Abstract
PPR proteins form a huge family in flowering plants and are involved in RNA maturation in plastids and mitochondria. These proteins are sequence-specific RNA-binding proteins that recruit the machinery of RNA processing. We summarize progress in the research on the functional mechanisms of divergent RNA maturation and on the mechanism by which RNA sequences are recognized. We further focus on two topics. RNA editing is an enigmatic process of RNA maturation in organelles, in which members of the PLS subfamily contribute to target site recognition. As the first topic, we speculate on why the PLS subfamily was selected by the RNA editing machinery. Second, we discuss how the regulation of plastid gene expression contributes to efficient photosynthesis. Although the molecular functions of PPR proteins have been studied extensively, information on the physiological significance of regulation by these proteins remains very limited.
Keywords: chloroplast, photosynthesis, PPR, RNA, RNA editing
Introduction
Plastids and mitochondria originated from symbiosis and, consequently, have their own genomes. During the evolution of eukaryotic phototrophs, the majority of genes of prokaryotic origin were transferred from the organelle genomes to the host genome; in flowering plants, approximately 120 genes remain in plastids and 60 genes remain in mitochondria. Because gene expression systems also have a bacterial origin, they retain prokaryotic features, especially in plastids. For example, one of two transcription systems in plastids depends on multi-subunit-type RNA polymerase, similar to the bacterial enzyme. Whereas core subunits of RNA polymerase are encoded by the plastid genome (therefore called plastid-encoded plastid RNA polymerase or PEP), genes encoding sigma factors have been transferred to the host genome and are used by the host nuclear genome to regulate gene expression in plastids.1 Another prokaryotic feature of the plastid gene expression system is that multiple genes are transcribed from a single promoter as polycistronic RNA, which is the same process as that in bacterial cells. Plastid gene expression is regulated mainly in the processes of the maturation of precursor RNAs rather than in transcription.2
Pentatricopeptide repeat (PPR) proteins play a central role in the post-transcriptional regulation of gene expression in plastids and mitochondria. In contrast to sigma factors, a member of the PPR family is part of the eukaryotic machinery for the regulation of gene expression. Some members of the PPR family were independently characterized in pioneering work as sequence-specific RNA binding proteins in Saccharomyces cerevisiae,3 Neurospora crassa4 and maize.5 However, the whole picture of the family was first recognized in a bioinformatics study on the basis of the huge size of the family in the genome of Arabidopsis thaliana.6 The Arabidopsis genome encodes approximately 450 members of the PPR family; in contrast, Chlamydomonas reinhardtii and Homo sapiens encode only 12 and six members, respectively.7
Due to the huge size of this family and the involvement of its members in fundamental cellular processes, PPR proteins have been identified in genetics studies that focused on divergent cellular functions, including photosynthesis,8,9 respiration (slow growth),10 chloroplast development,5,11 embryogenesis,12 cytoplasmic male sterility (CMS),13,14 isoprenoid biosynthesis15 and retrograde signaling.16 Some excellent reviews have already covered the molecular function of this family.6,7,17,18 To avoid overlapping with these reviews, we will briefly summarize the basic knowledge of the family and will then focus on functional diversity among family members and on the physiological significance of regulation in plastid gene expression.
Structure of PPR proteins
Members of the PPR family are characterized by tandem arrays (two to 30 repeats) of the degenerate 35-amino-acid motif (PPR motif). The PPR family is divided into P and PLS subfamilies according to the nature of the PPR motifs.7 Each subfamily accounts for approximately half of the PPR family in flowering plants.19 Typical members of the P subfamily consist simply of tandem arrays of PPR motifs, whereas those of the PLS subfamily contain long (L) and short (S) PPR-like motifs as well as canonical PPR motifs (P) in a pattern of P-L-S.19,20 Members of the PLS subfamily are further divided into PLS, E/E+ and DYW classes based on their C-terminal domains (Fig. 1).19 Human mitochondrial RNA polymerase contains two PPR motifs, and its structure has been resolved.21 In this structure, a PPR motif forms a pair of anti-parallel α helices, as predicted from its similarity with the structure of tetratricopeptide repeat (TPR) proteins.22 On the basis of the in vitro RNA-binding assay,11,23-29 the RIP-Chip assay25,27,30-32 and the RNA-bound structure predicted from evolutionary constraints,33 most PPR proteins are believed to be sequence-specific RNA-binding proteins. In silico analyses predicted that 64% and 16% of PPR proteins target mitochondria and plastids, respectively, in Arabidopsis;19 this result most likely underestimates the localization of PPR proteins in the two organelles in flowering plants. The predicted localization is consistent with the mutant phenotypes reported so far,7 indicating that PPR proteins are involved in various RNA maturation processes in mitochondria and in plastids.
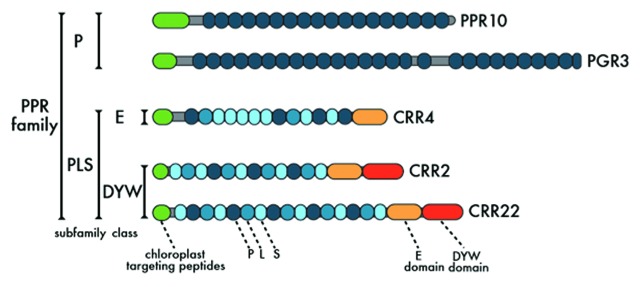
Figure 1. Schematics of representative PPR protein structures.
Members of the P subfamily are involved in divergent processes of RNA maturation in organelles, including RNA stabilization, splicing, intergenic RNA cleavage and translation.7 On the other hand, PLS subfamily members are closely associated with RNA editing.34 In plastids, gene expression is regulated mainly by post-transcriptional steps, in which PPR proteins play a central role. However, as discussed below, it is unlikely that RNA editing plays a role in the regulation of gene expression in plant organelles. Despite their conservation, in terms of structure and primary function as sequence-specific RNA-binding proteins, the P and PLS subfamilies may have distinct physiological functions.
General function of the P subfamily
The majority of P-subfamily members consist of tandem arrays of PPR motifs and do not contain any other motifs. Exceptions are observed in some PPR proteins, including GENOME UNCOUPLED 1 (GUN1),16 ATP431 and SUPPRESSOR OF VARIAGATION7 (SVR7),35 which contain a small mutS-related (SMR) domain. In our study, we focus on typical members of the P subfamily. The first question is how the simple arrays of PPR motifs help in diverse RNA maturation processes, including translation. Genetic analyses have clarified the function of several members of the P subfamily in RNA stabilization. Maize ppr10 mutants fail to accumulate transcripts that have 5′ or 3′ termini in the atpI-atpH intergenic or psaJ-rpl33 intergenic regions.25 The 3′ UTR of atpI mRNA shares 22–26 nucleotides with the 5′ UTR of atpH mRNA. The minimum 17 nucleotides needed for high-affinity binding by PPR10 is included in this region.29 A similar case is observed in the intergenic region between psaJ and rpl33.29 There is one simple way to explain this partial overlap of RNA termini between adjacent mature transcripts: PPR10 is a cap protein that protects target RNA from 5′→3′ and 3′→5′ exonucleases.25 This case is most likely not specific to PPR10 because RNA deep-sequencing has identified short RNAs (16–28 nucleotides) that are mapped to the plastid genome.36,37 Many of these RNAs correspond to the 5′ or 3′ ends of plastid transcripts, suggesting that these are the footprints of PPR proteins, as observed in the PPR10-binding regions.
The ppr10 mutants accumulate precursor RNAs that, unlike in the wild-type, are not processed in the atpI-atpH intergenic region; the levels of ATPase subunits are very low, indicating that PPR10 is required not only for RNA stabilization but also for translation.29 The PPR10 binding site and the translational initiation site of atpH are separated by 26 nucleotides, which include the putative Shine-Dalgarno (SD) sequence. The binding of PPR10 to the site may release the SD element from the predicted secondary structure, which would recruit ribosomes.29
A similar mode of function has been suggested for another P-subfamily member, PROTON GRADIENT REGULATION 3 (PGR3) (Fig. 2).9 PGR3 is required for the stabilization of petL operon RNA and the translation of petL and ndhA.28 Direct evidence of a translational defect in ndhA is still lacking, but recombinant PGR3 protein binds the 5′UTR of ndhA; this idea is consistent with the phenotype in the accumulation pattern of the chloroplast NADH dehydrogenase-like (NDH) complex in the mutants.28 Putative PGR3 footprints have been detected in the regions recognized by PGR3 in vitro.28,36 Whereas PPR10 binds the intergenic region in precursor RNAs, the PGR3 binding site localizes to the 5′UTR of petL, which is the most upstream gene of the operon. Consequently, PGR3 is required for the stabilization of the entire petL operon RNA but is not involved in the stabilization of the upstream region of RNA from 3′→5′ exonuclease.

Figure 2. Functions mediated by PGR3. The model is based on that of Cai et al.28 PGR3 binds the 5′UTR of petL operon RNA and protects RNA from 5′ → 3′ exonuclease (function 1). Simultaneously, PGR3 binding activates translation of petL, most likely modifying the secondary RNA structure of the region, including the SD element (function 2). PGR3 also activates translation of ndhA (function 3). All of the function of the pgr3-1 allele is affected. pgr3-2 is defective in functions 1 and 2, and pgr3-3 cannot translate petL and ndhA (functions 2 and 3).
