Significance
Influenza A virus is characterized by a large host range and rapid antigenic drift—features not common to many viruses. To begin to understand the molecular basis behind these features, we performed genome-wide mutagenesis to gauge the relative flexibility of the viral genome. We found that only two small regions in the genome (the hemagglutinin head domain and nonstructural protein 1) were highly tolerant to transposon-based insertional mutation. We believe that this analysis highlights an evolutionary strategy of influenza A virus: maintain highly plastic domains in proteins required for rapid host adaptation. This finding may help explain why influenza viruses possess certain traits not common to all RNA viruses.
Keywords: insertional mutagenesis, protein flexibility, viral evolution
Abstract
The molecular basis for the diversity across influenza strains is poorly understood. To gain insight into this question, we mutagenized the viral genome and sequenced recoverable viruses. Only two small regions in the genome were enriched for insertions, the hemagglutinin head and the immune-modulatory nonstructural protein 1. These proteins play a major role in host adaptation, and thus need to be able to evolve rapidly. We propose a model in which certain influenza A virus proteins (or protein domains) exist as highly plastic scaffolds, which will readily accept mutations yet retain their functionality. This model implies that the ability to rapidly acquire mutations is an inherent aspect of influenza HA and nonstructural protein 1 proteins; further, this may explain why rapid antigenic drift and a broad host range is observed with influenza A virus and not with some other RNA viruses.
Influenza A virus (IAV) is a segmented negative-sense, single-stranded RNA virus (1). Every year in the United States alone, IAV is thought to infect 5–20% of the population, leading to more than 200,000 hospitalizations for IAV-related disease (2). Despite this major burden to human health, and significant research efforts into viral pathogenesis, there are still major aspects of IAV biology that remain poorly understood.
Most RNA viruses, including IAV, encode an error-prone RNA-dependent RNA polymerase responsible for replicating the viral genome (1, 3). Despite the fact that all eight influenza viral segments have errors introduced during RNA replication (1), sequence analysis shows that the conservation of segments (and therefore proteins) range from highly variable to highly conserved (4–7). It is currently unclear as to whether this disparity arises purely from a lack of selective pressure (i.e., all proteins encoded by the virus tolerate a range of mutations, but only those that confer an evolutionary advantage become fixed in a population) or if there is an inherent difference in the ability of the viral proteins themselves to tolerate mutations. Currently, there is no direct experimental evidence to distinguish between these two possibilities.
One approach to directly provide evidence of the potential mutability of the viral genome is genome-wide insertional mutagenesis. This technique is one in which relatively small insertions are randomly introduced at all (or at least the majority of) possible sites across a genome. By determining the locations at which the viral genome will tolerate insertions, one can evaluate the intrinsic flexibility of the genome. Viruses with small genomes are excellent candidates for this type of analysis. Genome-wide insertional mutagenesis of two positive-stranded RNA viruses has been reported (8, 9); however, no such mutagenesis has been performed for a negative-sense RNA virus.
In this report, we performed near-saturating insertional mutagenesis of the IAV genome. Upon mapping tolerated insertion sites, we identified two regions of the viral genome that were highly enriched for mutations. The identified regions were in domains associated with host adaptation and immune response evasion (10, 11): the viral glycoprotein hemagglutinin (12, 13) and nonstructural protein 1 (NS1) (14, 15). These data suggest that although the majority of the viral genome is resistant to major mutation, two viral proteins contain extremely flexible proteins scaffolds. These flexible domains may allow for rapid adaptation to new or changing host environments.
Results
To determine the relative tolerance of the IAV proteins to mutation, we generated high-coverage mutant libraries of all eight segments of the human H1N1 strain influenza A/Puerto Rico/8/1934 (PR8). We mutagenized the genome in vitro using the bacteriophage Mu transposase and an artificial transposable element (16). After selection and removal of the transposable element, a 15-nt insertion that contains a unique 10-nt conserved molecular tag remains in the virus genome (Fig. 1A). The insertions can code for different amino acids, depending on the frame, but cannot code for a stop codon. We chose to use insertional mutagenesis over a strategy that would introduce point mutations to model a more significant lesion in the viral protein and thus identify the most flexible genomic locations.
Fig. 1.
