Abstract
Brain metastatic breast cancer (BMBC) is uniformly fatal and increasing in frequency. Despite its devastating outcome, mechanisms causing BMBC remain largely unknown. The mechanisms that implicate circulating tumor cells (CTCs) in metastatic disease, notably in BMBC, remain elusive. Here we characterize CTCs isolated from peripheral blood mononuclear cells (PBMCs) of patients with breast cancer, and also develop CTC lines from three of these patients. In epithelial cell adhesion molecule (EpCAM)–negative CTCs, we identified a potential signature of brain metastasis comprising “brain metastasis selected markers (BMSM)” HER2+/EGFR+/HPSE+/Notch1+. These CTCs—which are not captured by the CellSearch platform because of their EpCAM negativity—were analyzed for cell invasiveness and metastatic competency in vivo. CTC lines expressing the BMSM signature were highly invasive and capable of generating brain and lung metastases when xenografted in nude mice. Notably, increased brain metastatic capabilities, frequency, and quantitation were detected in EpCAM− CTCs overexpressing the BMSM signature. The presence of proteins of the BMSM CTC signature was also detected in the metastatic lesions of animals. Collectively, we provide evidence of isolation, characterization, and long-term culture of human breast cancer CTCs, leading to the description of a BMSM protein signature that is suggestive of CTC metastatic competency to the brain.
Introduction
The overwhelming majority of cancer deaths are due to metastasis (1). Brain metastatic breast cancer (BMBC) occurs in approximately 20% of all breast cancer cases, is uniformly fatal, and may be increasing in frequency in patients with a particular subtype. Occult brain metastasis are exceedingly common at autopsy (2). Circulating tumor cells (CTCs) represent the primary cause of intractable metastatic disease and are considered essential for metastasis formation (3-5). However, characterization of CTCs that induce metastasis remains elusive because platforms that capture CTCs are not comprehensive, owing to the phenotypic heterogeneity of CTCs. For example, the Veridex CellSearch platform—the only test approved by the U.S. Food and Drug Administration (FDA)—relies on the use of antibodies targeting the epithelial cell adhesion molecule (EpCAM); thus, it is only capable of capturing EpCAM-positive CTCs, but not CTCs that are EpCAM-undetectable or EpCAM-negative (both termed “EpCAM-” here). CellSearch is also unable to capture CTCs in 30 to 35% of metastatic breast cancer patients (3-5), and over 60% of patients with BMBC who have undetectable CTCs by CellSearch analyses (5). Therefore, new approaches to identify and characterize EpCAM− CTCs in breast cancer patients are needed.
EpCAM− CTCs are crucial to understanding cancer metastasis, with the ultimate goal of developing treatments to prolong patient survival. CTCs derived from breast cancer patients with clinically detectable BMBC rarely express EpCAM (5), and EpCAM− CTCs have been shown to positively correlate with the presence of brain metastases in a large cohort of patients (5). The objective of the current study was to detect, isolate, and characterize EpCAM− CTCs present in the blood of breast cancer patients. We were particularly interested in investigating biomarkers expressed by these CTCs and their abilities to metastasize to the brain. Previously described methods for CTC detection, isolation, and enrichment are based on procedures involving density, immunomagnetic, size-exclusion and/or flow manipulation in microfluidic devices (4-8). However, similar to CellSearch, most of these approaches are limited by the heterogeneous nature of CTCs, often “missing” the EpCAM− CTCs (9, 10). Furthermore, the fraction of CTCs that extravasate and are able to generate distant metastases may have lost EpCAM expression and are believed to have undergone the epithelial-mesenchymal transition (EMT), which results in the downregulation of epithelial cell markers, such as E-cadherin, claudins, cytokeratins, and EpCAM (10-13). Several groups have also reported that CTCs express stem cell and/or EMT-associated markers (8, 9, 13); however, it is unclear whether CTCs that no longer express EpCAM are metastasis-competent. The goal of the field, and scope of this work, is to move beyond CTC detection and enumeration, to demonstrate that human CTCs can be isolated, cause new tumors, and be characterized as organ-specific (brain) homing CTC subsets.
Although tumors can be formed in animals by cells from the peripheral blood of patients (14), and murine CTCs have been successfully cultured (15, 16), long-term culture of isolated human CTCs has not been reported. In the present study, we isolated EpCAM− CTCs from patients with metastatic breast cancer, developed CTC lines from 3 of these patients, and then characterized these EpCAM− CTCs for metastatic competence by selecting for the protein signature HER2+/EGFR+/HPSE+/Notch1+ (the “brain metastasis selected markers (BMSM) signature”). We show that CTCs selected for this signature possess high propensities to metastasize to the brain and lungs once injected in immunodeficient animals, whereas parental CTCs metastasize to lung. This work fosters a new understanding of BMBC biology and the role of CTCs in metastasis, and has clinical relevance by suggesting strategies for therapeutic intervention to suppress and/or prevent CTCs from colonizing in distant organs, notably the brain.
RESULTS
Cancer-associated circulating cells with EGFR amplification and overexpression of HPSE and ALDH1
To interrogate the EpCAM− CTC subpopulation, we employed the FICTION platform (BioView Ltd), which combines protein detection by immunofluorescence (IF) with gene amplification by fluorescence in situ hybridization (FISH) analyses. Automated quantification of the signal, according to specific IF/FISH patterns, is then achieved by scanning IF/FISH-analyzed slides using the BioView system. Epidermal growth factor receptor (EGFR) is present on cancer cells, and is a high-risk predictor for BMBC (2, 22). Accordingly, we examined peripheral blood mononuclear cells (PBMCs) isolated from 38 patients who had been diagnosed with breast cancer (Table 1) for the presence of EGFR amplification compared to normal PBMCs. Aberrant EGFR amplification was observed in 8 patients’ PBMCs (21% of cases) with normal ploidy (CEP10/q10 diploid content) (Fig. 1A). Heparanase (HPSE), a potent tumorigenic, angiogenic, and pro-metastatic molecule, was also overexpressed in these cells in the same 8 patients (Fig. 1B) (24-26). Further, 0.0025% of cancer-associated circulating cells (CACCs) analyzed from PBMCs of these 8 patients and possessing EGFR amplification were HPSE+ and expressed aldhehyde dehydrogenase 1 (ALDH1), a tumor-initiating stem cell marker (Fig. 1, C and D) (18, 36). Cells that are ALDH1-negative cannot form tumors (18); therefore, cells that were ALDH1− were not considered to be metastatic CTCs. Accordingly, we only considered EGFR-overexpressing/HPSE+/ALDH1+ CACCs as potentially metastatic CTCs.
Table 1.
