Background: Fascin is a pro-metastasis actin bundling protein overexpressed in basal-like breast cancer.
Results: GATA3 abrogates TGFβ and Smad4-mediated fascin overexpression by abolishing the binding of Smad4 to fascin promoter.
Conclusion: GATA3 is a novel suppressor of the canonical TGFβ-Smad signaling pathway.
Significance: These findings provide mechanistic insight into how TGFβ-mediated invasion and metastasis are differentially regulated in different subgroups of breast cancer.
Keywords: Breast Cancer, Cell Invasion, Cytoskeleton, GATA, Metastasis, SMAD Transcription Factor
Abstract
Transforming growth factor β (TGFβ) is a potent and context-dependent regulator of tumor progression. TGFβ promotes the lung metastasis of basal-like (but not the luminal-like) breast cancer. Here, we demonstrated that fascin, a pro-metastasis actin bundling protein, was a direct target of the canonical TGFβ-Smad4 signaling pathway in basal-like breast cancer cells. TGFβ and Smad4 induced fascin overexpression by directly binding to a Smad binding element on the fascin promoter. We identified GATA3, a transcription factor crucial for mammary gland morphogenesis and luminal differentiation, as a negative regulator of TGFβ- and Smad4-induced fascin overexpression. When ectopically expressed in basal-like breast cancer cells, GATA-3 abrogated TGFβ- and Smad4-mediated overexpression of fascin and other TGFβ response genes, invadopodium formation, cell migration, and invasion, suggesting suppression of the canonical TGFβ-Smad signaling axis. Mechanistically, GATA3 abrogated the canonical TGFβ-Smad signaling by abolishing interactions between Smad4 and its DNA binding elements, potentially through physical interactions between the N-terminal of GATA3 and Smad3/4 proteins. Our findings provide mechanistic insight into how TGFβ-mediated cell motility and invasiveness are differentially regulated in breast cancer.
Introduction
The transcriptomic output of the transforming growth factor β (TGFβ)-Smad signaling axis is dictated by the context of Smad co-suppressors and co-activators in the cell (1, 2). As a consequence, TGFβ-Smad4 signaling can either promote or suppress tumor progression depending on tumor type and stage (3). In estrogen receptor (ER)2-negative breast cancer patients the overexpression of the type II TGFβ receptor is associated with worse overall survival, and the up-regulation of TGFβ signature genes promotes lung metastasis (4, 5). On the other hand, TGFβ signaling has no effect on the prognosis among ER positive breast cancer patients (4, 5). It is not fully understood how TGFβ-mediated tumor metastasis is differentially regulated among breast cancer subtypes.
GATA3 is a member of the GATA family of zinc finger transcription factors required for the development and morphogenesis of the mammary gland (6–8). GATA3 expression levels are high in well differentiated, luminal breast cancer (ER- and/or progesterone receptor (PR)-positive, Her2 (human EGF receptor 2)-positive or -negative) but suppressed in poorly differentiated basal-like subgroup (ER, PR, and Her2 triple negative) (6, 9–11). The targeted deletion of GATA3 in mouse mammary gland results in the expansion of luminal progenitor cells, and the ectopic expression of GATA3 in mammary stem cells induces luminal differentiation (7), suggesting that GATA3 is critical to maintaining the differentiation of the luminal lineage. Ectopic expression of GATA3 in basal-like breast cancer cells caused reversal of epithelial-to-mesenchymal transition and suppresses the metastasis of breast cancer to the lung (12–15). There is increasing evidence suggesting that re-introduction of GATA3 in basal-like breast cancer cells induces differentiation to luminal-like phenotype (6, 10, 12, 16); however, the regulation of TGFβ-mediated invasion and metastasis by GATA3-mediated differentiation is not clear.
Fascin is an actin bundling protein that plays a critical role in lung metastasis of basal-like breast cancer (17, 18). Fascin promotes the metastasis of breast and other cancers by facilitating membrane protrusions such as filopodia and invadopodia during cancer cell migration and invasion (19–22). We recently reported that fascin expression is up-regulated by the canonical TGFβ-Smad3-Smad4 signaling pathways in poorly differentiated cancer cells but not in well differentiated polygonal-shaped cancer cells (19). TGFβ-mediated filopodia formation and cancer cell invasion were almost abrogated when fascin was depleted with shRNA, suggesting that fascin is critical for TGFβ-mediated invasion and metastasis. However, it is not clear how the differentiation state of cancer cells affects TGFβ-induced fascin expression. Here we demonstrate that Smad4 directly promotes fascin transcription by binding to a Smad binding site on the fascin promoter. The binding of Smad4 to the fascin promoter is abrogated by ectopic GATA3, potentially through direct interactions between GATA3 N-terminal and Smad3/4 proteins. Importantly, ectopic GATA3 abrogates Smad4-mediated invadopodia formation, Matrigel invasion, and the transcription of direct or indirect TGFβ response genes, suggesting that ectopic GATA3 inhibits the global response to the canonical TGFβ-Smad signaling axis. Our data imply that high expression levels of GATA3 in ER-positive, luminal-like breast cancer might be responsible for the lack of TGFβ-mediated metastasis in this subtype of breast cancer.
EXPERIMENTAL PROCEDURES
Cell Lines and Culture Medium
MDA-MB-231, MDA-MB-468, MCF-7, BT474, and HEK293 cells were all cultured in DMEM supplemented with 10% FBS and penicillin/streptomycin.
Antibodies
The following antibodies were used in this study: anti-fascin (#sc-21743), anti-Smad4 (#sc-7154), and anti-GATA3 (#sc-22206) were from Santa Cruz Biotechnology; anti-Smad3 (#9523) was from Cell Signaling; anti-HA (#SAB4300603) and anti-GAPDH (#G8795) were from Sigma. Anti-Cortactin (#05-180) was from Millipore.
Plasmids and DNA Constructs
pBabe-puro-GATA3 (plasmid #1286) and Lpcx-FLAG-Smad3 (plasmid #12638) were obtained from Addgene. LPCX-FLAG-Smad4 was generated by inserting PCR products between SalI and ClaI. pLPCX-HA-GATA3 was subcloned by inserting PCR products between BglII and EcoRI.
Retrovirus and Stable Cell Line Preparation
Vesicular stomatitis virus-G pseudotyped retroviruses were prepared and concentrated as described previously (23). Briefly, HEK293 cells (in 10-cm dishes) were co-transfected with retrovirus vector encoding desired cDNA (5 μg), retrovirus packaging plasmids encoding gag-pol (5 μg), and vesicular stomatitis virus-G (5 μg) using PEI reagent. The retroviruses in the supernatant were harvested and concentrated by centrifugation. To generate stable cell lines, MDA-MB-231 cells were infected with retrovirus and selected with appropriate antibiotics for 1–2 weeks before being used for experiments.
Luciferase Assay
The full-length human fascin promoter has been described previously (24). The luciferase reporter constructs were generated by inserting full-length or truncated human fascin promoter into pGL3 basic vector (Promega) between XhoI and HindIII. To perform dual luciferase reporter assay, 12,000 MDA-MB-231 cells were seeded in 12-well plates and cultured overnight. Cells were transfected with 1 μg/well fascin promoter reporter together with 100 ng/well Renilla luciferase construct (pRL-TK) using Lipofectamine 2000. 24 h after transfection, the cells were treated with 5 ng/ml TGFβ for 12 h before lysis. Cell lysates were subjected to dual reporter luciferase assays according to the manufacturer's instructions (Promega).