Construction of mutant libraries and rescue of viruses. (A) Influenza segments (blue) were cloned into a pGEM shuttling vector and mutagenized in vitro. The virus segment containing the transposon (red) was removed from the shuttling vector and ligated into the viral RNA expression plasmid pPolI. Finally, the body of the transposon was removed and the viral segment was ligated back together to leave the 15-nt insertion (red striped). (B) The mutant library for a given segment was transfected with the other seven WT segments into 293T cells. Rescued virus was used to infect MDCK cells. After 24 h, virus was collected and used to infect naïve MDCK cells. RNA was collected from the second set of MDCK cells and subjected to RT-PCR for the mutant segment. The input library was PCR-amplified from the expression plasmid. Both sets of amplified DNA were prepared for sequencing and submitted for Illumina next-generation sequencing.
To generate recombinant viruses, the mutant library for one segment was transfected with the seven other nonmutagenized segments into cells. To maximize the recovery of mutant viruses, multiple independent rescues were performed, and the rescued viral particles were pooled and used to infect the highly permissive Madin–Darby canine kidney (MDCK) cell line. Virus was allowed to replicate for 24 h (corresponding to approximately three replication cycles), and then infectious viral particles were passaged on new MDCK cells for another 24 h. Total RNA was then extracted and subjected to RT-PCR for the segment encoded by the mutant library. cDNA was then prepared for next-generation Illumina sequencing to identify locations in the viral genome that tolerate the insertions (Fig. 1B).
Sequencing of our input libraries revealed good genome coverage for all segments (Fig. 2). Despite segments being similarly mutagenized and virus being rescued under the same conditions, sequencing revealed that the vast majority of the viral genome (six of the eight segments) was highly refractory to tolerating insertions (Fig. 2 B–D and F–H; Dataset S1). Two segments, however—the segments coding for the viral hemagglutinin (HA) and nonstructural protein 1(NS1)/nuclear export protein (NEP)—were found to tolerate insertions in many locations; sites enriched greater than 10-fold over background are represented by green bars (Fig. 2 A and E; Dataset S1). To confirm our results, we independently repeated our rescues of the mutant libraries and sequenced plaquable virus clones. Though we could rescue at least one insertional mutant in all of the viral coding regions, the HA and NS segments again were the most easily recovered and overrepresented (Table 1).
Fig. 2.
Insertional mutagenesis of the IAV genome reveals unique plasticity of NS and HA segments. (A–H) Results of the Illumina sequencing analysis for all eight segments. For all panels, input libraries (Left) were sequenced from the DNA rescue plasmids to determine mutational coverage. Rescued virus was amplified via RT-PCR from MDCK cell culture (Right). The x axis corresponds to the nucleotide position along the viral segment. The y axis corresponds to raw number of reads for the input and fold increase above background for the output. For output samples, a cutoff of 10-fold enrichment is indicated with the dotted line. Green lines represent viruses that tolerate insertions at nucleotide positions greater than 10-fold above background. Red lines indicate viruses that are enriched less than 10-fold. Asterisks on A and B indicate values greater than the y-axis limit; actual values are denoted on the panel.
Table 1.
Verified transposon insertions sites of viruses recoverable by plaque purification
 |
Insertional mutant viruses rescued in 293T cells were amplified in eggs and plaque-purified on MDCK cells. Purified viral RNAs were sequenced individually to determine the location of insertion. The “translation of insertion” column is a reflection of into which translational frame the insertion fell. The insertions for the HA and NS segments are shaded in light blue.
Upon closer inspection, it became apparent that the HA and NS segments did not uniformly tolerate mutations. In fact, most of the identified insertion sites in the NS segment were in NS1 (approximately nucleotides 30–720) and excluded the vast majority of the NEP protein (Fig. 2A; Fig. S1). Similarly, the HA segment had the vast majority of insertion sites with greater than 10-fold enrichment over background in the HA1 region corresponding to the HA head domain (approximately nucleotides 50–1000; Fig. 2E). These data exclude the notion that the viral genome as a whole is tolerant to mutations, even under highly permissive conditions with minimal exogenous selection pressures. Further, only two small regions of the viral genome were able to tolerate insertions, identifying these domains as rare “islands” of genomic plasticity.
To further characterize the nature of our insertional mutants, we decided to focus on the HA protein. We first repeated our rescue experiments of the HA library, but instead of infecting MDCK cells, we inoculated the entire rescue into embryonated chicken eggs. For this experiment, instead of trying to preserve the diversity of rescued viruses, we wanted to introduce the mutant pool into a more competitive environment and identify the most well-tolerated insertion sites. Competition in eggs was expected due to differences in viral mutant growth rates as well as a more restrictive environment due to the presence of the chicken embryo innate immune system (17). After incubation in eggs for 48 h (a sufficient time for the virus to reach maximal potential titer), we collected virions, amplified the HA segment, and submitted the library for deep sequencing.
