Fragile X mental retardation protein (FMRP) is known to exist as several isoforms generated by alternative splicing. This paper shows that two specific isoforms bind the coding region of FMRP mRNA, which contains two G-quadruplexes with higher affinity than the major isoform.
Keywords: FMRP, G-quadruplex RNA, protein–RNA interactions, fluorescence spectroscopy
Abstract
Fragile X syndrome, the most common form of inherited mental impairment in humans, is caused by the absence of the fragile X mental retardation protein (FMRP) due to a CGG trinucleotide repeat expansion in the 5′-untranslated region (UTR) and subsequent translational silencing of the fragile x mental retardation-1 (FMR1) gene. FMRP, which is proposed to be involved in the translational regulation of specific neuronal messenger RNA (mRNA) targets, contains an arginine-glycine-glycine (RGG) box RNA binding domain that has been shown to bind with high affinity to G-quadruplex forming mRNA structures. FMRP undergoes alternative splicing, and the binding of FMRP to a proposed G-quadruplex structure in the coding region of its mRNA (named FBS) has been proposed to affect the mRNA splicing events at exon 15. In this study, we used biophysical methods to directly demonstrate the folding of FMR1 FBS into a secondary structure that contains two specific G-quadruplexes and analyze its interactions with several FMRP isoforms. Our results show that minor splice isoforms, ISO2 and ISO3, created by the usage of the second and third acceptor sites at exon 15, bind with higher affinity to FBS than FMRP ISO1, which is created by the usage of the first acceptor site. FMRP ISO2 and ISO3 cannot undergo phosphorylation, an FMRP post-translational modification shown to modulate the protein translation regulation. Thus, their expression has to be tightly regulated, and this might be accomplished by a feedback mechanism involving the FMRP interactions with the G-quadruplex structures formed within FMR1 mRNA.
INTRODUCTION
Fragile X syndrome (FXS) is the most common form of inherited intellectual disability in humans (Crawford et al. 2001), affecting ∼1 in 3000 males and 1 in 5000 females (Morton et al. 1997; Hawkins et al. 2011). In the vast majority of the cases, FXS is caused by the absence of a single protein named the fragile X mental retardation protein (FMRP) required for normal neural function (O'Donnell and Warren 2002; Jin et al. 2004). The gene encoding for this protein, FMR1, is located on the X chromosome and contains in its 5′-untranslated region (5′ UTR) a sequence of cytosine-guanine-guanine (CGG) repeats that is 20–60 repeats in healthy individuals, being expanded to more than 200 repeats in FXS patients (Jin and Warren 2000; O'Donnell and Warren 2002). The cytosines in the expanded region of CGG repeats become hypermethylated, causing the gene silencing of FMR1 and loss of expression of FMRP (Pieretti et al. 1991; Tassone et al. 1999).
FMRP is an RNA binding protein that contains two types of RNA binding domains: two K-homology domains and one arginine-glycine-glycine (RGG) box domain, as well as a nuclear localization signal (NLS) and a nuclear export signal (NES) (Fig. 1; Ashley et al. 1993; Siomi et al. 1993; O'Donnell and Warren 2002). FMRP is postulated to regulate the translation of specific neuronal messenger RNA (mRNA) targets (Darnell et al. 2001) as part of large ribonucleoprotein (mRNP) complexes, although the detailed mechanism of this regulatory process is not well understood. FMRP is post-translationally modified by arginine methylation within the RGG box domain and by serine phosphorylation in a region directly upstream of the RGG box domain (Siomi et al. 2002; Stetler et al. 2006). Additionally, several FMRP isoforms can be produced through alternative splicing events involving the inclusion/skipping of exons 12 and 14, as well as three acceptor sites at exon 15 and two at exon 17 (Verkerk et al. 1993; Sittler et al. 1996; Brackett et al. 2013). Figure 1 shows differences in the sequences of the FMRP isoforms 1 (ISO1, the largest FMRP isoform), 2 (ISO2), and 3 (ISO3) created using the three acceptor sites at exon 15 (and inclusive of exons 12 and 14).
FIGURE 1.

A schematic representation of the full-length FMRP, which shows the nuclear localization signal (NLS), the two K-homology domains (KH1 and KH2), the nuclear export signal (NES), the main site of phosphorylation (P), and the RGG box (RGG). An expansion of the sequence differences within isoforms 1–3, resulting from the alternative splicing at exon 15 of FMR1 mRNA, is also illustrated. The phosphorylation of serine 500 (arrow) has been shown to be biologically relevant. The bracket encompasses the amino acids encoded by the G-rich FBSsh region.
Several studies showed that the RGG box domain of FMRP binds with high affinity to mRNA targets that have been shown to adopt G-quadruplex structures (Brown et al. 2001; Darnell et al. 2001; Menon and Mihailescu 2007; Bole et al. 2008; Menon et al. 2008; Evans et al. 2012). G-quadruplexes consist of guanine tetrads held together by Hoogsteen base-pairing and stabilized by monocations, the potassium ion (K+) providing the most thermodynamic stability (Williamson et al. 1989; Williamson 1994).
FMRP has also been shown to interact with its own mRNA within a 100-nucleotide G-rich region named the FMRP binding sequence (FBS), which was proposed to fold into two distinct G-quadruplex structures (Schaeffer et al. 2001, 2003; Didiot et al. 2008). This finding prompted the hypothesis that FMRP might use an autoregulatory loop to regulate its own translation; however, a subsequent study has shown that the FMRP interactions with FBS do not affect the FMR1 mRNA stability and translation (Didiot et al. 2008). The purine-rich FBS region has been found instead to be a potent exonic splicing enhancer whose function is dependent on the presence of the G-rich region (Didiot et al. 2008). FBS is located in the proximity of the three different acceptor sites at exon 15, and FMRP binding to FBS has been found to control the splicing events at exon 15. An overexpression of FMRP ISO1 decreased the usage of exon 15 first acceptor site, concomitant with an increased usage of exon 15 second and third acceptor sites, whereas the absence of FMRP resulted in the absence of the minor isoforms created by the usage of exon 15 second and third acceptor sites (Didiot et al. 2008). This direct involvement of FMRP in regulating the production of its minor isoforms created by the usage of exon 15 acceptor sites 2 and 3 (which include FMRP ISO2 and ISO3) could add a new layer of regulation to the FMRP translation regulator function, as these isoforms lack the major phosphorylation site at position 500 (Fig. 1, arrow). The FMRP phosphorylation/dephosphorylation events have been shown to play a major role in the association of FMRP with the miRNA pathway (Muddashetty et al. 2011), as phosphorylated FMRP has been found associated with the RNA interference silencing complex (RISC), the microRNA miR-125a, and the postsynaptic density 95 (PSD95) mRNA; whereas its dephosphorylation triggered by synaptic input leads to the dissociation of the RISC complex from PSD95 mRNA, allowing for the translation of the PSD95 protein (Zalfa et al. 2007; Muddashetty et al. 2011). The fact that FMRP isoforms ISO2 and ISO3 lack the major site of phosphorylation raises the possibility that these minor isoforms could fine-tune the FMRP translation regulator function by controlling the duration different mRNA targets are undergoing translation. Thus, their production has to be tightly regulated, and this might be achieved by a feedback mechanism involving the FBS exonic splicing enhancer. In this study, we have first used biophysical methods to characterize the G-quadruplex structures proposed to form in the FMR1 mRNA FBS and subsequently analyzed the binding of several FMRP isoforms (ISO1, ISO2, ISO3), as well as of a mimic of phosphorylated FMRP ISO1 (ISO1 S500D) to a 42-nucleotide FBS fragment.
RESULTS AND DISCUSSION
Biophysical characterization of the FMR1 mRNA G-quadruplex structures
Moine and coworkers proposed that two G-quadruplex structures form in the C terminus of FMR1 mRNA in a 100-nt stretch (FBS) (Schaeffer et al. 2001, 2003; Didiot et al. 2008) based on potassium-dependent stops of reverse-transcription that disappeared when mutations were introduced in the G-rich region recognized by FMRP. In this study, we used biophysical methods to directly prove the existence and characterize the fold of the G-quadruplex structures in FMR1 mRNA. Initially, we truncated FBS to a 67-nt fragment (position 1590–1657 within the FMR1 gene), named FBS_67 RNA, that retained the G-rich region proposed to fold into the two G-quadruplex structures. FBS_67 RNA has been produced by in vitro transcription reactions off a synthetic DNA template and one dimensional (1D) 1H NMR spectroscopy has been used to analyze G-quadruplex formation within FBS_67 in the presence of KCl. A broad envelope of imino proton resonances centered around 11 ppm, which correspond to guanine imino protons involved in G-quartet formation, was observed even in the absence of K+ ions (Supplemental Fig. 1A). Resonances were also observed in the region 12–14.5 ppm, corresponding to guanine and uracil imino protons involved in Watson-Crick base-pair formation, indicating also the presence of a duplex region in the structure of FBS_67 RNA.