Chromatin Immunoprecipitation (ChIP) Assays
ChIP assays were performed according to a previously reported protocol with minor modification (25). 1 × 107 MDA-MB-231 Cells were treated with control medium or medium containing 5 ng/ml TGFβ for 5 h and then fixed with 1% formaldehyde (Sigma F8775) for 10 min at room temperature. The cells were scraped, washed in ice-cold PBS, and centrifuged at 1500 × g at 4 °C for 5 min. Subsequently, the pellet was resuspended in cell lysis buffer (44 mm Tris-HCl (pH 8.1), 1% SDS, and 1 mm EDTA (pH 8.0)). The cells were sonicated 3 times for 15 s each. Subsequently, the cell lysates were centrifuged at 10,000 × g at 4 °C for 15 min. An aliquot of the sheared chromatin was used as the input for the ChIP assay. The remainder of the chromatin was diluted with ChIP dilution buffer (16 mm Tris-HCl (pH 8.1), 250 mm NaCl, 0.1% SDS, 1% Triton-X-100, and 1.2 mm EDTA) and rotated at least 4 h with primary anti-Smad4 antibody at 4 °C with mouse IgG as control. 60 μl of 1:1 protein G-Sepharose were added to the immune complexes, and the mixture was rotated at 4 °C for 2 h. The beads were washed 5 times with ChIP dilution buffer and eluted with ChIP elution buffer (0.1 m sodium bicarbonate, 1% SDS, 5 mm NaCl). The cross-links were reversed by incubation at 65 °C for 4 h. DNA was isolated by ethanol precipitation. The associated proteins with the DNA were digested with 50 μg of proteinase K at 37 °C for 30 min. DNA was purified by the phenol:chloroform extraction method followed by ethanol precipitation. Purified DNA was resuspended in 30 μl of water and assayed with semi-quantitative PCR.
Immunoprecipitation
HEK293 cells were transfected with 2.0 μg of pLPCX-HA-GATA3 or LPCX-HA-gata3 and of LPCX-FLAG-SMAD3/4. 36 h after transfection, cells were harvested and lysed in a buffer containing 50 mm Tris (pH 8.0), 150 mm NaCl, 0.5% sodium deoxycholate, 1% Triton X-100, and 1 mm phenylmethylsulfonyl fluoride. The lysate was incubated with anti-FLAG antibody (M2)-conjugated agarose beads (Sigma) for 2 h at 4 °C. The beads were washed extensively in a buffer containing 50 mm Tris (pH 8.0), 150 mm NaCl, 1 mm PMSF, and 1% Nonidet P-40. Bound proteins were boiled in 1× protein sample buffer for 5 min and then resolved on SDS-PAGE and detected with Western blot assay.
For TGFβ treated coimmunoprecipitation experiment, MDA-MB-231 cells overexpressing Smad4 or Smad4 and GATA3 were seeded on 60-mm dishes overnight. The cells were treated with 5 ng/ml TGFβ for 6 h and lysed in a lysis buffer containing 1 mm NaVO4 and 5 mm NaF. The lysate were incubated with anti-FLAG antibody (M2)-conjugated agarose beads for 2 h at 4 °C. The beads were washed extensively, and the bound proteins were eluted by boiling in 1× SDS sample buffer for 5 min and then subjected to Western blotting.
Quantitative Real-time (qPCR)
Total RNA was extracted from cultured cells using TRIzol reagent (Invitrogen), and the RNA was treated with DNase for 15 min at 37 °C. The reverse transcription was performed using the iScript cDNA synthesis kit (Bio-Rad). The qRT-PCR assay was carried out with the Applied Biosystems 7900HT fast real-time PCR system using Applied Biosystems SYBR Green PCR master mix. Primers for qRT-PCR are shown in Table 1. All reactions were performed in triplicate, and the experiments were repeated at least three times.
TABLE 1.
Primers used for ChIP and q-PCR
F, forward; R, reverse.
| Primer names | Primer sequence |
|---|---|
| Fascin CHIP-443 F | 5′-GCCTCTTTCCCTCGTCTG-3′ |
| Fascin CHIP-282 R | 5′-CTAGGAGCAAGGACGAGC-3′ |
| p21 CHIP-F | 5′-GGTGGTGGTGAGCTAGAAG-3′ |
| p21 CHIP-R | 5′-CCTCTGCTTTCAGGCATTTC-3′ |
| Fascin qPCR-F | 5′-TGCTACTTTGACATCGAGTGG-3′ |
| Fascin qPCR-R | 5′-ATGAGCTTCATGAGGAAGAGC-3′ |
| E-cadherin qPCR-F | 5′-TCTCTGCTCGTGTTTGACTATG-3′ |
| E-cadherin qPCR-R | 5′-TGCCCCATTCGTTCAAGTAG-3′ |
| ANGPTL-4 qPCR-F | 5′-AGCCATTCCAACCTCAACG-3′ |
| ANGPTL-4 qPCR-R | 5′-ATGGGCTGGATCAACATGG-3′ |
| N-cadherin qPCR-F | 5′-GACCAGGACTATGACTTGAGC-3′ |
| N-cadherin qPCR-R | 5′-TGGGTCTTTCATCCATTCGTC-3′ |
| Vimentin qPCR-F | 5′-CGTGAATACCAAGACCTGCTC-3′ |
| Vimentin qPCR-R | 5′-GGAAAAGTTTGGAAGAGGCAG-3′ |
| p21 qPCR-F | 5′-TGTCACTGTCTTGTACCCTTG-3′ |
| p21 qPCR-R | 5′-ATCTGTCATGCTGGTCTGC-3′ |
| GAPDH qPCR-F | 5′-AGAAGGCTGGGGCTCATTTG-3′ |
| GAPDH qPCR-R | 5′-AGGGGCCATCCACAGTCTTC-3′ |
Invadopodia Assay
The invadopodia activity assay protocol was adapted according to Artym et al. (26) by plating cancer cells onto glass coverslips coated with a thin film of fluorescent gelatin. The immunofluorescence staining was performed as previously described (22, 27, 28). Briefly, 80,000 MDA-MB-231 cells were plated on Texas Red-labeled gelatin-coated glass coverslips (18 mm). After a 24-h incubation, cells were fixed in 4% paraformaldehyde and permeabilized with antibody diluting buffer (2% BSA, 0.1% Triton X-100 in PBS) and followed by incubation with Alexa Fluor 488 phalloidin for 30 min. An extensive wash was carried out between each step. The coverslips were then fixed onto slides and imaged using a Zeiss fluorescence microscope. To quantify gelatin degradation, fluorescent micrographs were taken from 3–5 random fields for each group. The total gelatin degradation area in each field was measured using ImageJ software by selecting regions of interest. The degradation area per cell for each field was derived by dividing total area with total number of cells present in the field.
To determine the effects on invadopodium formation, cells were plated on coverslips coated with unlabeled gelatin and stained for actin (using phalloidin) and cortactin (using anti-cortactin antibody, 1:1000 dilution). Invadopodia are defined as actin- and cortactin-rich dots on the ventral side of the cells. Cells with three or more invadopodia are defined as invadopodia-positive and otherwise as invadopodia-negative.
Cell Migration and Invasion Assay
Cells (1 × 105) suspended in starvation medium were added to the upper chamber of an insert (for migration assay) or a Matrigel-coated insert (for invasion assay), and the insert was placed in a 24-well dish containing medium with or without serum. Cell migration assays were carried out for 4 h, and invasion assays were carried out for 12 h. Cells were then fixed with 3.7% formaldehyde and stained with crystal violet-staining solution; cells on the upper side of the insert were removed with a cotton swab. Three randomly selected fields (10× objectives) on the lower side of the insert were photographed, and the cells on the lower surface of the insert were counted.
RESULTS
Fascin Is a Direct TGFβ-Smad Target Gene
When Smad3 or Smad4 was ectopically expressed in two basal-like breast cancer cells (MDA-MB-231 and MDA-MB-468), fascin protein levels increased from 2 to >30-fold, phenocopying the TGFβ-induced fascin overexpression in this poorly differentiated subtype of breast cancer (Fig. 1, A and B). In contrast, ectopic Smad4 had no detectable effects on fascin protein levels in three luminal-like breast cancer cell lines (MCF-7, BT-474, and T47D) (Fig. 1B). The lack of Smad4-induced fascin expression in luminal breast cancer cells suggested that Smad co-factors might play a role in this regulation.