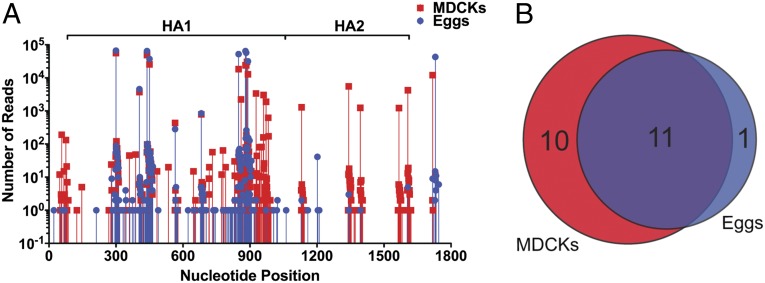
To our surprise, many of the insertion sites identified in MDCK cells were also identified in our egg competition analysis, indicating that, for the most part, diversity of the recovered viruses was preserved (Fig. 3). In fact, essentially every site identified in the HA head in MDCKs was also identified in the egg analysis. In contrast, however, we failed to recover mutations in the HA2 domain from the eggs (Fig. 3). We interpret this data to mean that insertions in the head domain are relatively well tolerated, whereas those in the stalk domain may lead to a loss of fitness.
Fig. 3.
The HA head domain tolerates insertional mutations. (A) The HA library rescued under noncompetitive conditions in MDCK cells (red) and under competitive conditions in embryonated chicken eggs (blue). The nucleotide position of the HA segment is indicated along the x axis. The HA1 (head) and HA2 (stalk) domains are denoted above the graph. (B) Venn diagram showing the overlap of the two rescue conditions from A.
It is well known that antibody-escape point mutations readily occur in five well-mapped antigenic sites in the HA head (18). We therefore overlaid the known antigenic sites with the locations of insertional mutants verified by plaque purification or common to both the MDCK and eggs rescues (Fig. 4A). Though our analysis overlapped with four of the five characterized sites, we also found insertions outside of the classical antigenic sites, indicating that the HA head can change in areas not typically associated with antibody escape. Upon highlighting these locations on the crystal structure of the HA protein [Protein Data Bank (PDB) ID code 1RU7] (19), we observe that the insertions all sit on the upper third of the protein structure, excluded from the stalk domain (Fig. 4 B and C). To confirm that a range of HA1 mutations are not associated with a loss of fitness, we randomly selected five of the transposon insertion sites, cloned them into the reverse genetic system, and rescued the viruses. Growth-curve analysis of the mutants compared with WT PR8 revealed no significant differences in growth under these conditions (Fig. 4D; Fig. S2).
Fig. 4.
Locations of transposon insertions in the HA head. (A) Diagram of the HA segment showing the locations of transposons (red and green triangles) recovered from both the MDCK and egg rescues as well as HA mutants verified by plaque purification. All insertional mutants in the HA1 domain were also denoted on the primary sequence of the HA1 domain (black triangles). Escape mutation locations defining antigenic sites are indicated by the colored boxes. Insertion locations from A are shown as red or green spheres on the crystal structure (PDB ID code 1RU7) of the HA monomer (B) and trimer (C). B Inset is an enlargement of the HA head domain. (D) Individual viruses (denoted by green in A–C) were cloned and rescued. MDCK cells were infected at an MOI of 0.001, and samples were plaqued at the indicated times.
Discussion
Our genome-wide transposon mutagenesis of IAV revealed that the viral genome, for the most part, tolerates very few insertions. For example, only a single insertion site in any of the three viral polymerase segments could be rescued. Similarly behaved were the NA, NP, and M segments with the exception of one small ∼10-nt region in the M2 protein. The most well tolerated M2 insertions, around amino acid 29, fall into the transmembrane helix (20). The translation of these insertions is such that a string of alanines is inserted into an existing run of alanines (Table 1); we suspect that this simply extends the helix, which has minor effects on structure and function. It will be important, however, to characterize some of the insertional mutants for impacts on ion channel activity and other M2 functions such as membrane scission (21). Overall, however, in contrast to the mostly intolerant genome, the vast majority of the identified insertion sites were found in the HA1 domain of the HA protein and in NS1, despite the fact that these domains together represent only ∼12% of the entire genome.