As the KCl concentration was increased in the range of 0–100 mM, the resonances corresponding to the imino protons involved in G-quartet formation increased in intensity and became sharper, whereas the intensity of the resonances corresponding to canonical base-pairing remained constant. These results indicate unambiguously that one or more G-quadruplex structures that are stabilized by K+ ions are present in FBS_67 RNA. Circular dichroism (CD) spectroscopy was used next to characterize the G-quadruplex fold of FBS_67 RNA, and a positive band whose intensity increased upon KCl titration was observed at 264 nm and a negative one at 240 nm, signatures of parallel-type G-quadruplex structures (Supplemental Fig. 1B; Williamson 1994; Dapic et al. 2003). Nondenaturing polyacrylamide gel electrophoresis (native PAGE) was also used to analyze FBS_67, several bands being observed at all RNA and KCl concentrations investigated in the range of 0–100 mM, which indicates that multiple conformations coexist in FBS_67 RNA (data not shown). In an attempt to solve this problem, we have produced by in vitro transcription reactions two shorter fragments of FBS: FBS_Q1 RNA (15 nt, position 1602–1616 within the FMR1 gene) and FBS_Q2 RNA (19 nt, position 1617–1635 within the FMR1 gene) whose sequences were predicted to adopt G-quadruplex structures (Table 1).
TABLE 1.
FMR1 oligonucleotide sequences used in this study

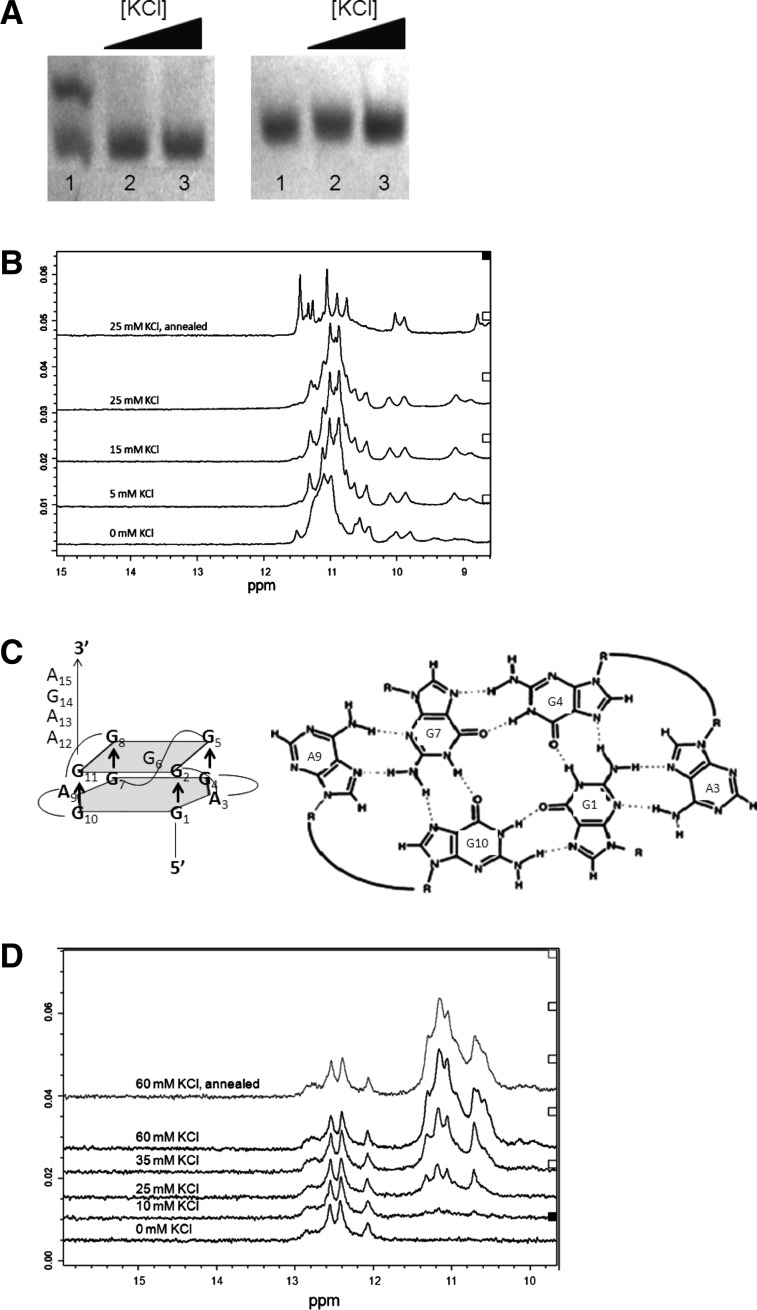
First, we used native PAGE to analyze the conformations of FBS_Q1 and FBS_Q2 in the presence of increasing KCl concentrations. Two bands exist for FBS_Q1 in the absence of KCl (Fig. 2A, left, lane 1), which collapse into a single band once KCl is titrated in the sample (Fig. 2A, left, lanes 2,3). In contrast, a single band exists for FBS_Q2 at all KCl concentrations investigated (Fig. 2A, right, lanes 1–3).
FIGURE 2.
(A) Native PAGE of FBS_Q1 RNA (left) and FBS_Q2 RNA (right) in the presence of increasing KCl concentrations: 10 µM RNA at 0 mM KCl (lane 1), 25 mM KCl (lane 2), and 50 mM KCl (lane 3). The gels were visualized by UV shadowing at 254 nm. (B) 1D 1H-NMR spectra of 1.1 mM FBS_Q1 RNA in 10 mM cacodylic acid, pH 6.5, in the presence of increasing KCl concentrations in the range of 0–25 mM. (C) Proposed structure of FBS_Q2 (left); A:(G:G:G:G):A hexad formed by two adenines that are in the plane with the G-quartet (right). The formation of a hexad stabilizes the amino protons that are involved in Hoogsteen base-pairing within the G quartet. (D) 1D 1H-NMR spectra of 1.1 mM FBS_Q2 RNA in 10 mM cacodylic acid, pH 6.5, in the presence of increasing KCl concentrations in the range of 0–60 mM.
Next, we used 1D 1H-NMR spectroscopy to investigate G-quadruplex formation within each of these two RNA sequences. For FBS_Q1 RNA, resonances corresponding to imino protons of guanines involved in G-quartet formation were observed in the region 10–11.5 ppm, even in the absence of KCl (Fig. 2B). Upon titration of KCl up to 25 mM, these imino proton resonances became sharper, indicating that the FBS_Q1 G-quadruplex is stabilized by K+ ions, and the annealing of the sample resulted in the sharpest resonances (Fig. 2B, top spectrum). Thus, all further experiments were performed with annealed FBS_Q1 RNA samples. No imino proton resonances corresponding to Watson-Crick base pairs were observed in the region 12–15 ppm, indicating the absence of an alternate duplex conformation. It is interesting to note the presence of two unusually sharp and downfield-shifted guanine amino resonances at 10.0 ppm and 9.9 ppm. A second set of sharp and downfield-shifted amino protons were observed at 8.8 and 8.6 ppm. Sharp and downfield-shifted guanine amino proton resonances observed in G-quadruplex forming sequences containing GGAGG stretches have been attributed to the presence of an A:(G:G:G:G):A hexad in the structure, in which two guanines have both their amino protons hydrogen bonded, one to a neighboring G in the G-tetrad and the second to the N7 of the adenine to form a hexad (Fig. 2C, right; Kettani et al. 2000; Liu et al. 2002; Mergny et al. 2006; Matsugami et al. 2008; Lipay and Mihailescu 2009). 1H–1H two-dimensional (2D) nuclear Overhauser enhancement NMR spectroscopy (NOESY) experiments revealed strong NOE cross peaks between the amino proton pairs at (10.0 ppm; 8.8 ppm) and (9.9 ppm; 8.6 ppm). Additionally, strong NOEs were observed between each of these sharp amino protons and their corresponding imino proton at (10.0 ppm; 11.5 ppm), (8.8 ppm; 11.5 ppm) and (9.9 ppm; 11.3 ppm), (8.6 ppm; 11.3 ppm), respectively (Supplemental Fig. 2A). This result is consistent with the presence of a hexad structure in which two guanines have both amino protons involved in hydrogen bonding. Next, we performed proton–deuterium exchange experiments, which revealed that the same set of imino protons (at 11.5 ppm and 11.3 ppm), and amino proton pairs (at 10.0 ppm, 8.8 ppm and 9.9 ppm, 8.6 ppm, respectively) for which we observed strong NOE cross peaks in the NOESY experiments, exchange very slowly (hours or days) compared to the rest of the imino protons present in the spectrum (minutes) (Supplemental Fig. 2B). This exchange with the solvent, which requires the base pairs opening, is slowed down considerably when both amino protons of some the guanines are involved in hydrogen bonds, as predicted in the hexad structure (Fig. 2C). Taken together, these results strongly suggest that the FBS_Q1 sequence adopts a G-quadruplex, which contains an A:(G:G:G:G):A hexad (Fig. 2C, left).
