Abstract
Metals are utilized for a variety of critical cellular functions and are essential for survival. However cells are faced with the conundrum of needing metals coupled with e fact that some metals, iron in particular are toxic if present in excess. Maintaining metal homeostasis is therefore of critical importance to cells. In this review we have systematically analyzed sequenced genomes of three members of the Rhodobacter genus, R. capsulatus SB1003, R. sphaeroides 2.4.1 and R. ferroxidans SW2 to determine how these species undertake iron homeostasis. We focused our analysis on elemental ferrous and ferric iron uptake genes as well as genes involved in the utilization of iron from heme. We also discuss how Rhodobacter species manage iron toxicity through export and sequestration of iron. Finally we discuss the various putative strategies set up by these Rhodobacter species to regulate iron homeostasis and the potential novel means of regulation. Overall, this genomic analysis highlights surprisingly diverse features involved in iron homeostasis in the Rhodobacter genus.
I. INTRODUCTION
The origin of oxygenic photosynthesis can be traced to ~2.9 billion years ago when cyanobacteria-driven photosynthesis created a Great Oxidizing Event that enriched atmospheric oxygen. Prior to photosynthetic oxidation of Earth, most iron was in a reduced ferrous state that is biologically available as it has a solubility of 0.1M at pH7. This form of iron is thought to been present in deep biotopes until ~1.8-1 billion years ago (Van Der Giezen and Lenton, 2012; Planavsky et al., 2011). Beyond that time the presence of atmospheric oxygen effectively oxidized most surface and oceanic iron to a ferric state that has an extremely low solubility (10−18M at pH 7) (Andrews et al., 2003). Consequently, the oxidation of Earth must have caused a crisis of iron availability necessitating that cells evolved a diverse array of ferrous and ferric iron uptake systems.
Iron is an important cofactor in many enzymes where it can form mono- or di-iron centers, or more complex iron-sulfur clusters. Iron is also bound to protoporphyrin IX to form heme that has an important role as a gas and electron carrier. Enzymes that utilize iron are involved in major biochemical processes such as photosynthesis, N2 fixation, methanogenesis, H2 production and consumption, respiration, the trichloroacetic acid cycle, oxygen transport, gene regulation and DNA biosynthesis. Iron is also an important actor in cellular events such as virulence, biofilm formation and quorum sensing (Vasil, 2007; Steele et al., 2012; Wen et al., 2012). The role of iron in so many systems indicates that Life evolved enzymes that utilized iron when it was readily available and as a result had to invent biochemical pathways to maintain iron homeostasis when Earth’s oxidation caused iron to become scarce. Extremely rare are the examples of organisms that solved the iron availability issue by circumventing the need for iron. This includes the lactobacilli and the Lyme disease agent (Archibald, 1983; Weinberg, 1997; Posey, 2000).
Bacteria have developed iron uptake systems to both the ferrous, ferric forms of elemental iron. They also evolved iron scavenging pathways by excreting and transporting siderophores that function as iron chelators. Bacteria have also developed means of transporting heme that is synthesized by other organisms as a salvage pathway (Wandersman and Delepelaire, 2004; Andrews et al., 2003; Braun and Hantke, 2011). These diverse iron transport systems must be tightly regulated, as excess cellular iron is toxic. Free iron can generate free hydroxyl radicals through Fenton’s chemistry (equations 1, 2, 3) that have deleterious effects on fatty acids and other biomacromolecules (Touati, 2000; Chiancone et al., 2004).
| (Eq. 1) |
| (Eq. 2) |
| (Eq. 3) |
Challenged with balancing a need for iron with iron’s toxicity, cells must maintain a tightly regulated iron homeostasis that controls the dynamic equilibrium between import, export and the storage of iron in proteins. Purple nonsulfur bacteria are facultative phototrophs distributed among the α-and β-subclasses of proteobacteria. They have an extremely versatile metabolism that utilizes iron in ways that allows growth under multiple environmental conditions. The use of iron by purple nonsulfur bacteria can be exemplified by such processes as: i) Aerobic respiration where terminal cytochrome oxidase cbb3 and b260 use heme as a cofactor; ii) Respiratory and photosynthesis electron transport where heme containing cytochromes cy, c2 and bc1 shuttle electrons to photosystem reaction centers as well as to respiratory terminal oxidases; iii) Enzymes such as coproporphyrinogen III oxidase that contains a Fe-S cluster involved in synthesis of heme; iv) Enzymes involved in bacteriochlorophyll synthesis that utilize iron-sulfur clusters (Sirijovski et al., 2007; Sarma et al., 2008); v) Purple nonsulfur bacteria are also capable of anaerobic oxidation of ferrous iron to facilitate phototrophic growth (Widdel et al., 1993; Ehrenreich and Widdel, 1994; Croal et al., 2007; Caiazza et al., 2007; Poulain and Newman, 2009). These are just a few representative examples of the many processes used by this group of bacteria that rely on the use of iron as a cofactor, and that illustrate their heavy need for this metal. In this chapter, we will discuss what is known about iron homeostasis in the Rhodobacter genus, focusing on iron homeostasis genes present in the genomes of R. capsulatus SB1003, R. sphaeroides 2.4.1 and R. ferroxidans SW2 (formerly known as R. sp. SW2 (Saraiva et al., 2012)). Despite numerous studies on photosynthesis, respiration and general physiology, there have been surprisingly few studies on their iron needs and how they regulate cellular iron homeostasis.
II. FERROUS IRON UPTAKE
A. FEO SYSTEM
The Feo iron transport system is widespread among bacteria and thus appears to be a major route of ferrous iron acquisition (Cartron et al., 2006; Perry et al., 2007). Since the first description of the Feo system in Escherichia coli in 1987 (Hantke, 1987), it has been shown to be involved in many iron-related phenotypes such as magnetosome formation (Rong et al., 2012) and virulence (Fetherston et al., 2012). The Feo system is present in many bacterial and in some archaeal genomes. Interestingly, the uptake Feo system has significant sequence similarity to eukaryal G-proteins and thus has been referred to as a “living fossil” of this family of eukariotic GTPases (Hantke, 2003).
Genes coding for the Feo system are present in each of the three genomes considered in this study, namely R. capsulatus SB1003, R. sphaeroides 2.4.1 and R. ferroxidans SW2. R. sphaeroides and R. ferroxidans contain only one feo gene cluster while R. capsulatus exhibits two clusters. It is not unusual to find two (or more) feo loci in one organism. In such cases, it is hypothesized that one of the Feo systems is specialized in manganese uptake and when proven to be involved in this process is thus called Meo (Cartron et al., 2006; He et al., 2006; Dashper et al., 2005). However, duplicate Feo systems can also be specialized in two different iron-related pathways, such as magnetosome formation and oxidative stress management (Rong et al., 2012).