A pathogen that causes persistent infection or relies on reinfection of the same individual must have an antigenic variation strategy to escape the host antimicrobial response (22). When genome size is not a limiting factor, antigenic variability can simply be encoded as is well characterized to occur with the antigenic pilin of some Neisseria species (23). In the case of the large DNA poxvirus Vaccinia, the viral genome can literally expand to increase the adaptive potential in response to a strong selective pressure (24). Viruses with small RNA genomes, like IAV, cannot use similar mechanisms, and thus face an inherent dilemma; they must encode multifunctional and highly optimized proteins to accomplish the conquest of host cells, but must also remain flexible to escape the host response.
The model that we propose here implies that the strategy of IAV is to evolve certain proteins or protein domains to be inherently plastic and mutable; this allows for the “viable and fit” pool of viral quasispecies generated via random replication error to be disproportionately increased rapidly in certain viral proteins due to their much higher inherent tolerances (Fig. S3). The HA head domain and NS1 may have evolved to exist as flexible scaffolds, which can perform a core set of functions while maintaining a large evolutionary potential. This potential must exist to rapidly adapt to antibody pressure and antagonize antiviral signaling in a changing host environment, respectively. NS1 has been described to interact with a multitude of host proteins (14, 25), which puts it in the center of genetic conflicts across a range of hosts. We mapped the predicted NS1 insertions on the crystal structure of NS1 and found that six of seven sites are surface-exposed (PDB ID code 3F5T) (26) (Fig. S4), and this is consistent with the idea of flexible viral domains interacting with host proteins. Our model suggests that the virus is inherently predisposed to rapidly acquire mutations in certain proteins but not in others.
This framework may have significant implications for translational medicine. For example, an area of major current work in the IAV field is the development of cross-reactive antibodies, which target the HA2 conserved stalk domain region of the IAV HA (27, 28). Our model, in addition to explaining why the stalk is less variable than the head, predicts that if stalk-reactive antibodies are elicited by vaccination, the virus has a lower probability of easily escaping neutralization due to the inherently less plastic stalk domain relative to the very flexible head, which tolerates mutations.
The data presented in this study represent an artificial and stringent test of protein mutability. Our analysis almost certainly did not reveal all potential insertion sites in the virus. The viral neuraminidase (NA) is known to tolerate changes to the length of the stalk region (29–31). Although our deep-sequencing analysis failed to recover highly fit insertional mutants in this region, the one rescued mutant virus was in fact in the NA stalk domain (Table 1), which is in general agreement with previous studies. Further, many sites in the genome that are known to tolerate point mutations were not able to tolerate insertions, a good example of which is the NA protein; despite being a surface glycoprotein that drifts over time (32), our analysis found it to be intolerant of many insertions. Although this finding was initially unexpected, it is well known that there is less sequence diversity relative to HA (i.e., NA drifts slower than HA) (33, 34). Further, the selective pressures on NA are thought to be different compared with HA. For example, antibodies against NA are thought to play a less important role in the neutralization of IAV (35, 36). Thus, these data illustrate that though NA certainly has the ability to drift, NS1 and HA contain exceptionally plastic domains whose flexibility may allow for a more rapid expansion of diversity or a larger potential range of diversity relative to proteins like NA.
In sum, our mutagenesis of the IAV genome agrees with the accepted notion that viruses with small genomes are highly optimized and encode little if any nonessential genetic material. By virtue of this fact, there is very little space that tolerates insertional mutations. However, our analysis has added that despite this optimized genome, there are two major regions, the HA head and NS1, that exist as highly plastic and mutable domains and are evolved to provide a neutral platform on which variability can be generated. This plasticity probably manifests itself by influencing the speed and range of possible mutations. Finally, it is tempting to speculate that the ability of these proteins to tolerate mutations may explain why IAV is observed to have such a rapid drift rate and large host range relative to other RNA viruses. We suspect that viruses such as measles virus, mumps virus, and poliovirus may encode inherently less-flexible scaffolds relative to IAV, which limits their drift rate or evolutionary potential; this may contribute to why single vaccination against these viruses leads to lifelong protection and why they exist as human-restricted pathogens. Although this theory remains to be tested, if true, this type of genome-wide mutagenesis may help predict and/or validate optimal vaccine epitopes for viruses such as IAV, which would be predisposed to evolve or mutate very slowly.
Materials and Methods
Cells.
The 293T and MDCK cells were propagated in DMEM supplemented with 10% (vol/vol) FBS, penicillin/streptomycin, and sodium bicarbonate.
Mutant Library Generation and Rescue of Viruses.