When FBS_Q2 was analyzed by 1D 1H-NMR spectroscopy, a set of resonances corresponding to the imino protons of Gs and Us involved in Watson-Crick base pairs were observed in the region 12–14 ppm at all KCl concentrations investigated, whereas the characteristic G-quadruplex imino proton resonances centered around 11 ppm were not observed until 25 mM KCl was titrated in the sample (Fig. 2D). The G-quadruplex imino proton resonances increased in intensity as KCl was titrated in FBS_Q2; however, the intensity of the imino proton resonances corresponding to Watson-Crick base pairs remained unaffected, suggesting that FBS_Q2 adopts a G-quadruplex structure stabilized by K+ ions but also an alternative conformation involving Watson-Crick base pairs.
To determine if the FMRP RGG box peptide binds to the G-quadruplex structures formed by FBS_Q1 and FBS_Q2 RNAs, native PAGE was performed on each sequence in the presence of KCl. Because the RGG box peptide has an overall positive charge due to its high arginine/lysine content, the RGG-RNA complex does not give rise to a tight band in the native PAGE; thus, the binding of RGG box peptide to RNA is monitored by the loss of the free RNA band in the gel. The FBS_Q1 free RNA band nearly disappears in the presence of a 2:1 ratio of RGG box: RNA (Fig. 3A, lane 4), whereas the intensity of the FBS_Q2 free RNA band does not change significantly in the presence of the same ratio of RGG box: RNA. This result indicates that the FMRP RGG box peptide does not have a strong binding affinity for FBS_Q2 RNA.
FIGURE 3.

(A) Native PAGE of 30 µM FBS_Q1 RNA in 25 mM KCl in the presence of increasing concentrations of FMRP RGG box peptide: 0 µM RGG box (lane 1), 15 µM RGG box (lane 2), 30 µM RGG box (lane 3), and 60 µM RGG box (lane 4). (B) Native PAGE of 30 µM FBS_Q2 RNA in 25 mM KCl in the presence of increasing concentrations of FMRP RGG box peptide: 0 µM RGG box (lane 1), 15 µM RGG box (lane 2), 30 µM RGG box (lane 3), and 60 µM RGG box (lane 4). The gels were visualized by UV shadowing at 254 nm.
FBSsh RNA forms a G-quadruplex structure
Next, we inquired if the A:(G:G:G:G):A hexad in FBS_Q1 was also formed when FBS_Q1 is located in the context of its larger surrounding sequence within FMR1 mRNA. Thus, we combined FBS_Q1 and FBS_Q2 RNAs to form a 42-nt fragment, named FBSsh RNA (nucleotides 1597–1638 within the FMR1 gene) (Table 1). The imino proton region of the 1D 1H-NMR spectrum of FBSsh RNA shows a broad resonance in the region between 10 and 11.5 ppm, corresponding to imino protons of guanines involved in G-quartet formation, even in the absence of KCl (Fig. 4A). Upon the addition of KCl up to 25 mM, more defined resonances develop under this broad envelope, indicating that the G-quadruplex structures of FBSsh RNA are further stabilized in the presence of K+ ions. Nonetheless, at all KCl concentrations investigated, the G-quadruplex imino proton resonances remain broad, which could be due to the presence of dynamic G-quadruplex structures in FBSsh. Interestingly, no sharp amino proton resonances, signatures of the A:(G:G:G:G):A hexad, were observed around 10 ppm, indicating that this structure is not formed in FBSsh but is likely induced in the isolated FBS_Q1 by the short length of the sequence. Additionally, in contrast to the 1D 1H-NMR spectra of FBS_Q2 RNA, no resonances appear in the Watson-Crick base-pair region of the 1H-NMR spectrum of FBSsh RNA, indicating that this sequence does not form any alternate conformations involving Watson-Crick base pairs.
FIGURE 4.
(A) 1D 1H-NMR spectra of 334 µM FBSsh RNA in 10 mM cacodylic acid, pH 6.5, in the presence of increasing KCl concentrations in the range of 0–25 mM. (B) Native PAGE of a fixed concentration of 10 µM FBSsh RNA in the presence of increasing concentrations of KCl in the range of 0–150 mM. (C) Native PAGE of increasing concentrations of FBSsh RNA in the range of 5–30 µM in the presence of 25 mM KCl. Gels were visualized by UV shadowing at 254 nm. (D) CD spectra of 10 µM FBSsh RNA in 10 mM cacodylic acid, pH 6.5, at 0 mM KCl, and 25 mM KCl.
Native PAGE was used next to analyze FBSsh RNA in the presence of different KCl concentrations. At 0 mM KCl, FBSsh RNA migrates in the gel as a single band (Fig. 4B, lane 1), which changes position with increasing salt concentrations up to 150 mM KCl (Fig. 4B, lanes 2–7). Similarly, a single band was observed when several concentrations of FBSsh RNA in the range of 5–30 µM were analyzed by native PAGE in the presence of a fixed concentration of 25 mM KCl (Fig. 4C, lanes 1–5).
To gain additional information about the fold of the G-quadruplexes within FBSsh RNA, we used CD spectroscopy. The CD spectrum of FBSsh RNA shows a negative band at 240 nm and a positive band at 265 nm that increase in intensity upon the titration of KCl from 0 to 25 mM (Fig. 4D). This result, which indicates that one or more parallel G-quadruplex structures are present in FBSsh RNA, is consistent with the 1H-NMR spectra of FBSsh RNA, which showed an increase in the intensities of the G-quartet imino proton resonances upon the titration of K+ ions.