In the analyzed Rhodobacter genomes, a putative four-gene operon is present in each strain, consisting of two feoA genes followed by feoB and feoC loci (feoA1A2BC) (table 1). Other cases of multiple feoA genes in a feo operons have been reported (Cartron et al., 2006). Another putative operon, named feo2AB, is unique to R. capsulatus SB1003 that displays only one feoA gene followed by an feoB (table 1). In ~80% of the genomes where a feo locus is present, it consists of a small feoA gene followed by a larger feoB gene in an feoAB operonal organization. Occasionally an additional feoC ORF is also present, particularly in the γ-proteobacteria phylum, which forms an feoABC operon. Alignment of translated feoA1, feoA2 and feo2A genes from these three Rhodobacter species highlights a higher sequence similarity/identity for FeoA1 representatives with homologues from the other Rhodobacter species than to gene paralogues present in their own genome (table 2). For example, R. capsulatus FeoA2 shows 69.51% and 68.29% similarity with FeoA2 from R. sphaeroides and R. ferroxidans, respectively. This is contrasted by FeoA2 from R. capsulatus exhibiting only 40.22% and 36.56% homology with FeoA1 and Feo2A, that is present in its own genome (table 2). The same pattern occurs when comparing FeoB from these three species: FeoB sequences, translated from the feoA1A2BC operons, exhibit more homology with other species than to FeoB present in other operons (table 2). Taken together, gene organization and sequence homology indicate that the feoA1A2BC operon may be a general features of the Rhodobacter genus, while the feo2B operon may be specific to R. capsulatus.
Table 1.
Ferrous iron uptake genes identified in the genomes of R. capsulatus SB1003, R. sphaeroides 2.4.1 and R. ferroxidans SW2.
| Putative operon name | Gene name | COG | Label Rcap SB1003a | Label Rsph 2.4.1a | Label Rfer SW2a | Product |
|---|---|---|---|---|---|---|
| efeUOB | efeUO | 2822, 0672 | 03065 | N.D. | N.D. | Ferrous iron permease EfeU |
| efeB | 2837 | 03066 | N.D. | N.D. | Dyp-type peroxidase family protein | |
| efeM | 2822 | 03067 | N.D. | N.D. | Protein of unknown function DUF451 | |
|
| ||||||
| feoA1A2BC | feoA1 | 1918 | 00090 | 6020 | 3172 | Ferrous iron transport protein A |
| feoA1 | 1918 | 00091 | 1819 | 3171 | Ferrous iron transport protein A | |
| feoB | 0370 | 00092 | 1818 | 3170 | Ferrous iron transport protein B | |
| feoC | 00093 | 1817 | 3169 | Fe-S dependent transcriptional regulator | ||
|
| ||||||
| feo2AB | feo2A | 1918 | 02028 | N.D. | N.D. | Ferrous iron transport protein A |
| feo2B | 0370 | 02029 | N.D. | N.D. | Ferrous iron transport protein B | |
N.D.: Not detected
R. capsulatus SB1003, R. sphaeroides 2.4.1 and R. ferroxidans SW2 ORFs are accessible in the Integrated Microbial Genomes of the DOE Joint Genome Institute (http://img.jgi.doe.gov/cgi-bin/w/main.cgi) using the following locus tag RCAP_rcc#####, RSP_#### and Rsw2DRAFT_####, respectively, where #### and ##### are the 4 or 5 digits indicated in the table. Regarding R. capsulatus numbering, genes on the plasmid are references as RCAP_rcp##### on the JGI database and p##### in this study.
Table 2.
Identity and similarity in FeoA, FeoB and FeoC from R. capsulatus SB1003, R. shaeroides 2.4.1 and R. ferroxidans SW2. Similarity is indicated using parenthesis.
| FeoA | Rcap FeoA1 | Rcap FeoA2 | Rcap Feo2A | Rsph FeoA1 | Rsph FeoA2 | Rfer FeoA1 | Rfer FeoA2 |
|---|---|---|---|---|---|---|---|
| Rcap FeoA1 | 100 (100) | ||||||
| Rcap FeoA2 | 19.57 (40.22) | 100 (100) | |||||
| Rcap Feo2A | 15.05 (36.56) | 19.57 (36.96) | 100 (100) | ||||
| Rsph FeoA1 | 30.00 (42.22) | 27.59 (36.78) | 18.89 (37.78) | 100 (100) | |||
| Rsph FeoA2 | 18.48 (38.04) | 57.32 (69.51) | 19.10 (37.08) | 22.99 (35.63) | 100 (100) | ||
| Rfer FeoA1 | 31.11 (48.89) | 22.09 (38.37) | 23.86 (40.91) | 52.44 (68.29) | 22.09 (38.37) | 100 (100) | |
| Rfer FeoA2 | 20.65 (42.39) | 50.00 (68.29) | 15.73 (35.96) | 18.39 (34.48) | 65.82 (78.48) | 19.77 (37.21) | 100 (100) |
| FeoB | Rcap FeoB | Rcap Feo2B | Rsph FeoB | Rfer FeoB |
|---|---|---|---|---|
| Rcap FeoB | 100 (100) | |||
| Rcap Feo2B | 30.05 (45.64) | 100 (100) | ||
| Rsph FeoB | 55.56 (71.84) | 29.13 (44.64) | 100 (100) | |
| Rfer FeoB | 56.94 (72.73) | 29.99 (46.76) | 71.08 (82.62) | 100 (100) |
| FeoC | Rcap FeoC | Rsph FeoC | Rfer FeoC |
|---|---|---|---|
| Rcap FeoC | 100 (100) | ||
| Rsph FeoC | 30.95 (41.67) | 100 (100) | |
| Rfer FeoC | 33.33 (44.05) | 35.71 (52.38) | 100 (100) |
Sequences were aligned using the CLC Sequence Viewer software (CLC Bio, Denmark). Similarity and identity scores were calculated based on alignments using the “Ident and Sim” utility (http://www.bioinformatics.org/sms2/ident_sim.html) (Stothard, 2000).
Even though no experimental evidence exists for the role of Feo gene products in Rhodobacter species, it is likely that the general model for the Feo system will prevail. If this is the case then FeoB codes for a membrane bound ferrous iron permease with weak GTPase activity that is enhanced by FeoA. FeoA has a SH3 domain that is thought to be involved in protein-protein interactions. In addition, feoA mutants have been shown to have hampered ferrous iron uptake (Kammler et al., n.d.; Kim et al., 2012). In operons that contain FeoC, this protein is hypothesized to be a transcription factor involved in controlling the feoABC operon. However experimental confirmation of the role of FeoC is still lacking (Fetherston et al., 2012; Guo et al., 2011). Multiple control of the FeoB permease by two different FeoA GTPase enhancers, a local transcription factor FeoC and a global regulator such as Fur might be related to potential toxicity of iron and the need to maintain finely tuned regulation (Kammler et al., n.d.; Cartron et al., 2006). A closer look at the FeoB protein sequences from Rhodobacter species confirm the presence of the GTPase domain as 4 out of the 5 G-motifs are conserved (Dashper et al., 2005). The only major divergence may be Feo2B from R. capsulatus where the G1 motif exhibits a GPPNCG sequence instead of GNPNCG. The hydrophobic C-terminal domain of FeoB, which consists of two GATE motifs in opposite orientation in the membrane, is similar to the Ftrp1 yeast iron permease. Good conservation of the Rhodobacter Gate motifs occurs especially with the presence of two key Cys residues located in the segment IV of Gates 1 and 2, which are potentially involved in iron binding. Once again, the only striking substitution is observed for Feo2B from R. capsulatus where a consensus PC is changed into QC in the Gate 2 segment IV. Preliminary transcription data with R. capsulatus show that both Feo systems are controlled by iron availability, but in a different manner. Indeed, while FeoA1A2BC is overexpressed ~70- and 200-fold under mild and harsh iron stress, respectively; Feo2AB is overexpressed 3-fold and repressed ~4-fold under the same respective conditions (Zappa and Bauer, in preparation). Generation of a dendrogram based on FeoB sequence alignment highlights the occurrence of two major clades: one containing the Rhodobacter FeoB sequences with the exception of R. capsulatus Feo2B (fig. 1). Interestingly, two FeoB loci from Porphyromonas gingivalis are distributed in each clade. In P. gingivalis FeoB1 was shown to be involved in iron uptake, while FeoB2 is shown to be the major manganese transporter (Dashper et al., 2005; He et al., 2006). R. capsulatus Feo2B may therefore be involved in manganese homeostasis.