CTC selection and culture. Peripheral blood samples from a population of 38 patients with invasive breast cancer were initially analyzed by CellSearch (EpCAM+ CTCs). Undetectable CTC levels (CTC = 0) were found in 15 patients’ samples. Analyses of these patient PMBCs showed 8 patients to have EGFR gain/ALDH+/HPSE+ CACCs. PMBCs isolated from these patients underwent multi-parametric FACS analysis (CD45–/ALDH1+/EpCAM selection). Successful FACS selection for potential CTCs was achieved in 3 patients. Only EpCAM– CACCs could survive and grow in culture to establish CTC lines CTC-1, CTC-2, and CTC-3 (in bold).
| Patient ID | CTC detection by CellSearch | EGFR gain/ALDH1+/HPSE+ | CTC capture by FACS and viable culture | |
|---|---|---|---|---|
| n = 38 | n = 15 | n = 23 | n = 8 | n = 8 |
| 1 | Undetectable | + | Nonviable CTC | |
| 2 | + | |||
| 3 | + | |||
| 4 | + | |||
| 5 | Undetectable | |||
| 6 | Undetectable | + | Nonviable CTC | |
| 7 | + | |||
| 8 | + | |||
| 9 | Undetectable | |||
| 10 | Undetectable | + | Nonviable CTC | |
| 11 | Undetectable | |||
| 12 | + | |||
| 13 | + | |||
| 14 | Undetectable | |||
| 15 | + | |||
| 16 | Undetectable | + | Viable CTC (CTC-1) | |
| 17 | + | |||
| 18 | Undetectable | |||
| 19 | + | |||
| 20 | Undetectable | + | Nonviable CTC | |
| 21 | + | |||
| 22 | Undetectable | + | Viable CTC (CTC-2) | |
| 23 | Undetectable | + | Viable CTC (CTC-3) | |
| 24 | + | |||
| 25 | + | |||
| 26 | + | |||
| 27 | Undetectable | |||
| 28 | + | |||
| 29 | + | |||
| 30 | + | |||
| 31 | + | |||
| 32 | Undetectable | + | Nonviable CTC | |
| 33 | + | |||
| 34 | + | |||
| 35 | Undetectable | |||
| 36 | + | |||
| 37 | + | |||
| 38 | + | |||
Fig. 1. EGFR amplification and nuclear HPSE expression in PBMCs from metastatic breast cancer patients.
(A) Representative image of FISH analyses for EGFR amplification in PBMCs isolated from BMBC patients. EGFR amplification (white arrow) was compared to CEP10/10q signal for ploidy content. (B) Representative FISH and IF analyses of nuclei of CACCs showing EGFR and heparanase (HPSE) expression. EGFR 7p12 was also overlayed with CEP7 to show ploidy content. DAPI staining is in blue. (C) Representative images of IF analyses showing simultaneous expression of HPSE and aldehyde dehydrogenase 1 (ALDH1) in nucleated PBMCs from the blood of patients (n = 8), in PBMCs from patients without breast cancer (n = 8), and in the MDA-MB-231BR cell line. MDA-MB-231BR cells and PBMCs were used as ALDH1-positive and - negative controls, respectively. All scale bars, 20 μm. (D) EGFR-amplified PBMCs isolated from BMBC patients (n = 3) were analyzed for ALDH1 and HPSE expression. The system randomly scanned ~5.0 × 103 cells for each marker per slide sample. Data are averages ± S.D.
Selection and isolation of metastatic CTCs
We hypothesized that brain-homing CTCs are present within the PMBC subset EGFR+/HPSE+/ALDH1+ described above. To capture CTCs, we developed strategies to isolate EpCAM-negative neoplastic cells within the EGFR+/HPSE+/ALDH1+ PBMC population of the 8 breast cancer patients (Table 1). The initial selection of CTCs relied on ALDH1. Cells that are ALDH1-negative or CD44+/CD24-/lin- cannot form tumors when transplanted into the mammary fat pad of nude mice (18, 36); as such, they were used as negative selection criteria. PBMCs were isolated from blood, then multiparametric FACS was applied to select CD45-/ALDH1+ cells that were both EpCAM+ and EpCAM− from breast cancer patients’ blood (n = 8), but not from PBMCs of healthy donors of the same race and age (n = 8) (Table 1).
EpCAM+/ALDH1+/CD45- or EpCAM-/ALDH1+/CD45- circulating cell subsets were obtained from only 3 out of the 8 patients because this is the subset from which we were able to successfully apply FACS CTC selection (Table 1; Table 2). These CACCs were cultured for further characterization. Captured cells were used to surviving in blood under suspension, therefore we provided a period of transition using stem cell culture medium for the first week, followed by a medium for epithelial cells (Methods). Cells were monitored for survival and growth over 28 days, and colonies were initiated starting from a single cell (fig. S1). Percentages of EpCAM+ and EpCAM− CACCs from the original CD45-/ALDH1+ population were 1.2 × 10-2 and 5.3 × 10-2 respectively (table S1). Further, long-term culture of EpCAM+ CACCs could not be established since they did not survive beyond 14 days of culturing. However, thirteen colonies of EpCAM− CACCs were observed in the culture from Patient 1 by day 21 of cell culture, 7 from Patient 2, and 11 from Patient 3. CTC lines were established as CTC-1, CTC-2, and CTC-3 for Patients 1 to 3 (Supplementary Methods), respectively (table S1).
Table 2. Increased frequency of metastasis by BMSM CTCs in a xenograft model.
Comparative analysis of metastatic capabilities of primary CTCs versus BMSM CTCs in xenografts of brain and lung tumors. An equal number of cells (0.5 × 106 cells) were injected into each mouse. PBMCs from Patient 1 were used as a negative control. Data are averages from 10 mice per cell line (n = 70 total).
| Cells injected | Pathology | Tumor incidence in lung (%) | P-value for lung | Tumor incidence in brain (%) | P-value for brain |
|---|---|---|---|---|---|
| CTC-1 | Spindle perivascular Epitheloid micro-metastases | 80 | -- | 20 | -- |
| CTC-2 | Spindle perivascular | 60 | -- | 0 | -- |
| CTC-3 | Spindle perivascular | 60 | -- | 0 | -- |
| BMSM CTC-1 | Spindle perivascular Epitheloid micro-metastases | 100 | <0.001 (compared to CTC-1) | 80 | <0.001 (compared to CTC-1) |
| BMSM CTC-2 | Spindle perivascular Epitheloid micro-metastases | 80 | <0.001 (compared to CTC-2) | 60 | <0.001 (compared to CTC-2) |
| BMSM CTC-3 | Spindle perivascular | 80 | <0.001 (compared to CTC-3) | 60 | <0.001 (compared to CTC-3) |
| PBMC | No tumor | 0 | -- | 0 | -- |
Characterization of CTC lines
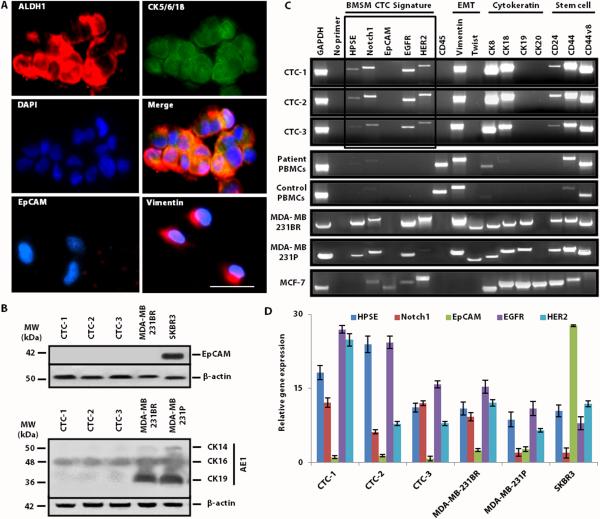
Surviving CTCs (CTC-1, CTC-2, and CTC-3 lines generated from respective patients) were evaluated for expression of ALDH1, EpCAM, and tumor cell markers cytokeratin (CK) 5, CK6, and CK18. Both ALDH1 and CK5/6/18 were expressed in cultured EpCAM− cell lines, as shown by immunofluorescence staining (Fig. 2A). CK16 was also detected by Western blot and its expression levels were similar to human breast cancer cell lines MDA-MB-231 parental and the brain metastatic MDA-MB-231BR variant (Fig. 2B). However, different levels of CK expression were observed between CTCs and MDA-MB-231 cell lines (Fig. 2B), suggesting that CTCs have metastatic breast cancer features and share markers with MDA-MB-231 cells.