The identification of three mutant alleles of pgr3 has helped us to clarify three distinct functions of PGR3 (Fig. 2).9 In the pgr3-1 allele, which has the strongest phenotype, almost all of the PGR3 functions are disturbed. The pgr3-2 allele does not accumulate petL operon RNA, but the accumulation of chloroplast NDH is unaffected. The third allele, pgr3-3, was identified in a different mutant screening that focused on chlororespiratory reduction (crr) mutants defective specifically in NDH activity.8 Despite the typical crr phenotype of pgr3-3 in photosynthetic electron transport, the mutant also does not accumulate PetL protein.9 This phenotype was unexpected because the petL operon RNA is stabilized and because the linear electron transport that reflects the level of the cytochrome (Cyt) b6f complex is unaffected in pgr3-3.9 This puzzling phenotype is explained by the concept that PetL, which is not translated in pgr3-3, is not required for the stability or activity of the Cyt b6f complex, whereas PetG is essential for both. The petG gene is localized downstream of petL and is transcribed from the same promoter with petL (Fig. 2). Because petL operon RNA is stable in pgr3-3, petG is likely translated. It is also likely that PGR3 is required for petL translation.
PGR3 consists of 27 PPR motifs, and the pgr3-1 and pgr3-2 alleles contain amino acid alterations in the 15th and 12th motifs, respectively (Fig. 1).9 The final PPR motif is incomplete and includes an amino acid alteration in pgr3-3. An in vitro RNA-binding assay that used mutant versions of PGR3 demonstrated that the 15th PPR motif is required for the binding of both target RNAs, whereas the 12th PPR motif is required only for petL 5′UTR.28 The recombinant PGR3 protein with the pgr3-3 mutation can bind both targets. Based on the specific phenotype of pgr3-3, we can separate the function of PGR3 in translation from that in RNA stabilization. The final incomplete PPR motif is not necessary for binding the target RNAs, but it is essential for translation; this motif most likely modifies the secondary structure of ribosome-binding sites in target RNAs in a manner similar to that of PPR10.29
How do simple arrays of PPR motifs help the divergent RNA processing steps in PPR10 or PGR3? The Alice Barkan group has proposed a simple passive model in which PPR proteins bind target RNA in a sequence-specific manner, thus imparting site-specific barrier and RNA remodeling effects.37 This scenario explains the function of PPR proteins in RNA stabilization, protein translation and even RNA splicing. The following question remains: to what extent is this function conserved among PPR family members?
Function of the PLS subfamily in RNA editing
In land plants, RNA editing converts specific C residues to U residues in RNA molecules; this editing was first discovered in mitochondria38-40 and then in plastids.41 In Arabidopsis, approximately 450 sites in mitochondrial RNA42 and 34 sites in plastid RNA are edited.43 This high frequency of RNA editing in plant organelles is believed to be closely related to the huge size of the PPR family.18 To determine the specific C site to be edited, a short sequence surrounding the target C is recognized by the RNA editing machinery; this sequence is called the cis-element.44 The use of both a technique for plastid transformation in tobacco45 and of an in vitro RNA editing assay46 contributed substantially to early progress in this field. Because RNA editing generally occurs in the protein-coding regions encoding different amino acid sequences, cis-elements cannot be highly conserved among sites. One question was asked: how did the RNA editing machinery recognize approximately 500 editing sites with independent cis-elements? The answer was that a PPR protein is a trans-factor that recognizes a cis-element.23,47
CRR4 is a member of the PLS subfamily that was discovered through the analysis of Arabidopsis mutants defective in RNA editing in plastids (Fig. 1). crr4 mutants were isolated for their specific defect in chloroplast NDH activity.47 The plastid ndhD gene encodes a subunit of the chloroplast NDH complex, and its translational initiation codon is encoded by ACG in the Arabidopsis plastid genome. The codon is edited to AUG in the RNA molecules. crr4 mutants are specifically defective in this RNA editing; consequently, they cannot translate ndhD.47 Recombinant CRR4 protein binds the 36 nucleotides surrounding the target site (25 upstream and 10 downstream nucleotides) in vitro, providing evidence that the PPR protein is the trans-factor required for the site-recognition of RNA editing in plastids.23 This discovery has been followed by the identification of approximately 30 PLS members that function in the site-recognition of RNA editing in Arabidopsis, rice and Physcomitrella patens.48 The site-recognition model that was first established in plastids has now also been accepted in mitochondrial RNA editing.49
The discovery of CRR4 was followed by the identification of CRR2150 and CLB1951 in plastids. All of these proteins belong to the E class. However, it is now estimated that approximately half of all trans-factors are members of the DYW class, indicating that there was an accidental bias toward the E class in the order of discovery. For this reason, we focused on the function of the DYW domain in RNA editing52 after the discovery of trans-factors with a further C-terminal extension in the form of the DYW domain.53 Special interest has been paid to this domain because the DYW domain is proposed to still have unidentified C-to-U conversion activity for the following two reasons:54 (1) the DYW domain contains the C/HxE(x)nPCxxC signature, which is highly conserved in C deaminase, and (2) the phylogenetic distribution of DYW members is strictly restricted to organisms with C-to-U-type RNA editing.
CRR22 and CRR28 are members of the DYW class (Fig. 1), and their mutants have also been isolated on the basis of defects in NDH activity.52 crr22 mutants cannot edit the ndhB-7, ndhD-5, and rpoB-3 sites, whereas crr28 cannot edit the ndhB-2 and ndhD-3 sites. Editing at the rpoB-3 site is not required for the function of PEP, and the phenotype of crr22 is specific to chloroplast NDH. To analyze the function of the DYW domain in CRR22 and CRR28, the sequences encoding the DYW domain were deleted from both genes, and the truncated genes were introduced into the corresponding mutants. The transformation fully complemented the function of trans-factors in RNA editing, indicating that the DYW domain is not required for the function of CRR22 and CRR28.52 Similar results have been reported in OTP82.55 In contrast to the DYW domain, the E domain is essential for the function of CRR22 and CRR28.52 This phenomenon is consistent with the result that the E domain is also essential for the function of CRR21 (E class), which is required for RNA editing at the ndhD-2 site.50 The E domain is exchangeable between CRR4 and CRR21 without affecting the function of either protein.50 In contrast, the E domain is not exchangeable between mitochondrial and plastid PPR proteins.56 The N-terminal PPR motifs (PLS motifs) of E- and DYW-class members likely help to recognize the different cis-elements. For this reason, the amino acid sequences are not highly conserved in the PLS domain. However, the C-terminal E domain is highly conserved, at least in plastids, as it is exchangeable between CRR4 and CRR21 without affecting the function of each PPR protein.50 However, divergence is observed between the E domains of mitochondria and plastids, implying the functional unequivalence of the E domains in the two organelles.56 We proposed a two-component model in which the E domain recruits the unidentified editing enzyme, which then mediates C-to-U conversion.34
Although the DYW domain is not essential for the function of trans-factors, this domain is still the best candidate for the unidentified editing enzyme; the domain is at least believed to be closely associated with the editing reaction. If the DYW domain is required for C-to-U conversion, how do E class members that lack the DYW domain edit their target sites? crr4 mutants are specifically defective in the RNA editing of the ndhD-1 site.47 Surprisingly, dyw1 mutants exhibit exactly the same phenotype as crr4.57 The mutants are defective in the gene encoding DYW1, which has no PPRs but has a DYW domain. The crr4 dyw1 double mutant is complemented by a CRR4-DYW1 fusion protein, indicating that the two proteins function together in the RNA editing of the ndhD-1 site.57 In contrast, the DYW domain is not essential for this function in some trans-factors. One of the possible explanations for the role of E-class members is that molecules other than trans-factors supply the DYW domain in the RNA-editing machinery. DYW1 is the C deaminase in this scenario.57 The following question remains: what is the origin of the DYW domain for other editing sites? Approximately half of trans-factors lack the DYW domain (Fig. 1), and the dyw1 phenotype is specific to the ndhD-1 site.
Why do plants edit RNA?
The molecular function of P-subfamily members can be simply explained by their ability to bind targets to protect the region from exonuclease attack. In addition, they modify RNA secondary structures to recruit the machinery of translation or other RNA maturation steps.37 For this function, tandem arrays of PPR motifs are likely sufficient. However, RNA editing requires the function of C-terminal domains (at least the E domain and, most likely, the DYW domain), although their exact function is unclear. The PLS subfamily is specific to land plants,18,58 and the majority of members are predicted to be involved in RNA editing. In the PLS subfamily, N-terminal PLS motifs function in RNA sequence recognition, as do canonical PPR motifs.23,52 Members of the P family with the C-terminal extension of E (+DYW) domain(s) from the PLS subfamily may be able to act as trans-factors in RNA editing. However, this was not the case in the evolution of land plants. Why did the RNA editing machinery select the PLS subfamily as trans-factors? Before answering this question, we need to take into account the physiological function of RNA editing.
Although members of the P subfamily are involved in the post-transcriptional regulation of gene expression in plastids and mitochondria, a regulatory function is unlikely for the PLS members involved in RNA editing. CRR4 determines the efficiency of the RNA editing that generates the translational initiation codon of ndhD. This phenomenon is the most probable case for the regulation of gene expression by RNA editing in plastids.59 However, the efficiency of RNA editing does not limit translation—and, consequently, the accumulation of the NDH complex—in vivo, even in Nicotiana tomentosiformis, in which the site is edited in only 15% of transcripts.60 We also cannot find any physiological reasons for the existence of other RNA editing sites. Many, but not all, editing sites should be precisely edited to encode functional proteins, but the sites can be edited to T in the genome. T already encodes the ndhD-1 site in the genome in monocots,61 suggesting that translational regulation via RNA editing is not essential for this site.