Fig 1.

Dendrogram of FeoB from various organisms, based on protein sequence alignment. Alignment and dendrogram were built using CLC Sequence Viewer software (CLC Bio, Denmark). The bootstrap analysis algorithm was used, with 100 replicates. Bootstrap values are indicated at each knots and substitution rate at the bottom. Sequences were retrieved from the Integrated Microbial Genomes of the DOE Joint Genome Institute (http://img.jgi.doe.gov/cgi-bin/w/main.cgi). Ecoli: Escherichia coli DH1, Legpneu: Legionella pneumophila Paris, Methaja: Methanocaldococcus jannaschii DSM 2661, Porgin: Porphyromonas gingivalis ATCC 33227, Rcap: Rhodobacter capsulatus SB1003, Rsph: Rhodobacter sphaeroides 2.4.1, Rfer: Rhodobacter ferroxidans SW2, Staphepi: Staphylococcus epidermidis ATCC 12228, Syn: Synechocystis sp. PCC 6803. Accession numbers: Ecoli FeoB, BAJ45144; Legpneu FeoB, YP_125016; Methaja FeoB, NP_247545; Porgin FeoB1, YP_001929201; Porgin FeoB2, YP_001929425; Rcaps FeoB, YP_003576264; Rcaps Feo2B, YP_003578180; Rsph FeoB, ; YP_351866 Rsw2 FeoB, ZP_05845183; Staphepi FeoB1, NP_763744; Staphepi FeoB2, NP_765669; Syn FeoB, NP_440528.
B. EFEUOB(M) SYSTEM
The Elemental Ferrous iron (EfeUOB) system was recently identified as a highly specific to ferrous iron transporter (Grosse et al., 2006; Cao et al., 2007). Orthologues of E. coli EfeOUB, formerly called YcdNOB, can be found in many bacterial genomes {(Rajasekaran, Nilapwar, et al., 2010b). Ferrous iron uptake systems similar but distinct to EfeUOB (called EfeUOB-like) have also been identified in various bacteria such as the P19-Ftr1P system in Campilobacter jejuni (van Vliet et al., 1998), FetMP in E. coli (Koch et al., 2011) and FtrABCD in Bordetella species (Brickman and Armstrong, 2012), making the EfeUOB-type transporter a widely utilized iron uptake strategy among microorganisms. Among the Rhodobacter genomes, R. capsulatus display an EfeUOB operon but no EfeUOB-like system (table 1, fig. 2a). In R. sphaeroides and in R. ferroxidans SW2 there is a surprising absence of EfeUOB and EfeUOB-like operons. Moreover, R. capsulatus also encodes a putative EfeU-EfeO fusion protein (fig. 2b). Although very unusual, a similar EfeU-EfeO fusion has been reported previously (Rajasekaran, Nilapwar, et al., 2010b).
Fig 2.
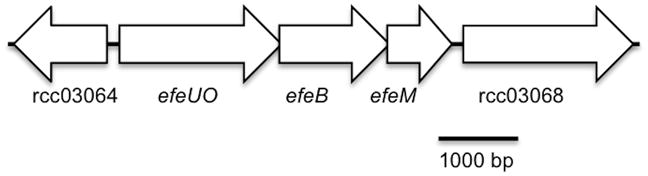
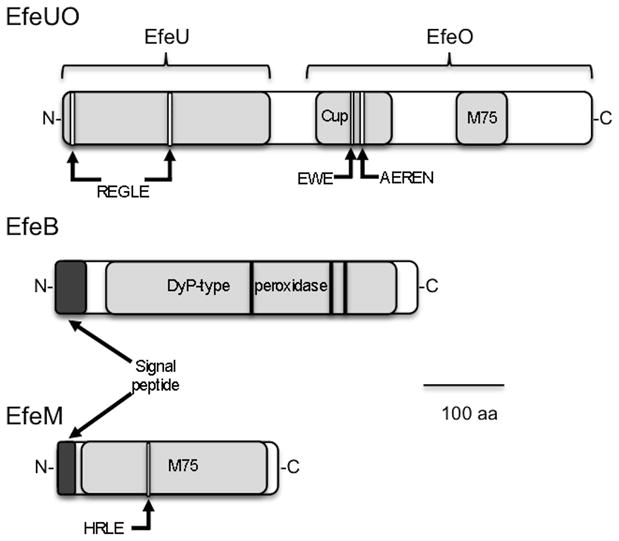
Genetic organization (a) and protein architecture (b) of R. capsulatus EfeUOBM system. Flanking genes orf 03064 and orf 03068 are predicted to encode a membrane protein (HPP family/CBS domain) and a cache sensor protein, respectively. Arrows indicate conserved sequences putatively involved in metal binding. Cup: cupredoxin domain; M75: M75 metallopeptidase domain.
In E. coli, transcription of the efeUOB operon is known to be induced under iron starvation, low pH or in the presence of exogenous copper. The transcription factor Fur and the phosphorelay CpxAR are involved in the iron-and pH-dependent expression, respectively (Cao et al., 2007). Interestingly, expression occurs under aerobic conditions (Cao et al., 2007). In addition to E. coli a few other studies indicate that efeUOB homologues from other species are controlled by iron availability. For example, efeUOB homologues from Bacillus subtilis, (Baichoo et al., 2002; Ollinger et al., 2006), Neisseria menigitidis (Grifantini et al., 2003), the magnetotactic bacterium strain MV-1 (Dubbels et al., 2004) and Magnetospirillum magneticum AMB-1 (Suzuki et al., 2006) are known to be regulated in response to iron.
Only a few experimental studies on the function of efeUOB proteins have been reported so a large part of its understanding comes from sequence analyses. The EfeU protein is homologous to the yeast iron permease Ftr1p, with seven transmembrane helices (TMH). Two of these helices, TMH-I and TMH-IV, contain a iron transport REXXE motif (Grosse et al., 2006; Rajasekaran, Nilapwar, et al., 2010b). These motifs are conserved in the R. capsulatus EfeU domain of the EfeU-EfeO fusion protein (fig. 2b).
