Abstract
We have isolated two genes from yeast encoding 3-hydroxy-3-methylglutaryl-coenzyme A reductase [hydroxymethylglutaryl-coenzyme A reductase (NADPH); HMG-CoA reductase; EC 1.1.1.34], the rate-limiting enzyme of sterol biosynthesis. These genes, HMG1 and HMG2, were identified by hybridization to a cDNA clone encoding hamster HMG-CoA reductase. DNA sequence analysis reveals homology between the amino acid sequence of the proteins encoded by the two yeast genes and the carboxyl-terminal half of the hamster protein. Cells containing mutant alleles of both HMG1 and HMG2 are unable to undergo spore germination and vegetative growth. However, cells containing a mutant allele of either HMG1 or HMG2 are viable but are more sensitive to compactin, a competitive inhibitor of HMG-CoA reductase, than are wild-type cells. Assays of HMG-CoA reductase activity in extracts from hmg1- and hmg2- mutants indicate that HMG1 contributes at least 83% of the activity found in wild-type cells.
Full text
PDF




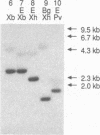
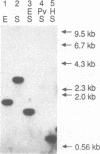
Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Barnes G., Hansen W. J., Holcomb C. L., Rine J. Asparagine-linked glycosylation in Saccharomyces cerevisiae: genetic analysis of an early step. Mol Cell Biol. 1984 Nov;4(11):2381–2388. doi: 10.1128/mcb.4.11.2381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bolivar F., Rodriguez R. L., Greene P. J., Betlach M. C., Heyneker H. L., Boyer H. W., Crosa J. H., Falkow S. Construction and characterization of new cloning vehicles. II. A multipurpose cloning system. Gene. 1977;2(2):95–113. [PubMed] [Google Scholar]
- Bradford M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976 May 7;72:248–254. doi: 10.1016/0003-2697(76)90527-3. [DOI] [PubMed] [Google Scholar]
- Brown D. A., Simoni R. D. Biogenesis of 3-hydroxy-3-methylglutaryl-coenzyme A reductase, an integral glycoprotein of the endoplasmic reticulum. Proc Natl Acad Sci U S A. 1984 Mar;81(6):1674–1678. doi: 10.1073/pnas.81.6.1674. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown M. S., Goldstein J. L. Multivalent feedback regulation of HMG CoA reductase, a control mechanism coordinating isoprenoid synthesis and cell growth. J Lipid Res. 1980 Jul;21(5):505–517. [PubMed] [Google Scholar]
- Chang T. Y., Limanek J. S. Regulation of cytosolic acetoacetyl coenzyme A thiolase, 3-hydroxy-3-methylglutaryl coenzyme A synthase, 3-hydroxy-3-methylglutaryl coenzyme A reductase, and mevalonate kinase by low density lipoprotein and by 25-hydroxycholesterol in Chinese hamster ovary cells. J Biol Chem. 1980 Aug 25;255(16):7787–7795. [PubMed] [Google Scholar]
- Chin D. J., Gil G., Russell D. W., Liscum L., Luskey K. L., Basu S. K., Okayama H., Berg P., Goldstein J. L., Brown M. S. Nucleotide sequence of 3-hydroxy-3-methyl-glutaryl coenzyme A reductase, a glycoprotein of endoplasmic reticulum. Nature. 1984 Apr 12;308(5960):613–617. doi: 10.1038/308613a0. [DOI] [PubMed] [Google Scholar]
- Dente L., Cesareni G., Cortese R. pEMBL: a new family of single stranded plasmids. Nucleic Acids Res. 1983 Mar 25;11(6):1645–1655. doi: 10.1093/nar/11.6.1645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Edwards P. A., Lan S. F., Tanaka R. D., Fogelman A. M. Mevalonolactone inhibits the rate of synthesis and enhances the rate of degradation of 3-hydroxy-3-methylglutaryl coenzyme A reductase in rat hepatocytes. J Biol Chem. 1983 Jun 25;258(12):7272–7275. [PubMed] [Google Scholar]
- Endo A., Kuroda M., Tanzawa K. Competitive inhibition of 3-hydroxy-3-methylglutaryl coenzyme A reductase by ML-236A and ML-236B fungal metabolites, having hypocholesterolemic activity. FEBS Lett. 1976 Dec 31;72(2):323–326. doi: 10.1016/0014-5793(76)80996-9. [DOI] [PubMed] [Google Scholar]
- Gibson D. M., Parker R. A., Stewart C. S., Evenson K. J. Short-term regulation of hydroxymethylglutaryl coenzyme A reductase by reversible phosphorylation: modulation of reductase phosphatase in rat hepatocytes. Adv Enzyme Regul. 1982;20:263–283. doi: 10.1016/0065-2571(82)90020-6. [DOI] [PubMed] [Google Scholar]