Fig 2. EpCAM– CTC characterization and culture from three patients.
(A) Immunofluorescence staining of potential CTCs (EpCAM–/ALDH1+/CD45–) for ALDH1, CK5/6/18, and vimentin expression. Merge panel consists of ALDH1/CK/DAPI–positive IF staining. Scale bar, 50 μm. (B) Western blot for EpCAM and CK14/16/19 in CTC lines. The brain-metastatic MDA-MB-231BR and non-brain-metastatic SKBR3 cell lines were used as positive and negative controls, respectively. β-Actin was the loading control. (C) RT-PCR analyses of CTCs. Selected genes were classified into four groups: BMSM signature, EMT, cytokeratin, and stemness. MDA-MB-231BR (HER2-transfected), MDA-MB-231 parental, and MCF-7 cell lines were used as controls as brain metastatic, poorly brain metastatic, and non-metastatic breast cancer cell lines, respectively. PBMCs isolated from patients with (patient) or without (control) breast cancer were used as negative controls for the BMSM CTC signature. GAPDH was the loading control. Further, CTCs did not express the mesenchymal stem cell (MSC) marker triplet (fig. S2). (D) Real-time PCR analysis of the BMSM gene expression. All cell lines were analyzed and normalized to the expression levels of GAPDH per cell line. Data are means ± SEM (n = 3, four independent experiments). Data displayed in fig. 2C (RT-PCR) were not quantified because the best quantification for markers of the BMSM CTC signature was achieved employing real-time PCR. However, we have quantified the RT-PCR data for CTCs at passage 20 (fig. S3).
We hypothesized that a unique gene signature for metastasis was present in CTCs derived from the 3 breast cancer patients. We therefore developed a specific semi-quantitative RT-PCR assay (table S2) for the parallel detection of multiple genetic biomarkers—the BMSM signature—which was selected because of their proven relevance to BMBC onset (3, 4, 21-28): HPSE+/Notch1+/EGFR+/HER2+ (Fig. 2C). FACS-isolated CTCs from a symptomatic BMBC patient and two non-symptomatic breast cancer patients (Table 1) resulting in the generation of CTC lines (CTC-1, CTC-2, CTC-3) were analyzed by RT-PCR assays. In addition, we used PBMCs collected from each patient as an internal control that did not undergo FACS selection for CTCs. PBMCs were also isolated from healthy subjects (n = 8) of the same race, similar age, and medical and socioeconomic backgrounds as negative controls. Further, CTC lines were analyzed by CellSearch by performing dose-dependent experiments spiking cells into aliquots of blood from healthy donors. Cells could not be captured by CellSearch, confirming their EpCAM– status and the validity of FACS procedures (table S3).
Next, we examined other known markers of tumor-initiating cells, such as CD24low/CD44high—a typical pattern of tumor-initiating cells—and CD44 variant 8 (CD44v8). CD24low/CD44high was observed in cultured CTCs but not in PBMC controls (Fig. 2C). The average ratio of CD24low/CD44high expression in these CTC lines was 0.7±0.000873 (SD; n = 9 independent RT-PCR analyses). Markers of potential epithelial– mesenchymal transition (EMT), such as vimentin and Twist, were assessed. High vimentin expression was observed in cultured CTCs whereas Twist expression was negligible (Fig. 2C) (41). We observed higher levels of CK8 and CK18 transcripts in cultured CTCs versus PBMCs (Fig. 2C), whereas CK19 and CK20 genes were below the detection limit in all CTC samples analyzed except for cell lines. This suggests that the absence of these cytokeratins may not affect CTC-mediated breast cancer metastasis, and/or that differential CK transcript levels relate to the particular stage of tumor progression because the expression of several CKs is known to change during metastasis (12). Gene transcripts for markers of neoplasticity were also detected in CTC lines 1 to 3, and included urokinase plasminogen activator receptor (uPAR), mucin 1 (Muc1), and caveolin 1 (fig. S2).
The brain metastatic MDA-MB-231BR breast cancer cell line expressed all genes of the BMSM signature with patterns and levels similar to CTCs (Fig. 2C). Of note, HPSE was nearly absent in the non-metastatic MCF-7 breast cancer cell line (Fig. 2C), substantiating notions that HPSE is a critical player in metastasis and BMBC mechanisms. We next evaluated the specific transcript levels for each BMSM CTC signature gene by real-time PCR, relating them to the known expression of these markers in human breast cancer cells with differing metastatic propensities (Fig. 2D).
To confirm that isolated CTCs did not represent hematopoietic or non-CTC cell populations, we performed RT-PCR analyses for markers of circulating endothelial cells (CD31+), bone marrow hematopoietic cells (CD34+), or mesenchymal stem cells (MSCs) (CD105+/CD73+/CD90+ but negative for CD45, CD34, and CD14) (33). The three CTC lines selected from patients were all negative for these non-CTC marker patterns (fig. S2). PBMCs isolated from healthy control patients were positive for the MSC triplet CD105/CD73/CD90 as well as for CD31 (fig. S3), consistent with previous reports showing the presence of these markers in breast cancer cells (33, 34). Lastly, we showed that CTCs maintained the BMSM signature expression in long-term culture (more than 20 cell passages), along with the other markers of neoplasticity and cell stemness described above (fig. S3). The BMSM CTC signature was validated by analyses for the presence of respective proteins and the absence of EpCAM expression (fig. S4).
CTC genotyping analyses
To confirm the establishment of CTC lines, we performed comprehensive genotyping analyses. First, short tandem repeat (STR) DNA fingerprinting for 16 loci was employed. Data from CTC lines and breast cancer cell lines were compared to the database of the Characterized Cell Line Core of MD Anderson Cancer Center (www.mdanderson.org/education-and-research/resources-for-professionals/scientific-resources/core-facilities-and-services/characterized-cell-line-core-facility/index.html). The STR profiles of CTCs were distinct in eight of the 16 loci analyzed from known DNA fingerprinting profiles of MDA-MB231 parental and MDA-MB-231BR cells (table S4). The three CTC lines possessed STR DNA fingerprinting profiles among CTC lines which were distinct from either established highly brain metastatic MDA-MB-231BR and MDA-MB-435 lines, the poorly brain metastatic (but overall metastatic) MDA-MB-231 parental line, or the non-metastatic (MCF-7) breast cancer cell line (table S4). Further, we could also identify differences in STR DNA fingerprinting profiles among CTC lines, suggesting that these lines are distinct despite sharing the CTC signature. Second, the neoplastic nature of the CTC lines was evaluated by analyzing known somatic mutations (> 200) for hallmark cancer genes: mutations for BRAF (1391G>T), KRAS (38G>A), and TP53 (839G>A) were detected. Mutational analyses for heterozygosity of these mutations aligned well with breast cancer cell lines (table S5) (29).
BMSM CTCs are invasive and metastasize
To assess the biological relevance of the CTC signature in metastasis, parental EpCAM− CTCs were first sorted using anti-Notch1 antibody, relative to control MCF-7 cells. EpCAM-/Notch1+ CTCs were then expanded and sorted again (sequential FACS) using anti-HER2 and anti-EGFR antibodies to obtain EpCAM-/Notch1+ CTCs that overexpressed HER2 and EGFR relative to control SKBR3 cells (Fig. 3A; fig. S5). Captured EpCAMCTCs overexpressing HPSE/Notch1/EGFR/HER2 have been termed “BMSM CTCs”. EGFR, HER2, and Notch1 protein expression in these EpCAM− BMSM CTCs was confirmed by immunofluorescence (Fig. 3B; fig. S6). HPSE activity was high in both cell lysates and supernatants of BMSM CTCs compared with MCF-7 cells (Fig. 3C). There were also elevated levels of activated Notch1 in the nuclei of BMSM CTCs compared to parental CTCs (Fig. 3B) (38).