RNA editing restores the T-to-C mutation that occurs in the genome at the RNA level.62,63 This idea is scientifically acceptable because it describes what plants actually do, and it does not provide any reason why plants prefer RNA editing over correcting the genome. We will speculate further on these reasons here, although our speculation is not fully supported by experimental evidence. During the evolution of RNA editing in land plants, the T-to-C mutations were tentatively stored in the genome as long as they were corrected by RNA editing. The mutations are deleterious in general and, in the end, need to be corrected in the genome. In actuality, editing sites, at least in plastids, are considered to be on the way out during evolution.64,65 However, some mutations may have been beneficial to plants. RNA editing may have provided more chances for evolution in plastid or mitochondrial genes. RNA editing sites are concentrated into 11 ndh genes in plastids. Compared with the other fundamental machinery of photosynthesis (photosystems, the Cyt b6f complex, ATPase and ribulose-1,5-bisphosphate carboxylase/oxygenase [Rubisco]) and with the system of gene expression, the chloroplast NDH complex has experienced more drastic evolution in land plants.66-68 RNA editing may have contributed to this evolution.
Evolutionary models such as cryptic genetic variation (CGV) may be applied to explain potential benefits of RNA editing. The fundamental idea of these theories is as follows: robustness of phenotypes under various environmental conditions can accumulate mutations, and advantages of these CGVs can be exposed when facing against challenging or new conditions. As observed in the previous studies, the robustness of the plastid translation system tolerates and produces the same amount of proteins in vivo, even when RNA editing efficiency is deviated.60 In theory, unintended RNA editing by leaky RNA recognition of PPR proteins may raise the ratio of transcripts harboring CGV in the plastid RNA pool, and some beneficial mutations could have been fixed by natural selection and, thereby, giving birth to the new RNA editing site.
Although it has been stated in many cases, it is not entirely true that RNA editing is essential for the function of proteins encoded by the target RNA: defects in RNA editing do not necessarily result in a mutant protein function phenotype. OTP82 is a DYW member and is required for RNA editing at the ndhB-9 and ndhG-1 sites.55 Despite the clear phenotypes in RNA editing, the otp82 mutants do not exhibit any defects in chloroplast NDH stability, activity or interaction with photosystem I (PSI). Furthermore, the sites are partially edited even in the wild-type, implying that both edited and unedited RNAs are translated and that the NDH complex may include different versions of NdhB and NdhG in the wild-type.55 We do not eliminate the possibility that proteins translated from edited RNA are superior to those translated from unedited RNA under certain conditions. However, the case of OTP82 is unlikely to be an exception because defects in eight editing events among 34 editing sites in Arabidopsis plastids did not cause the mutant phenotype that can be detected in chlorophyll fluorescence analysis.62 However, plants correct the mutations that occur in their genomes by RNA editing, even though the correction is not always physiologically necessary.
Flexibility of the PLS subfamily in RNA recognition
Why does the RNA editing machinery often correct T-to-C mutations despite the absence of strong selection pressure? To answer this question, we can speculate on the flexibility of trans-factors in RNA sequence recognition. Flowering plants contain approximately 500 editing sites in their plastids and mitochondria, but a set of PLS members encoded by the genome may recognize more RNA sequences, including those not associated with RNA editing. A PPR protein often recognizes multiple RNA sequences. A well-studied example is PPR10, which recognizes short sequences conserved in the atpI-atpH and psaJ-rpl33 intergenic regions.25 Among 17 or 18 nucleotides (the psaJ-rpl33 site has one nucleotide insertion) recognized by PPR10, 13 are conserved. For some members of the P family, a PPR protein often recognizes multiple sequences with relatively high identity.30 However, in PLS subfamily members involved in RNA editing, the sites recognized by a single PPR protein are often not highly conserved.24,52,55 To explain this ambiguous rule of sequence recognition in the PLS subfamily, Hammani et al.69 proposed a model in which a trans-factor generally distinguishes pyrimidines from purines. However, this model still cannot explain CRR22, which recognizes unrelated sequences present at the ndhB-7, ndhD-5 and rpoB-3 sites.52 Does CRR22 function as a genuine sequence-recognizing trans-factor for all the sites? Alternatively, CRR22 may interact with other PPR proteins that determine the sequence specificity for some sites; in this way, CRR22 may play an essential role other than site recognition in RNA editing. The latter possibility may explain the puzzling fact that many trans-factors, including CRR4, lack the DYW domain, whereas CRR4 requires DYW1 in RNA editing.57 However, recombinant CRR22 binds to unrelated sequences present at the ndhB-7 and ndhD-5 sites with similar affinity.24 We can partially explain the molecular code by which a tandem array of PPR motifs recognizes specific RNA sequences,70 but we still cannot explain the flexibility of RNA recognition observed in some PLS members. In many cases, the target sites of trans-factors in RNA editing become clear through an examination of the mutant phenotypes. Although the trans-factors can recognize other sites, these sites have not been detected experimentally by genetics. Extensive in vitro binding surveys may be effective in testing how divergent sequences can be recognized by a single PLS member. We speculate that the flexibility of RNA sequence recognition is one of the reasons why the RNA editing machinery selected the PLS subfamily as a trans-factor. As discussed later, the L motif of PPR variants may not contribute to RNA sequence recognition.70 This phenomenon may be the reason for the flexibility of the PLS subfamily in RNA sequence recognition. More than 200 members of the PLS subfamily likely manage approximately 500 RNA editing sites, but the same set of proteins could potentially recognize many more sites. This flexibility in RNA sequence recognition may explain why some sites are recognized by trans-factors even without strong selection pressure for RNA editing. The putative cryptic binding sites of the PLS family may give rise to novel RNA editing sites. The fixation of an RNA editing site could be enabled by the flexibility of PLS PPR proteins, in agreement with the genetic drift hypothesis.63
Many footprints of PPR proteins have been detected in the plastid transcriptome.36 However, this is not the case for the cis-elements recognized by trans-factors of RNA editing. Members of the P subfamily are likely to stably associate with the target RNA, which is protected from endonucleases.37 However, many cis-elements for RNA editing are present in the protein-coding regions, and the stable binding of trans-factors may interfere with translation. Once a site is edited, trans-factors should be released from the site. Deamination from C to U by RNA editing may modify the affinity of cis-elements for trans-factors, but this is not the case with CRR4.23 Alternatively, trans-factors may only transiently interact with target RNA in vivo. PPR10 binds target sequences with a Kd of 0.1 nM,29 but the affinity is approximately 100 times lower in CRR4,23 OTP82 and CRR22.24 Affinity may not absolutely depend on the subfamily: the Kd values of PGR3 (P subfamily) are 6.4 and 9.5 nM for the petL and ndhA target sequences, respectively,28 although PGR3 maintains a clear footprint.36 Different modes of target binding may be required between RNA editing and other RNA maturation steps, including stabilization and translation. This requirement may be another reason why the PLS subfamily was selected by the RNA editing machinery as a trans-factor.
Non-PPR factors required for RNA editing
Members of the PLS subfamily are trans-factors that recruit an editing enzyme to the target sites. Consistent with this model, the mutant phenotypes of trans-factors are specific to one or several RNA editing sites.48 If the editing enzyme catalyzing the C-to-U conversion is shared by multiple editing sites, the mutation in the enzyme should cause a global defect in RNA editing in plastids or mitochondria. The Arabidopsis multiple organellar RNA editing factor (MORF) has been identified in mutants exhibiting defects in more than 40 mitochondrial RNA editing events.71 In Arabidopsis, the MORF forms a small family with nine members. MORF2 and MORF9 target plastids, and a defect in them affects nearly all of the RNA editing sites in plastids. Both proteins form a homo- or heterodimer, as do mitochondrial members. MORF8 is dual-targeting to plastids and mitochondria and has been independently identified as an RNA-editing factor interacting protein 1 (RIP1) through its affinity with a trans-factor, RARE1 (a DYW member), that is required for the RNA editing of the accD-1 site.72 The structure of MORF/RIP does not suggest that the family members possess C-deaminase activity.