EfeB is a periplasmic homodimeric heme containing DyP-type peroxidase (Sturm et al., 2006; X. Liu et al., 2011). EfeB has also been proposed to act as a deferrochelatase, providing iron to E. coli by extracting iron from heme (Létoffé et al., 2009). However, a study with the related protein YfeX could not confirm such a role (Dailey et al., 2011). The R. capsulatus EfeB sequence shows the presence of a conserved TAT signal (Bendtsen et al., 2005) indicating that it is likely exported to the periplasm (fig. 2b). Moreover, alignment with other DyP-type peroxidases (not shown) show conservation of residues involved in heme binding: D235, H330, R347 (Sugano, 2009).
The role of EfeO (COG 2822) is even less clear. EfeO proteins can present different domain organization with the most common one consisting of a N-terminal cupredoxin domain followed by a C-terminal peptidase M75 domain. However EfeOs comprised of only a cupredoxin domain or a peptidase M75 domain are also frequently found in genomes (Rajasekaran, Nilapwar, et al., 2010b). Although nomenclature is still being undefined the trend is to name cupredoxin-containing members as EfeO and cupredoxin-less members that consist of a solo peptidase M75 domain as EfeM (Rajasekaran, Nilapwar, et al., 2010b; Rajasekaran, Mitchell, et al., 2010a). The R. capsulatus putative operon shows two representative of the EfeO/M family (COG 2822), bringing the gene organization to an efeUOBM operon where efeU and efeO are fused (table 1, fig. 2a). The first one is fused to the EfeU domain mentioned above and has the signature of a EfeO protein, i.e. showing both the cupredoxin and M75 domains (fig. 2b). On the other hand the second copy (orf 03067) is a typical M75 containing only EfeM. Motifs potentially involved in copper and iron binding (EWE, EEREN) are conserved in the EfeUO cupredoxin domain (fig. 2b). The EfeM peptide also contains the putative HXXE iron binding sequence while EfeUO M75 domain does not, highlighting probable functional differences between these two EfeO/M like proteins (Rajasekaran, Nilapwar, et al., 2010b; Rajasekaran, Mitchell, et al., 2010a). Finally, EfeO/M are thought to be periplasmic proteins (Sturm et al., 2006; Rajasekaran, Nilapwar, et al., 2010b) as a signal peptide signature sequence is predicted to be present on EfeM (Petersen et al., 2011). The cellular location of EfeO is less clear, nevertheless, export of the EfeO domain in the periplasm could be achieved during folding of EfeUO but this has to be confirmed experimentally.
A proposed mechanism of ferrous iron uptake by the EfeUOB system is based on homology with the yeast permeation/ferroxidation Ftr1p/Fet3p system (Stearman et al., 1996; Kosman, 2003; Rajasekaran, Nilapwar, et al., 2010b). Briefly, ferrous iron in the periplasm binds to the M75 domain of EfeO and is subsequently oxidized to ferric iron by the copper center of cupredoxin domain. The ferric iron is then transferred first to the EfeO cupredoxin domain and then to the permease EfeU. The copper center is finally regenerated by the EfeB peroxidase (Rajasekaran, Nilapwar, et al., 2010b). A similar mechanism could be possible in R. capsulatus, involving both the EfeO domain of EfeUO and EfeM instead of a unique EfeO. Moreover, according to the hypothetic mechanism, EfeU and EfeO are interacting. The fusion of these proteins in R capsulatus is compatible with such a hypothesis. In summary, R. capsulatus is unique among Rhodobacter species in that it seems to have an intact ferrous iron uptake system with analysis of the sequence indicating that it likely functional (fig. 2a and 2b).
III. FERRIC IRON UPTAKE
The major ferric iron uptake system involves mediation by siderophores. This iron uptake pathway has been extensively studied and reviewed (Köster, 2001; Krewulak and Vogel, 2008; Sandy and Butler, 2009; Chu et al., 2010; Hider and Kong, 2010; Krewulak and Vogel, 2011). Briefly, siderophores are small molecules secreted by bacteria, fungi and graminaceous plants that can solubilize ferric iron in aerobic environments, due to their high binding affinity for Fe(III) (10−20 M). Siderophore transporters consist of a tonB-dependent outer membrane ferrisiderophore receptor and an ABC transporter cassette (a periplasmic siderophore binding protein, a permease and an ATPase). Such organization is very well conserved among bacteria and archaea with similar systems used to import heme and vitamin B12. The specificity of the outer-membrane receptor is usually very high compared the specific of the ABC cassette. This sometimes enables the cross-use of the same ABC cassette to uptake different siderophores that have been imported into the periplasm by different outer-membrane receptors.
According to Rhodobacter genome annotations, R. capsulatus and R. sphaeroides display an exbB-exbD-tonB operon while R. ferroxidans does not (table 3). The outer membrane bound ExbB-ExbD-TonB complex enables transfer of protomotive force from the cytoplasmic membrane to the outer-membrane. As such, it drives the energy needed to import the ferrisiderophore from its outer-membrane receptor to the periplasm where the cytoplasmic membrane bound ABC transporter then imports the ferrisiderophore into the cytoplasm. Given that the ExbB-ExbD-TonB complex is absent in R. ferroxidans it is not surprising that no siderophore uptake system is annotated in the R. ferroxidans SW2 genome. On the other hand, 8 and 5 complete siderophore uptake systems are present in the R. capsulatus SB1003 and R. sphaeroides 2.4.1 genomes, respectively. Looking closer at the sequences and the genomic environment indicates that the respective TonB-dependent uptake systems orf 3358 to orf 3362 and orf 2102 to orf 2105 in R. capsulatus and R. sphaeroides, respectively, are more likely to be involved in vitamin B12 uptake rather than siderophore transport. These strains still potentially encoding 7 and 4 siderophore uptake systems as detailed in table 3. Interestingly, no siderophore synthesis gene cluster is present in either genome, meaning that R. capsulatus and R. sphaeroides likely scavenge siderophores from other species (termed xenosiderophores) to fulfill their iron needs. Such situation is not unusual as some species even rely on xenosiderophores to provide enough iron for growth (D'Onofrio et al., 2010). This situation happens between bacteria as well as between bacteria and fungi (Kosman, 2003). Moreover, it has been shown that bacteria can use “improbable siderophores” such as α-keto acids instead of classical siderophores such as catecholates and hydroxamates (Reissbrodt et al., 1997).
Table 3.