Next, we used thermodynamic methods to determine if the FBSsh RNA folds into an intermolecular or intramolecular conformation. Specifically, the melting temperatures of the FBSsh G-quadruplex structures at various RNA concentrations in the presence of 25 mM KCl were measured by UV thermal denaturation. The melting temperature, Tm, for a species containing n number of strands depends on the total RNA concentration, cT (Equation 1; Hardin et al. 2000):
 |
(1) |
For intramolecular species, n = 1, and Tm is independent of cT (Equation 2):
 |
(2) |
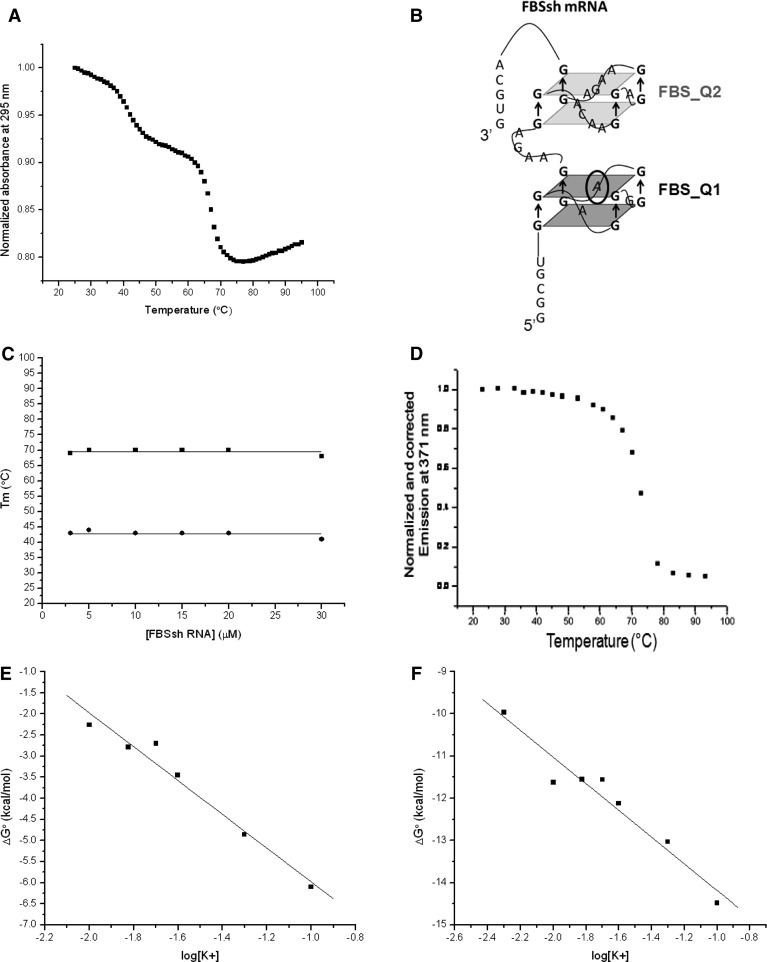
The experiments were recorded at 295 nm, a wavelength shown to be sensitive to G-quadruplex denaturation (Mergny et al. 1998). The UV thermal denaturation of FBSsh RNA at 295 nm in the presence of 25 mM KCl revealed two distinct hypochromic transitions characteristic of the denaturing of a G-quadruplex with melting temperatures of Tm ∼ 42°C and Tm ∼ 69°C, respectively (Fig. 5A), consistent with the presence of two distinct G-quadruplex structures in FBSsh RNA. Figure 5B shows a model of the two individual G-quadruplexes proposed to form within FBSsh RNA as the combination of FBS_Q1 RNA and FBS_Q2 RNA.
FIGURE 5.
(A) UV thermal denaturation curve of 10 µM FBSsh RNA in 10 mM cacodylic acid, pH 6.5, containing 25 mM KCl. (B) Model of FBSsh RNA showing the combination of the two G-quadruplexes formed by FBS_Q1 RNA and FBS_Q2 RNA. (C) Melting temperatures of both G-quadruplex structures of FBSsh RNA plotted against the RNA concentration. (D) Fluorescence spectroscopy thermal denaturation of 150 nM FBSsh_14AP RNA in 10 mM cacodylic acid, pH 6.5, and 25 mM KCl. (E) Plot of ΔG° as a function of the logarithm of K+ ion concentration for FBS_Q2. (F) Plot of ΔG° as a function of the logarithm of K+ ion concentration for FBS_Q1.
To determine if the FBSsh G-quadruplexes are intramolecular, increasing concentrations of FBSsh RNA in the range of 3–30 µM were UV thermally denatured in the presence of 25 mM KCl. As Figure 5C shows, the Tm of each transition remained constant at 42°C and 69°C, respectively, independent of the RNA concentration, indicating that each of the two G-quadruplex structures formed in FBSsh RNA is intramolecular (Fig. 5B,C; Equation 2).
Based solely on the UV thermal denaturation experiments, it is not possible to assign the two hypochromic transitions to the FBS_Q1 and FBS_Q2 G-quadruplexes. Thus, we constructed a fluorescently labeled RNA by replacing the adenine at position 14 of FBSsh RNA with 2-aminopurine (2AP) (FBSsh_14AP RNA, circled in Fig. 5B), which reports only on the melting of the first quadruplex in the sequence, FBS_Q1 RNA. 2AP is a highly fluorescent analog of adenine, which is sensitive to changes in its microenvironment (Serrano-Andres et al. 2006; Bharill et al. 2008). In contrast to UV thermal denaturation in which the change in absorbance as a function of temperature has contributions from all nucleotides in the sequence, thermal denaturation using fluorescence spectroscopy monitors only the changes in the steady-state fluorescence of the 2AP reporter, hence only the melting of the FBS_Q1 quadruplex. FBSsh_14AP RNA was thermally denatured in the presence of 25 mM KCl (Fig. 5D), each temperature point of the steady-state fluorescence being corrected to account for the dependence of the free fluorophore emission upon temperature. As expected, a single transition was observed for FBSsh_14AP, which was fitted with Equation 3, which assumes a two-state model:
 |
(3) |
yielding a Tm of 69.6 ± 0.1°C (Supplemental Fig. 3A). This result is in very good agreement with the Tm value of 68.8 ± 1.0°C determined when the UV thermal denaturation for transition 2 was fitted with Equation 3 (Supplemental Fig. 3B). Thus, we assign transition 2 to the melting of the FBS_Q1 G-quadruplex and transition 1 to the melting of the FBS_Q2 G-quadruplex within FBSsh RNA. These results are consistent with the literature, as it has been shown that the length of the loops connecting the G-quartet planes affects the stability of the G-quadruplex structure, with shorter loops forming tighter, more thermodynamically stable structures (Guedin et al. 2010). FBS_Q1 has single nucleotide connecting loops, and thus this structure was expected to have a higher melting point than FBS_Q2, which has longer connecting loops on two sides of its structure (Fig. 5B).
Next, the number of K+ ions coordinating each of the two G-quadruplexes was determined by performing UV thermal denaturation experiments at a fixed FBSsh RNA concentration of 10 µM and variable KCl concentrations in the range of 5–150 mM. The ΔG° of each of the two transitions, calculated using the thermodynamic parameters obtained by fitting the data at each KCl concentration with Equation 3, was plotted as a function of log[K+] according to Equation 4 (Fig. 5E,F):
 |
(4) |
The number of K+ ions coordinated to each G-quadruplex was determined from the negative slope of the plots (Fig. 5E,F) to be ∼3 K+ ions for each G-quadruplex in FBSsh RNA. It is well documented in the literature that K+ ions are present in the central ion channel of G-quadruplex structures at the center of two successive G quartets (Wei et al. 2012). However, more recently it has been shown that these ions also bind specifically to and stabilize the G-quadruplex loops (Gray and Chaires 2011; Wei et al. 2012). Since each of the FBS_Q1 and FBS_Q2 G-quadruplex structures are proposed to contain two G-quartet planes (Fig. 5B), it is possible that one K+ ion is present in the central channel of each structure, whereas the other two coordinate the surrounding loops. Alternately, the FBS_Q1 and FBS_Q2 structures might stack upon each other, creating an additional binding site for K+ within the central channel.
Taken together, our 1D 1H NMR spectroscopy, UV thermal denaturation, and CD spectroscopy results prove that FBSsh RNA folds into two intramolecular parallel G-quadruplex structures, in agreement with the proposal that two G-quadruplexes exist within the FBS region of the FMR1 mRNA (Schaeffer et al. 2001).
FBSsh RNA interactions with different FMRP isoforms
FMRP ISO1 has been shown to influence the FMR1 mRNA alternative splicing events at exon 15 by increasing the production of the minor isoforms created by the usage of the second and third acceptor site. Alternately, in the absence of FMRP ISO1, the mRNAs encoding for these minor isoforms are not detected in FMR1−/− mice that do not produce FMRP but still produce FMR1 mRNA (Didiot et al. 2008). These results clearly indicate that FMRP ISO1 regulates the alternative splicing of FMR1 through its interactions with the G-quadruplex forming FBS, but the exact mechanism by which this is accomplished has not been elucidated. In an effort to gain additional insight about this mechanism, we used fluorescence spectroscopy to obtain quantitative information about the interactions of FMRP with FBSsh RNA. Several FMRP isoforms were analyzed—FMRP ISO1, ISO2, and ISO3—created by the usage of the three different acceptor sites at exon 15 located in the close proximity of FBS, all of which contain an identical RGG box, the FMRP domain shown to bind with high affinity to G-quadruplex RNA (Darnell et al. 2001; Bole et al. 2008; Evans et al. 2012; Brackett et al. 2013). Additionally, since phosphorylation of FMRP has been shown to be important in mediating its translation regulator function (Ceman et al. 2003; Muddashetty et al. 2011), we inquired if this post-translational modification may also play a role in the regulation of the alternative splicing of FMR1 mRNA by analyzing the interactions with FBS of an isomimetic of phosphorylated FMRP, FMRP ISO1 S500D, created by mutating the serine at position 500 by aspartic acid (Tarrant and Cole 2009; Coffee et al. 2011; Muddashetty et al. 2011). Recombinant FMRP isoforms ISO1, ISO2, ISO3, and ISO1 S500D were expressed in E. coli and purified by nickel affinity column purification as described (Laggerbauer et al. 2001; Evans and Mihailescu 2010). In these studies, we have used FBSsh_14AP RNA, constructed by replacing the adenine at position 14 located within the FBS_Q1 G-quadruplex by 2AP (circled in Fig. 5B) described above.