FIGURE 1.
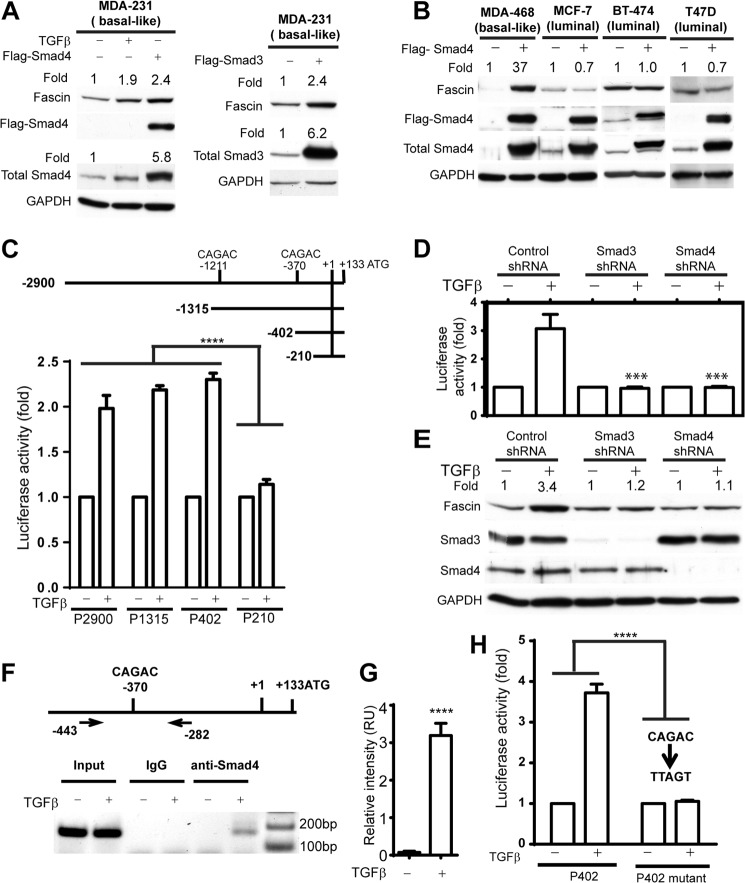
Fascin is a direct target gene of Smad4 in basal-like breast cancer cells. A, Western blot showing that TGFβ treatment (5 ng/ml, 72-h treatment in growth medium) and ectopically expressed FLAG-Smad4 and FLAG-Smad3 increased fascin protein levels in basal-like MDA-MB-231 cells. B, Western blot showing that ectopic Smad4 induced fascin up-regulation in basal-like MDA-MB-468 cells but not in luminal-like MCF-7, BT474, and T47D cells. The increase in total Smad4 and Smad3 levels after enforced expression were determined through immunoblotting using respective antibodies, as shown in A and B. C, luciferase assay indicating TGFβ-activated fascin promoter. Upper panel, schematic diagram of full-length and truncated fascin promoter used in the experiment; −1211 and −370 Smad binding sites are indicated. Lower panel, activation of full-length or truncated fascin promoter by TGFβ treatment, as determined by luciferase activity assay. D and E, effects of Smad3 and Smad4 knockdown on the transactivation of fascin promoter (P402) by TGFβ, as determined by luciferase assay (D) or on TGFβ (5 ng/ml, 72-h treatment)-induced fascin overexpression, as determined by Western blotting (E). The knockdown of Smad3 and Smad4 protein levels in MDA-MB-231 cells was determined through Western blotting, as shown in E. F, ChIP assay using anti-Smad4 antibody showed that direct binding of Smad4 to fascin promoter in MDA-MB-231 cells was increased after TGFβ (5 ng/ml) treatment. G, quantification of the results in F through densitometry using ImageJ software. H, luciferase assay showing that mutation of the −370 Smad binding site abolished the activation of fascin promoter by TGFβ. p < 0.001 (***) and p < 0.0001 (****), respectively, as determined by one-way ANOVA (C and D) or Student's t test. The relative expression levels of fascin and/or Smad3/4 in A, B, and E were determined through densitometry using ImageJ software. All the expression level changes were relative to the untreated control within the same group (as indicated by the black line).
There are two potential Smad binding sites (at −1211 and −370, respectively) on the fascin promoter (Fig. 1C). To determine whether Smad4 directly regulates fascin expression by binding to the fascin promoter, we constructed a series of luciferase reporters containing full-length (P2900) or truncated fascin promoters (P1315, P402, and P210) (Fig. 1C). The truncation of the promoter region containing the −1211 Smad binding element had no noticeable effect on the activation of fascin promoter by TGFβ (Fig. 1C). However, the activation of fascin promoter by TGFβ was abolished in the P210 reporter, which contains the core fascin promoter elements (29) but neither of the two CAGAC Smad binding elements (Fig. 1C). To investigate whether Smad3 and Smad4 were required for the transactivation of fascin promoter by TGFβ, we knocked down Smad3 and Smad4 in MDA-MB-231 cells using shRNA. Smad3 and Smad4 knockdown abrogated the activation of P402 luciferase reporter by TGFβ as well as TGFβ-induced overexpression of fascin protein (Fig. 1, D and E).
Our luciferase reporter experiments suggested that the Smad transcription complex might promote fascin expression by directly binding to the −370 Smad binding sites. To determine if this was the case, we performed ChIP experiments using anti-Smad4 antibody. The Smad4 antibody successfully precipitated the fascin promoter in TGFβ-treated MDA-MB-231 cells but not in the control cells, suggesting that Smad4 directly interacted with the −370 Smad binding element upon TGFβ activation (Fig. 1, F and G). To further confirm that the −370 site is required for TGFβ to activate fascin promoter activity, we mutated the −370 Smad binding element from CAGAC to TTAGT in the P402 reporter. The mutation almost completely abrogated TGFβ-induced luciferase expression. Taken together, our data suggest that fascin is a novel direct target gene of the canonical TGFβ-Smad signaling pathway. The activation of fascin expression by TGFβ is dependent on the binding of Smad transcription complex to the −370 CAGAC site on the fascin promoter.
GATA3 Negatively Correlates with Fascin Expression in Breast Cancer Patients
We noted that TGFβ and Smad4 only induced fascin overexpression in basal-like but not in luminal-like breast cancer cells (Fig. 1, A and B). It was also noted that most of the cancer cells that respond to TGFβ-induced fascin transcription also have mesenchymal-like morphology, whereas the non-responsive cells mostly adopt epithelial-like polygonal shapes (19). These observations led us to postulate that epithelial-to-mesenchymal (EMT) transcription factors might be involved in regulating the TGFβ-mediated fascin overexpression in breast cancers. We examined the correlation between fascin levels and the expression levels of a panel of EMT transcription factors (including SNAI1, SNAI2, ZEB1, ZEB2, KLF8, TWIST1, TWIST2, TCF4, SIX1, FOXC2, GRHL2, ELF3, ELF5, GATA1–6, and HIF1A) in two cohorts of breast cancer samples (the MSKCC cohort and the Stockholm cohort) (18, 30). Pearson correlation coefficient (r) and probability (p) values of the correlation between fascin probe sets and each of the probe sets for the EMT transcription factors were calculated. The SNAI2 and GATA3 probe sets significantly correlate with the two fascin probe sets in both cohorts (supplemental Table 1, Fig. 2A), suggesting that these two EMT transcription factors might be involved in regulating fascin expression in breast cancer patients. We decided to focus on GATA3 due to its robust correlation with fascin. When the 99 breast cancer patients in the MSKCC cohort were sorted according to GATA3 expression levels, fascin expression levels were higher in patients with low GATA3 expression (Fig. 2B). Indeed, when patients were stratified into “GATA3 low” (GATA3 levels at or below the median level) or “GATA3 high” (above median) groups, the average fascin expression levels in the GATA3 low patients were >2-fold higher than the GATA3 high group (Fig. 2C).