Organelle RRM protein 1 (ORRM1) is also required for RNA editing at 12 sites in Arabidopsis and nine sites in maize; this protein has a pair of truncated RIP domains and an RNA recognition motif (RRM).73 ORRM1 interacts with PLS PPR proteins through its duplicated RIP moiety. In addition to ORRM1, another RRM protein, CP31, was suggested to be involved in RNA editing in plastids.46,74 The editing machinery in plastids may be more complicated than we previously estimated in the two-component model.34 In addition to a trans-factor, the MORF2/MORF9 heterodimer is also necessary for almost all editing sites in plastids. RRM proteins are likely to play an additional role. Furthermore, the DYW domain may be provided from the fourth molecule in the editosome; this domain is DYW1 for the ndhD-1 site. If the DYW domain does not have C-deaminase activity, then we will need to predict additional protein(s) with this enzyme activity. In trypanosomes, RNA editing is also performed by a multitude of proteins that still need to be fully characterized.75
CRR2 is an important exception
CRR2 is the first member of the DYW class that has been characterized in detail (Fig. 1). The Arabidopsis crr2 mutants were isolated because of their specific defects in chloroplast NDH activity.8 Because crr2 is also the first well-characterized mutant defective in chloroplast NDH in Arabidopsis, it has been used extensively in studies of the physiology of cyclic electron transport around PSI.76-78 crr2 mutants fail to accumulate two transcripts in which the 5′ and 3′ ends are localized to the rps7-ndhB intergenic region.8 As discussed above, the DYW motif is considered to be involved in RNA editing. CRR2 may edit RNA, which is the process that is necessary prior to intergenic RNA cleavage, but we failed to find any RNA editing defect in the intergenic region in crr2 mutants. Furthermore, the DYW domains of some PPR proteins, including CRR2, have endonuclease activity in vitro rather than C deaminase activity,52,79 suggesting that CRR2 is a restriction ribonuclease. RNA PROCESSING FACTOR3 and 5 are RF-like PPR proteins that are also required for the RNA cleavages that generate the 5′ ends of RNAs in mitochondria.80,81 CRR2 is unique because even a purified recombinant PPR protein might be involved in RNA cleavage.52,79
Notably, CRR2 truncated in the DYW domain does not complement the crr2 mutant, in contrast with the fact that the DYW domain is not required for RNA editing, as shown in some DYW members.23,52 Chimeric CRR22 and CRR28 with the DYW domain originating from CRR2 do not complement the crr22 and crr28 mutants, respectively, whereas CRR22 and CRR28 without the DYW domains complement the respective phenotypes. This result suggests that the DYW domain of CRR2 has a dominant-negative effect.52 We cannot eliminate the possibility that the DYW domain is required for the stability of CRR2 in vivo. However, based on all of the above-mentioned results, we still hesitate to either accept or reject the idea that the DYW domain is responsible for C-deaminase activity in the RNA editing machinery.
Based on the crr2 mutant phenotype in transcripts, we concluded that CRR2 is involved in intergenic RNA cleavage between rps7 and ndhB.8 This phenomenon would be the simplest explanation for the mutant phenotype. However, the new model established in the P family suggests that the 5′ and 3′ ends of RNA are not directly generated by endonuclease.37 We can imagine that CRR2 binds the intergenic region between rps7 and ndhB and protects the 5′ end of ndhB and the 3′ end of rps7 from 5′→3′ and 3′→5′ exonucleases, respectively. CRR2 may also modify the secondary structure of RNA near the binding site and recruit ribosomes for ndhB translation. We accept the idea that the translational initiation codon was incorrectly annotated in our report8 and that the real initiation codon should be the downstream in-frame ATG.36 This alternative model of CRR2 function is supported by the discovery of RNA footprints located at the predicted binding site of CRR2.36 The binding sequence of CRR2 needs to be determined experimentally in vitro. In general, members of the PLS subfamily do not leave footprints;36 this phenomenon may be due to the transient interaction of PLS members with target RNAs. Information is still limited on PLS members that are not involved in RNA editing. In the case of DYW members functioning as trans-factors in RNA editing, the distance between their binding sites and the target C is highly conserved.70,82 On the basis of this information, the CRR2 target sequence is predicted to be slightly more upstream than the reported footprint, where in silico prediction70 has found a reasonable candidate for the CRR2 binding sequence (data not shown). Usually, PLS binding sites are presumed to terminate with an edited C, but in this case, emphasis is placed on placing the PPR close to the 5′ end of mature ndhB mRNA.
In any case, CRR2 is an important exception, and further research is needed. Another exception to the DYW class is PpPPR_43, which is required for the splicing of the group II intron of cox1 in Physcomitrella patens.83 Both the E domain and the DYW domain are not essential for the function of PpPPR_43. In contrast, the DYW domain is essential for CRR2. The PLS domain may play a role similar to that of P-subfamily members in RNA splicing. If this is the case, we raise the following question: what is the difference between the exceptional PLS motifs included in CRR2 or PpPPR_43 and those included in the trans-factors for RNA editing?
Mechanism of RNA sequence recognition
Almost 10 y after the discovery of the PPR family, the mode of RNA recognition by PPR proteins has only begun to be elucidated. As predicted from their similarity to TPR proteins,6,33,84 crystallized PPR fragments form a modular alignment of α-solenoid helices.21,85,86 Notably, the active RNA-binding status of PPR has been solved as its monomer form,70 not its homodimer form.11,25 Considering this information, although direct experimental support by the RNA–PPR co-structure is still missing, it would be fair to assume that each PPR tract recognizes a single RNA base. The mechanism of RNA sequence recognition by PPR should resemble that of transcription-activator-like effectors (TALEs). TALEs are modular repeat proteins that bind to specific DNA sequences on the basis of a predictable code.87,88 Two particular consecutive amino acids within 30–40 amino acid units of TALEs (known as repeat-variable diresidues: RVDs) are responsible for determining DNA recognition specificity.87,88 Similarly, several recent studies have attempted to solve the “code” of RNA recognition by PPR proteins.
The amino acids that serve as major determinants of RNA recognition have been highlighted by an evolutionary systems biological approach.33 A group of mitochondrial PPR proteins that were under a much higher selection pressure than all of the other members of the PPR family were selected by comprehensive phylogenetic analysis.33,58 The genes encoding these proteins are known as fertility restorer (Rf) genes and are counter-evolved against mitochondrial cytoplasmic male sterility (CMS). CMS has been widely observed in > 150 plant species, and the rapid evolutionary turnover of the mitochondrial gene that is responsible for CMS has been well characterized.89,90 Similar to disease-resistance genes, Rf-encoding PPR genes (Rf-PPR) show a high degree of diversifying selection in all tested plant species.33 A cumulative calculation of positive selection probability suggests that positions 1, 3 and 6 within 35 amino acids of the PPR motif are hypervariable residues in Rf-PPR (Fig. 3).33 It has been speculated that these residues act as RVDs in TALEs. An independent experiment has also found that substitutions by alanine at positions 1, 3, 4, 6, 10 and 14 impairs the RNA-binding ability of the minimally functional PPR protein that consists of two PPR motifs.91

Figure 3. The X-ray structure of two consecutive PPR motifs in human mitochondrial RNA polymerase (PDB 3SPA).21 Residues known to be involved in RNA binding are highlighted in color. Positions 1 (brown) and 6 (yellow) are the most important components of the PPR code.70
Based on the accumulated information on trans-factors (PLS members) and their corresponding RNA editing sites, a statistical study was performed to elucidate the principal components of the PPR code; the study was also based on the research that had focused on the amino acid residues involved in RNA recognition in PPRs.70,92 In summary, a strong correlation was observed between the combination of positions 6 and 1 on the following PPR (position 1’) and the corresponding RNA base (Fig. 3). It has also been suggested that position 3 acts in synergy to specify the RNA base.92 The strongest determinant was located at position 6. In the P and S motifs of PPR variants, N at this position is correlated with the pyrimidine base in the RNA target, whereas S/T is correlated with purine, in agreement with the results of a prior investigation on the purine/pyrimidine distinguishing basis of PPR proteins.69 The largest caveat here is that these codes do not apply to L motifs, which are PPR variants with slightly longer (36 amino acid) consensus sequences,20 and only a weak correlation has been observed between the L motif and the RNA sequence.92 It remains to be determined whether these L motifs are non-functional base-skip regions or are ruled by a distinct code. Furthermore, it must be noted that not all of the RNA binding sites of trans-factors (i.e., PLS members) are consistent with the consensus code.70 For instance, the predicted RNA-binding alignments of CRR4 and CRR21 (chloroplastic) or MEF9 and SLG1 (mitochondrial) are plausible with the current code. However, the alignments of a few RNA editing factors are relatively poor, especially in the case of those that mediate RNA editing at multiple sites (CRR22, MEF11, etc.). A fine functional dissection of the additional amino acid residues implicated in RNA binding (positions 3, 4, 10 and 14) is necessary to address this issue. For example, structural predictions have indicated that basic regions, including position 14 at the bottom of the PPR central groove, may interact with RNA phosphate.84 Alternatively, the RNA editing activity of these trans-factors might be governed by some other factors beyond the PPR code, such as by collaborative RNA recognition, a tandem PPR-PPR interaction or external molecules, such as MORF/RIP.71-73,93
Physiological function of the post-transcriptional regulation of plastid gene expression
In the final section, we focused on the physiological function of regulation that may be mediated by PPR proteins. During chloroplast development, the switching of the transcription system from being dependent on nuclear-encoded RNA polymerase (NEP) to being dependent on PEP plays a crucial role in regulating global gene expression.94 However, once chloroplasts are developed, the regulation of gene expression is shifted to post-transcriptional levels,95 at which PPR proteins play a central role. The examination of mutant phenotypes clearly shows that PPR proteins are essential for gene expression in plastids, but these results do not necessarily mean that regulation via PPR proteins is physiologically significant in the wild-type. Notably, we failed to find any physiological function of RNA editing, although mutants defective in RNA editing are often lethal. Numerous RNA editing sites are specific to some plant lineages, suggesting that the sites have been corrected to T at the genome level during evolution. The majority of RNA maturation processes mediated by PPR proteins may be solely for debugging the genomes: P class members are also trans-acting factors that suppress the mutations that occur in the gonome.96 In actuality, the information on the physiological significance of the regulation via PPR protein is very limited: the RNA maturation process limits the accumulation of product, and the PPR protein is regulated by a signal. Because many organelle genes encode subunits of large protein complexes that are encoded by both nuclear and organelle genomes, it is unclear whether the subunit limits the accumulation of the complex. An exception is MCA1, which determines the level of the Cyt b6f complex by regulating the stability of petA mRNA.97
Although the exact molecular mechanism is unclear, the plastid gene expression needs to be regulated. Regulation via transcription is unlikely to have sufficient complexity to respond to the physiological and developmental requirements. In NADP-malic enzyme-type C4 plants, rbcL should be specifically expressed in bundle sheath cells (BSCs).98 In contrast, plastid-encoded PSII subunit genes (psb genes) are preferentially expressed in mesophyll cells (MCs) to avoid O2 evolution in BSCs.99 Despite the cell type-specific accumulation of proteins, the transcriptional activity of rbcL and psb do not differ drastically between BSCs and MCs in maize, suggesting that there is post-transcriptional regulation.99 MRL1 is a PPR protein that was discovered in Chlamydomonas and is required for the stabilization of rbcL mRNA.100 MRL1 is conserved in flowering plants, but the expression of its maize ortholog does not show tissue specificity, at least at transcript levels.101 Additional factors are needed to explain the BSC-specific accumulation of Rubisco in maize.