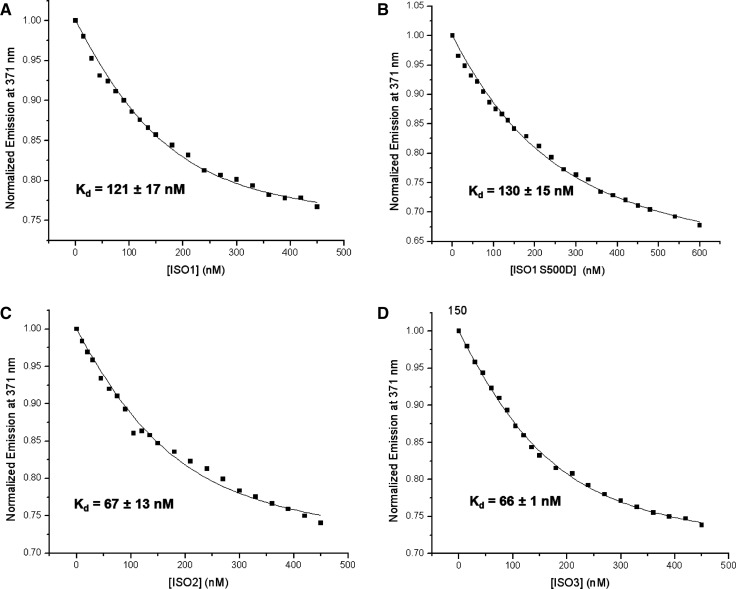
Each FMRP isoform—ISO1, ISO2, ISO3, and ISO1 S500D—was titrated into a fixed concentration of FBSsh_14AP RNA, monitoring the changes in the steady state fluorescence intensity of the 2AP reporter located within the FBS_Q1 quadruplex (Fig. 6A–D), and the dissociation constant, Kd, of the complex formed by each FMRP isoform with FBSsh_14AP RNA was determined from the fits of the binding curves with Equation 5 (Table 2; Materials and Methods).
FIGURE 6.
(A) FMRP ISO1 titrated into 150 nM FBSsh_14AP RNA in 10 mM cacodylic acid, pH 6.5, 750 nM BSA, and 25 mM KCl. (B) FMRP ISO1 S500D titrated into 150 nM FBSsh_14AP RNA in 10 mM cacodylic acid, pH 6.5, 750 nM BSA, and 25 mM KCl. (C) FMRP ISO2 titrated into 150 nM FBSsh_14AP RNA in 10 mM cacodylic acid, pH 6.5, 750 nM BSA, and 25 mM KCl. (D) FMRP ISO3 titrated into 150 nM FBSsh_14AP RNA in 10 mM cacodylic acid, pH 6.5, 750 nM BSA, and 25 mM KCl. Figures show a representative curve of the experiments performed in triplicate. The x-axis represents the total protein concentration [P]t from Equation 5 (Materials and Methods) for each FMRP isoform investigated.
TABLE 2.
FMRP isoform–FBSsh_14AP RNA complex dissociation constants

All FMRP isoforms investigated bind with low nM Kd values to FBSsh_14AP RNA (Table 2), consistent with previous findings that FMRP binds G-quadruplex forming RNA with high affinity (Evans et al. 2012). For example, the Kd = (120 ± 17) nM determined for FMRP ISO1 binding to FBSsh_14AP RNA is within error from the Kd = (104 ± 11) nM value determined for the binding of FMRP ISO1 to the G-quadruplex forming semaphorin 3F RNA, another FMRP target (Evans et al. 2012). FMRP ISO1 and ISO1 S500D bind with similar affinity to FBSsh_14AP RNA [Kd = (120 ± 17) nM versus Kd = (130 ± 15) nM], indicating that the binding of FMRP ISO1 to its own mRNA sequence is not affected by the mutation of S500D, which mimics the post-translational modification of phosphorylation. The phosphorylation of Ser500 has been shown to be important in the exertion of the FMRP translation regulator function on some of its target mRNAs (Ceman et al. 2003; Muddashetty et al. 2011); however, no quantitative data are available regarding the binding of phosphorylated/dephosphorylated FMRP ISO1 to such G-quadruplex forming mRNA targets. Thus, it remains to be seen whether the S500D mutation does not affect the RNA binding properties of FMRP only in the context of its function of splicing factor or if this trend is also valid for G-quadruplex forming mRNAs whose translation is regulated directly by FMRP. It is possible that the protein phosphorylation does not play a role in the context of the FMRP function as a splicing factor or, alternatively, that FMRP phosphorylation might be important for FMRP function as a translation regulator and/or splicing factor by regulating its interactions with protein partners rather than with the RNA. Another possibility that has to be considered is that although the S500D has been used to mimic the translation regulator function of phosphorylated FMRP, the D mutation might not be a good substitution for a phosphorylated serine in the context of RNA binding due to the charge difference.
The Kd values for FMRP ISO2 and ISO3 binding to FBSsh_14AP RNA were also within error of each other, yet lower than those measured for ISO1 and ISO1 S500D (Table 2), indicating that the truncation of the protein in the proximity of the RGG box results in higher affinity binding for FMR1 mRNA. The same trend was also reported for the FMRP binding to the G-quadruplex forming semaphorin 3F mRNA, an mRNA target whose translation is postulated to be regulated by FMRP (Evans et al. 2012). All FMRP isoforms investigated in this study have an identical RGG box; thus, the difference in binding affinity of FMRP ISO1, ISO1 S500D, and ISO2, ISO3 to FBSsh_14AP RNA could be due to subtle conformational changes in the region proximal to the RGG box, which might result in its better orientation for binding G-quadruplex RNA.
FMRP ISO1 binding to the G-rich region within FMR1 has been shown to influence the splicing events at exon 15 by increasing the production of the isoforms ISO2 and ISO3, which lack the major phosphorylation site at position 500, and decreasing the production of FMRP ISO1 (Didiot et al. 2008). It has also been shown that FMR1−/− knockout mice, which produce FMR1 mRNA but not any FMRP protein, do not produce FMR1 mRNA that has been alternatively spliced at exon 15 (Didiot et al. 2008). Our findings that FMRP ISO2 and ISO3 isoforms bind FBSsh RNA with higher affinity than FMRP ISO1 suggest the existence of a feedback autoregulatory loop for the production of the FMRP ISO2 and ISO3: once sufficient amounts of FMRP ISO2 and ISO3 are produced by FMRP ISO1 binding to the FBS RNA sequence, they can compete with FMRP ISO1 for binding, possibly shutting down their own production. This might have implications for the translation regulator function of FMRP. In its phosphorylated state, FMRP has been shown to be associated with stalled polyribosomes (Ceman et al. 2003), and in the case of the PSD-95 mRNA target, with the RISC complex (Muddashetty et al. 2011), suggesting that in this state FMRP prevents the translation of its mRNA targets. The FMRP dephosphorylation in response to synaptic input has been shown to rescue the PSD-95 mRNA translation (Muddashetty et al. 2011). FMRP ISO2 and ISO3 isoforms do not have the ability to be regulated by phosphorylation/dephosphorylation events. Our findings that they have higher affinity for G-quadruplex forming FMRP targets than FMRP ISO1 (this study; Evans et al. 2012) suggest a mechanism by which FMRP ISO2 and/or FMRP ISO3 could prolong the “on” state for translation of mRNA targets by competing with FMRP ISO1, which can turn “off” translation by becoming rephosphorylated. Thus, the levels of the different FMRP isoforms have to be tightly regulated in the cell as the expression of specific isoforms could control the timing of the translation of different FMRP mRNA targets.