Ferric-siderophore uptake genes identified in the genomes of R. capsulatus SB1003, R. sphaeroides 2.4.1 and R. ferroxidans SW2.
| Gene product | COG | Label Rcap SB1003a | Label Rsph 2.4.1a | Label Rfer SW2a |
|---|---|---|---|---|
| ExbB | 0811 | 02375 | 0920 | N.D. |
| ExbD | 0848 | 02376 | 0921 | N.D. |
| TonB | 0810 | 02377 | 0922 | N.D. |
|
| ||||
| Uptake element | Label in Rcap SB1003a | Comments from annotations | ||
|
| ||||
| TOMR | 4773 | 00111 (fhuE) | Ferrichrome uptake system | |
| PBP | 4607 | 00108 (fhuD1) | Presence of a putative esterase (orf00110) | |
| IMP | 4605 | 00106 (fhuB1) | ||
| 4606 | 00107 (fhuB2) | |||
| ATPase | 4604 | 00105 (fhu-1) | ||
|
| ||||
| TOMR | 4774 | 01049 | Outer membrane receptor for monomeric catechols | |
| PBP | 0614 | 01047 | Presence of a nearby enterochelin esterase (orf 01050) and an AraC-like regulator (orf 01048) | |
| IMP | 0609 | 01046 | ||
| ATPase | 1120 | 01045 | ||
|
| ||||
| TOMR | 1629 | 01429 | Presence of a protein from the major facilitator superfamily: potential role of PBP? | |
| PBP | N.D. | N.D. | ||
| IMP/ATPase | 1132 | 01426/01427 | ||
|
| ||||
| TOMR | 4774 | 01433 | Enterobactin uptake systeme | |
| PBP | 0614 | 01434 (fepB1) | Presence of a nearby siderophore interacting protein (orf 01438) and two AraC-like regulators (orf 01431 and 01432) | |
| IMP | 0609 | 01435 (fepD1) | ||
| 4779 | 01436 (fepG1) | |||
| ATPase | 1120 | 01437 (fepC1) | ||
|
| ||||
| TOMR | 1629 | 01445 | Enterobactin uptake systeme | |
| PBP | 4592 | 01444 (fepB2) | Presence of a nearby siderophore interacting protein (orf 01438) | |
| IMP | 0609 | 01443 (fepD2) | ||
| 4779 | 01442 (fepG2) | |||
| ATPase | 1120 | 01441 (fepC2) | ||
|
| ||||
| TOMR | 1629 | p00051 | Presence of one TOMR with two ABC transport systems | |
| PBP | 0614 | p00046 | ||
| p00050 | ||||
| IMP | 0609 | p00045 | ||
| p00049 | ||||
| ATPase | 1120 | p00044 | ||
| p00048 | ||||
|
| ||||
| TOMR | 4774 | p00112 (fhuA) | Ferrichrome uptake systeme | |
| PBP | 0614 | p00111 (fhuD2) | Presence of a nearby AraC-like regulator (orf p00113) | |
| IMP | 0609 | p00110 (fhuB3) | ||
| 1132 | p00108 | |||
| ATPase | 1132 | p00108 | ||
| 1120 | p00109 (fhuC2) | |||
|
| ||||
| Uptake element | Label in Rsph 2.4.1a | Comments from annotations | ||
|
| ||||
| TOMR | 4774 | 1440 | Hydroxamate – type siderophore uptake system orf 1438 is a fused subunit permease | |
| PBP | 0614 | 1439 | ||
| IMP | 0609 | 1438 | ||
| 4779 | ||||
| ATPase | 1120 | 1437 | ||
|
| ||||
| TOMR | 4771 | 3223 | Enterochelin/colicin uptake system | |
| PBP | N.D. | A periplamic protein, annotated as histidine kinase, is in the same operon (orf 03225): a potential PBP? | ||
| IMP | 4606 | 3220 | ||
| 4605 | 3221 | |||
| ATPase | 4604 | 3222 | ||
|
| ||||
| TOMR | 4774 | 3417 | Ferrichrome uptake system | |
| PBP | 0614 | 3416 | ||
| IMP | 0609 | 3415 | ||
| 4779 | 3414 | |||
| ATPase | 1120 | 3413 | ||
|
| ||||
| TOMR | 4773 | 4273 (fhuA) | Ferric coprogen or ferric rhotorulic acid uptake system. In the same putative operon of a FecRI system (orf 4274 and 4275) | |
| PBP | 06014 | 4272 (fhuD) | ||
| IMP | 0609 | 4271 (fhuB) | ||
| ATPase | 1120 | 7397 | ||
N.D.: Not detected; TOMBR: TonB-dependent outer-membrane receptor; PBP: Periplasmic binding protein; IMP: Inner-membrane protein.
See table 1.
Once a ferrisiderophore complex had entered the cytoplasm, its fate is not well known. One of the most studied siderophore processing events is the involvement of the enterobactin esterase in E. coli. This enzyme hydrolyses enterobactin, producing trimers of dihydroxybenzoylserine that weakens the bond with ferric iron. Enterobactin esterase also seems to act as an enterobactin-specific reductase that reduces the ferric iron into a soluble ferrous iron which triggers iron release from the siderophore (Andrews et al., 2003; Rudolph et al., 2006). Importantly, two putative esterases both located in siderophore uptake gene clusters are annotated in the R. capsulatus genome, orf 00110 and orf 01050. The latter is actually annotated as an enterobactin (enterochelin) esterase. The presence of this gene is very unusual among α-proteobacteria (Rudolph et al., 2006).
IV. IRON ABC TRANSPORTERS
Some bacteria also contain metal-ABC transporters that exhibit specificity for iron. Unlike classical siderophore-based iron uptake systems, the metal-ABC transporters are not dependent on an outer-membrane receptor for iron or siderophore transport into the periplasmic space. The best characterized version of this class of transporters is the FbpABC system studied in Neisseria gonorrhoeae, Neisseria meningitidis, Campilobacter jejuni, Bordetella pertussis, Marinobacter species, Vibrio cholerae, Pasteurella multocida and Actinobacillus pleuropneumoniae (called sometimes AfuABC) (Chin et al., 1996; Khun et al., 1998; Paustian et al., 2001; Tom-Yew et al., 2005; Wyckoff et al., 2006; Brickman et al., 2011; Strange et al., 2011; Amin et al., 2012). This transporter is specific to ferric iron and consists in a Fe(III)-binding periplasmic protein, a membrane permease and an ATPase. Also, in Neisseria and Bordetella, this transporter also undergoes tonB-independent uptake of endogenous siderophores or xenosiderophores (Brickman et al., 2011; Strange et al., 2011). Similar transporters have been characterized in Serratia marcescens and Haemophilus influenzae under the respective names SfuABC and HitABC (Angerer et al., 1992; Sanders et al., 1994). The cyanobacterium Synechocystis sp. PCC6803 also contains this transport system, with the particularity of having two Fe(III)-binding proteins instead of one (Katoh, Hagino, Grossman, et al., 2001b; Katoh, Hagino and Ogawa, 2001a; Badarau et al., 2008).
Present in the annotated R. sphaeroides genome is a putative homologue of FbpA, the periplasmic substrate-binding protein of the FbpABC system (Tom-Yew et al., 2005). Looking closer at the genome reveals that this ORF (orf 2913) is not part of an FbpABC operon. On the other hand, a second ORF matching FbpA’s COG 1840 appears to share an operonal organization with three ORF’s that subunits of an ABC transporter: two inner membrane proteins and an ATPase subunits. These four ORFs (orf 0346 to orf 0349) may thus constitute an FbpABC transport system in R. sphaeroides. Such a transport system is also present in R. ferroxidans and R. capsulatus where a similar gene organization is found (table 4).
Table 4.