FIGURE 2.
GATA3 expression levels negatively correlated with fascin in breast cancer patients. A, correlation between fascin (FSCN1) and GATA3 probe sets in two cohorts of breast cancer patients (the MSKCC cohort and the Stockholm cohort). Pearson correlation coefficients (r) and p values are as indicated. B, mRNA expression levels of GATA3 (lower panel) and fascin (upper panel) in 99 breast cancer patients in the MSKCC cohort; the patients were sorted according to GATA3 express levels from low (left) to high (right). C, fascin mRNA levels were >2-fold higher in the GATA3 low group (n = 50, GATA3 levels at or below the median) when compared with the GATA3 high group (n = 49, GATA3 levels above the median) in the MSKCC cohort. ****, p < 0.0001 (two-tailed Students t test). D and E, 50 breast caner patients in the MSKCC cohort with high TGFβ1 expression levels (TGF1 at or above median) were further stratified into two subgroups based on fascin (D) or GATA3 (E) expression levels. The probability values (p) and hazard ratios (HR) of lung or bone metastasis free survival were calculated by log-rank tests.
GATA3 and Fascin Are Critical for Breast Cancer Lung Metastasis
We previously reported that the expression levels of fascin were about 2-fold higher in “TGFβ high” breast cancer patients than in “TGFβ low” patients (19). Breast cancer patients with high levels of fascin are more prone to developing lung metastasis (17, 18). Intriguingly, it was recently suggested that TGFβ also promoted lung metastasis in ER-negative breast cancer patients (5). To evaluate whether fascin and GATA3 played a role in TGFβ-mediated breast cancer lung metastasis, we stratified the TGFβ high breast cancer patients in the MSKCC cohort (19) to different groups based on fascin or GATA3 expression levels. As shown in Fig. 2D, TGFβ high breast cancer patients with high fascin levels (or low GATA3 levels in Fig. 2E) were remarkably more susceptible to developing lung metastasis (p = 0.003, HR = 10.7 for fascin and p = 0.004, HR = 9.4 for GATA3) than those patients with low fascin levels (or high GATA3 levels in Fig. 2E). In contrast, it appeared that fascin or GATA3 expression levels had no significant impact on the bone metastasis in this group of breast cancer patients (Fig. 2, D and E). Taken together, our data indicated that both fascin and GATA3 are critical for breast cancer lung metastasis.
Ectopic Expression of GATA3 Abrogates TGFβ and Smad4-mediated Gene Transcription
To determine the role of GATA3 in TGFβ and Smad4-mediated fascin transcription, we stably expressed GATA3 in MDA-MB-231 and MDA-MB-468 cells. The ectopic expression of GATA3 in the spindle-shaped MDA-MB-231 cells induced a morphology change to epithelial-like polygonal shape, which is consistent with the previous observation that ectopic GATA3 reversed the epithelial-to-mesenchymal transition in basal-like breast cancer cells (12). When ectopically expressed in the two basal-like breast cancer cell lines, GATA3 only had a very modest inhibitory effect on fascin protein expression (∼10–30% reduction in protein levels according to Western blotting) (Fig. 3, A and B). However, GATA3 almost abrogated the TGFβ and Smad4-mediated overexpression of fascin (Fig. 3, A and B). GATA3 also abrogated TGFβ and Smad4-mediated increase in fascin mRNA levels, suggesting that GATA3 might inhibit the TGFβ- and Smad4-mediated transcription (Fig. 3C). The robust inhibition of TGFβ- and Smad4-mediated fascin overexpression in basal-like breast cancer cells was not due to un-physiologically high levels of ectopic GATA3, as the levels of ectopically expressed GATA3 in MDA-MB-231 and MDA-MB-468 cells were <10% that of the endogenous GATA3 protein levels in the luminal breast cancer cell lines (MCF-7 and T47D). It is also worth noting that even at such low levels ectopic GATA3 was sufficient to exert a robust inhibitory effect on TGFβ- and Smad4-mediated responses. We also sought to determine whether GATA3 knockdown in luminal breast cancer cells would make them responsive to TGFβ-mediated fascin overexpression. Despite successful reduction of endogenous GATA3 protein by >80% through shRNA, the residual GATA3 levels in MCF-7 and T47D cells were still two to three times higher than the ectopic GATA3 levels in basal-like cells with enforced expression (Fig. 3D). Consequently, TGFβ-induced fascin expression was unremarkable in these luminal breast cancer cells even after GATA3 knockdown (Fig. 3E).
FIGURE 3.
Ectopic GATA3 abrogated TGFβ- and Smad4-mediated transcription of fascin in basal-like breast cancer cells. A and B, ectopic GATA3 abrogated TGFβ (A) and Smad4-induced (B) overexpression of fascin protein in MDA-MB-231 and MDA-MB-468 cells. Basal-like breast cancer cells stably expressing control vector or ectopic GATA3 were treated with TGFβ1 (5 ng/ml, 72 h) (A) or used for the enforced expression of FLAG-Smad4 (B). *, nonspecific band detected by the GATA3 antibody (A). C, ectopic GATA3 abrogated TGFβ and Smad4-mediated transcription of fascin mRNA in MDA-MB-231 cells. MDA-MB-231 cells stably expressing control vector or ectopic GATA3 were treated as described in A and B; fascin mRNA levels were determined through qPCR. D, expression levels of endogenous GATA3, GATA3 levels after enforced expression (in MDA-MB-231 and MDA-MB-468 cells), or shRNA knockdown (in MCF-7 and T47D cells). E, effects of GATA3 knockdown on responsiveness to TGFβ-induced fascin expression in T47D and MCF-7 cells. F and G, ectopic GATA3 abrogated the TGFβ- or Smad4-mediated transcription of five TGFβ response genes (E-cadherin, ANGPTL-4, N-cadherin, vimentin, and p21) in MDA-MB-231 cells. MDA-MB-231 cells stably expressing control vector or ectopic GATA3 were treated as described in A and B; mRNA levels of TGFβ response genes were determined through qPCR. The relative protein expression levels of fascin in A, B, and E were quantified as described in Fig. 1. ***, p < 0.001, as determined by two-tailed Student's t test.
To determine whether the inhibition of TGFβ and Smad4-mediated gene transcription is specific to fascin, we used quantitative PCR to assess the GATA3 effects on the transcription of a panel of five additional TGFβ response genes, including three genes directly regulated by the Smad transcriptional complexes (ANGPTL4, vimentin, and p21) and two genes indirectly regulated by TGF-Smad signaling (E-cadherin and N-cadherin). Gata3 modestly increased the expression levels of E-cadherin and decreased the levels of N-cadherin and vimentin (Fig. 3, F and G), which is consistent with the luminal differentiation and reversal of EMT phenotypes induced by GATA3 (7, 31). Strikingly, ectopically expressed GATA3 abrogated TGFβ- and Smad4-mediated transcription of all five TGFβ response genes, suggesting that GATA3 might globally inhibit the signaling of the canonical TGFβ-Smad signaling pathway (Fig. 3, F and G).