Fig. 3. Sorting and characterization of EpCAM– CTCs overexpressing Notch1, EGFR, and HER2 (BMSM CTCs).
(A) Sequential cell sorting for EpCAM–/Notch1+ followed by EGFR+/HER2+ to obtain BMSM CTCs. The percentage of positive cells for each sorting is indicated. MCF-7 and SKBR3 cells were used as positive controls for EpCAM and EGFR/HER2, respectively (see also fig. S5). (B) Immunofluorescence staining of cells isolated in (A) for BMSM proteins HPSE, Notch1, EGFR, and HER2. Inset shows ZR75 human breast cancer cells (luminal sub-type) as positive control for EpCAM expression. Scale bar, 100 μm. Fluorescence signal was quantified as percentage of positive cells/microscopy field (8-10 fields/slide). Data are means ± SEM (n = 3)(see also fig. S4 and S6). (C) Fluorescence signal was quantified as percentage of positive cells/microscopy field (8-10 fields/slide). Data are means ± SEM (n = 3) (see also fig. S4 and S6). (D) HPSE activity was measured in BMSM CTCs from (A). Data are means ± SEM (n = 3). Human MCF-7 and MDA MB-231BR cell lines were used as negative and positive controls for HPSE activity, respectively.
To examine whether BMSM CTCs were capable of generating tumors, we injected them either intracardiacally or into the tail vein of immunodeficient mice, and monitored for metastasis formation in the brain and lungs. All BMSM CTCs metastasized to lung and brain (Fig. 4, A and B) by 6 weeks after injection. OKFurther, tumor cell quantification for brain and lung metastasis for both BMSM CTCs and parental CTCs was performed (Fig. 4C). Although all CTC lines (CTC-1, CTC-2, CTC-3) were capable of forming lung metastasis, we found that only CTC-1 promoted relevant brain colonization. However, and further, all BMSM CTC lines possessed higher values for BMBC onset cThese findings were even more compelling after the evaluation of CTC-induced tumor burden. This was assessed in all BMSM CTC lines, and quantified at a single tumor-cell level. Representative images of CTC-induced BMBC and adjacent brain tissue and concurrent tumor burden quantification of lung and brain metastasis are shown in Fig. 4, A and B, and fig. S7.
Fig. 4. BMSM CTCs are invasive and metastasize to brain and lung.
(A) Representative images of BMBC (inserts) induced by BMSM CTCs when injected intracardiacally into mice. BMBC tumor sections underwent analyses for tumor burden by the CRi Vectra Intelligent system which highlights and enumerates single tumor cells (in green). (B) Representative lung images from BMSM CTC–injected mice in (A) are shown. Lung samples were analyzed concurrently with BMBC and under the same conditions. (C) Quantification of only tumor cells for brain- and lung-metastatic breast cancer arising from both BMSM CTCs and parental CTCs (CTC-1, CTC-2, and CTC-3 cell lines). Data are means ± SEM (n = 8 sections/mouse; 10 mice per CTC line). ***P < 0.001, ANOVA. (D) Chemoinvasion of parental CTCs and BMSM CTCs by Matrigel chamber assays. Invasion was quantified and compared with breast cancer cell lines of known invasive capacity. Data are means ± SEM (n = 8-10 fields/cell line, four independent experiments. *P < 0.05, **P < 0.01 by ANOVA. All scale bars, 100 μm.
To evaluate the invasive and metastatic capacities of BMSM CTCs, we performed in vitro chemoinvasion assays and related CTC invasive values to ones using various metastatic and non-metastatic breast cancer cell lines. BMSM CTCs were highly invasive compared to controls (Fig. 4D). The CTC-1 line was the most invasive, with approximately 25% more cells penetrating the Matrigel barrier than the highly invasive MDAMB-231BR cells, likely explained by these cells having been isolated from a “triple negative” breast cancer patient (ER–/PR–/HER2low)—a subtype characterized by an aggressive neoplastic behavior. The invasive capability of BMSM CTC-1 was significantly increased compared to parental CTC-1 cells (P < 0.05, ANOVA). Conversely, CTC-2 and CTC-3 were 68% and 72% less invasive compared to MDA-MB-231BR control cells (P < 0.01, ANOVA). However, the invasive capabilities of BMSM CTC-2 and BMSM CTC-3 were significantly augmented in relation to their parental CTC counterparts (P < 0.01, ANOVA) (Fig. 4D).
CTC-induced brain metastases in xenografts also presented a typical branching pattern of tumor growth (Fig. 5B), which is suggestive of spread along pre-existing blood vessels, a hallmark of BMBC (Fig. 5B). Further, BMBC cell morphology in xenografts closely resembled one from pathological BMBC tissue of Patient 1 whose blood was analyzed and CTCs isolated from (CTC-1 cell line) (Fig. 5B).
Fig. 5. Histological analyses of CTC tumor xenografts and BMSM CTC signature proteins.
This is shown in Figure S8, panel B. Two animals subgroups (n = 5 each) were injected intracardiacally in parallel with either MDA-MB-231BR cells or PBMCs isolated from patients (positive and negative controls for BMBC, respectively). (A) CTCs (parental CTC-1) metastasize to brain in immunocompromised mice (tumor xenografts). Representative images show BMBC macro-metastasis (upper panel) and micro-metastasis (lower panel) surrounded by neuroglial tissue. (B) CTC-induced BMBC displayed a branching pattern indicative of tumor growth along the preexisting vasculature, similar to the clinical presentation of BMBC (upper panel). Further, tumor cell morphology of CTC-induced BMBC closely resembled one of pathologically assessed BMBC of donor patient whose blood was analyzed and CTC isolated from (lower panel). (D) Representative image of BMBC generated from MDA-MB-231BR xenografts (positive control; upper panel). Conversely, no BMBC was observed following the injection of patient PBMCs (negative control, lower panel). (C) Representative image of BMBC from BMSM CTC xenografts and adjacent neuroglial tissue showing an aberrant mitotic figure (yellow arrow, upper panel). Aberrant mitotic figures like starbust mitosis (inset) were detected as hallmark of neoplasticity, and indicated with yellow arrows (lower panel). (E and F) Representative images from murine brain (E) and lung (F) sections from animals injected with BMSM CTC-1 and analyzed for BMSM signature protein expression by IHC (n = 8-10 images/section, 20 sections/mouse)(see also fig. S7 and S8). Graph displays the quantification of staining scores in E of combined BMSM signature protein levels per CTC line per mouse (n = 10 sections per mouse; 10 mice per BMSM CTC line). **p < 0.001. Scale bars, 100 μm.
Of note, mitotic aberrancy is a histopathological hallmark of malignant tumors that correlates with genetic instability. Mitotic activity was prominent in the lung and brain metastases arising from BMSM CTCs (Fig. 5, A to D). Some of the mitoses were morphologically normal with a symmetrical alignment of chromosomes, but a significant number of abnormal mitotic figures were also present including “starburst” mitoses and mitoses in which the chromosomes were misaligned (Fig. 5C).
Lastly, to assess whether the expression of the BMSM CTC signature was present in CTC-induced experimental brain metastasis, we examined the expression of signature BMSM proteins in mouse tumors. Brain tumor tissues displayed the presence of BMSM proteins HPSE, Notch1, EGFR, and HER2, indicating an expression of these marker proteins specifically elevated in brain vs. lung metastasis and suggesting the relevance of the BMSM CTC signature to BMBC (Fig. 5, E and F; fig. S7 and S8).