To generate insertional mutants of all viral segments, the entire viral segment was amplified from viral rescue plasmids with the following primers: CATCGCTCTTCTGCCAGCAAAAGCAGG (forward) or CATCGCTCTTCTGCCAGCGAAAGCAGG, depending on the segment, and GGCCGGCTCTTCTATTAGTAGAAACAAGG (reverse). Viruses were subcloned into the pGEM-T shuttling vector flanked by SapI sites encoded in primers. The sequences of the viral segments corresponded to GenBank accession nos.: AF389115, AF389116, AF389117, AF389118, AF389119, AF389120, AF389121, and AF389122). The PB1 and PB2 segments contain SapI sites, so we therefore made silent mutations to eliminate those sites. For PB1, we used the QuikChange mutagenesis kit (Agilent) according to the manufacturers’ instructions with the following primers: CTGTTCCACCATTGAGGAGCTCAGACGGCAA (forward) and TTGCCGTCTGAGCTCCTCAATGGTGGAACAG (reverse). For PB2, we digested the WT pGEM-T clone with NcoI, which cuts in the vector and in the middle of segment. We amplified the WT 5′ end of PB2 with the following primers: CGCATGCTCCCGGCCGCCATGGCCGC (forward) and AGCAATAATCAAGCTTTGATCAACAT (reverse). We then synthesized and amplified a 456-nt region (with the SapI sites silently mutated) to complete the sequence with the following primers: CAAAGCTTGATTATTGCTGCT (forward) and ATCCTCTTGTGAAAATACCATG (reverse). We then performed a three-piece recombination (InFusion HD; Clontech) with the two PCR products and the rest of the segment in the digested pGEM vector to restore the complete viral segment.
All eight segments in pGEM-T were then subjected to in vitro transposon mutagenesis using the Mutation Generation System (MGS F701; Fisher Scientific) according to the manufacturer’s instructions. Two independent mutagenesis reactions were performed on 200 ng of target DNA each and then pooled for transformation. Mutagenized DNA was introduced into XL10-Gold Ultracompetent Escherichia coli (Agilent), and then plated on ten 15-cm plates of LB agar supplemented with ampicillin and kanamycin to select for the plasmid and the transposon. After ∼36 h of growth at 30 °C, bacteria were scraped and pooled, and DNA was extracted via MaxiPrep kit (Invitrogen). The purified DNA was then digested with SapI to release the viral segment (containing the transposon). After gel extraction, the segment was ligated into the viral RNA expression plasmid pPol1, which was linearized with SapI via overnight 16 °C incubation with T4 DNA ligase (NEB). The ligation was precipitated with sodium acetate and ethanol, resuspended in water, and again introduced into XL10-Gold E. coli and plated on ten 15-cm LB plates containing ampicillin and kanamycin. After ∼36 h of growth at 30 °C, bacteria were scraped and pooled, and DNA was isolated via MaxiPrep. Purified DNA was then digested with NotI to remove the transposon. After gel purification, the linear DNA vector was ligated back together via overnight incubation at 16 °C with T4 DNA ligase. The ligation was again precipitated with sodium acetate and ethanol, resuspended in water, and introduced into XL10-Gold E. coli and plated on ten 15-cm LB plates containing kanamycin (because the transposon containing KanR is now removed). Bacteria were scraped and pooled, and DNA was purified via MaxiPrep. The purified DNA from this final step represented the viral segment library of 15-nt insertions.
Virus was rescued via standard protocols (37, 38), with a few modifications. The mutant library for a given segment was introduced into 293T cells along with the seven other WT rescue plasmids. Protein expression plasmids expressing WT proteins from the influenza strain A/WSN/33 were also introduced when appropriate. Multiple independent rescues were performed for each library, with between 6 and 12 × 106 cells transfected for each library. At 24 h after transfection, the supernatant was collected, spun to pellet any cells or debris, and used to infect a confluent 15-cm dish of MDCK cells in the presence of tosyl phenylalanyl chloromethyl ketone (TPCK) treated trypsin to allow for viral spread. Because the rescue contains WT protein, and all viruses in the initial infection may not reflect true viable mutants, we passaged virus to reduce potential background. At 24 h after the initial infection, supernatant was removed, spun to remove debris, and used to infect a second 15-cm dish of MDCK cells again in the presence of trypsin. In the case where viruses were rescued in eggs, the rescue from 293T cells was directly injected into 10-d-old embryonated chicken eggs and incubated at 37 °C for 48 h.
RT-PCR and Next-Generation Sequencing.