GATA3 Abrogates Smad4-mediated Invadopodium Formation and Invasion
Invadopodia are adhesive membrane protrusions that coordinate ECM degradation and invasion in cancer cells (32, 33). Invadopodia share many protein components and similar regulatory mechanisms with filopodia and are considered “invasive filopodia” in metastatic cancer cells (20, 21, 34). It was recently reported that fascin promoted invadopodium formation by stabilizing the actin core of invadopodia (20). We sought to investigate the role of TGFβ and fascin in invadopodium regulation in basal-like breast cancer cells. When stained for F-actin and cortactin, ∼30% of MDA-MB-231 cells contained round actin and cortactin-positive dots on the ventral side of the cell (Fig. 4A). When plated on glass coverslips coated with fluorescence-labeled gelatin, these actin protrusions were able to degrade gelatin, leaving dark spots on a bright background, suggesting that those were invadopodia. Treatment with TGFβ or overexpression of Smad4 increased the percentage of invadopodia positive cells from ∼30% (47 of 159) to ∼90% (139 of 151) and ∼50% (79 of 159), respectively (Fig. 4, A and B). Smad3 and Smad4 knockdown almost abolished TGFβ-mediated invadopodium formation without significant effects on basal levels of invadopodia positive cells (Fig. 4, C and D), suggesting that TGFβ promotes invadopodium formation through the canonical Smad-dependent pathway. To investigate the role of fascin in TGFβ-mediated invadopodium formation, we employed shRNA to knock down fascin expression in MDA-MB-231 cells. Fascin knockdown decreased the proportion of invadopodia-positive cells from ∼26% (39 of 151) to ∼10% (15 of 152) (Fig. 4, C and D). Although TGFβ treatment in fascin knockdown cells still increased invadopodium formation (about 20% of the cells were positive for invadopodia after TGFβ treatment), the increase was remarkably lower than in control shRNA expressing cells, suggesting that fascin is critical for TGFβ-mediated invadopodia formation.
FIGURE 4.
GATA3 abrogated Smad4-mediated invadopodium formation and invasion. A, actin (red) and cortactin (green) staining revealed that TGFβ treatment (5 ng/ml) and ectopic Smad4 promoted the formation of invadopodia in MDA-MB-231 cells. Cells were allowed to attach to gelatin-coated coverslips for 24 h before PFA fixation. For TGFβ treatment, cells were pretreated with 5 ng/ml TGFβ before being plated onto coverslips. B, quantification of invadopodia-positive and -negative cells in A. C and D, MDA-MB-231 cells expressing control and targeting shRNAs were treated with TGFβ as described in A and stained with actin and cortactin. C shows representative images, and D shows quantification of invadopodia-positive and -negative cells in each group. E–G, effects of ectopic GATA3 expression in MDA-MB-231 cells on invadopodium formation and ECM degradation. Representative images showing GATA3 abolished Smad4-mediated invadopodia formation and gelatin degradation are presented in E. Quantification of invadopodia positive versus negative cells is shown in F and, quantification of gelatin degradation area per cell is shown in G. Insets in A, C, and E are magnified views of the boxed area in the main images. Scale bars are 10 μm. H and I, effects of ectopic GATA3 on Smad4-mediated MDA-MB-231 cell migration (H) and invasion (I). J, effects of fascin knockdown on Smad4-mediated MDA-MB-231 invasion. K, ectopic expression of fascin partially rescued the inhibition of Smad4-mediated invasion by GATA3. *, p < 0.05 and ****, p < 0.0001, respectively (determined by two-tailed Fisher's exact test in B, D, and F or by Student's t test in H–K).
Because our data indicated that GATA3 abrogated the global response to TGFβ-Smad4 signaling, we further investigated the effects of GATA3 in Smad4-mediated invadopodium formation and ECM degradation. The ectopic expression of GATA3 remarkably decreased the invadopodium-positive MDA-MB-231 cells from ∼30% to <10% (14 of 150) and inhibited the gelatin degradation activity of the breast cancer cells (Fig. 4, E–G). Importantly, unlike in the control MDA-MB-231 cells, overexpression of Smad4 in MDA-MB-231-GATA3 cells failed to increase either the proportion of invadopodium-positive cells or the degradation of gelatin (Fig. 4, E–G). Next, we investigated the effects of GATA3 on Smad4-mediated migration and invasion of MDA-MB-231 cells through Boyden chamber assay. Smad4 promoted the motility and invasiveness of MDA-MB-231 cells by >2.5-fold. In MDA-MB-231-GATA3 cells the pro-migration and pro-invasion activity of Smad4 was dramatically diminished (Fig. 4, H–I) despite similar levels of Smad4 protein expression in the control cells and GATA3 cells (Fig. 3B). Fascin knockdown was able to significantly inhibit Smad4-mediated invasion (Fig. 4J), and ectopic overexpression of fascin was able to partially rescue the inhibition of Smad4-mediated invasion by GATA3 (Fig. 4K). Taken together, our data indicated that re-introduction of GATA3 into the basal-like MDA-MB-231 breast cancer cells abrogated the ability of TGFβ-Smad signaling pathway to promote invadopodium formation, ECM degradation, and Matrigel invasion at least partially through abrogating the TGFβ- and Smad4-mediated fascin overexpression.
GATA3 Abrogates the Binding of Smad4 to Fascin Promoter
To understand the molecular mechanisms by which GATA3 regulates Smad4-mediated fascin transcription, we inspected the fascin promoter for GATA3 binding elements and identified three potential GATA3 sites at −1869, −1707, and −1114 (Fig. 5A). To determine whether these GATA3 binding elements were required for GATA3 to inhibit TGFβ and Smad4-mediated fascin transcription, we used the P402 luciferase reporter to investigate the activation of this truncated reporter by TGFβ. Surprisingly, although the P402 reporter did not contain any of the three GATA3 binding elements, ectopic GATA3 still abolished the activation of P402 truncated promoter by TGFβ.
FIGURE 5.
GATA3 abrogated the interaction between Smad4 and its binding sites on fascin and p21 promoters. A, the upper panel shows potential GATA3 binding sites on fascin promoter, and the lower panel shows luciferase assay results, showing that ectopic GATA3 abrogated the activation of P402 fascin promoter by TGFβ in MDA-MB-231 cells. B, control and GATA3-overexpressing MDA-MB-231 cells were treated with TGFβ (5 ng/ml) for the various time (as indicated); the phosphor-Smad3 and total Smad3 levels in these cells were determined by Western blotting. C, physical interactions between GATA3 and Smad3 and between GATA3 Smad4 were determined by immunoprecipitation (IP). HEK293 cells were transient transfected with HA-GATA3 alone or HA-GATA3 together with FLAG-Smad3 or FLAG-Smad4. FLAG-tagged Smads were precipitated with M2 beads, and co-precipitated GATA3 was detected with Western blotting (IB). D, MCF-7 cells with or without enforced expression of FLAG-Smad3 were treated with or without TGFβ and used for immunoprecipitation as described in C. IgG heavy-chain bands from M2 beads that migrated in close proximity to GATA3 bands are indicated. F and G, GATA3 ectopically expressed in MDA-MB-231 cells abrogated the binding of Smad4 to the fascin promoter and p21 promoter, as determined by ChIP assay with anti-Smad4 antibody. The fascin and p21 promoter precipitated by the anti-Smad4 antibody were detected through semi-quantitative PCR. G, quantification of the results in F through densitometry using ImageJ. RU, relative units.
We next investigated whether GATA3 abolished TGFβ-mediated fascin expression by inhibiting Smad3 phosphorylation. Ectopically expressed GATA3 only modestly decreased basal and TGFβ-stimulated phosphor-Smad3 levels in MDA-MB-231 cells (Fig. 5B), suggesting that this is not likely to be a major mechanism. To determine whether GATA3 might directly regulate the activity of the Smad transcriptional complex by binding to Smad proteins, we expressed HA-GATA3 alone or together with FLAG-Smad3 or FLAG-Smad4, respectively, in HEK293 cells. The FLAG-tagged Smads were immunoprecipitated with M2 anti-FLAG beads, and the presence of GATA3 bound to the beads was assayed through anti-HA immunoblotting. As shown in Fig. 5C, M2 beads precipitated GATA3 when GATA3 was co-expressed with either FLAG-Smad3 or with FLAG-Smad4, but not when expressed alone, suggesting that GATA3 directly interacts with Smad transcription factors (Fig. 5C). To investigate whether Smad3 might interact with endogenous GATA3 and whether such interaction might be regulated by TGFβ, we expressed FLAG-Smad3 in MCF-7 cells. As shown in Fig. 5D, M2 anti-FLAG beads successfully co-immunoprecipitated endogenous GATA3 with FLAG-Smad3 in MCF-7 cells with or without TGFβ treatment, suggesting that the Smad3-GATA3 complex might not be affected by TGFβ treatment.