The exact mechanism(s) by which FMRP ISO1 binding to the FBS region within FMR1 mRNA increases the usage of the second and third acceptor sites at exon 15 is not known. Several mechanisms that are not necessarily mutually exclusive could be envisioned: FMRP ISO1 might interact directly with splicing factors, recruiting them to FMR1 mRNA, or it might modulate the mRNA structure in such a way to make accessible/unaccessible the RNA binding sites for various splicing factors. We analyzed the entire 100-nt FBS sequence (1557–1658 within FMR1 mRNA, which contains FBSsh) with the SFmap software, which predicts and maps known splicing factor binding sites on RNA sequences (Akerman et al. 2009; Paz et al. 2010), identifying binding sites for several splicing factors (Supplemental Table 1). These splicing factors are members of the family of heterogeneous nuclear ribonucleoproteins (hnRNPs: hnRNP A1, hnRNP F, and hnRNP H1), or of the family of serine/arginine-rich proteins (SR) (SF2ASF and 9G8) or SR-like proteins (Tra2α and Tra2β). Both families of proteins are important in determining the splice site; in general, the hnRNP proteins antagonize SR and SR-like proteins’ function. In many cases, these splicing factors have overlapping RNA binding sites, competing for binding the mRNA, and the FBS site within FMR1 mRNA is no exception (Supplemental Table 1). The finding that some of these splicing binding sites are embedded within the G-quadruplex structure formed by FBSsh (Supplemental Table 1, bolded sequences) adds a new layer of complexity to the splicing mechanism, as when folded in this structure these sites are not available for binding by the corresponding splicing factors. An extensive search of the literature did not reveal any evidence about direct protein–protein interactions between FMRP and any of the splicing factors identified by the SFmap software (Supplemental Table 1), suggesting that FMRP might function to control the accessibility of the splicing factors to their RNA binding sites. Supporting this hypothesis are the findings that the binding of hnRNP F to its G-rich RNA binding site is limited by the stability of a G-quadruplex structure adopted by this sequence (Samatanga et al. 2012). Specifically, it has been shown that hnRNP F does not bind to the G-quadruplex structure formed by its G-rich binding site, and there is a competition between the formation of the G-quadruplex structure and of the hnRNP F protein–RNA complex. This situation could be similar in the case of the FMR1 mRNA, as the binding sites for hnRNP F are located within the FBS_Q1 quadruplex. Thus, in vivo, in the absence of FMRP, this sequence could adopt alternate structures (single-stranded versus G-quadruplex), whereas in the presence of FMRP, which binds G-quadruplex RNA, the G-quadruplex conformation could be favored, limiting the access of the hnRNP F splicing factor and ultimately influencing the splicing events in the proximity of this site within FMR1 mRNA.
MATERIALS AND METHODS
In vitro RNA synthesis
Unlabeled FBS_67 RNA, FBSsh RNA, FBS_Q1 RNA, and FBS_Q2 RNA (Table 1) oligonucleotides were synthesized in vitro off synthetic DNA templates (Trilink Biotechnologies, Inc.) using T7 RNA polymerase produced in-house (Milligan and Uhlenbeck 1989). Oligonucleotides were purified by 15% or 20% 8 M urea denaturing polyacrylamide gel electrophoresis (PAGE), eluted using electrophoretic elution, and extensively dialyzed against 10 mM cacodylic acid, pH 6.5 (Milligan and Uhlenbeck 1989). A fluorescent oligonucleotide was designed for FBSsh in which the highly fluorescent adenine analog 2AP replaced the adenine at position 14 of FBSsh RNA to construct FBSsh_14AP (Dharmacon, Inc.). Samples of FBS_67, FBS_Q1, and FBS_Q2 RNA sequences were annealed by heating at 95°C in the presence or absence of KCl and slow cooling for 30 min to 25°C. All samples of FBSsh and FBSsh_14AP RNA were prepared by incubation in the presence or absence of KCl for 20 min at 25°C.
Expression of recombinant FMRP isoforms
The recombinant pET21a-FMRP plasmid encoding FMRP isoform 1 (ISO1) fused with a C-terminal 6X histidine tag was a kind gift from Dr. Bernhard Laggerbauer (Institute of Pharmacology and Toxicology, Technische Universität München, Munich, Germany). The truncations of the FMR1 gene encoding for FMRP ISO1 to create the genes encoding for ISO2 and ISO3 were performed by GenScript USA, Inc. and confirmed by sequencing at the University of Pittsburgh Genomics and Proteomics Core (Evans et al. 2012). In order to recombinantly express, purify, and dialyze ISO2 and ISO3, we used previously developed protocols (Laggerbauer et al. 2001; Evans and Mihailescu 2010). In brief, plasmids were transformed into Rosetta2(DE3)pLysS E. coli cells. All media used in the cell growth consisted of Luria-Bertani (LB; Fisher Scientific) media containing 200 µg/mL ampicillin (AMP; MP Biomedical), and 15 µg/mL chloramphenicol (CHL; MP Biomedical). Cells were incubated at 37°C until the target O.D. of 0.8–1.0 was reached, and protein expression was induced by adding 1 mM isopropyl β-D-1-thiogalactopyranoside (IPTG) and incubating cells at 250 rpm for 12 h at 25°C. FMRP was purified using Ni-NTA Superflow resin (Qiagen) as described (Evans and Mihailescu 2010). Purified proteins were concentrated using dialysis tubing filled with polyethylene glycol (PEG) 20,000 and dialyzed into a buffer devoid of K+, Na+, or imidazole. Final protein buffer consisted of 20 mM Hepes, 5% glycerol, 1 mM EDTA, and 300 mM LiCl. The concentration of FMRP isoforms was determined at A280 by using the molar extinction coefficients of 46,370 M−1 cm−1 for ISO1 and ISO2 and 40,680 M−1 cm−1 for ISO3 (Evans and Mihailescu 2010).
FMRP ISO1 has been shown to be phosphorylated at three serine sites in exon 15, serine 500 being shown to be biologically relevant (Coffee et al. 2011; Muddashetty et al. 2011). We used a phosphomimetic version of FMRP ISO1 in which serine 500 was replaced by aspartic acid in order to mimic phosphorylation on serine 500 (Tarrant and Cole 2009). This phosphomimetic FMRP ISO1 S500D mutation was performed by GenScript USA, Inc. The concentration of FMRP ISO1 S500D was determined at A280 by using the molar extinction coefficients of 46,370 M−1 cm−1. The presence of the FMRP isoforms was analyzed using a 10% tris-glycine sodium dodecylsulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and visualized by Coomassie blue staining.
UV spectroscopy
The melting curves of the RNA oligonucleotides were collected on a Varian Cary 3E spectrophotometer outfitted with a peltier temperature control cell holder. Experiments were carried out in a 10-mm path length 200-µL quartz cuvette (Starna Cells) using samples prepared as follows in 10 mM cacodylic acid, pH 6.5 up to a final volume of 200 µL. FBS_67 RNA, FBS_Q1 RNA, and FBS_Q2 RNA were all prepared by boiling for 10 min and cooling to 25°C for 30 min. Samples of FBSsh RNA were prepared and incubated for 30 min at 25°C. Samples were heated from 25°C to 95°C at a rate of 0.2°C per minute with points recorded every 1°C. Samples and reference cells were covered with 200 µL mineral oil to prevent evaporation of aqueous solutions at high temperatures. Spectral absorbance changes were monitored at either 295 nm or 305 nm, wavelengths that have been identified as being sensitive to G-quadruplex dissociation, depending on the absorbance of the RNA at different concentrations (Mergny et al. 1998). All UV thermal denaturation experiments were performed in duplicate. In order to determine the inter- or intramolecular conformation of the G-quadruplex, melting temperatures of various concentrations of RNA between 3 and 50 µM were recorded.
Circular dichroism spectroscopy
CD experiments were performed on a J-810 spectrapolarimeter at 25°C using a 200-µL quartz cuvette with a 1-mm path length (Starna Cells). Parallel G-quadruplex formation of the RNA oligonucleotides at 10 µM in 10 mM cacodylic acid, pH 6.5, was observed as KCl was titrated in increasing concentrations from a 2 M stock, by monitoring the change in molar elipticity at 240 nm and 265 nm. Each spectrum was scanned seven times from 200 to 350 nm with a 1-sec response time and a 2-nm bandwidth.
1H NMR spectroscopy
The one-dimensional (1D) 1H-NMR spectra of the RNA oligonucleotides were acquired at 25°C on a 500 MHz Bruker AVANCE spectrometer. Water suppression was carried out by using the Watergate pulse sequence (Piotto et al. 1992). RNA oligonucleotides were prepared in 10 mM cacodylic acid, pH 6.5, in a 90:10 ratio of H2O:D2O. G-quadruplex formation was observed by titrating increasing concentrations of KCl form a 2 M stock to each sample and allowing samples to equilibrate for 10 min at 25°C. The D2O exchange experiment was carried out by lyophilizing the FBS_Q1 RNA sample and resuspending it in 250 µL pure D2O, with spectra measured in the interval 10 min to 5 d at 25°C.