Discussion
CTCs represent the “seeds” of cancer metastasis and a promising alternative to tumor biopsies to detect, investigate, and monitor solid tumors. Enumerating CTCs has been shown to independently prognosticate tumor progression (3-9, 16). Thus far, only one platform—CellSearch (Veridex)—has been cleared by the FDA for clinical analysis of CTCs, and CellSearch only captures EpCAM+ CTCs. The purpose of this study was to develop a novel approach to identify and characterize EpCAM– CTCs present in breast cancer patients. CTCs are a heterogeneous population, but we hypothesized that there exists a multiplexed biomarker that would identify a brain-metastatic CTC subset. We specifically investigated EpCAM– CTCs isolated from breast cancer patients that could not be captured by the CellSearch platform.
We applied CD45+/ALDH1+ as initial selection markers for possible EpCAM+ and EpCAM– CTCs by sorting PBMCs isolated from breast cancer patients. However, we could not exclude having captured normal stem cells since they also express ALDH1 (18). Accordingly, we aimed to detect markers of MSCs: expression of CD105, CD73, and CD90, but not CD45, CD34, and CD14 (31). Our CTC lines did not meet these criteria (fig. S2). Further, we detected the presence of CD105 and CD73 transcripts in CTCs which aligns well for presence of these markers in breast cancer cells (33, 34).
Surprisingly, CTCs derived from a triple-negative (ER–/PR–/HER2 low) BMBC patient expressed EGFR and HER2 at both mRNA and protein levels. This supports similar findings that HER2 status is altered from the primary tumor to metastatic CTCs (9, 23). Although the precise mechanisms accountable for this HER2 variability are unknown, we provide further evidence that CTCs may develop a HER2 content differential from the primary tumor over the course of neoplastic progression. These observations can have high clinical relevance and support notions that HER2– breast cancers may become HER2+ by one of two mechanisms: the selection of rare HER2-amplified clones that have metastatic potential or an upregulation of HER2 by cells that are not amplified during the metastatic process (39, 40).
Heparanase is another component of the BMSM CTC signature and a potent tumorigenic, angiogenic, and pro-metastatic molecule. High HPSE activity has been reported in cells selected in vivo for highest propensities to colonize the brain, regardless of the cancer type or model system used (25, 40). Recent findings have also demonstrated that the latent form of HPSE can promote cell adhesion, augment EGFR phosphorylation, and act as a signal transducer (26, 32). HPSE overexpression and functionalities may be central to the initial events of brain metastasis and the cross-talk between CTCs and the brain vasculature. Thus, therapeutic disruption of HPSE may prevent tumor cell adhesion, extravasation, and metastasis, particularly to the brain (25).
After identifying a BMSM CTC signature, we isolated and established human EpCAM− BMSM CTC lines to confirm their metastatic competence in vivo. The injection of animals with three putative CTC lines (CTC-1, CTC-2, and CTC-3) resulted in variability towards the BMBC phenotype: only CTC-1 promoted brain metastasis, whereas no brain metastases were observed in CTC-2 and CTC-3 lines. A possible reason might be that the CTC-1 line was derived from a triple-negative BMBC patient, the most aggressive cancer subtype. However, EpCAM– CTCs sorted for Notch1, EGFR, and HER2 (BMSM CTCs; HPSE largely localizes in cytoplasm) increased the incidence of CTC-1 - derived brain and lung metastases (60–100%) when xenografted in mice, as well as the number of brain micro- and macro-metastasis. These results suggest that the BMSM CTC signature could be used to predict which circulating cancer cells are metastatic.
Notably, we could not detect brain metastases in some animals (20%) after the injection of BMSM CTCs (table S6). Reasons for this are unclear, but may involve CTC dormancy and/or the presence of occult brain micro-metastasis (35). The cellular localization of the BMSM CTC signature proteins might also vary among these CTC lines, resulting in a differential metastatic capacity; for example, Notch1 mostly resides in the cell membrane whereas HPSE is localized in the cytoplasm, although we observed Notch1 in the nuclei of some CTCs (Fig. 3B). Another limitation of our study is that the concordance between the BMSM immunofluorescence/immunohistochemistry and FACS data was not 100%. This, too, could be explained by the subcellular localization of the proteins (e.g., Notch1, EGFR, HER2) encompassing the BMSM CTC signature. Previous studies have demonstrated that HPSE can localize in the nuclear/nucleolar fraction, with nucleolar HPSE directly regulating DNA Topoisomerase I activity and enhancing tumor cell proliferation (24). It is unknown whether the localization of HPSE in the nucleolus affects metastasis. Accordingly, the relationship between localization and functionality of the BMSM CTC signature proteins and metastasis in vivo should be investigated further.
A third limitation was finite resources. Although primary cancers for patients 1-3 were assessed for EGFR, HER2, and the presence of other breast cancer markers, we could not verify the expression of the BMSM CTC signature in these primary cancers, nor did we conduct extensive investigations for its presence in animal tissues other than brain or lung. The BMSM CTC signature has the potential to predict which circulating cells are metastatic, but it will need to be validated in a larger number of clinical samples from breast cancer patients.
Identification of a BMSM CTC signature in patient cells and in xenografts is the first step towards improved understanding of metastatic breast cancer. foster furtheronly/ortumor progression towards the metastatic phenotypeThe BMSM marker profile reported here is based on two major paradigms: first, the study provides a critical evaluation of currently used markers for CTCs; and demonstrates that several markers presently used in the clinic cannot accurately identify tumor cells within the blood stream of breast cancer patients with brain metastasis, thus unlikely to predict BMBC. Second, this group of markers are termed as a brain-metastasis-selected signature to identify CTCs capable of colonizing the brain. From here, cellular mechanisms of CTC homing to distant locations such as brain and lung can be explored to better understand tumor cell dormancy versus cell proliferation as well as ways to treat brain metastatic disease. These aspects have not been investigated in the context of a crosstalk between CTCs and the brain microenvironment. With the development of CTC lines that possess various signature proteins, these mechanisms can now be explored in detail.
MATERIALS AND METHODS
Patient samples and blood collection
Samples were collected with consent from 38 patients with metastatic/non-metastatic breast cancer according to a protocol approved by the Institutional Review Board at MD Anderson Cancer Center (Table 1). Peripheral blood volumes (20-45 ml) were collected in CellSave tubes (Veridex, LLC) in sterile conditions. Blood was obtained at the middle of vein puncture after the first 5 ml of blood were discarded to avoid contamination by normal epithelial cells. All samples were provided immediately to the laboratory for CTC analysis. Only patients starting a new line of therapy were included to the study. Patients with concurrent disease(s) were excluded. Patients were required to have clinical and radiological evidence of progressive breast cancer and underwent systemic therapy as appropriate for their malignancy irrespective of CTC status. Patient data regarding age, tumor histology, estrogen/progesterone (ER/PR), and HER2 receptor status, type, and number of metastatic sites, and systemic therapy were recorded. PBMCs were isolated by Ficoll-Hypaque gradient, as described (17), and used for FACS analyses or FISH/IF determinations following cytospins and slide preparation.