Iron-ABC transporter genes identified in the genomes of R. capsulatus SB1003, R. sphaeroides 2.4.1 and R. ferroxidans SW2.
| ABC transporter type | Gene name | COG | Label Rcap SB1003a | Label Rsph 2.4.1a | Label Rfer SW2a | Product |
|---|---|---|---|---|---|---|
| FbpABC | 1840 | 01369 | 0346 | 1135 | ABC transporter, substrate binding protein | |
| 4132 | 01370 | 0347 | 1134 | ABC transporter, inner membrane subunit | ||
| 1177 | 01371 | 0348 | 1135 | ABC transporter, inner membrane subunit | ||
| 3842 | 01371 | 0348 | 1135 | ABC transporter, ATPase subunit | ||
|
| ||||||
| SitABCD | sitA | 0803 | N.D. | 0904 | N.D. | ABC Mn+2/Fe+2 transporter, periplasmic substrate-binding protein SitA |
| sitB | 1121 | N.D. | 0905 | N.D. | ABC Mn+2/Fe+2 transporter, ATPase subunit SitB | |
| sitC | 1108 | N.D. | 0906 | N.D. | ABC Mn+2/Fe+2 transporter, inner membrane subunit SitC | |
| sitD | 1108 | N.D. | 0907 | N.D. | ABC Mn+2/Fe+2 transporter, inner membrane subunit SitD | |
N.D.: Not detected
See table 1.
Another metal-ABC transporter system, YfeABCD/SitABCD, is thought to be involved in iron uptake, although its specificity between iron and manganese is not clear. Indeed, the YfeABCD system has been shown to transport both Fe(III) and Mn(II) in Yersinia pestis and Photorhabdus luminescens, being involved in virulence mechanisms in both cases (Bearden and Perry, 1999; Watson et al., 2010). The SitABCD system has also been shown to transport both Fe(II), instead of Fe(III), and Mn(II) in Sinorhizobium meliloti and Shigella flexneri (Chao et al., 2004; Platero:2004gi Fisher et al., 2009). In an E. coli avian pathogenic strains, the transport of Fe(II), Fe(III) or Mn(II) was varying as a function of the strain genetic background (Sabri et al., 2006). Finally, SitABCD in Salmonella enterica Serovar Typhimurium was shown to be able to uptake both Fe(II) and Mn(II) but with an affinity for Mn(II) that is stronger and physiologically more relevant (Janakiraman and Slauch, 2000; Kehres et al., 2002). An orthologous SitABCD transporter is annotated in the genome of R. sphaeroides 2.4.1, but not in R. capsulatus SB1003 and R. ferroxidans SW2 genomes (table 4). However, there is no sequence signature in the SitABCD system in general and in the R. sphaeroides SitABCD cluster in particular, that enables prediction of its involvement in either iron or/and manganese transport. Nevertheless, R. sphaeroides does not seem to have an NRAMP-type manganese dedicated transporter MntH, while R. capsulatus and R. ferroxidans SW2 do contain the MntHR Mn transport system (table 5). Assuming that R. sphaeroides has regular manganese needs, and that no other unknown manganese transporters are present, it is reasonable to hypothesize that the SitABCD system in R. sphaeroides is dedicated to manganese uptake rather than to iron.
Table 5.
Putative Ferrous iron efflux pumps (COG 0053) identified in the genomes of R. capsulatus SB1003, R. sphaeroides 2.4.1 and R. ferroxidans SW2, with the sequence corresponding to Fe/Zn binding sites dimerization in the E. coli FieF (Lu and Fu, 2007; Nies, 2011).
| Organism/gene locus | Z1 | Fe/Zn binding site Z2 dimerization | Z3/Z4 |
|---|---|---|---|
| E. coli K12 FieFa | D45X3D49-H153X3D157 | D68DNHX3H75 | H232-H261-H283XD285 |
| Rcaps/orf 00522 | D47X3D51-H156X3D160 | D70DDHX3H77 | H236-H265-H287XD299 |
| Rsph/orf 0463 | E40X3N44-H148X3D152 | D63ANHX3H70 | H229-H258-H280XE282 |
| Rfe/orf 1374 | D50X3D54-H160X3D164 | D73EDHX3H80 | H240-H269-H291XD293 |
| Rfe/orf 2021 | H88X3D92-H196X3D200 | S111RTFX3L118 | H247-L303-H419XV421 |
E. coli K12 Fief accession number NP_418350
numbering relative to each sequence is given. X: random amino acid. Xn: succession of n random amino acids.
V. HEME IRON USAGE
Heme is often a crucial iron source for pathogens that scavenge it from their host. However, heme uptake is also frequently an iron acquisition mechanism that occurs in beneficial symbiotic bacteria (Nienaber et al., 2001; Runyen-Janecky et al., 2010; Anzaldi and Skaar, 2010; Septer et al., 2011). A major hemin uptake system is the PhuRSTUVW-type heme transporter as described in Pseudomonas aeruginosa, where PhuR is a TonB-dependent outer-membrane heme receptor. The PhuT subunit of the PhuRSTUVW heme transporter is the periplamic heme-binding protein while PhuUVW is an ABC transporter and PhuS is a cytoplasmic protein (Anzaldi and Skaar, 2010). Extensive characterization of similar systems has been undertaken in the pathogenic bacteria Bordetella pertussis, Yersinia pestis, Yersinia enterolitica, Shigella dysenteriae, Vibrio cholera, Campylobacter jejuni, Bartonella quintana and E. coli O157:H7 (Anzaldi and Skaar, 2010) with the transporters names with varying terminology. A homologue has described in the symbiotic Rhizobiales Bradyrhizobium japonicum, Rhizobium leguminosarum and Sinorhizobium meliloti where it is called HmuQR-HmuTUV, HmuPSTUV and ShmR-HmuPSTUV, respectively (Nienaber et al., 2001; Wexler et al., 2001; Anzaldi and Skaar, 2010; Amarelle et al., 2010; Amarelle et al., 2008).
R. capsulatus displays an orthologous gene cluster where it is annotated as hmuRSTUV (loci from orf 00094 to orf 00098). Interestingly, R. sphaeroides and R. ferroxidans do not seem to code for this heme transporter. Thus, unlike other Rhodobacter representatives, R. capuslatus is likely to be able to use heme from de novo synthesis as well as form exogenous sources. Unlike Bradyrhizobium japonicum and Rhizobium leguminosarum, the R. capsulatus heme uptake gene cluster does not have a nearby TonB-ExBD subunit. It may therefore rely on a homologous gene located on a different region of the genome (table 3) or contain an unidentified protein that provides this function. Also, the R. capsulatus Hmu system lacks the HmuP regulator that has been identified in Bradyrhizobium japonicum and Sinorhizobium meliloti that functions as a co-activator of this heme uptake system along with Irr (Amarelle et al., 2010; Escamilla-Hernandez and O'Brian, 2012). A BLAST analysis on the Rhodobacter clade indicates possible candidates for an HmuP transcription factor in R. capsulatus (orf 01112) and in R. sphaeroides (orf 6006) but not in R. ferroxidans. However, it should be cautioned that these putative HmuP homologues have poor sequence conservation, and no genome context with heme transporters, so their actual function needs to be experimentally confirmed.