Plastid gene expression also needs to be regulated in C3 plants. A possible mechanism is the regulation of the excitation balance of two photosystems. State transition is believed to be involved in short-term adjustments of the excitation balance by controlling the distribution of the mobile part of the light harvesting complex II (LHCII) between two photosystems.102 However, in the longer term of excitation imbalance, plants remodel the stoichiometry of photosystems by a process in which chloroplast gene expression is regulated at both the transcriptional and the post-transcriptional levels.103,104 How is the excitation balance disturbed in steady-state photosynthesis, especially in land plants, because light quality does not change drastically in terrestrial environments? The Arabidopsis stn7 mutant defective in state transition is sensitive to fluctuating light conditions due to photodamage to PSI.105,106 However, at a constant light intensity, stn7 mutants decrease the PSII to PSI ratio to balance the excitation of the two photosystems in the absence of state transition.107 This is the case in the mutants, but it suggests that the physiological significance of the process tunes the excitation balance exactly.
Because Rubisco is the most abundant leaf protein due to its low rate of catalysis, plants have to optimize their Rubisco content for nitrogen economy, as well as for maximum photosynthesis under fluctuating light conditions. In flowering plants, Rubisco consists of eight large subunits (LS) encoded by the plastid rbcL gene and eight small subunits (SS) encoded by the small family of nuclear rbcS genes. Antisense rbcS tobacco plants accumulate normal levels of rbcL mRNA in their chloroplasts, but their translation of rbcL is downregulated.108 The translation of rbcL is under the control of the assembly mediated regulation of translation.109 How does the reduction in the Rubisco level affect photosynthesis? In antisense rbcS rice plants, in which the Rubisco level is reduced to 65% of the wild-type level, the CO2 fixation rate is reduced to 80% of the wild-type rate under saturating light in air (36 Pa CO2).110 In contrast, the rate is 5–15% higher at elevated partial pressures of CO2 (80–90 Pa). This high rate of CO2 fixation is explained by more efficient nitrogen utilization: plants contain slightly higher levels of chlorophyll, Cyf and ATPase.110 An important message from this result is that even a mild alteration (35% decline) in the Rubisco content leads to clear photosynthesis phenotypes in a leaf. This range of protein level changes is usually not the subject of molecular biology. However, we need to precisely detect slight changes and to evaluate the consequent phenotypes in photosynthesis to link the molecular biology of chloroplasts and the physiology of photosynthesis; the regulation of photosynthesis is related to nitrogen economy because the machinery of photosynthesis is very abundant in proteins.
Future prospects
PPR proteins are part of the eukaryotic machinery for regulating organelle gene expression in plants. A dramatic expansion of the family has occurred not only in the PLS subfamily but also in the P subfamily, suggesting that the presence of this huge family cannot be solely explained by the high frequency of RNA editing in flowering plants. Host cells need to regulate gene expression in plastids precisely, especially in land plants. In this decade, our understanding of this protein family has dramatically improved. However, our questions have mainly been answered in molecular biological terms; the answers have linked the gene and the RNA phenotypes but have not elucidated the physiological reasons why a eukaryotic regulatory system is required for the regulation of plastid function. It is also puzzling that more than half of the PPR family members target mitochondria. Half of these PPR proteins in the mitochondria may mediate RNA editing, but we still cannot imagine that the remaining half is required for the regulation of approximately 60 genes in the mitochondria. As discussed in this review article, the phenotypes of knockout mutants do not necessarily inform us of the physiological significance of this regulation. Fundamental questions in molecular biology remain; for example, the identity of the editing enzyme is still unclear. However, our focus in future research needs to move on to the link with physiology so we can gain an understanding of the evolutionary strategy of symbiosis.
Acknowledgments
TS was supported by Grants 22114509 and 22247005 from the Ministry of Education, Culture, Sports, Science and CREST. SF was supported by the Grant-in-Aid for Japan Society for the Promotion of Science fellows.
Glossary
Abbreviations:
- BSCs
bundle sheath cells
- CGV
cryptic genetic variation
- CMS
cytoplasmic male sterility
- Cyt
cytochrome
- LHCII
light harvesting complex II
- LS
Rubisco large subunit
- MCs
mesophyll cells
- NDH
NADH dehydrogenase-like (complex)
- NEP
nuclear-encoded RNA polymerase
- NPQ
non-photochemical quenching of chlorophyll fluorescence
- PEP
plastid-encoded plastid RNA polymerase
- PPR
pentatricopeptide repeat
- PQ
plastoquinone
- PSI(II)
photosystem I(II)
- Rf
fertility restorer
- Rubisco
ribulose-1,5-bisphosphate carboxylase/oxygenase
- RVDs
repeat-variable diresidues
- SD
Shine-Dalgarno
- SS
Rubisco small subunit
- TALEs
Transcription-Activator-Like Effectors
Disclosure of Potential Conflicts of Interest
No potential conflicts of interest were disclosed.
Footnotes
Previously published online: www.landesbioscience.com/journals/rnabiology/article/25207
References
- 1.Shiina T, Tsunoyama Y, Nakahira Y, Khan MS. Plastid RNA polymerases, promoters, and transcription regulators in higher plants. Int Rev Cytol. 2005;244:1–68. doi: 10.1016/S0074-7696(05)44001-2. [DOI] [PubMed] [Google Scholar]
- 2.Stern DB, Goldschmidt-Clermont M, Hanson MR. Chloroplast RNA metabolism. Annu Rev Plant Biol. 2010;61:125–55. doi: 10.1146/annurev-arplant-042809-112242. [DOI] [PubMed] [Google Scholar]
- 3.Manthey GM, McEwen JE. The product of the nuclear gene PET309 is required for translation of mature mRNA and stability or production of intron-containing RNAs derived from the mitochondrial COX1 locus of Saccharomyces cerevisiae. EMBO J. 1995;14:4031–43. doi: 10.1002/j.1460-2075.1995.tb00074.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Coffin JW, Dhillon R, Ritzel RG, Nargang FE. The Neurospora crassa cya-5 nuclear gene encodes a protein with a region of homology to the Saccharomyces cerevisiae PET309 protein and is required in a post-transcriptional step for the expression of the mitochondrially encoded COXI protein. Curr Genet. 1997;32:273–80. doi: 10.1007/s002940050277. [DOI] [PubMed] [Google Scholar]
- 5.Barkan A, Walker M, Nolasco M, Johnson D. A nuclear mutation in maize blocks the processing and translation of several chloroplast mRNAs and provides evidence for the differential translation of alternative mRNA forms. EMBO J. 1994;13:3170–81. doi: 10.1002/j.1460-2075.1994.tb06616.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Small ID, Peeters N. The PPR motif - a TPR-related motif prevalent in plant organellar proteins. Trends Biochem Sci. 2000;25:46–7. doi: 10.1016/S0968-0004(99)01520-0. [DOI] [PubMed] [Google Scholar]
- 7.Schmitz-Linneweber C, Small I. Pentatricopeptide repeat proteins: a socket set for organelle gene expression. Trends Plant Sci. 2008;13:663–70. doi: 10.1016/j.tplants.2008.10.001. [DOI] [PubMed] [Google Scholar]
- 8.Hashimoto M, Endo T, Peltier G, Tasaka M, Shikanai T. A nucleus-encoded factor, CRR2, is essential for the expression of chloroplast ndhB in Arabidopsis. Plant J. 2003;36:541–9. doi: 10.1046/j.1365-313X.2003.01900.x. [DOI] [PubMed] [Google Scholar]
- 9.Yamazaki H, Tasaka M, Shikanai T. PPR motifs of the nucleus-encoded factor, PGR3, function in the selective and distinct steps of chloroplast gene expression in Arabidopsis. Plant J. 2004;38:152–63. doi: 10.1111/j.1365-313X.2004.02035.x. [DOI] [PubMed] [Google Scholar]
- 10.Sung TY, Tseng CC, Hsieh MH. The SLO1 PPR protein is required for RNA editing at multiple sites with similar upstream sequences in Arabidopsis mitochondria. Plant J. 2010;63:499–511. doi: 10.1111/j.1365-313X.2010.04258.x. [DOI] [PubMed] [Google Scholar]