- Gil G., Faust J. R., Chin D. J., Goldstein J. L., Brown M. S. Membrane-bound domain of HMG CoA reductase is required for sterol-enhanced degradation of the enzyme. Cell. 1985 May;41(1):249–258. doi: 10.1016/0092-8674(85)90078-9. [DOI] [PubMed] [Google Scholar]
- Hanahan D. Studies on transformation of Escherichia coli with plasmids. J Mol Biol. 1983 Jun 5;166(4):557–580. doi: 10.1016/s0022-2836(83)80284-8. [DOI] [PubMed] [Google Scholar]
- Harwood H. J., Jr, Brandt K. G., Rodwell V. W. Allosteric activation of rat liver cytosolic 3-hydroxy-3-methylglutaryl coenzyme A reductase kinase by nucleoside diphosphates. J Biol Chem. 1984 Mar 10;259(5):2810–2815. [PubMed] [Google Scholar]
- Hinnen A., Hicks J. B., Fink G. R. Transformation of yeast. Proc Natl Acad Sci U S A. 1978 Apr;75(4):1929–1933. doi: 10.1073/pnas.75.4.1929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liscum L., Finer-Moore J., Stroud R. M., Luskey K. L., Brown M. S., Goldstein J. L. Domain structure of 3-hydroxy-3-methylglutaryl coenzyme A reductase, a glycoprotein of the endoplasmic reticulum. J Biol Chem. 1985 Jan 10;260(1):522–530. [PubMed] [Google Scholar]
- Luskey K. L., Faust J. R., Chin D. J., Brown M. S., Goldstein J. L. Amplification of the gene for 3-hydroxy-3-methylglutaryl coenzyme A reductase, but not for the 53-kDa protein, in UT-1 cells. J Biol Chem. 1983 Jul 10;258(13):8462–8469. [PubMed] [Google Scholar]
- Messing J., Vieira J. A new pair of M13 vectors for selecting either DNA strand of double-digest restriction fragments. Gene. 1982 Oct;19(3):269–276. doi: 10.1016/0378-1119(82)90016-6. [DOI] [PubMed] [Google Scholar]
- Mosley S. T., Brown M. S., Anderson R. G., Goldstein J. L. Mutant clone of Chinese hamster ovary cells lacking 3-hydroxy-3 -methylglutaryl coenzyme A reductase. J Biol Chem. 1983 Nov 25;258(22):13875–13881. [PubMed] [Google Scholar]
- Norrander J., Kempe T., Messing J. Construction of improved M13 vectors using oligodeoxynucleotide-directed mutagenesis. Gene. 1983 Dec;26(1):101–106. doi: 10.1016/0378-1119(83)90040-9. [DOI] [PubMed] [Google Scholar]
- Payne G. S., Schekman R. A test of clathrin function in protein secretion and cell growth. Science. 1985 Nov 29;230(4729):1009–1014. doi: 10.1126/science.2865811. [DOI] [PubMed] [Google Scholar]
- Reynolds G. A., Basu S. K., Osborne T. F., Chin D. J., Gil G., Brown M. S., Goldstein J. L., Luskey K. L. HMG CoA reductase: a negatively regulated gene with unusual promoter and 5' untranslated regions. Cell. 1984 Aug;38(1):275–285. doi: 10.1016/0092-8674(84)90549-x. [DOI] [PubMed] [Google Scholar]
- Rine J., Hansen W., Hardeman E., Davis R. W. Targeted selection of recombinant clones through gene dosage effects. Proc Natl Acad Sci U S A. 1983 Nov;80(22):6750–6754. doi: 10.1073/pnas.80.22.6750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roitelman J., Shechter I. Regulation of rat liver 3-hydroxy-3-methylglutaryl coenzyme A reductase. Evidence for thiol-dependent allosteric modulation of enzyme activity. J Biol Chem. 1984 Jan 25;259(2):870–877. [PubMed] [Google Scholar]
- Rothstein R. J. One-step gene disruption in yeast. Methods Enzymol. 1983;101:202–211. doi: 10.1016/0076-6879(83)01015-0. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Servouse M., Mons N., Baillargeat J. L., Karst F. Isolation and characterization of yeast mutants blocked in mevalonic acid formation. Biochem Biophys Res Commun. 1984 Sep 17;123(2):424–430. doi: 10.1016/0006-291x(84)90247-x. [DOI] [PubMed] [Google Scholar]
- Skalnik D. G., Simoni R. D. The nucleotide sequence of Syrian hamster HMG-CoA reductase cDNA. DNA. 1985 Dec;4(6):439–444. doi: 10.1089/dna.1985.4.439. [DOI] [PubMed] [Google Scholar]
- Struhl K., Davis R. W. A physical, genetic and transcriptional map of the cloned his3 gene region of Saccharomyces cerevisiae. J Mol Biol. 1980 Jan 25;136(3):309–332. doi: 10.1016/0022-2836(80)90376-9. [DOI] [PubMed] [Google Scholar]
- Yanisch-Perron C., Vieira J., Messing J. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene. 1985;33(1):103–119. doi: 10.1016/0378-1119(85)90120-9. [DOI] [PubMed] [Google Scholar]
- Young R. A., Davis R. W. Efficient isolation of genes by using antibody probes. Proc Natl Acad Sci U S A. 1983 Mar;80(5):1194–1198. doi: 10.1073/pnas.80.5.1194. [DOI] [PMC free article] [PubMed] [Google Scholar]