The heme uptake gene cluster in R. capsulatus also contains hmuS that in other organisms has been shown to be involved in heme degradation. The mechanism of action of HmuS orthologues is not clear but seems eclectic: i) Some HmuS were shown to enzymatically degrade heme; ii) Some seem to operate via a non-enzymatic process with H2O2; iii) Some may store and/or traffic heme to a heme oxygenase (Anzaldi and Skaar, 2010; M. Liu et al., 2012; O'Neill et al., 2012; Barker et al., 2012).
Based on COG analysis, it is notable that R. capsulatus contains a second putative HmuS locus (orf 03488) that is located next to a putative coproporphyrinogen oxidase III encoding gene. R. sphaeroides only has one HmuS homolog (orf 0228), while R. ferroxidans has no HmuS annotated. Presence of a putative heme degrading system is thought to be crucial to use heme as an iron source as well as to prevent heme toxicity by ensuring that there is no pool of unbound free heme in the cell(Frankenberg-Dinkel, 2004; Anzaldi and Skaar, 2010). Moreover, a typical heme oxygenase (BphO-like) is not present in either R. capsulatus or R. ferroxidans, while there are two annotated heme oxygenase genes annotated in R. sphaeroides. These are associated with bacteriophytochrome encoding genes (orf 4191, orf 7212). As shown recently in Pseudomonas aeruginosa, bacteriophytochrome-associated BphO is not involved in heme degradation from exogenous heme uptake, but only in holo-bacteriophytochrome synthesis from de novo synthesized heme (Barker et al., 2012). Atypical IsdG-like heme oxygenases HmuQ and HmuD, were recently characterized in Bradyrhizobium japonicum (Puri and O'Brian, 2006; Skaar et al., 2006). A BLAST analysis against the Rhodobacter clades reveals the presence of a potential HmuQ/D-like protein in R. sphaeroides (orf 0826) however further work is needed to confirm this identity. Finally, as detailed in chapter II.B, the EfeUOB system has been shown to be able to extract iron from heme by a deferrochelatase activity, although such activity is debated (Létoffé et al., 2009; Dailey et al., 2011). Such an activity could provide iron to R. capsulatus, the only representative of the Rhodobacter genus displaying the efeUOB gene cluster (table 1).
A final point regarding heme usage is the presence of “heme exporter” gene clusters in the annotation of the three Rhodobacter strains of interest. Closer look reveals they corresponds to CcmABCDG system used for c-type cytochrome synthesis. It has been shown with R. capsulatus that a mutation that disrupts cytochrome c maturation results in massive secretion of porphyrins (S. W. Biel and A. J. Biel, 1990). The authors did not confirm if a ccmABCDG operon mutation was truly causing this phenotype but did speculate that there is a connection between iron homeostasis, heme and cytochrome synthesis (LASCELLES, 1956; S. W. Biel and A. J. Biel, 1990)
VI. MANAGING IRON TOXICITY
As stated in the introduction, free iron can be extremely toxic. Thus, once cellular iron needs are fulfilled, being able to counterbalance the intake to avoid overload is definitely of interest for a cell. There are two ways are available in bacteria to counterbalance iron intake. One mechanism consists in reversing the uptake process by excreting excess iron using iron efflux pumps. The second process involves the binding or sequestration of iron in a dedicated peptide where it is stored in a harmless state. These peptides are called ferritins and/or bacterioferritins.
A. IRON EFFLUX PUMP
The efflux of iron in bacteria is not well characterized although the last decade has highlighted a prominent role of Cation Diffusion Facilitators (CDF). The CDF family is ubiquitous in the three domains of life and grouped together as heavy metal transporters. They are classified into three subfamilies according to their metal substrate specificity: Zn-CDF, Fe/Zn-CDF and Mn-CDF (Montanini et al., 2007; Nies, 2011). Despite having a “favorite” metal substrate, they usually are capable of transporting a wide array of metals (Munkelt et al., 2004; Montanini et al., 2007). One of the most studied representatives is the ferrous iron efflux protein (FieF, also known as YiiP). Its presence was shown to increase E. coli’s tolerance to iron and to lower the total iron cellular content (Grass et al., 2005). Although FieF was biochemically well characterized in vitro using zinc as a substrate, its main substrate in vivo is ferrous iron and, as such, is likely to an important role in iron homeostasis (Nies, 2007). An ORF was annotated as FieF in R. capsulatus SB1003 (orf 00522). Using the same COG number (COG 0053), candidates were found in the two other Rhodobacter genomes: orf 0463 in R. sphaeroides 2.4.1, and orf 1374 and orf 2021 in R. ferroxidans SW2. E. coli FieF has four metal binding sites named Z1 through Z4 (Nies, 2011; Lu and Fu, 2007). Alignment of the putative Rhodobacter FieF sequences shows good conservation at most of these sites, with the exception of orf 2021 in R. ferroxidans. Other CDF genes were also found in these Rhodobacter genomes, under COG 1230, but their sequences align better with the E. coli zinc transporter ZitB then with the iron transporter Fief (data not shown). However, given the relaxed substrate specificity of CDFs, it is worth noting that the R. capsulatus ZitB homologue (orf 00089) is located next to the feoA1A2BC putative ferrous iron uptake system.
B. STORAGE AND DETOXIFICATION
The ferritin family consists of three types of protein that form distinct phylogenetical clades: the ferritins (Ftn), the bacterioferritins (Bfr) and the DNA-binding proteins from starved cells (DPS). Ftn and Bfr have strong structural homology with each consisting of homooligomers of 24 subunits. DPS are comprised of 12-mers of the same subunit (Bou-Abdallah, 2010; Andrews, 2010). Ftn and Bfr form ball-shaped complexes with an outer diameter of ~120Å that can store up to 4500 iron atoms in a ~80Å diameter cavity. The iron in this cavity consists of either amorphous iron with inorganic phosphate or crystalline ferrihydrite. DPS also form a ball like structure only they have an outer diameter of ~95Å that can handle up to ~500 molecules of iron (Bou-Abdallah, 2010; Carrondo, 2003; Andrews, 2010). Cellular iron concentration is estimated to ~10−4M, which is far above the solubility of this metal. This concentration is reached do to the presence of these iron sequestration storage proteins that concentrate and store the metal in non-reactive form (Theil and Goss, 2009). By isolating the toxic iron from cellular machinery, storing it and releasing upon needs, ferritins functions almost as cellular organelles.