- 11.Meierhoff K, Felder S, Nakamura T, Bechtold N, Schuster G. HCF152, an Arabidopsis RNA binding pentatricopeptide repeat protein involved in the processing of chloroplast psbB-psbT-psbH-petB-petD RNAs. Plant Cell. 2003;15:1480–95. doi: 10.1105/tpc.010397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Cushing DA, Forsthoefel NR, Gestaut DR, Vernon DM. Arabidopsis emb175 and other ppr knockout mutants reveal essential roles for pentatricopeptide repeat (PPR) proteins in plant embryogenesis. Planta. 2005;221:424–36. doi: 10.1007/s00425-004-1452-x. [DOI] [PubMed] [Google Scholar]
- 13.Kazama T, Toriyama K. A pentatricopeptide repeat-containing gene that promotes the processing of aberrant atp6 RNA of cytoplasmic male-sterile rice. FEBS Lett. 2003;544:99–102. doi: 10.1016/S0014-5793(03)00480-0. [DOI] [PubMed] [Google Scholar]
- 14.Wang Z, Zou Y, Li X, Zhang Q, Chen L, Wu H, et al. Cytoplasmic male sterility of rice with boro II cytoplasm is caused by a cytotoxic peptide and is restored by two related PPR motif genes via distinct modes of mRNA silencing. Plant Cell. 2006;18:676–87. doi: 10.1105/tpc.105.038240. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kobayashi K, Suzuki M, Tang J, Nagata N, Ohyama K, Seki H, et al. Lovastatin insensitive 1, a Novel pentatricopeptide repeat protein, is a potential regulatory factor of isoprenoid biosynthesis in Arabidopsis. Plant Cell Physiol. 2007;48:322–31. doi: 10.1093/pcp/pcm005. [DOI] [PubMed] [Google Scholar]
- 16.Koussevitzky S, Nott A, Mockler TC, Hong F, Sachetto-Martins G, Surpin M, et al. Signals from chloroplasts converge to regulate nuclear gene expression. Science. 2007;316:715–9. doi: 10.1126/science.1140516. [DOI] [PubMed] [Google Scholar]
- 17.Nakamura T, Yagi Y, Kobayashi K. Mechanistic insight into pentatricopeptide repeat proteins as sequence-specific RNA-binding proteins for organellar RNAs in plants. Plant Cell Physiol. 2012;53:1171–9. doi: 10.1093/pcp/pcs069. [DOI] [PubMed] [Google Scholar]
- 18.Fujii S, Small I. The evolution of RNA editing and pentatricopeptide repeat genes. New Phytol. 2011;191:37–47. doi: 10.1111/j.1469-8137.2011.03746.x. [DOI] [PubMed] [Google Scholar]
- 19.Lurin C, Andrés C, Aubourg S, Bellaoui M, Bitton F, Bruyère C, et al. Genome-wide analysis of Arabidopsis pentatricopeptide repeat proteins reveals their essential role in organelle biogenesis. Plant Cell. 2004;16:2089–103. doi: 10.1105/tpc.104.022236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Rivals E, Bruyère C, Toffano-Nioche C, Lecharny A. Formation of the Arabidopsis pentatricopeptide repeat family. Plant Physiol. 2006;141:825–39. doi: 10.1104/pp.106.077826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Ringel R, Sologub M, Morozov YI, Litonin D, Cramer P, Temiakov D. Structure of human mitochondrial RNA polymerase. Nature. 2011;478:269–73. doi: 10.1038/nature10435. [DOI] [PubMed] [Google Scholar]
- 22.Das AK, Cohen PW, Barford D. The structure of the tetratricopeptide repeats of protein phosphatase 5: implications for TPR-mediated protein-protein interactions. EMBO J. 1998;17:1192–9. doi: 10.1093/emboj/17.5.1192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Okuda K, Nakamura T, Sugita M, Shimizu T, Shikanai T. A pentatricopeptide repeat protein is a site recognition factor in chloroplast RNA editing. J Biol Chem. 2006;281:37661–7. doi: 10.1074/jbc.M608184200. [DOI] [PubMed] [Google Scholar]
- 24.Okuda K, Shikanai T. A pentatricopeptide repeat protein acts as a site-specificity factor at multiple RNA editing sites with unrelated cis-acting elements in plastids. Nucleic Acids Res. 2012;40:5052–64. doi: 10.1093/nar/gks164. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Pfalz J, Bayraktar OA, Prikryl J, Barkan A. Site-specific binding of a PPR protein defines and stabilizes 5′ and 3′ mRNA termini in chloroplasts. EMBO J. 2009;28:2042–52. doi: 10.1038/emboj.2009.121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Williams-Carrier R, Kroeger T, Barkan A. Sequence-specific binding of a chloroplast pentatricopeptide repeat protein to its native group II intron ligand. RNA. 2008;14:1930–41. doi: 10.1261/rna.1077708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Schmitz-Linneweber C, Williams-Carrier RE, Williams-Voelker PM, Kroeger TS, Vichas A, Barkan A. A pentatricopeptide repeat protein facilitates the trans-splicing of the maize chloroplast rps12 pre-mRNA. Plant Cell. 2006;18:2650–63. doi: 10.1105/tpc.106.046110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Cai W, Okuda K, Peng L, Shikanai T. PROTON GRADIENT REGULATION 3 recognizes multiple targets with limited similarity and mediates translation and RNA stabilization in plastids. Plant J. 2011;67:318–27. doi: 10.1111/j.1365-313X.2011.04593.x. [DOI] [PubMed] [Google Scholar]
- 29.Prikryl J, Rojas M, Schuster G, Barkan A. Mechanism of RNA stabilization and translational activation by a pentatricopeptide repeat protein. Proc Natl Acad Sci USA. 2011;108:415–20. doi: 10.1073/pnas.1012076108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Schmitz-Linneweber C, Williams-Carrier R, Barkan A. RNA immunoprecipitation and microarray analysis show a chloroplast pentatricopeptide repeat protein to be associated with the 5′ region of mRNAs whose translation it activates. Plant Cell. 2005;17:2791–804. doi: 10.1105/tpc.105.034454. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Khrouchtchova A, Monde RA, Barkan A. A short PPR protein required for the splicing of specific group II introns in angiosperm chloroplasts. RNA. 2012;18:1197–209. doi: 10.1261/rna.032623.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Zoschke R, Kroeger T, Belcher S, Schöttler MA, Barkan A, Schmitz-Linneweber C. The pentatricopeptide repeat-SMR protein ATP4 promotes translation of the chloroplast atpB/E mRNA. Plant J. 2012;72:547–58. doi: 10.1111/j.1365-313X.2012.05081.x. [DOI] [PubMed] [Google Scholar]
- 33.Fujii S, Bond CS, Small ID. Selection patterns on restorer-like genes reveal a conflict between nuclear and mitochondrial genomes throughout angiosperm evolution. Proc Natl Acad Sci USA. 2011;108:1723–8. doi: 10.1073/pnas.1007667108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Shikanai T. RNA editing in plant organelles: machinery, physiological function and evolution. Cell Mol Life Sci. 2006;63:698–708. doi: 10.1007/s00018-005-5449-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Liu X, Yu F, Rodermel S. An Arabidopsis pentatricopeptide repeat protein, SUPPRESSOR OF VARIEGATION7, is required for FtsH-mediated chloroplast biogenesis. Plant Physiol. 2010;154:1588–601. doi: 10.1104/pp.110.164111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Ruwe H, Schmitz-Linneweber C. Short non-coding RNA fragments accumulating in chloroplasts: footprints of RNA binding proteins? Nucleic Acids Res. 2012;40:3106–16. doi: 10.1093/nar/gkr1138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Zhelyazkova P, Hammani K, Rojas M, Voelker R, Vargas-Suárez M, Börner T, et al. Protein-mediated protection as the predominant mechanism for defining processed mRNA termini in land plant chloroplasts. Nucleic Acids Res. 2012;40:3092–105. doi: 10.1093/nar/gkr1137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Covello PS, Gray MW. RNA editing in plant mitochondria. Nature. 1989;341:662–6. doi: 10.1038/341662a0. [DOI] [PubMed] [Google Scholar]
- 39.Gualberto JM, Lamattina L, Bonnard G, Weil JH, Grienenberger JM. RNA editing in wheat mitochondria results in the conservation of protein sequences. Nature. 1989;341:660–2. doi: 10.1038/341660a0. [DOI] [PubMed] [Google Scholar]
- 40.Hiesel R, Wissinger B, Schuster W, Brennicke A. RNA editing in plant mitochondria. Science. 1989;246:1632–4. doi: 10.1126/science.2480644. [DOI] [PubMed] [Google Scholar]
- 41.Hoch B, Maier RM, Appel K, Igloi GL, Kössel H. Editing of a chloroplast mRNA by creation of an initiation codon. Nature. 1991;353:178–80. doi: 10.1038/353178a0. [DOI] [PubMed] [Google Scholar]
- 42.Giegé P, Brennicke A. RNA editing in Arabidopsis mitochondria effects 441 C to U changes in ORFs. Proc Natl Acad Sci USA. 1999;96:15324–9. doi: 10.1073/pnas.96.26.15324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Chateigner-Boutin AL, Small I. A rapid high-throughput method for the detection and quantification of RNA editing based on high-resolution melting of amplicons. Nucleic Acids Res. 2007;35:e114. doi: 10.1093/nar/gkm640. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Chaudhuri S, Maliga P. Sequences directing C to U editing of the plastid psbL mRNA are located within a 22 nucleotide segment spanning the editing site. EMBO J. 1996;15:5958–64. [PMC free article] [PubMed] [Google Scholar]