After 24-h incubation with the second passage of rescued virus, cells were washed with PBS, and RNA was isolated via TRIzol (Invitrogen) extraction per the manufacturer’s instructions (6 mL per 150-cm dish). RNA was resuspended in RNase free water and subjected to RT-PCR with the appropriate primer set (sequences described in Table S1) via SuperScript III One-Step RT-PCR System with Platinum Taq as per the manufacturer’s instructions. RT-PCR primers were designed to overlap to ensure complete coverage of the segment. Amplified cDNA was gel purified, sheared via Corvaris sonication, and prepared for sequencing using the TruSeq DNA Sample Preparation Kit (Illumina) as per the manufacturer’s instructions. Samples were barcoded, multiplexed, and sequenced on the HiSeq2000 machine via 100-nt single-end run.
Analysis of Sequencing Datasets.
Raw sequencing data were first filtered for reads containing the molecular tag sequence introduced by the transposon (TGCGGCCGCA). Positive reads were then trimmed to remove the transposon and mapped against a reference genome using Bowtie. Mapping the end of the trimmed read identified the location of the insertion. Matching read locations were aggregated and counted to give counts per potential insertion location. All software was run on Mount Sinai's high-performance computing cluster.
In our output sequencing, large regions of the genome were absent of any mapped transposon insertion sites; to ensure this was a reflection of biology (i.e., no viable viruses with insertions in these areas) rather than lack of sequencing coverage, we performed a depth of coverage analysis on all reads that mapped to the genome regardless of whether they contained the transposon. We limited our counting tool to 8,000 reads for a given sequencing file, and we reached that threshold for most positions. In fact, our analysis revealed that all possible positions in all segments were covered by our deep sequencing (Fig. S5). We therefore have confidence that regions that had no recoverable transposons were not a reflection of technical error.
As a control for potential PCR bias in our sequencing, we took the individual reads from major hits from three segments and mapped the start site for each read containing the transposon sequence. Because our cDNA shearing was random, we expect to have a random distribution of read start sites containing a transposon in a given position. In contrast, if a hit is a reflection of PCR bias during library preparation, we would expect only a few or a single clonal DNA fragment to contain the transposon. Our analysis revealed that the distribution was fairly even across DNA fragment start sites, indicating that PCR bias was probably minimal in our library preparations (Fig. S6).
Because we biased our RT-PCR to be extremely sensitive, we also amplified highly attenuated viruses and defective viral RNAs, which increased our sequencing background noise. To set a cutoff for background noise in our analysis, we ordered the data for each segment by number of reads at a given position. We averaged the number of reads for ordered positions 21–40 to generate a stringent threshold defining where real signal could be discriminated from background while not penalizing a dataset for having a few dominant viruses. This background threshold was used to normalize each output sequencing file against itself.
Cloning and Characterization of Insertional Mutants.
To rescue viruses with transposon insertions in the positions identified by our deep-sequencing analysis, we amplified the HA segment in two pieces, encoding the transposon sequence in the 3′ end of the 5′ PCR and the 5′ end of the 3′ PCR with the primers indicated in Table S2. A three-piece recombination into the viral expression plasmid pDZ was performed via InFusion HD kit (Clontech) as per manufacturer’s instructions. Clones were sequence-validated and rescued in 293T cells with the seven other WT rescue plasmids. Rescued virus was plaque purified and sequenced to ensure correct insertion of the transposon. For growth-curve analysis, MDCK cells were infected at a multiplicity of infection (MOI) of 0.001 and incubated in Opti-MEM (Invitrogen) with TPCK trypsin. At the indicated time points, samples were harvested and plaqued via standard protocols (38, 39).
Supplementary Material
Acknowledgments
We thank the Mount Sinai Genomics Core Facility and Omar Jabado for their expertise with the Illumina sequencing; Ravi Sachidanandam for helpful discussions; and Paul Leon for experimental assistance. N.S.H. is a Merck Fellow of the Life Sciences Research Foundation. P.P. acknowledges partial funding from Center for Research on Influenza Pathogenesis Grant HHSN266200700010C, National Institutes for Health Program Project 1P01AI097092-01A1, and Program for Appropriate Technology in Health (PATH).
Footnotes
The authors declare no conflict of interest.
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1320524110/-/DCSupplemental.