CTC selection by FACS
Isolated patient PBMCs were analyzed and sorted using the BD FACS Aria II 3 Laser high-speed sorting flow cytometer (Becton Dickinson, Inc.) equipped with 12 independent fluorescence channel capabilities and DIVA acquisition software (multiparametric flow cytometry) (30). PBMCs staining for each patient included single-color controls to facilitate rigorous instrument set-up and compensation. Between 5.0 × 105 and 2.0 × 106 events were collected per list mode data file. For the primary FACS selection, markers used were CD45, ALDH1, and EpCAM. Collected cells were divided into two groups: CD45-/ALDH1+/EpCAM+ or CD45-/ALDH1+/EpCAM-. Antibodies and reagents as well as protocols used for flow cytometry and cell sorting are in Supplementary Methods.
CTC culture
CD45–/ALDH1+ CACCs were collected from FACS and cultured in stem cell culture medium (DMEM/F12 containing 5 mg/ml insulin, 0.5 mg/ml hydrocortisone, 2% B27, 20 ng/ml EGF, and 20 ng/ml FGF-2) for the first seven days, then switched to EpiCult-C medium from day 8 (STEMCELL Technologies Inc.) with 10% FBS and 1% penicillin/streptomycin at 37°C, 5% CO2, and continued to grow in this medium until day 21. Single colonies of both EpCAM+ and EpCAM– CACCs from the original CD45–/ALDH1+ population were transferred into 24- or 6-well plates for further growth, and subsequently into T75 tissue culture flasks for culture expansion. The medium used from day 22 on was DMEM/F12 plus 10% FBS and 1% penicillin/streptomycin solution (Regular M) (table S2).
Human MDA-MB-231 parental and the brain metastatic MDA-MB231BR cell variant (19, 21) transfected with HER2 were obtained from P. Steeg (National Cancer Institute). CTC clones were obtained at early passage, DNA-fingerprinted, analyzed for somatic mutations (homozygous for TP53 G839A, heterozygous for KRAS G38A, and BRAF G1391T), and tested for continued and consistent in vivo abilities to metastasize to brain over the course of 2 years (test every 20 passages). Cells were cultured in Dulbecco's Modified Eagle Medium plus F12 (DMEM/F12) (Invitrogen) supplemented with 10% FBS (Invitrogen). All other cell lines used in this study were obtained from the American Tissue Culture Collection (ATCC), DNA fingerprinted, and cultured under prescribed conditions. Cells were harvested using trypsin/EDTA (GIBCO) for spiking experiments using blood from healthy donors (disease-free) or for IF and/or flow cytometry.
CTC cultures were DNA fingerprinted (Supplementary Methods) to ensure tumor cell fidelity and grown to generate primary CTC lines CTC-1, CTC-2, and CTC-3, which were used for analyses at early passage. Blood and cell culture DNA Midi kit (Qiagen Inc.) was used to isolate DNA from the various cell sources. Cells were grown in DMEM/F12 supplemented with 10% FBS (Invitrogen) in a humidified, 5% CO2 atmosphere at 37oC, and were assessed as pathogen-free by periodic testing for Mycoplasma contamination. They were employed only at p < 20 passage and if Mycoplasma-negative.
Real-time PCR of patient circulating cells
Total RNA from PBMCs was isolated using the RNeasy Plus Mini Kit with QIAshredder (Qiagen) according to manufacturer's instructions. RNA isolation and PCR amplification protocols are described in Supplementary Methods.
Experimental animal metastasis
Female athymic nude mice (nu/nu, 4-5 weeks old) were purchased from Harlan Sprague Dawley Inc., and maintained at the accredited animal facility of Baylor College of Medicine (BCM). All studies were conducted according to NIH animal use guidelines and a protocol approved by the BCM Animal Care Committee. CTC lines (n = 10 mice per CTC line per treatment group) were injected either intracardiacally (left ventricle) or into the tail vein of nude mice (0.5 × 106 cells per mouse) after animals were anesthesized with isofluorane. For intracardiac cell injections, we inserted a 30-gauge needle on a tuberculin syringe into the second intercoastal space 3 mm to the left of the stemnum and centrally aimed. The spontaneous and continuous entrance of pulsating blood into the transparent needle hub indicated the proper positioning of the needle into the left ventricle of the mouse heart. Further, cells were injected in animals over a strict 20-40 sec time window. Mice were euthanized by CO2 asphyxiation when they showed signs of neurological impairment (usually 4-6 weeks), the whole lungs and brains removed, and fixed in Bouin's solution. Serial sections were obtained by cutting every 300 μm2 area. Small metastases (micro-metastasis) were defined as lesions ≤ 50 μm2. Three independent experiments were performed per CTC line, per treatment group (e.g., BMSM CTCs), per injection route. Data were the pooled and analyzed for statistical significance.
Statistical analyses
All data were analyzed using ANOVA or Student's t test, and represent the mean ± SEM of at least triplicate samples or the average ± S.D. of independent analyses, as indicated. P < 0.05 was considered statistically significant. Statistical tests were performed with SAS statistical software (version 9.1; SAS Institute).
Supplementary Material
Acknowledgments
We thank T. Zaidi, W. He, and R. Katz (MD Anderson Cancer Center) for FICTION/BioView analyses, expert pathological assistance, and the use of the BioView platform. We thank the FACS and Characterized Cell Line (CCLC) Core facilities at MD Anderson Cancer Center. We finally thank the nurses and the breast cancer patients for donating samples that made these investigations possible. Funding: Support for STR DNA fingerprinting and mutation analyses (Sequenom system) Core was provided by the Cancer Center Support Grant-funded Characterized Cell Line Core (NCI CA016672). This study was supported by NIH grant 1R01 CA160335, the Department of Defense-CDMRP IDEA Award (W81XWH-11-1-0315), and The Breast Cancer Award from the Avon Foundation for Women to D.M. The project was supported by the Human Tissue Acquisition and Pathology (HTAP) Core at Baylor College of Medicine with funding from the NIH NCI P30-CA125123.
Footnotes
Author Contributions: L.Z. performed experiments, analyzed data, and wrote the manuscript; L.D.R. designed and completed RT-PCR, and designed and analyzed data from FACS experiments; M.A.W. completed the isolation of PBMCs and performed CTC analyses (CellSearch and FACS); J.N. assisted in FACS and real-time PCR/RT-PCR experiments, and cell culture; W.J. assisted in performing RT-PCR and cell culture; D.K. assisted in performing IHC analyses; J.C.G. provided pathologic assessment of tumor metastasis in xenografts; M.D.G. provided patient samples and clinical information; D.M. designed experiments, revised and edited the manuscript, and supervised the research. NOT at present.
Competing interests: D.M., L.Z., and L.R. are inventors on patent application PCT/US12/66868, titled “A CTC Biomarker Assay to Combat Breast Cancer Brain Metastasis,” associated with this work and owned by Baylor College of Medicine.
None.
