234. doi: 10.1371/journal.pbio.0050234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Perry M.W., Boettiger A.N., Bothma J.P., Levine M. Shadow enhancers foster robustness of Drosophila gastrulation. Curr. Biol. 2010;20:1562–1567. doi: 10.1016/j.cub.2010.07.043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Frankel N., Davis G.K., Vargas D., Wang S., Payre F., Stern D.L. Phenotypic robustness conferred by apparently redundant transcriptional enhancers. Nature. 2010;466:490–493. doi: 10.1038/nature09158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Ravi V., Bhatia S., Gautier P., Loosli F., Tay B.H., Tay A., Murdoch E., Coutinho P., van Heyningen V., Brenner S. Sequencing of Pax6 loci from the elephant shark reveals a family of Pax6 genes in vertebrate genomes, forged by ancient duplications and divergences. PLoS Genet. 2013;9:e1003177. doi: 10.1371/journal.pgen.1003177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Wolf L.V., Yang Y., Wang J., Xie Q., Braunger B., Tamm E.R., Zavadil J., Cvekl A. Identification of pax6-dependent gene regulatory networks in the mouse lens. PLoS ONE. 2009;4:e4159. doi: 10.1371/journal.pone.0004159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Suster M.L., Kikuta H., Urasaki A., Asakawa K., Kawakami K. Transgenesis in zebrafish with the tol2 transposon system. Methods Mol. Biol. 2009;561:41–63. doi: 10.1007/978-1-60327-019-9_3. [DOI] [PubMed] [Google Scholar]
- 38.Wang S., Zhao Y., Leiby M., Zhu J. A new positive/negative selection scheme for precise BAC recombineering. Mol. Biotechnol. 2009;42:110–116. doi: 10.1007/s12033-009-9142-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