References
- 1.Shaw ML, Palese P. Orthomyxoviruses. In: Knipe DM, Howley PM, editors. Fields Virology. Philadelphia: Lippincott Williams & Wilkins; 2013. pp. 1151–1185. [Google Scholar]
- 2.Thompson WW, et al. Influenza-associated hospitalizations in the United States. JAMA. 2004;292(11):1333–1340. doi: 10.1001/jama.292.11.1333. [DOI] [PubMed] [Google Scholar]
- 3.Drake JW. Rates of spontaneous mutation among RNA viruses. Proc Natl Acad Sci USA. 1993;90(9):4171–4175. doi: 10.1073/pnas.90.9.4171. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Fitch WM, Bush RM, Bender CA, Cox NJ. Long term trends in the evolution of H(3) HA1 human influenza type A. Proc Natl Acad Sci USA. 1997;94(15):7712–7718. doi: 10.1073/pnas.94.15.7712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Plotkin JB, Dushoff J, Levin SA. Hemagglutinin sequence clusters and the antigenic evolution of influenza A virus. Proc Natl Acad Sci USA. 2002;99(9):6263–6268. doi: 10.1073/pnas.082110799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Shu LL, Bean WJ, Webster RG. Analysis of the evolution and variation of the human influenza A virus nucleoprotein gene from 1933 to 1990. J Virol. 1993;67(5):2723–2729. doi: 10.1128/jvi.67.5.2723-2729.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Sevilla-Reyes EE, Chavaro-Pérez DA, Piten-Isidro E, Gutiérrez-González LH, Santos-Mendoza T. Protein clustering and RNA phylogenetic reconstruction of the influenza a virus NS1 protein allow an update in classification and identification of motif conservation. PLoS ONE. 2013;8(5):e63098. doi: 10.1371/journal.pone.0063098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Arumugaswami V, et al. High-resolution functional profiling of hepatitis C virus genome. PLoS Pathog. 2008;4(10):e1000182. doi: 10.1371/journal.ppat.1000182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Beitzel BF, Bakken RR, Smith JM, Schmaljohn CS. High-resolution functional mapping of the Venezuelan equine encephalitis virus genome by insertional mutagenesis and massively parallel sequencing. PLoS Pathog. 2010;6(10):e1001146. doi: 10.1371/journal.ppat.1001146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.dos Reis M, Tamuri AU, Hay AJ, Goldstein RA. Charting the host adaptation of influenza viruses. Mol Biol Evol. 2011;28(6):1755–1767. doi: 10.1093/molbev/msq317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Taubenberger JK, Kash JC. Influenza virus evolution, host adaptation, and pandemic formation. Cell Host Microbe. 2010;7(6):440–451. doi: 10.1016/j.chom.2010.05.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Blackburne BP, Hay AJ, Goldstein RA. Changing selective pressure during antigenic changes in human influenza H3. PLoS Pathog. 2008;4(5):e1000058. doi: 10.1371/journal.ppat.1000058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Wiley DC, Skehel JJ. The structure and function of the hemagglutinin membrane glycoprotein of influenza virus. Annu Rev Biochem. 1987;56:365–394. doi: 10.1146/annurev.bi.56.070187.002053. [DOI] [PubMed] [Google Scholar]
- 14.Hale BG, Randall RE, Ortín J, Jackson D. The multifunctional NS1 protein of influenza A viruses. J Gen Virol. 2008;89(Pt 10):2359–2376. doi: 10.1099/vir.0.2008/004606-0. [DOI] [PubMed] [Google Scholar]
- 15.García-Sastre A, et al. Influenza A virus lacking the NS1 gene replicates in interferon-deficient systems. Virology. 1998;252(2):324–330. doi: 10.1006/viro.1998.9508. [DOI] [PubMed] [Google Scholar]
- 16.Haapa S, Taira S, Heikkinen E, Savilahti H. An efficient and accurate integration of mini-Mu transposons in vitro: A general methodology for functional genetic analysis and molecular biology applications. Nucleic Acids Res. 1999;27(13):2777–2784. doi: 10.1093/nar/27.13.2777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Dar A, et al. CpG oligodeoxynucleotides activate innate immune response that suppresses infectious bronchitis virus replication in chicken embryos. Avian Dis. 2009;53(2):261–267. doi: 10.1637/8560-121808-Reg.1. [DOI] [PubMed] [Google Scholar]
- 18.Caton AJ, Brownlee GG, Yewdell JW, Gerhard W. The antigenic structure of the influenza virus A/PR/8/34 hemagglutinin (H1 subtype) Cell. 1982;31(2 Pt 1):417–427. doi: 10.1016/0092-8674(82)90135-0. [DOI] [PubMed] [Google Scholar]
- 19.Gamblin SJ, et al. The structure and receptor binding properties of the 1918 influenza hemagglutinin. Science. 2004;303(5665):1838–1842. doi: 10.1126/science.1093155. [DOI] [PubMed] [Google Scholar]