REFERENCES AND NOTES
- 1.Talmadge JE, Fidler I. AACR Centennial Series: The biology of cancer metastasis. Cancer Res. 2010;70(14):5649–69. doi: 10.1158/0008-5472.CAN-10-1040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Lin NU, Winer EP. Brain metastasis: the HER2 paradigm. Clin. Cancer Res. 2007;13(6):1648–55. doi: 10.1158/1078-0432.CCR-06-2478. [DOI] [PubMed] [Google Scholar]
- 3.Cristofanilli M, Budd GT, Ellis MJ, Stopeck A, Matera J, Miller MC, Reuben JM, Doyle GV, Allard WJ, Terstappen LW, Hayes DF. Circulating tumor cells, disease progression and survival in metastatic breast cancer. New Eng J Med. 2004;351:781–91. doi: 10.1056/NEJMoa040766. [DOI] [PubMed] [Google Scholar]
- 4.Alix-Panabieres C, Schwarzenbach H, Pantel K. Circulating tumor cells and circulating tumor DNA. Annu. Rev. Med. 2012;63:199–215. doi: 10.1146/annurev-med-062310-094219. [DOI] [PubMed] [Google Scholar]
- 5.Mego M, De Giorgi U, Dawood S, Wang X, Valero V, Andreopoulou E, Handy B, Ueno NT, Reuben JM, Cristofanilli M. Characterization of metastatic breast cancer patients with nondetectable circulating tumor cells. Int J Cancer. 2011;129:417–23. doi: 10.1002/ijc.25690. [DOI] [PubMed] [Google Scholar]
- 6.Yu M, Bardia A, Wittner BS, Stott SL, Smas ME, Ting DT, Isakoff SJ, Ciciliano JC, Wells MN, Shah AM, Concannon KF, Donaldson M, Sequist LV, Brachtel E, Sgroi D, Baselga J, Ramaswamy S, Toner M, Haber DA, Maheswaran S. Circulating breast tumor cells exhibit dynamic changes in epithelial and mesenchymal composition. Science. 2013;339:580–4. doi: 10.1126/science.1228522. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Yu M, Stott S, Toner M, Maheswaran S, Haber B. Circulating tumor cells: approaches to isolation and characterization. J. Cell Biol. 2011;192(3):373–82. doi: 10.1083/jcb.201010021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Pecot CV, Bischoff FZ, Mayer JA, Wong KL, Pham T, Bottsford-Miller J, Stone RL, Lin YG, Jaladurgam P, Roh JW, Goodman BW, Merritt WM, Pircher TJ, Mikolajczyk SD, Nick AM, Celestino J, Eng C, Ellis LM, Deavers MT, Sood AK. A novel platform for detection of CK+ and CK− CTCs. Cancer Discovery. 2011;1(7):580–6. doi: 10.1158/2159-8290.CD-11-0215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Sieuwerts AM, Kraan J, Bolt J, van der Spoel P, Elstrodt F, Schutte M, Martens JW, Gratama JW, Sleijfer S, Foekens JA. Anti-epithelial cell adhesion molecule antibodies and the detection of circulating normal-like breast tumor cells. J Natl Cancer Inst. 2009;101:61–6. doi: 10.1093/jnci/djn419. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Königsberg R, Obermayr E, Bises G, Pfeiler G, Gneist M, Wrba F, de Santis M, Zeillinger R, Hudec M, Dittrich C. Detection of EpCAM positive and negative circulating tumor cells in metastatic breast cancer patients. Acta Oncol. 2011;50:700–10. doi: 10.3109/0284186X.2010.549151. [DOI] [PubMed] [Google Scholar]
- 11.Harrell JC, Prat A, Parker JS, Fan C, He X, Carey L, Anders C, Ewend M, Perou CM. Genomic analysis identifies unique signatures predictive of brain, lung, and liver relapse. Breast Cancer Res Treat. 2012;132(2):523–35. doi: 10.1007/s10549-011-1619-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Joosse SA, Hannemann J, Spotter J, Bauche A, Andreas A, Muller V, Pantel K. Changes in keratin expression during metastatic progression of breast cancer: Impact on the detection of circulating tumor cells. Clin Cancer Res. 2012;18(4):993–1003. doi: 10.1158/1078-0432.CCR-11-2100. [DOI] [PubMed] [Google Scholar]
- 13.Mego M, Mani SA, Cristofanilli M. Molecular mechanisms of metastasis in breast cancer clinical applications. Nat Rev Clin Oncol. 2010;7:693–701. doi: 10.1038/nrclinonc.2010.171. [DOI] [PubMed] [Google Scholar]
- 14.Pretlow TG, Schwartz S, Giaconia JM, Wright AL, Grimm HA, Edgehouse NL, Murphy JR, Morkowitz SD, Jamison JM, Summers JL, Hamlin CR, MacLennan GT, Resnick MI, Pretlow TP, Connell CF. Prostate cancer and other xenografts from cells in peripheral blood of patients. Cancer Res. 2000;60:4033–6. [PubMed] [Google Scholar]
- 15.Ameri K, Luong R, Zhang H, Powell AA, Montgomery KD, Espinosa I, Bouley DM, Harris AL, Jeffrey SS. Circulating tumor cells demonstrate an altered response to hypoxia and an aggressive phenotype. Br J Cancer. 2010;102:561–9. doi: 10.1038/sj.bjc.6605491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Maheswaran S, Sequist LV, Nagrath S, Ulkus L, Brannigan B, Collura CV, Inserra E, Diederichs S, Iafrate AJ, Bell DW, Digumarthy S, Muzikansky A, Irimia D, Settleman J, Tompkins RG, Lynch TJ, Toner M, Haber DA. Detection of mutations in EGFR in circulating lung-cancer cells. New Engl. J Med. 2008;359:366–77. doi: 10.1056/NEJMoa0800668. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Katz RL, He W, Khanna A, Fernandez RL, Zaidi TM, Krebs M, Caraway NP, Zhang HZ, Jiang F, Spitz MR, Blowers DP, Jimenez CA, Mehran RJ, Swisher SG, Roth JA, Morris JS, Etzel CJ, El-Zein R. Genetically abnormal circulating cells in lung cancer patients: an antigen independent fluorescence in situ hybridization – based control study. Clin Cancer Res. 2010;16(15):3976–87. doi: 10.1158/1078-0432.CCR-09-3358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Ginestier C, Hur MH, Charafe-Jauffret E, Monville F, Dutcher J, Brown M, Jacquemier J, Viens P, Kleer CG, Liu S, Schott A, Hayes D, Birnbaum D, Wicha MS, Dontu G. ALDH1 is a marker of normal and malignant human mammary stem cells and a predictor of poor clinical outcome. Cell Stem Cell. 2007;1:555–67. doi: 10.1016/j.stem.2007.08.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.McGowan PM, Simedrea C, Ribot EJ, Foster PJ, Palmieri D, Steeg PS, Allan AL, Chambers AF. Notch1 inhibition alters the CD44hi/CD24lo population and reduces formation of brain metastases from breast cancer. Mol Cancer Res. 2011;9(7):834–44. doi: 10.1158/1541-7786.MCR-10-0457. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Hirose H, Ishii H, Mimori K, Ohta D, Ohkuma M, Tsujii H, Saito T, Sekimoto M, Doki Y, Mori M. Notch pathway as candidate therapeutic target in Her2/Neu/ErbB2 receptor-negative breast tumors. Oncol Rep. 2010;1:35–43. [PubMed] [Google Scholar]
- 21.Palmieri D, Bronder JL, Herring JM, Yoneda T, Weil RJ, Stark AM, Kurek R, Vega-Valle E, Feigenbaum L, Halverson D, Vortmeyer AO, Steinberg SM, Aldape K, Steeg PS. Her-2 overexpression increases the metastatic outgrowth of breast cancer cells in the brain. Cancer Res. 2007;67:4190–98. doi: 10.1158/0008-5472.CAN-06-3316. [DOI] [PubMed] [Google Scholar]