1H-1H two-dimensional (2D) homonuclear NOESY experiments with a mixing time of 50 msec were acquired at 25°C in 90%:10% H2O:D2O with an RNA concentration of 1.1 mM in 10 mM cacodylic acid, pH 6.5 (Piotto et al. 1992; Sklenar et al. 1993). Data sets were processed using XWIN-NMR (Bruker).
RGG box peptide synthesis
The FMRP RGG box peptide with the sequence N-RRGDGRRRGGGGRGQGGRGRGGGFKGNDDHSR-C was chemically synthesized by the Peptide Synthesis Unit at the University of Pittsburgh Center for Biotechnology and Bioengineering.
Native polyacrylamide gel electrophoresis
FBS_67 RNA, FBS_Q1 RNA, and FBS_Q2 RNA were prepared by heating for 5 min at 95°C and allowing samples to cool for 30 min to 25°C. Samples of FBSsh RNA were prepared by incubation for 20 min at 25°C. Gels were run at 4°C from 40 to 75 V for 4 to 6 h. Gels (15% for FBSsh and FBS_67 RNA and 20% for FBS_Q1 and FBS_Q2 RNA) were visualized by UV shadowing at 254 nm (Hendry and Hannan 1996) or staining in ethidium bromide or SYBR gold and visualized on an AlphaImager (AlphaInnotech). FMRP RGG box binding experiments were performed by preparing the RNA sequences as described above and then incubating samples with RGG box peptide in different ratios for an additional 30 min at 25°C.
Fluorescence spectroscopy
Steady-state fluorescence spectroscopy experiments of FBSsh_14AP were performed on a Horiba Jobin Yvon Fluoromax-3 and accompanying software fitted with a 150 W ozone-free xenon arclamp. Experiments were performed in a 150-µL sample volume, 3-mm path-length quartz cuvette (Starna Cells). Excitation wavelength was set to 310 nm, the emission spectrum was recorded in the range of 330–450 nm, and the bandpass for excitation and emission monochromators were both set to 3 nm. For binding experiments, the temperature was set to 25°C, and increasing concentrations of FMRP isoforms (ISO1, ISO2, ISO3, and ISO1 S500D) were titrated in 15 nM increments to a fixed RNA concentration of 150 nM in 10 mM cacodylic acid, pH 6.5, and 25 mM KCl. A ratio of 5:1 BSA:RNA was added to the RNA sample before titration to reduce nonspecific binding. Emission values were corrected for free protein emission of both BSA and FMRP isoforms, and the data were normalized to free RNA fluorescence intensity monitored at 371 nm. Experiments were performed in triplicate, and the resulting averages of normalized intensity were plotted as a function of the FMRP isoform concentration and fitted to Equation 5. IF and IB represent steady-state fluorescence intensities of free and bound RNA, respectively; [RNA]t is the total fixed RNA concentration; and [P]t is the total FMRP isoform concentration. The protein–RNA complex dissociation constant, Kd, was determined for each experiment by fitting the binding curve with Equation 5:
 |
(5) |
These experiments were performed in triplicate for each FMRP isoform, and the reported errors represent the standard deviations of the dissociation constants determined from independent fits to the three measurements.
SUPPLEMENTAL MATERIAL
Supplemental material is available for this article.
ACKNOWLEDGMENTS
We thank Dr. Bernhard Laggerbauer (Institute of Pharmacology and Toxicology, Technische Universität München, Munich, Germany) for the plasmid pET21a-FMRP encoding for FMRP ISO1. This work was supported by the NIH Grants 2R15GM074660-02A1 and 9R15HD078017-03A1 to M.R.M.
REFERENCES
- Akerman M, David-Eden H, Pinter RY, Mandel-Gutfreund Y 2009. A computational approach for genome-wide mapping of splicing factor binding sites. Genome Biol 10: 30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ashley CT, Sutcliffe JS, Kunst CB, Leiner HA, Eichler EE, Nelson DL, Warren ST 1993. Human and murine FMR-1: Alternative splicing and translational initiation downstream of the CGG-repeat. Nat Genet 4: 244–251 [DOI] [PubMed] [Google Scholar]
- Bharill S, Sarkar P, Ballin JD, Gryczynski I, Wilson GM, Gryczynski Z 2008. Fluorescence intensity decays of 2-aminopurine solutions: Lifetime distribution approach. Anal Biochem 377: 141–149 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bole M, Menon L, Mihailescu MR 2008. Fragile X mental retardation protein recognition of G quadruplex structure per se is sufficient for high affinity binding to RNA. Mol Biosyst 4: 1212–1219 [DOI] [PubMed] [Google Scholar]
- Brackett DM, Qing F, Amieux PS, Sellers DL, Horner PJ, Morris DR 2013. Fmr1 transcript isoforms: Association with polyribosomes; regional and developmental expression in mouse brain. PLoS One 8: e58296. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown V, Jin P, Ceman S, Darnell JC, O'Donnell WT, Tenenbaum SA, Jin X, Feng Y, Wilkinson KD, Keene JD, et al. 2001. Microarray identification of FMRP-associated brain mRNAs and altered mRNA translational profiles in fragile X syndrome. Cell 107: 477–487 [DOI] [PubMed] [Google Scholar]
- Ceman S, O'Donnell WT, Reed M, Patton S, Pohl J, Warren ST 2003. Phosphorylation influences the translation state of FMRP-associated polyribosomes. Hum Mol Genet 12: 3295–3305 [DOI] [PubMed] [Google Scholar]
- Coffee RL Jr, Williamson AJ, Adkins CM, Gray MC, Page TL, Broadie K 2011. In vivo neuronal function of the fragile X mental retardation protein is regulated by phosphorylation. Hum Mol Genet 21: 900–915 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Crawford DC, Acuna JM, Sherman SL 2001. FMR1 and the fragile X syndrome: Human genome epidemiology review. Genet Med 3: 359–371 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dapic V, Abdomerovic V, Marrington R, Peberdy J, Rodger A, Trent JO, Bates PJ 2003. Biophysical and biological properties of quadruplex oligodeoxyribonucleotides. Nucleic Acids Res 31: 2097–2107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Darnell JC, Jensen KB, Jin P, Brown V, Warren ST, Darnell RB 2001. Fragile X mental retardation protein targets G quartet mRNAs important for neuronal function. Cell 107: 489–499 [DOI] [PubMed] [Google Scholar]
- Didiot MC, Tian Z, Schaeffer C, Subramanian M, Mandel JL, Moine H 2008. The G-quartet containing FMRP binding site in FMR1 mRNA is a potent exonic splicing enhancer. Nucleic Acids Res 36: 4902–4912 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Evans TL, Mihailescu MR 2010. Recombinant bacterial expression and purification of human fragile X mental retardation protein isoform 1. Protein Expr Purif 74: 242–247 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Evans TL, Blice-Baum AC, Mihailescu MR 2012. Analysis of the Fragile X mental retardation protein isoforms 1, 2 and 3 interactions with the G-quadruplex forming semaphorin 3F mRNA. Mol Biosyst 8: 642–649 [DOI] [PubMed] [Google Scholar]
- Gray RD, Chaires JB 2011. Linkage of cation binding and folding in human telomeric quadruplex DNA. Biophys Chem 159: 205–209 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guedin A, Gros J, Alberti P, Mergny JL 2010. How long is too long? Effects of loop size on G-quadruplex stability. Nucleic Acids Res 38: 7858–7868 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hardin CC, Perry AG, White K 2000. Thermodynamic and kinetic characterization of the dissociation and assembly of quadruplex nucleic acids. Biopolymers 56: 147–194 [DOI] [PubMed] [Google Scholar]