To determine the effects of GATA3 on the formation of Smad3-Smad4 transcriptional complex, we investigated the Smad3-Smad4 interaction in the MDA-MB-231 control cells and the GATA3 stable line using FLAG-Smad4 to immunoprecipitate endogenous Smad3. A very low Smad3-Smad4 interaction was detected in the control cells and GATA3 cells before TGFβ stimulation (Fig. 5E). Stimulation with 5 ng/ml TGFβ increased the amount of Smad3 precipitated FLAG-Smad4 in both cell lines; however, the formation of Smad3-Smad4 complex was inhibited by ∼70% in the GATA3 cells when compared with the control cells (Fig. 5E).
We postulated that the direct interaction between GATA3 and Smad3/4 and the reduced formation of Smad3-Smad4 complex might contribute synergistically to inhibit the binding of Smad4 to the promoter of TGFβ response genes and thus abrogated Smad4-mediated invasion in basal-like breast cancer cells. To examine this possibility we investigated the interaction between Smad4 and the promoters of fascin and p21 through ChIP. Only very small amounts of the fascin or the p21 promoter were immunoprecipitated by anti-Smad4 antibody in the MDA-MB-231 control cells or the GATA3 cells (Fig. 5, F and G). TGFβ increased the immunoprecipitation of p21 and fascin promoter by anti-Smad4 antibody by 5–30-fold (Fig. 5, F and G), consistent with the notion that these two genes are directly regulated by the canonical TGFβ-Smad4 signaling. However, when GATA3 was ectopically expressed in MDA-MB-231 cells, TGFβ was not able to increase the binding of Smad4 to either the fascin promoter or the p21 promoter, suggesting that GATA3 abrogated the TGFβ-mediated activation of Smad4 transcriptional activity.
GATA3 N-terminal Interacts with Smad3 and Smad4
GATA3 is a 443-residue protein containing two transactivation domains (TA1 and TA2) on the N terminus and two DNA binding Zinc-finger domains (ZnF1 and ZnF2) on the C terminus (Fig. 6A). To understand the structural determinant for the interactions between GATA3 and Smad3/4, we constructed a series of HA-tagged GATA3 fragments (Fig. 6A). These fragments were co-expressed with FLAG-Smad3 or FLAG-Smad4, and their interactions were determined through co-immunoprecipitation (Fig. 6, B–E). Smad3 and Smad4 interacted strongly with both the N1 (1–259) and the N2 (1–295) fragments and weakly with the C1 (259–443) fragment, suggesting that the interactions with Smad3/4 mainly involved the N-terminal region containing the two transactivation domains (Fig. 6, B and C). It appeared that the N2 fragment (N1 fragment plus ZnF1) interactions with Smad3/4 were stronger than with the N1 fragment, suggesting that the zinc finger domain 1 might contribute to strengthening the interactions between Smad3/4 and GATA3 N-terminal and might account for the residual interaction activity in the C1 fragment. To test this possibility we constructed a C2 fragment containing residue 295–443 (C1 minus ZnF1) (Fig. 6A). The C2 fragment appeared to be prone to degradation, but we were able to obtain high expression levels of this protein fragment after inhibition of the proteasome pathway with MG132. The MG132 treatment did not affect the interaction between full-length GATA3 and Smad3/4; however, despite the high expression levels in the MG132 treated cells, the C2 fragment failed to interact with either Smad3 or Smad4 (Fig. 6, D and E). Taken together, our data suggest that the interactions between GATA3 and Smad3/4 are mainly mediated by GATA3 N-terminal and further strengthened by ZnF1 domain.
FIGURE 6.
The physical interactions between Smad3/4 and GATA3 were mediated by its N-terminal and ZnF1 domain. A, schematic illustration of the structural organization of GATA3 domains and truncated GATA3 fragments used in this study; TA1, TA2, ZnF1, and ZnF2 are transactivation domains 1 and 2 and zinc finger domains 1 and 2, respectively. The beginning and ending residue numbers for each domain/region/fragments are as indicated. B–E, physical interactions between full-length GATA3, GATA3 fragments, and Smad3 and Smad4 were determined by immunoprecipitation (IP). HEK293 cells were transiently transfected with HA-tagged GATA3 and GATA3 fragments alone or together with FLAG-Smad3 or FLAG-Smad4 as indicated in the respective panels. FLAG-tagged Smads were precipitated with M2 beads, and co-precipitated GATA3 fragments were detected with Western blotting (IB). Cells in D and E were treated with 2 μm MG132 for 12 h before cell lysis and immunoprecipitation. The two asterisks in D mark IgG heavy chain from M2 beads that co-migrate with FLAG-Smad3.
DISCUSSION
Fascin is a pro-metastasis actin bundling protein overexpressed in all of the carcinomas examined to date (35). In breast cancer patients, fascin expression levels are significantly higher in the basal-like subgroup when compared with the luminal subgroup or to normal breast tissues (17, 36). There is emerging evidence suggesting that cytokines and growth factors in the tumor microenvironment, such as TGFβ, IL-6, and EGF, may promote fascin overexpression in cancer cells (19, 24, 37–39). We previously reported that TGFβ promoted fascin overexpression in breast and lung cancer cells through a Smad3- and Smad4-dependent but MAPK-independent pathway (19); however, Fu et al. (38) reported that TGFβ mediated fascin overexpression through Smad-independent but MAPK-dependent pathway in gastric cancer. It is not immediately clear whether the discrepancy is due to different types of cancer cells used in the two studies. Nonetheless, our data here further indicate that fascin is a direct target gene of the canonical TGFβ-Smad signaling pathway, at least in basal-like breast cancer. TGFβ activates the transcription of fascin gene by promoting the binding of Smad4 to the −370 Smad binding sites on the fascin promoter. Our findings together with previous reports on the regulation of fascin expression by Stat3 and NFκB (37, 40) suggest that signaling pathways downstream of the inflammatory cytokines (e.g. TGFβ, IL-6, TNFα, etc.) might be responsible for fascin overexpression in metastatic cancers. The effects of inflammatory tumor microenvironment on fascin overexpression warrant further exploration in the future.
Our earlier data suggested that the differentiation state of the cancer cells might affect the response to TGFβ-mediated fascin overexpression (19). Interestingly, the up-regulation of fascin by TGFβ and Smad4 was only observed in basal-like breast cancer cells but not in the luminal-like cells. By examining two breast cancer microarray datasets, we identified GATA3, a master regulator of mammary morphogenesis and luminal differentiation (6–9, 31) as a potential regulator of fascin expression. Indeed, ectopically expressed GATA3 abrogated the TGFβ and Smad4-mediated transcription and overexpression of fascin in basal-like breast cancer cells. Our data further indicated that ectopically expressed GATA3 might globally suppress the transcriptional activity of the canonical TGFβ-Smad signaling and abrogate the ability of Smad4 to promote invadopodium formation, cell migration, and invasion in MDA-MB-231 cells. The abrogation of Smad4-mediated responses by GATA3 was probably due to the blockade of the interaction between Smad4 and its DNA binding sites.