- 22.Rimawi MF, Shetty PB, Weiss HL, Schiff R, Osborne K, Chamness GC, Elledge RM. EGFR expression in breast cancer association with biologic phenotype and clinical outcomes. Cancer. 2010;116(5):1234–46. doi: 10.1002/cncr.24816. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Fehm T, Müller V, Aktas B, Janni W, Schneeweiss A, Stickeler E, Lattrich C, Löhberg CR, Solomayer E, Rack B, Riethdorf S, Klein C, Schindlbeck C, Brocker K, Kasimir-Bauer S, Wallwiener D, Pantel K. HER2 status of circulating tumor cells in patients with metastatic breast cancer: a prospective, multicenter trial. Br Cancer Res and Treat. 2010;124(2):403–12. doi: 10.1007/s10549-010-1163-x. [DOI] [PubMed] [Google Scholar]
- 24.Zhang L, Sullivan P, Suyama J, Marchetti D. Epidermal growth factor-induced heparanase nucleolar localization augments DNA topoisomerase I activity in brain metastatic breast cancer. Mol Cancer Res. 2010;8(2):278–90. doi: 10.1158/1541-7786.MCR-09-0375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Zhang L, Sullivan PS, Goodman JC, Gunaratne PH, Marchetti D. MicroRNA-1258 suppresses breast cancer brain metastasis by targeting heparanase. Cancer Res. 2011;71(3):645–54. doi: 10.1158/0008-5472.CAN-10-1910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Ridgway LD, Wetzel M, Epstein A, Marchetti D. Heparanase - induced GEF-H1 signaling regulates the cytoskeletal dynamics of brain metastatic breast cancer cells. Mol Cancer Res. 2012;10(6):689–702. doi: 10.1158/1541-7786.MCR-11-0534. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Li X, Lewis MT, Huang J, Gutierrez C, Osborne KC, Wu M, Hilsenbeck SG, Pavlick A, Zhang X, Chamness GC, Wong H, Rosen J, Chang JC. Intrisic resistance of tumorigenic breast cancer cells to chemotherapy. J Natl Cancer Inst. 2008;100:672–9. doi: 10.1093/jnci/djn123. [DOI] [PubMed] [Google Scholar]
- 28.Tarin D. Inappropriate gene expression in human cancer and its far-reaching biological and clinical significance. Cancer Metastasis Rev. 2012;31(1-2):21–39. doi: 10.1007/s10555-011-9326-8. [DOI] [PubMed] [Google Scholar]
- 29.Ikediobi ON, Davies H, Bignell G, Edkins S, Stevens C, O'Meara S, Santarius T, Avis T, Barthorpe S, Brackenbury L, Buck G, Butler A, Clements J, Cole J, Dicks E, Forbes S, Gray K, Halliday K, Harrison R, Hills K, Hinton J, Hunter C, Jenkinson A, Jones D, Kosmidou V, Lugg R, Menzies A, Mironenko T, Parker A, Perry J, Raine K, Richardson D, Shepherd R, Small A, Smith R, Solomon H, Stephens P, Teague J, Tofts C, Varian J, Webb T, West S, Widaa S, Yates A, Reinhold W, Weinstein JN, Stratton MR, Futreal PA, Wooster R. Mutation analysis of 24 known cancer genes in the NCI-60 cell line set. Mol. Cancer Ther. 2006;5(11):2606–12. doi: 10.1158/1535-7163.MCT-06-0433. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Rhim AD, Mirek ET, Aiello NM, Maitra A, Bailey JM, McAllister F, Reichert M, Beatty GL, Rustgi AK, Vonderheide RH, Leach SD, Stanger BZ. EMT and dissemination precede pancreatic tumor formation. Cell. 2012;148:349–61. doi: 10.1016/j.cell.2011.11.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Dominici M, Blanc Le K, Mueller I, Slaper-Cortenbach I, Marini FC, Krause DS, Deans RJ, Keating A, Prockop DJ, Horwitz EM. Minimal criteria for defining multipotent mesenchymal stromal cells. The International Society for Cellular Therapy position statement. Cytotherapy. 2006;8(4):315–7. doi: 10.1080/14653240600855905. [DOI] [PubMed] [Google Scholar]
- 32.Cohen-Kaplan V, Dowek I, Naroditsky I, Vlodavski I, Ilan N. Heparanase augments epidermal growth factor receptor phosphorylation: correlation with head and neck tumor progression. Cancer Res. 2008;68(24):10077–85. doi: 10.1158/0008-5472.CAN-08-2910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Fonsatti E, Jekunen AP, Kairemo KJA, Coral S, Snellman M, Nicotra MR, Natali PG, Altomonte M, Maio M. Endoglin is a suitable target for efficient imaging of solid tumors: in vivo evidence in a canine mammary carcinoma model. Clin Cancer Res. 2000;6:2037–43. [PubMed] [Google Scholar]
- 34.Ostapkowicz A, Inai K, Smith L, Kreda S, Spychala J. Lipid rafts remodeling in estrogen receptor-negative breast cancer is reversed by histone deacetylase inhibitor. Mol Cancer Ther. 2006;5(2):238–45. doi: 10.1158/1535-7163.MCT-05-0226. [DOI] [PubMed] [Google Scholar]
- 35.Bos DP, Zhang XH-F, Nadal C, Shu W, Gomis RR, Nguyen DX, Minn AJ, van de Vijer MJ, Gerald WL, Foekens JA. Massague J Genes that mediate breast cancer metastasis to brain. Nature. 2009;459(7249):1–8. doi: 10.1038/nature08021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Charafe-Jauffret E, Ginestier C, Iovino F, Tarpin C, Diebel M, ESterni B, Houvenaeghel G, Extra J-M, Bertucci F, Jacquemier J, Xerri L, Dontu G, Stassi G, Xiao Y, Barsky SH, Birnbaum D, Viens P, Wicha MS. Aldehyde dehydrogenase 1-positive cancer stem cells mediate metastasis and poor clinical outcome in inflammatory breast cancer. Clin Cancer Res. 2009;16(1):45–55. doi: 10.1158/1078-0432.CCR-09-1630. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Yatim A, Benne C, Sobhian B, Laurent-Chabalier L, Deas O, Judde J-G, Lilere J-D, Levy Y, Benkirane M. NOTCH1 nuclear interactome reveals key regulators of its transcriptional activity and oncogenic function. Cell. 2012;48:445–58. doi: 10.1016/j.molcel.2012.08.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Paik S, Kim C, Wolmark N. HER2 status and benefit from adjuvant trastuzumab in breast cancer. N Engl J Med. 2008;358:1409–11. doi: 10.1056/NEJMc0801440. [DOI] [PubMed] [Google Scholar]
- 39.Perez EA, Reinholz MM, Hillman DW, Hillman DW, Tenner KS, Schroeder MJ, Davidson NE, Martino S, Sledge GW, Harris LN, Gralow JR, Dueck AC, Ketterling RP, Ingle JN, Lingle WL, Kaufman PA, Visscher DW, Jenkins RB. HER2 and chromosome 17 effect on patient outcome in the N9831 adjuvant trastuzumab trial. J Clin Oncol. 2010;28:4307–15. doi: 10.1200/JCO.2009.26.2154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Vreys V, David G. Mammalian heparanase: what is the message? J Cell Mol Med. 2007;11(3):427–52. doi: 10.1111/j.1582-4934.2007.00039.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Brabletz T. EMT and MET in metastasis: Where are the cancer stem cells? Cancer Cell. 2012;22:699–701. doi: 10.1016/j.ccr.2012.11.009. [DOI] [PubMed] [Google Scholar]
- 42.Mina LA, Sledge GW., Jr. Rethinking the metastatic cascade as a therapeutic target. Nature Rev Clin Onc. 2011;8:325–32. doi: 10.1038/nrclinonc.2011.59. [DOI] [PubMed] [Google Scholar]
- 43.Romano P, Manniello A, Aresu O, Armento M, Cesaro M, Parodi B. Cell line data base: structure and recent improvements towards molecular authentication of human cell lines. Nucl Ac Res. 2009;37:D925–32. doi: 10.1093/nar/gkn730. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.