- Hawkins M, Boyle J, Wright KE, Elles R, Ramsden SC, O'Grady A, Sweeney M, Barton DE, Burgess T, Moore M, et al. 2011. Preparation and validation of the first WHO international genetic reference panel for Fragile X syndrome. Eur J Hum Genet 19: 10–17 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hendry P, Hannan G 1996. Detection and quantitation of unlabeled nucleic acids in polyacrylamide gels. Biotechniques 20: 258–264 [DOI] [PubMed] [Google Scholar]
- Jin P, Warren ST 2000. Understanding the molecular basis of fragile X syndrome. Hum Mol Genet 9: 901–908 [DOI] [PubMed] [Google Scholar]
- Jin P, Zarnescu DC, Ceman S, Nakamoto M, Mowrey J, Jongens TA, Nelson DL, Moses K, Warren ST 2004. Biochemical and genetic interaction between the fragile X mental retardation protein and the microRNA pathway. Nat Neurosci 7: 113–117 [DOI] [PubMed] [Google Scholar]
- Kettani A, Gorin A, Majumdar A, Hermann T, Skripkin E, Zhao H, Jones R, Patel DJ 2000. A dimeric DNA interface stabilized by stacked A·(G·G·G·G)·A hexads and coordinated monovalent cations. J Mol Biol 297: 627–644 [DOI] [PubMed] [Google Scholar]
- Laggerbauer B, Ostareck D, Keidel EM, Ostareck-Lederer A, Fischer U 2001. Evidence that fragile X mental retardation protein is a negative regulator of translation. Hum Mol Genet 10: 329–338 [DOI] [PubMed] [Google Scholar]
- Lipay JM, Mihailescu MR 2009. NMR spectroscopy and kinetic studies of the quadruplex forming RNA r(UGGAGGU). Mol Biosyst 5: 1347–1355 [DOI] [PubMed] [Google Scholar]
- Liu H, Matsugami A, Katahira M, Uesugi S 2002. A dimeric RNA quadruplex architecture comprised of two G:G(:A):G:G(:A) hexads, G:G:G:G tetrads and UUUU loops. J Mol Biol 322: 955–970 [DOI] [PubMed] [Google Scholar]
- Matsugami A, Mashima T, Nishikawa F, Murakami K, Nishikawa S, Noda K, Yokoyama T, Katahira M 2008. Structural analysis of r(GGA)4 found in RNA aptamer for bovine prion protein. Nucleic Acids Symp Ser (Oxf) 52: 179–180 [DOI] [PubMed] [Google Scholar]
- Menon L, Mihailescu MR 2007. Interactions of the G quartet forming semaphorin 3F RNA with the RGG box domain of the fragile X protein family. Nucleic Acids Res 35: 5379–5392 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Menon L, Mader SA, Mihailescu MR 2008. Fragile X mental retardation protein interactions with the microtubule associated protein 1B RNA. RNA 14: 1644–1655 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mergny JL, Phan AT, Lacroix L 1998. Following G-quartet formation by UV-spectroscopy. FEBS Lett 435: 74–78 [DOI] [PubMed] [Google Scholar]
- Mergny JL, De Cian A, Amrane S, Webba da Silva M 2006. Kinetics of double-chain reversals bridging contiguous quartets in tetramolecular quadruplexes. Nucleic Acids Res 34: 2386–2397 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Milligan JF, Uhlenbeck OC 1989. Synthesis of small RNAs using T7 RNA polymerase. Methods Enzymol 180: 51–62 [DOI] [PubMed] [Google Scholar]
- Morton JE, Bundey S, Webb TP, MacDonald F, Rindl PM, Bullock S 1997. Fragile X syndrome is less common than previously estimated. J Med Genet 34: 1–5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Muddashetty RS, Nalavadi VC, Gross C, Yao X, Xing L, Laur O, Warren ST, Bassell GJ 2011. Reversible inhibition of PSD-95 mRNA translation by miR-125a, FMRP phosphorylation, and mGluR signaling. Mol Cell 42: 673–688 [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Donnell WT, Warren ST 2002. A decade of molecular studies of fragile X syndrome. Annu Rev Neurosci 25: 315–338 [DOI] [PubMed] [Google Scholar]
- Paz I, Akerman M, Dror I, Kosti I, Mandel-Gutfreund Y 2010. SFmap: A web server for motif analysis and prediction of splicing factor binding sites. Nucleic Acids Res 38: 281–285 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pieretti M, Zhang FP, Fu YH, Warren ST, Oostra BA, Caskey CT, Nelson DL 1991. Absence of expression of the FMR-1 gene in fragile X syndrome. Cell 66: 817–822 [DOI] [PubMed] [Google Scholar]
- Piotto M, Saudek V, Sklenar V 1992. Gradient-tailored excitation for single-quantum NMR spectroscopy of aqueous solutions. J Biomol NMR 2: 661–665 [DOI] [PubMed] [Google Scholar]
- Samatanga B, Dominguez C, Jelesarov I, Allain FH 2012. The high kinetic stability of a G-quadruplex limits hnRNP F qRRM3 binding to G-tract RNA. Nucleic Acids Res 41: 2505–2516 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schaeffer C, Bardoni B, Mandel JL, Ehresmann B, Ehresmann C, Moine H 2001. The fragile X mental retardation protein binds specifically to its mRNA via a purine quartet motif. EMBO J 20: 4803–4813 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schaeffer C, Beaulande M, Ehresmann C, Ehresmann B, Moine H 2003. The RNA binding protein FMRP: New connections and missing links. Biol Cell 95: 221–228 [DOI] [PubMed] [Google Scholar]
- Serrano-Andres L, Merchan M, Borin AC 2006. Adenine and 2-aminopurine: Paradigms of modern theoretical photochemistry. Proc Natl Acad Sci 103: 8691–8696 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Siomi H, Siomi MC, Nussbaum RL, Dreyfuss G 1993. The protein product of the fragile X gene, FMR1, has characteristics of an RNA-binding protein. Cell 74: 291–298 [DOI] [PubMed] [Google Scholar]
- Siomi MC, Higashijima K, Ishizuka A, Siomi H 2002. Casein kinase II phosphorylates the fragile X mental retardation protein and modulates its biological properties. Mol Cell Biol 22: 8438–8447 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sittler A, Devys D, Weber C, Mandel JL 1996. Alternative splicing of exon 14 determines nuclear or cytoplasmic localisation of fmr1 protein isoforms. Hum Mol Genet 5: 95–102 [DOI] [PubMed] [Google Scholar]
- Sklenar V, Peterson RD, Rejante MR, Feigon J 1993. Two- and three-dimensional HCN experiments for correlating base and sugar resonances in 15N,13C-labeled RNA oligonucleotides. J Biomol NMR 3: 721–727 [DOI] [PubMed] [Google Scholar]
- Stetler A, Winograd C, Sayegh J, Cheever A, Patton E, Zhang X, Clarke S, Ceman S 2006. Identification and characterization of the methyl arginines in the fragile X mental retardation protein Fmrp. Hum Mol Genet 15: 87–96 [DOI] [PubMed] [Google Scholar]
- Tarrant MK, Cole PA 2009. The chemical biology of protein phosphorylation. Annu Rev Biochem 78: 797–825 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tassone F, Hagerman RJ, Iklé DN, Dyer PN, Lampe M, Willemsen R, Oostra BA, Taylor AK 1999. FMRP expression as a potential prognostic indicator in fragile X syndrome. Am J Med Genet 84: 250–261 [PubMed] [Google Scholar]
- Verkerk AJ, de Graaff E, De Boulle K, Eichler EE, Konecki DS, Reyniers E, Manca A, Poustka A, Willems PJ, Nelson DL, et al. 1993. Alternative splicing in the fragile X gene FMR1. Hum Mol Genet 2: 1348. [DOI] [PubMed] [Google Scholar]
- Wei D, Parkinson GN, Reszka AP, Neidle S 2012. Crystal structure of a c-kit promoter quadruplex reveals the structural role of metal ions and water molecules in maintaining loop conformation. Nucleic Acids Res 40: 4691–4700 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williamson JR 1994. G-quartet structures in telomeric DNA. Annu Rev Biophys Biomol Struct 23: 703–730 [DOI] [PubMed] [Google Scholar]
- Williamson JR, Raghuraman MK, Cech TR 1989. Monovalent cation-induced structure of telomeric DNA: The G-quartet model. Cell 59: 871–880 [DOI] [PubMed] [Google Scholar]
- Zalfa F, Eleuteri B, Dickson KS, Mercaldo V, De Rubeis S, di Penta A, Tabolacci E, Chiurazzi P, Neri G, Grant SG, et al. 2007. A new function for the fragile X mental retardation protein in regulation of PSD-95 mRNA stability. Nat Neurosci 10: 578–587 [DOI] [PMC free article] [PubMed] [Google Scholar]