Intriguingly, ectopic expression of GATA3 in MDA-MB-231 cells, although at a relative low level when compared with luminal cancer cells, remarkably inhibited invadopodium formation. Such inhibition is unlikely through fascin, as GATA3 only very modestly reduced basal expression levels of fascin in MDA-MB-231 cells. It was recently reported that transcription factor or TGFβ-mediated EMT significantly promoted invadopodium formation in breast cancer cells (41). It is well established that ectopic expression of GATA3 in mesenchymal-like cancer cells induces epithelial-like phenotypes. It is possible that ectopic expression of GATA3 inhibits invadopodium formation by reversing EMT.
It was previously reported that Smad3 and GATA3 physically interact with each other in T cells to enable the regulation of GATA3 target genes by TGFβ signaling (42). It was not clear, however, whether such interactions affect the canonical Smad-mediated signaling pathway. Our data indicate that, when ectopically expressed in basal-like breast cancer cells, GATA3 physically interacts with Smad3 and Smad4 and interferes with the formation of Smad3-Smad4 transcription complex. It is tempting to hypothesize that GATA3 is a novel co-suppressor of Smad transcription factors; the physical interactions between GATA3 and Smad3/4 might block the TGFβ-Smad signaling by abrogating the interactions between Smad transcription factors and their DNA binding elements. The interaction between Smad3/4 and GATA3 was mainly mediated by its N-terminal region and strengthened by the ZnF1 domain. It is interesting to note that the ZnF1 domain was previously implicated in the interactions between GATA3 and FOG1 and FOG2 (43).
The effects of TGFβ on cancer progression are highly context-dependent (3, 44). Although the core components of the canonical TGF signaling pathways are preserved in most breast cancers, TGFβ signaling promotes lung metastasis only in ER-negative, but not the ER-positive, breast cancer (4, 5, 44). Our data suggest that high expression levels of GATA3 in the ER-positive, luminal-like breast cancer might contribute to suppression of TGFβ-mediated metastasis in this subgroup of breast cancer by abrogating Smad4-mediated invadopodium formation, ECM degradation, cell migration, and invasion. Indeed, TGFβ high breast cancer patients in the MSKCC cohort with low fascin expression or high GATA3expression were much less likely to develop lung metastasis, lending further evidence to the notion that these two genes are critical players in TGFβ-mediated breast cancer metastasis.
Acknowledgments
We thank Drs. Minjung Kim and Jin Q. Cheng for reagents and cell lines, the Analytic Microscopy Core and Molecular Genomics core at Moffitt Cancer center for assistance with image acquisition and qPCR, and Rasa Hamilton for editorial assistance. The Analytic Microscopy Core and Molecular Genomics Core at Moffitt Cancer Center were supported in part by NCI, National Institutes of Health Grant P30-CA76292-14.
This work was supported in part by Institutional Research Grant IRG-93-032-16 from the American Cancer Society and a Milestone Award from Miles for Moffitt Foundation (to S. Y).

This article contains supplemental Table 1.
- ER
- estrogen receptor
- ECM
- extracellular matrix
- EMT
- epithelial-to-mesenchymal transition
- qRT
- quantitative real-time.
REFERENCES
- 1. Siegel P. M., Massagué J. (2003) Cytostatic and apoptotic actions of TGF-β in homeostasis and cancer. Nat. Rev. Cancer 3, 807–821 [DOI] [PubMed] [Google Scholar]
- 2. Kang Y. (2006) Pro-metastasis function of TGFβ mediated by the Smad pathway. J. Cell Biochem. 98, 1380–1390 [DOI] [PubMed] [Google Scholar]
- 3. Derynck R., Akhurst R. J., Balmain A. (2001) TGF-β signaling in tumor suppression and cancer progression. Nat. Genet. 29, 117–129 [DOI] [PubMed] [Google Scholar]
- 4. Buck M. B., Fritz P., Dippon J., Zugmaier G., Knabbe C. (2004) Prognostic significance of transforming growth factor β receptor II in estrogen receptor-negative breast cancer patients. Clin. Cancer Res. 10, 491–498 [DOI] [PubMed] [Google Scholar]
- 5. Padua D., Zhang X. H., Wang Q., Nadal C., Gerald W. L., Gomis R. R., Massagué J. (2008) TGFβ primes breast tumors for lung metastasis seeding through angiopoietin-like 4. Cell 133, 66–77 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Chou J., Provot S., Werb Z. (2010) GATA3 in development and cancer differentiation. cells GATA have it! J. Cell Physiol. 222, 42–49 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Asselin-Labat M. L., Sutherland K. D., Barker H., Thomas R., Shackleton M., Forrest N. C., Hartley L., Robb L., Grosveld F. G., van der Wees J., Lindeman G. J., Visvader J. E. (2007) Gata-3 is an essential regulator of mammary-gland morphogenesis and luminal-cell differentiation. Nat. Cell Biol. 9, 201–209 [DOI] [PubMed] [Google Scholar]
- 8. Kouros-Mehr H., Slorach E. M., Sternlicht M. D., Werb Z. (2006) GATA-3 maintains the differentiation of the luminal cell fate in the mammary gland. Cell 127, 1041–1055 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Kouros-Mehr H., Kim J. W., Bechis S. K., Werb Z. (2008) GATA-3 and the regulation of the mammary luminal cell fate. Curr. Opin. Cell Biol. 20, 164–170 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Usary J., Llaca V., Karaca G., Presswala S., Karaca M., He X., Langerød A., Kåresen R., Oh D. S., Dressler L. G., Lønning P. E., Strausberg R. L., Chanock S., Børresen-Dale A. L., Perou C. M. (2004) Mutation of GATA3 in human breast tumors. Oncogene 23, 7669–7678 [DOI] [PubMed] [Google Scholar]
- 11. Mehra R., Varambally S., Ding L., Shen R., Sabel M. S., Ghosh D., Chinnaiyan A. M., Kleer C. G. (2005) Identification of GATA3 as a breast cancer prognostic marker by global gene expression meta-analysis. Cancer Res. 65, 11259–11264 [DOI] [PubMed] [Google Scholar]
- 12. Yan W., Cao Q. J., Arenas R. B., Bentley B., Shao R. (2010) GATA3 inhibits breast cancer metastasis through the reversal of epithelial-mesenchymal transition. J. Biol. Chem. 285, 14042–14051 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Chou J., Lin J. H., Brenot A., Kim J. W., Provot S., Werb Z. (2013) GATA3 suppresses metastasis and modulates the tumour microenvironment by regulating microRNA-29b expression. Nat. Cell Biol. 15, 201–213 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Chu I. M., Michalowski A. M., Hoenerhoff M., Szauter K. M., Luger D., Sato M., Flanders K., Oshima A., Csiszar K., Green J. E. (2012) GATA3 inhibits lysyl oxidase-mediated metastases of human basal triple-negative breast cancer cells. Oncogene 31, 2017–2027 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Dydensborg A. B., Rose A. A., Wilson B. J., Grote D., Paquet M., Giguère V., Siegel P. M., Bouchard M. (2009) GATA3 inhibits breast cancer growth and pulmonary breast cancer metastasis. Oncogene 28, 2634–2642 [DOI] [PubMed] [Google Scholar]