- 20.Schnell JR, Chou JJ. Structure and mechanism of the M2 proton channel of influenza A virus. Nature. 2008;451(7178):591–595. doi: 10.1038/nature06531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Rossman JS, Jing X, Leser GP, Lamb RA. Influenza virus M2 protein mediates ESCRT-independent membrane scission. Cell. 2010;142(6):902–913. doi: 10.1016/j.cell.2010.08.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Janeway C, Travers P, Walport M. Failures of Host Defense Mechanisms. Immunobiology: The Immune System in Health and Disease. 5th Ed. New York: Garland Science; 2001. Chap 11. [Google Scholar]
- 23.Criss AK, Kline KA, Seifert HS. The frequency and rate of pilin antigenic variation in Neisseria gonorrhoeae. Mol Microbiol. 2005;58(2):510–519. doi: 10.1111/j.1365-2958.2005.04838.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Elde NC, et al. Poxviruses deploy genomic accordions to adapt rapidly against host antiviral defenses. Cell. 2012;150(4):831–841. doi: 10.1016/j.cell.2012.05.049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.de Chassey B, et al. The interactomes of influenza virus NS1 and NS2 proteins identify new host factors and provide insights for ADAR1 playing a supportive role in virus replication. PLoS Pathog. 2013;9(7):e1003440. doi: 10.1371/journal.ppat.1003440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Bornholdt ZA, Prasad BV. X-ray structure of NS1 from a highly pathogenic H5N1 influenza virus. Nature. 2008;456(7224):985–988. doi: 10.1038/nature07444. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Krammer F, Palese P. Influenza virus hemagglutinin stalk-based antibodies and vaccines. Curr Opin Virol. 2013;3(5):521–530. doi: 10.1016/j.coviro.2013.07.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Subbarao K, Matsuoka Y. The prospects and challenges of universal vaccines for influenza. Trends Microbiol. 2013;21(7):350–358. doi: 10.1016/j.tim.2013.04.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Luo G, Chung J, Palese P. Alterations of the stalk of the influenza virus neuraminidase: Deletions and insertions. Virus Res. 1993;29(3):321. doi: 10.1016/0168-1702(93)90069-y. [DOI] [PubMed] [Google Scholar]
- 30.Li J, Zu Dohna H, Cardona CJ, Miller J, Carpenter TE. Emergence and genetic variation of neuraminidase stalk deletions in avian influenza viruses. PLoS ONE. 2011;6(2):e14722. doi: 10.1371/journal.pone.0014722. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Castrucci MR, Kawaoka Y. Biologic importance of neuraminidase stalk length in influenza A virus. J Virol. 1993;67(2):759–764. doi: 10.1128/jvi.67.2.759-764.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Xu X, Cox NJ, Bender CA, Regnery HL, Shaw MW. Genetic variation in neuraminidase genes of influenza A (H3N2) viruses. Virology. 1996;224(1):175–183. doi: 10.1006/viro.1996.0519. [DOI] [PubMed] [Google Scholar]
- 33.Kilbourne ED, Johansson BE, Grajower B. Independent and disparate evolution in nature of influenza A virus hemagglutinin and neuraminidase glycoproteins. Proc Natl Acad Sci USA. 1990;87(2):786–790. doi: 10.1073/pnas.87.2.786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Sandbulte MR, et al. Discordant antigenic drift of neuraminidase and hemagglutinin in H1N1 and H3N2 influenza viruses. Proc Natl Acad Sci USA. 2011;108(51):20748–20753. doi: 10.1073/pnas.1113801108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Nayak B, et al. Contributions of the avian influenza virus HA, NA, and M2 surface proteins to the induction of neutralizing antibodies and protective immunity. J Virol. 2010;84(5):2408–2420. doi: 10.1128/JVI.02135-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Marcelin G, Sandbulte MR, Webby RJ. Contribution of antibody production against neuraminidase to the protection afforded by influenza vaccines. Rev Med Virol. 2012;22(4):267–279. doi: 10.1002/rmv.1713. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Fodor E, et al. Rescue of influenza A virus from recombinant DNA. J Virol. 1999;73(11):9679–9682. doi: 10.1128/jvi.73.11.9679-9682.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Hai R, et al. Influenza viruses expressing chimeric hemagglutinins: Globular head and stalk domains derived from different subtypes. J Virol. 2012;86(10):5774–5781. doi: 10.1128/JVI.00137-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Steel J, et al. Live attenuated influenza viruses containing NS1 truncations as vaccine candidates against H5N1 highly pathogenic avian influenza. J Virol. 2009;83(4):1742–1753. doi: 10.1128/JVI.01920-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.