- 45.Svab Z, Maliga P. High-frequency plastid transformation in tobacco by selection for a chimeric aadA gene. Proc Natl Acad Sci USA. 1993;90:913–7. doi: 10.1073/pnas.90.3.913. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Hirose T, Sugiura M. Involvement of a site-specific trans-acting factor and a common RNA-binding protein in the editing of chloroplast mRNAs: development of a chloroplast in vitro RNA editing system. EMBO J. 2001;20:1144–52. doi: 10.1093/emboj/20.5.1144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Kotera E, Tasaka M, Shikanai T. A pentatricopeptide repeat protein is essential for RNA editing in chloroplasts. Nature. 2005;433:326–30. doi: 10.1038/nature03229. [DOI] [PubMed] [Google Scholar]
- 48.Castandet B, Araya A. RNA editing in plant organelles. Why make it easy? Biochemistry (Mosc) 2011;76:924–31. doi: 10.1134/S0006297911080086. [DOI] [PubMed] [Google Scholar]
- 49.Zehrmann A, Verbitskiy D, Härtel B, Brennicke A, Takenaka M. PPR proteins network as site-specific RNA editing factors in plant organelles. RNA Biol. 2011;8:67–70. doi: 10.4161/rna.8.1.14298. [DOI] [PubMed] [Google Scholar]
- 50.Okuda K, Myouga F, Motohashi R, Shinozaki K, Shikanai T. Conserved domain structure of pentatricopeptide repeat proteins involved in chloroplast RNA editing. Proc Natl Acad Sci USA. 2007;104:8178–83. doi: 10.1073/pnas.0700865104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Chateigner-Boutin AL, Ramos-Vega M, Guevara-García A, Andrés C, de la Luz Gutiérrez-Nava M, Cantero A, et al. CLB19, a pentatricopeptide repeat protein required for editing of rpoA and clpP chloroplast transcripts. Plant J. 2008;56:590–602. doi: 10.1111/j.1365-313X.2008.03634.x. [DOI] [PubMed] [Google Scholar]
- 52.Okuda K, Chateigner-Boutin AL, Nakamura T, Delannoy E, Sugita M, Myouga F, et al. Pentatricopeptide repeat proteins with the DYW motif have distinct molecular functions in RNA editing and RNA cleavage in Arabidopsis chloroplasts. Plant Cell. 2009;21:146–56. doi: 10.1105/tpc.108.064667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Zhou W, Cheng Y, Yap A, Chateigner-Boutin AL, Delannoy E, Hammani K, et al. The Arabidopsis gene YS1 encoding a DYW protein is required for editing of rpoB transcripts and the rapid development of chloroplasts during early growth. Plant J. 2009;58:82–96. doi: 10.1111/j.1365-313X.2008.03766.x. [DOI] [PubMed] [Google Scholar]
- 54.Salone V, Rüdinger M, Polsakiewicz M, Hoffmann B, Groth-Malonek M, Szurek B, et al. A hypothesis on the identification of the editing enzyme in plant organelles. FEBS Lett. 2007;581:4132–8. doi: 10.1016/j.febslet.2007.07.075. [DOI] [PubMed] [Google Scholar]
- 55.Okuda K, Hammani K, Tanz SK, Peng L, Fukao Y, Myouga F, et al. The pentatricopeptide repeat protein OTP82 is required for RNA editing of plastid ndhB and ndhG transcripts. Plant J. 2010;61:339–49. doi: 10.1111/j.1365-313X.2009.04059.x. [DOI] [PubMed] [Google Scholar]
- 56.Chateigner-Boutin AL, Colas des Francs-Small C, Fujii S, Okuda K, Tanz SK, Small I. The E domains of pentatricopeptide repeat proteins from different organelles are not functionally equivalent for RNA editing. Plant J. 2013 doi: 10.1111/tpj.12180. In press. [DOI] [PubMed] [Google Scholar]
- 57.Boussardon C, Salone V, Avon A, Berthomé R, Hammani K, Okuda K, et al. Two interacting proteins are necessary for the editing of the NdhD-1 site in Arabidopsis plastids. Plant Cell. 2012;24:3684–94. doi: 10.1105/tpc.112.099507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.O’Toole N, Hattori M, Andres C, Iida K, Lurin C, Schmitz-Linneweber C, et al. On the expansion of the pentatricopeptide repeat gene family in plants. Mol Biol Evol. 2008;25:1120–8. doi: 10.1093/molbev/msn057. [DOI] [PubMed] [Google Scholar]
- 59.Hirose T, Sugiura M. Both RNA editing and RNA cleavage are required for translation of tobacco chloroplast ndhD mRNA: a possible regulatory mechanism for the expression of a chloroplast operon consisting of functionally unrelated genes. EMBO J. 1997;16:6804–11. doi: 10.1093/emboj/16.22.6804. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Okuda K, Habata Y, Kobayashi Y, Shikanai T. Amino acid sequence variations in Nicotiana CRR4 orthologs determine the species-specific efficiency of RNA editing in plastids. Nucleic Acids Res. 2008;36:6155–64. doi: 10.1093/nar/gkn629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Tsudzuki T, Wakasugi T, Sugiura M. Comparative analysis of RNA editing sites in higher plant chloroplasts. J Mol Evol. 2001;53:327–32. doi: 10.1007/s002390010222. [DOI] [PubMed] [Google Scholar]
- 62.Shikanai T. Why do plants edit RNA in plant organelles? In: Bullerwell CE, ed Organelle Genetics 2012:381-97. [Google Scholar]
- 63.Covello PS, Gray MW. On the evolution of RNA editing. Trends Genet. 1993;9:265–8. doi: 10.1016/0168-9525(93)90011-6. [DOI] [PubMed] [Google Scholar]
- 64.Tillich M, Lehwark P, Morton BR, Maier UG. The evolution of chloroplast RNA editing. Mol Biol Evol. 2006;23:1912–21. doi: 10.1093/molbev/msl054. [DOI] [PubMed] [Google Scholar]
- 65.Hayes ML, Mulligan RM. Pentatricopeptide repeat proteins constrain genome evolution in chloroplasts. Mol Biol Evol. 2011;28:2029–39. doi: 10.1093/molbev/msr023. [DOI] [PubMed] [Google Scholar]
- 66.Peng L, Fukao Y, Fujiwara M, Takami T, Shikanai T. Efficient operation of NAD(P)H dehydrogenase requires supercomplex formation with photosystem I via minor LHCI in Arabidopsis. Plant Cell. 2009;21:3623–40. doi: 10.1105/tpc.109.068791. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Peng L, Shikanai T. Supercomplex formation with photosystem I is required for the stabilization of the chloroplast NADH dehydrogenase-like complex in Arabidopsis. Plant Physiol. 2011;155:1629–39. doi: 10.1104/pp.110.171264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Ueda M, Kuniyoshi T, Yamamoto H, Sugimoto K, Ishizaki K, Kohchi T, et al. Composition and physiological function of the chloroplast NADH dehydrogenase-like complex in Marchantia polymorpha. Plant J. 2012;72:683–93. doi: 10.1111/j.1365-313X.2012.05115.x. [DOI] [PubMed] [Google Scholar]
- 69.Hammani K, Okuda K, Tanz SK, Chateigner-Boutin AL, Shikanai T, Small I. A study of new Arabidopsis chloroplast RNA editing mutants reveals general features of editing factors and their target sites. Plant Cell. 2009;21:3686–99. doi: 10.1105/tpc.109.071472. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Barkan A, Rojas M, Fujii S, Yap A, Chong YS, Bond CS, et al. A combinatorial amino acid code for RNA recognition by pentatricopeptide repeat proteins. PLoS Genet. 2012;8:e1002910. doi: 10.1371/journal.pgen.1002910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Takenaka M, Zehrmann A, Verbitskiy D, Kugelmann M, Härtel B, Brennicke A. Multiple organellar RNA editing factor (MORF) family proteins are required for RNA editing in mitochondria and plastids of plants. Proc Natl Acad Sci USA. 2012;109:5104–9. doi: 10.1073/pnas.1202452109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Bentolila S, Heller WP, Sun T, Babina AM, Friso G, van Wijk KJ, et al. RIP1, a member of an Arabidopsis protein family, interacts with the protein RARE1 and broadly affects RNA editing. Proc Natl Acad Sci USA. 2012;109:E1453–61. doi: 10.1073/pnas.1121465109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Sun T, Germain A, Giloteaux L, Hammani K, Barkan A, Hanson MR, et al. An RNA recognition motif-containing protein is required for plastid RNA editing in Arabidopsis and maize. Proc Natl Acad Sci USA. 2013;110:E1169–78. doi: 10.1073/pnas.1220162110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Tillich M, Hardel SL, Kupsch C, Armbruster U, Delannoy E, Gualberto JM, et al. Chloroplast ribonucleoprotein CP31A is required for editing and stability of specific chloroplast mRNAs. Proc Natl Acad Sci USA. 2009;106:6002–7. doi: 10.1073/pnas.0808529106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Stuart KD, Schnaufer A, Ernst NL, Panigrahi AK. Complex management: RNA editing in trypanosomes. Trends Biochem Sci. 2005;30:97–105. doi: 10.1016/j.tibs.2004.12.006. [DOI] [PubMed] [Google Scholar]
- 76.Shikanai T. Cyclic electron transport around photosystem I: genetic approaches. Annu Rev Plant Biol. 2007;58:199–217. doi: 10.1146/annurev.arplant.58.091406.110525. [DOI] [PubMed] [Google Scholar]