- 16. Chu I. M., Lai W. C., Aprelikova O., El Touny L. H., Kouros-Mehr H., Green J. E. (2013) Expression of GATA3 in MDA-MB-231 triple-negative breast cancer cells induces a growth inhibitory response to TGFß. PLoS ONE 8, e61125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Chen L., Yang S., Jakoncic J., Zhang J. J., Huang X. Y. (2010) Migrastatin analogues target fascin to block tumour metastasis. Nature 464, 1062–1066 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Minn A. J., Gupta G. P., Siegel P. M., Bos P. D., Shu W., Giri D. D., Viale A., Olshen A. B., Gerald W. L., Massagué J. (2005) Genes that mediate breast cancer metastasis to lung. Nature 436, 518–524 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Sun J., He H., Xiong Y., Lu S., Shen J., Cheng A., Chang W. C., Hou M. F., Lancaster J. M., Kim M., Yang S. (2011) Fascin protein is critical for transforming growth factor β protein-induced invasion and filopodia formation in spindle-shaped tumor cells. J. Biol. Chem. 286, 38865–38875 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Li A., Dawson J. C., Forero-Vargas M., Spence H. J., Yu X., König I., Anderson K., Machesky L. M. (2010) The actin-bundling protein fascin stabilizes actin in invadopodia and potentiates protrusive invasion. Curr. Biol. 20, 339–345 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Machesky L. M., Li A. (2010) Fascin. Invasive filopodia promoting metastasis. Commun. Integr. Biol. 3, 263–270 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Yang S., Huang F. K., Huang J., Chen S., Jakoncic J., Leo-Macias A., Diaz-Avalos R., Chen L., Zhang J. J., Huang X. Y. (2013) Molecular mechanism of fascin function in filopodial formation. J. Biol. Chem. 288, 274–284 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Yang S., Zhang J. J., Huang X. Y. (2012) Mouse models for tumor metastasis. Methods Mol. Biol. 928, 221–228 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Lu X. F., Li E. M., Du Z. P., Xie J. J., Guo Z. Y., Gao S. Y., Liao L. D., Shen Z. Y., Xie D., Xu L. Y. (2010) Specificity protein 1 regulates fascin expression in esophageal squamous cell carcinoma as the result of the epidermal growth factor/extracellular signal-regulated kinase signaling pathway activation. Cell. Mol. Life Sci. 67, 3313–3329 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Pillai S., Dasgupta P., Chellappan S. P. (2009) Chromatin immunoprecipitation assays. Analyzing transcription factor binding and histone modifications in vivo. Methods Mol. Biol. 523, 323–339 [DOI] [PubMed] [Google Scholar]
- 26. Artym V. V., Zhang Y., Seillier-Moiseiwitsch F., Yamada K. M., Mueller S. C. (2006) Dynamic interactions of cortactin and membrane type 1 matrix metalloproteinase at invadopodia. Defining the stages of invadopodia formation and function. Cancer Res. 66, 3034–3043 [DOI] [PubMed] [Google Scholar]
- 27. Yang S., Huang X. Y. (2005) Ca2+ influx through L-type Ca2+ channels controls the trailing tail contraction in growth factor-induced fibroblast cell migration. J. Biol. Chem. 280, 27130–27137 [DOI] [PubMed] [Google Scholar]
- 28. McGarrigle D., Shan D., Yang S., Huang X. Y. (2006) Role of tyrosine kinase Csk in G protein-coupled receptor- and receptor tyrosine kinase-induced fibroblast cell migration. J. Biol. Chem. 281, 10583–10588 [DOI] [PubMed] [Google Scholar]
- 29. Bros M., Ross X. L., Pautz A., Reske-Kunz A. B., Ross R. (2003) The human fascin gene promoter is highly active in mature dendritic cells due to a stage-specific enhancer. J. Immunol. 171, 1825–1834 [DOI] [PubMed] [Google Scholar]
- 30. Pawitan Y., Bjöhle J., Amler L., Borg A. L., Egyhazi S., Hall P., Han X., Holmberg L., Huang F., Klaar S., Liu E. T., Miller L., Nordgren H., Ploner A., Sandelin K., Shaw P. M., Smeds J., Skoog L., Wedrén S., Bergh J. (2005) Gene expression profiling spares early breast cancer patients from adjuvant therapy. Derived and validated in two population-based cohorts. Breast Cancer Res. 7, R953–R964 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Kouros-Mehr H., Bechis S. K., Slorach E. M., Littlepage L. E., Egeblad M., Ewald A. J., Pai S. Y., Ho I. C., Werb Z. (2008) GATA-3 links tumor differentiation and dissemination in a luminal breast cancer model. Cancer Cell 13, 141–152 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Gimona M., Buccione R., Courtneidge S. A., Linder S. (2008) Assembly and biological role of podosomes and invadopodia. Curr. Opin. Cell Biol. 20, 235–241 [DOI] [PubMed] [Google Scholar]
- 33. Murphy D. A., Courtneidge S. A. (2011) The “ins” and “outs” of podosomes and invadopodia. Characteristics, formation. and function. Nat. Rev. Mol. Cell Biol. 12, 413–426 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Schoumacher M., Goldman R. D., Louvard D., Vignjevic D. M. (2010) Actin, microtubules, and vimentin intermediate filaments cooperate for elongation of invadopodia. J. Cell Biol. 189, 541–556 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Hashimoto Y., Skacel M., Adams J. C. (2005) Roles of fascin in human carcinoma motility and signaling. Prospects for a novel biomarker? Int. J. Biochem. Cell Biol. 37, 1787–1804 [DOI] [PubMed] [Google Scholar]
- 36. Yoder B. J., Tso E., Skacel M., Pettay J., Tarr S., Budd T., Tubbs R. R., Adams J. C., Hicks D. G. (2005) The expression of fascin, an actin-bundling motility protein, correlates with hormone receptor-negative breast cancer and a more aggressive clinical course. Clin. Cancer Res. 11, 186–192 [PubMed] [Google Scholar]
- 37. Snyder M., Huang X. Y., Zhang J. J. (2011) Signal transducers and activators of transcription 3 (STAT3) directly regulates cytokine-induced fascin expression and is required for breast cancer cell migration. J. Biol. Chem. 286, 38886–38893 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Fu H., Hu Z., Wen J., Wang K., Liu Y. (2009) TGF-β promotes invasion and metastasis of gastric cancer cells by increasing fascin1 expression via ERK and JNK signal pathways. Acta Biochim. Biophys. Sin. 41, 648–656 [DOI] [PubMed] [Google Scholar]
- 39. Keshamouni V. G., Jagtap P., Michailidis G., Strahler J. R., Kuick R., Reka A. K., Papoulias P., Krishnapuram R., Srirangam A., Standiford T. J., Andrews P. C., Omenn G. S. (2009) Temporal quantitative proteomics by iTRAQ 2D-LC-MS/MS and corresponding mRNA expression analysis identify post-transcriptional modulation of actin-cytoskeleton regulators during TGF-β-Induced epithelial-mesenchymal transition. J. Proteome Res. 8, 35–47 [DOI] [PubMed] [Google Scholar]
- 40. Kress A. K., Kalmer M., Rowan A. G., Grassmann R., Fleckenstein B. (2011) The tumor marker Fascin is strongly induced by the Tax oncoprotein of HTLV-1 through NF-κB signals. Blood 117, 3609–3612 [DOI] [PubMed] [Google Scholar]
- 41. Eckert M. A., Lwin T. M., Chang A. T., Kim J., Danis E., Ohno-Machado L., Yang J. (2011) Twist1-induced invadopodia formation promotes tumor metastasis. Cancer Cell 19, 372–386 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Blokzijl A., ten Dijke P., Ibáñez C. F. (2002) Physical and functional interaction between GATA-3 and Smad3 allows TGF-β regulation of GATA target genes. Curr. Biol. 12, 35–45 [DOI] [PubMed] [Google Scholar]
- 43. Nesbit M. A., Bowl M. R., Harding B., Ali A., Ayala A., Crowe C., Dobbie A., Hampson G., Holdaway I., Levine M. A., McWilliams R., Rigden S., Sampson J., Williams A. J., Thakker R. V. (2004) Characterization of GATA3 mutations in the hypoparathyroidism, deafness, and renal dysplasia (HDR) syndrome. J. Biol. Chem. 279, 22624–22634 [DOI] [PubMed] [Google Scholar]
- 44. Padua D., Massagué J. (2009) Roles of TGFβ in metastasis. Cell Res 19, 89–102 [DOI] [PubMed] [Google Scholar]