- 77.Munekage Y, Hashimoto M, Miyake C, Tomizawa K, Endo T, Tasaka M, et al. Cyclic electron flow around photosystem I is essential for photosynthesis. Nature. 2004;429:579–82. doi: 10.1038/nature02598. [DOI] [PubMed] [Google Scholar]
- 78.Okegawa Y, Kobayashi Y, Shikanai T. Physiological links among alternative electron transport pathways that reduce and oxidize plastoquinone in Arabidopsis. Plant J. 2010;63:458–68. doi: 10.1111/j.1365-313X.2010.04252.x. [DOI] [PubMed] [Google Scholar]
- 79.Nakamura T, Sugita M. A conserved DYW domain of the pentatricopeptide repeat protein possesses a novel endoribonuclease activity. FEBS Lett. 2008;582:4163–8. doi: 10.1016/j.febslet.2008.11.017. [DOI] [PubMed] [Google Scholar]
- 80.Jonietz C, Forner J, Hildebrandt T, Binder S. RNA PROCESSING FACTOR3 is crucial for the accumulation of mature ccmC transcripts in mitochondria of Arabidopsis accession Columbia. Plant Physiol. 2011;157:1430–9. doi: 10.1104/pp.111.181552. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Hauler A, Jonietz C, Stoll B, Stoll K, Braun HP, Binder S. RNA PROCESSING FACTOR 5 is required for efficient 5′ cleavage at a processing site conserved in RNAs of three different mitochondrial genes in Arabidopsis thaliana. Plant J. 2013;74:593–604. doi: 10.1111/tpj.12143. [DOI] [PubMed] [Google Scholar]
- 82.Shikanai T, Obokata J. Machinery of RNA editing in plant organelles. In: RNA and DNA editing: Molecular mechanisms and their integration into biological system. 2008:99-119. [Google Scholar]
- 83.Ichinose M, Tasaki E, Sugita C, Sugita M. A PPR-DYW protein is required for splicing of a group II intron of cox1 pre-mRNA in Physcomitrella patens. Plant J. 2012;70:271–8. doi: 10.1111/j.1365-313X.2011.04869.x. [DOI] [PubMed] [Google Scholar]
- 84.Delannoy E, Stanley WA, Bond CS, Small ID. Pentatricopeptide repeat (PPR) proteins as sequence-specificity factors in post-transcriptional processes in organelles. Biochem Soc Trans. 2007;35:1643–7. doi: 10.1042/BST0351643. [DOI] [PubMed] [Google Scholar]
- 85.Gobert A, Pinker F, Fuchsbauer O, Gutmann B, Boutin R, Roblin P, et al. Structural insights into protein-only RNase P complexed with tRNA. Nat Commun. 2013;4:1353. doi: 10.1038/ncomms2358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Howard MJ, Lim WH, Fierke CA, Koutmos M. Mitochondrial ribonuclease P structure provides insight into the evolution of catalytic strategies for precursor-tRNA 5′ processing. Proc Natl Acad Sci USA. 2012;109:16149–54. doi: 10.1073/pnas.1209062109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Boch J, Scholze H, Schornack S, Landgraf A, Hahn S, Kay S, et al. Breaking the code of DNA binding specificity of TAL-type III effectors. Science. 2009;326:1509–12. doi: 10.1126/science.1178811. [DOI] [PubMed] [Google Scholar]
- 88.Moscou MJ, Bogdanove AJ. A simple cipher governs DNA recognition by TAL effectors. Science. 2009;326:1501. doi: 10.1126/science.1178817. [DOI] [PubMed] [Google Scholar]
- 89.Chase CD. Cytoplasmic male sterility: a window to the world of plant mitochondrial-nuclear interactions. Trends Genet. 2007;23:81–90. doi: 10.1016/j.tig.2006.12.004. [DOI] [PubMed] [Google Scholar]
- 90.Hanson MR, Bentolila S. Interactions of mitochondrial and nuclear genes that affect male gametophyte development. Plant Cell. 2004;16(Suppl):S154–69. doi: 10.1105/tpc.015966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Kobayashi K, Kawabata M, Hisano K, Kazama T, Matsuoka K, Sugita M, et al. Identification and characterization of the RNA binding surface of the pentatricopeptide repeat protein. Nucleic Acids Res. 2012;40:2712–23. doi: 10.1093/nar/gkr1084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Yagi Y, Hayashi S, Kobayashi K, Hirayama T, Nakamura T. Elucidation of the RNA recognition code for pentatricopeptide repeat proteins involved in organelle RNA editing in plants. PLoS One. 2013;8:e57286. doi: 10.1371/journal.pone.0057286. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Härtel B, Zehrmann A, Verbitskiy D, van der Merwe JA, Brennicke A, Takenaka M. MEF10 is required for RNA editing at nad2-842 in mitochondria of Arabidopsis thaliana and interacts with MORF8. Plant Mol Biol. 2013;81:337–46. doi: 10.1007/s11103-012-0003-2. [DOI] [PubMed] [Google Scholar]
- 94.Hanaoka M, Kanamaru K, Fujiwara M, Takahashi H, Tanaka K. Glutamyl-tRNA mediates a switch in RNA polymerase use during chloroplast biogenesis. EMBO Rep. 2005;6:545–50. doi: 10.1038/sj.embor.7400411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Barkan A, Goldschmidt-Clermont M. Participation of nuclear genes in chloroplast gene expression. Biochimie. 2000;82:559–72. doi: 10.1016/S0300-9084(00)00602-7. [DOI] [PubMed] [Google Scholar]
- 96.Maier UG, Bozarth A, Funk HT, Zauner S, Rensing SA, Schmitz-Linneweber C, et al. Complex chloroplast RNA metabolism: just debugging the genetic programme? BMC Biol. 2008;6:36. doi: 10.1186/1741-7007-6-36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Boulouis A, Raynaud C, Bujaldon S, Aznar A, Wollman FA, Choquet Y. The nucleus-encoded trans-acting factor MCA1 plays a critical role in the regulation of cytochrome f synthesis in Chlamydomonas chloroplasts. Plant Cell. 2011;23:333–49. doi: 10.1105/tpc.110.078170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Patel M, Berry JO. Rubisco gene expression in C4 plants. J Exp Bot. 2008;59:1625–34. doi: 10.1093/jxb/erm368. [DOI] [PubMed] [Google Scholar]
- 99.Sheen J. C4 Gene Expression. Annu Rev Plant Physiol Plant Mol Biol. 1999;50:187–217. doi: 10.1146/annurev.arplant.50.1.187. [DOI] [PubMed] [Google Scholar]
- 100.Johnson X, Wostrikoff K, Finazzi G, Kuras R, Schwarz C, Bujaldon S, et al. MRL1, a conserved pentatricopeptide repeat protein, is required for stabilization of rbcL mRNA in Chlamydomonas and Arabidopsis. Plant Cell. 2010;22:234–48. doi: 10.1105/tpc.109.066266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Li P, Ponnala L, Gandotra N, Wang L, Si Y, Tausta SL, et al. The developmental dynamics of the maize leaf transcriptome. Nat Genet. 2010;42:1060–7. doi: 10.1038/ng.703. [DOI] [PubMed] [Google Scholar]
- 102.Rochaix JD. Role of thylakoid protein kinases in photosynthetic acclimation. FEBS Lett. 2007;581:2768–75. doi: 10.1016/j.febslet.2007.04.038. [DOI] [PubMed] [Google Scholar]
- 103.Pfannschmidt T, Nilsson A, Allen JF. Photosynthetic control of chloroplast gene expression. Nature. 1999;397:625–8. doi: 10.1038/17624. [DOI] [Google Scholar]
- 104.Rochaix JD. Redox regulation of thylakoid protein kinases and photosynthetic gene expression. Antioxid Redox Signal. 2013;18:2184–201. doi: 10.1089/ars.2012.5110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Bellafiore S, Barneche F, Peltier G, Rochaix JD. State transitions and light adaptation require chloroplast thylakoid protein kinase STN7. Nature. 2005;433:892–5. doi: 10.1038/nature03286. [DOI] [PubMed] [Google Scholar]
- 106.Tikkanen M, Grieco M, Kangasjärvi S, Aro EM. Thylakoid protein phosphorylation in higher plant chloroplasts optimizes electron transfer under fluctuating light. Plant Physiol. 2010;152:723–35. doi: 10.1104/pp.109.150250. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Grieco M, Tikkanen M, Paakkarinen V, Kangasjärvi S, Aro EM. Steady-state phosphorylation of light-harvesting complex II proteins preserves photosystem I under fluctuating white light. Plant Physiol. 2012;160:1896–910. doi: 10.1104/pp.112.206466. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Rodermel S, Haley J, Jiang CZ, Tsai CH, Bogorad L. A mechanism for intergenomic integration: abundance of ribulose bisphosphate carboxylase small-subunit protein influences the translation of the large-subunit mRNA. Proc Natl Acad Sci USA. 1996;93:3881–5. doi: 10.1073/pnas.93.9.3881. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Wostrikoff K, Stern D. Rubisco large-subunit translation is autoregulated in response to its assembly state in tobacco chloroplasts. Proc Natl Acad Sci USA. 2007;104:6466–71. doi: 10.1073/pnas.0610586104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Makino A, Shimada T, Takumi S, Kaneko K, Matsuoka M, Shimamoto K, et al. Does decrease in ribulose-1,5-bisphosphate carboxylase by antisense rbcS lead to a higher N-use efficiency of photosynthesis under conditions of saturating CO2 and light in rice plants? Plant Physiol. 1997;114:483–91. doi: 10.1104/pp.114.2.483. [DOI] [PMC free article] [PubMed] [Google Scholar]