The ferritin family is ubiquitous with Ftn found in all three domains of life while Bfr and DPS are specific to Bacteria and Archaea (Andrews, 2010). Their activity relies on ferroxidase centers consisting in intrasubunit di-iron centers in Ftn and Bfr and intersubunit di-iron center in DPS (Andrews, 2010). These centers channel Fe(II) into the cavity by oxidizing it into Fe(III) which generates the insoluble oxidized form of iron stored inside the complex core. It seems that Ftn and Bfr uses mostly O2 as the iron oxidant while DPS use H2O2 (Bou-Abdallah, 2010; Andrews, 2010). Ftn and Bfr main role is for iron storage while the function of DPS is thought to be as a detoxifier that protects DNA from redox stress generated by the iron mediated Fenton reaction (equations 1–3) (Andrews, 2010). It is quite common for bacteria to have each ferritin types as well as several copies of one type of ferritin. For example, E. coli is typical with the presence of two Ftn, one Bfr and one DPS (Andrews et al., 2003; Chiancone et al., 2004). The major difference between Ftn and Bfr is the presence of 12 b-type heme moieties in the Bfr complex (Cobessi et al., 2001; Bou-Abdallah, 2010; Carrondo, 2003; Andrews, 2010). After decades of mystery about the role of the heme groups, it has recently been shown that they are involved in the mobilization of iron back to the cytosol. Thus while ferroxidase centers oxidize the iron during the mineralization process that internalizes the iron into the Bfr, the heme moieties reduce the core iron during its export into the cytoplasm. Ferroxidase centers and heme moieties functions independently (Yasmin et al., 2011).
Analysis of the Rhodobacter genomes reveals the presence of Ftn and Bfr proteins (table 6). R. sphaeroides has a membrane-bound ferritin (orf 0850). A BLAST analysis on the genome of R. capsulatus also reveals the presence of a homologue in this species (orf 03466). Interestingly, no homologues were found in the R. ferroxidans genome in the NCBI database but an ORF with the same COG number could be found in the JGI database (orf 0178). It is annotated as rubrerythrin, which is likely to be the ancestral form of the ferritin-like protein family (Andrews, 2010). The annotation is probably incorrect as the three putative Ftn share strong homology (more than 70% identity and more than 80% similarity, as shown in table 7). Indeed, they are all 325 amino acid long while Bfr are in the 160 amino acid range. Conserved domains analysis showed that they all consist in an Ftn-like N-terminal domain of ~140–150 amino acids followed by a C-terminal CCC1 domain of ~120–140 amino acids (not shown). CCC1 domains are involved in iron and manganese transport. In yeast, CCC1 is a vacuole transmembrane protein responsible for iron and manganese accumulation in vacuoles. Finally, these genes share similarity in terms of genomic organization, as seen on figure 3, indicating an ancient gene cluster that was conserved during evolution.
Table 6.
Ferritin-family proteins identified in the genomes of R. capsulatus SB1003, R. sphaeroides 2.4.1 and R. ferroxidans SW2.
| Ftn type | Gene name | COG | Label Rcap SB1003a | Label Rsph 2.4.1a | Label Rfer SW2a | Comments |
|---|---|---|---|---|---|---|
| Ftn | mbfA | 1633 | 03466 | 0850 | 0178 | Membrane bound ferritin |
| Bfr | bfr | 2193 | 00913 | 1546 | N.D. | |
| bfr2 | 2193 | 3342 | ||||
| DPS | 0783 | N.D. | N.D. | N.D. |
N.D.: Not detected
See table 1.
Table 7.
Identity and similarity in MbfA and Bfr from R. capsulatus SB1003, R. shaeroides 2.4.1 and R. ferroxidans SW2. Similarity is indicated using parenthesis.
| MbfA | Rcap | Rsph | Rfer |
|---|---|---|---|
| Rcap | 100 (100) | ||
| Rsph | 73.54 (81.85) | 100 (100) | |
| Rfer | 79.08 (86.15) | 73.54 (82.46) | 100 (100) |
|
| |||
| Bfr | Rcap | Rsph Bfr | Rsph Bfr2 |
|
| |||
| Rcap | 100 (100) | ||
| Rsph Bfr | 67.70 (79.50) | 100 (100) | |
| Rsph Bfr2 | 66.67 (78.79) | 82.42 (87.27) | 100 (100) |
Sequences were aligned using the CLC Sequence Viewer software (CLC Bio, Denmark). Similarity and identity scores were calculated based on alignments using the “Ident and Sim” utility (http://www.bioinformatics.org/sms2/ident_sim.html) (Stothard, 2000).
Fig 3.
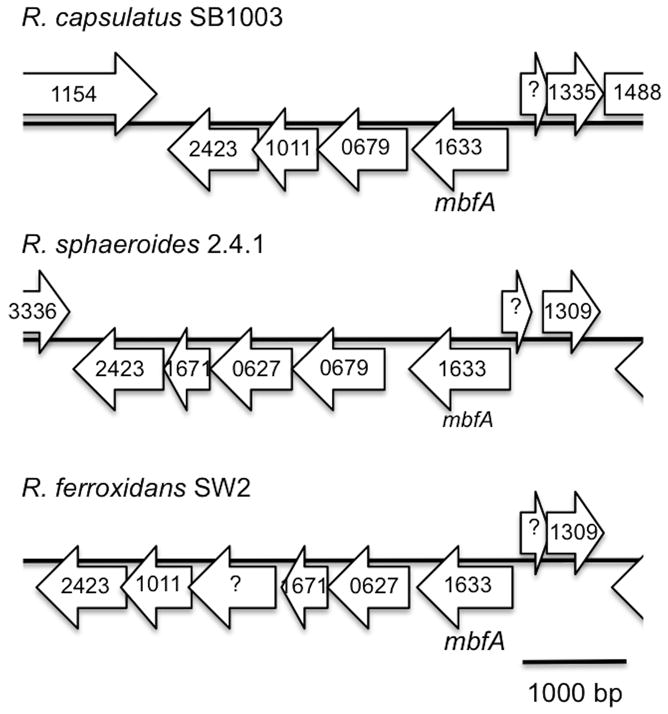
Genomic environment of membrane-bound ferritin encoding genes, mbfA, in R. capsulatus SB1003, R. sphaeroides 2.4.1, R. ferroxidans SW2. COG numbers of each reading frame are indicated.
Beside a membrane-bound ferritin, Bfr homologues are present in the genomes of R. capsulatus and R. sphaeroides, but not in R. ferroxidans (table 6). In fact, R. sphaeroides displays two Bfr, one on each chromosome, while R. capsulatus has only one. While R. capsulatus Bfr share almost 80% similarity with each of the R. sphaeroides Bfr, the latters show a similarity score to each other that is close to 90% similarity (table 7). This suggests that a single bfr gene duplicated in R. sphaeroides. Unlike membrane-bound ferritin, Bfr show weaker genomic conservation. One of R. sphaeroides Bfr (orf 1446) is organized in a putative operon with a Bfr-associated-ferredoxin (orf 1447), an iron-regulated protein (orf 1448) and a hypothetical protein (orf 6006). Interestingly, the latter shows homology with HmuP, a heme uptake regulator described in Bradyrhizobium japonicum and Sinorhizobium meliloti as reported in section V. No iron homeostasis genes are found in the vicinity of the other R. sphaeroides bfr gene (not shown). Regarding the R. capsulatus Bfr, it seems to be part of an operon that contains a hypothetical protein that aligns well with R. sphaeroides Bfr-associated-ferredoxin (orf 1447). Association of a bfr gene with a ferrodoxin is very common in bacteria (Rodionov et al., 2006). This connotation may suggest that orf 1446 was the “original” bfr in R. sphaeroides before being duplicated without the ferredoxin gene.
Some experimental work has been undertaken with the Rhododacter Bfr. The first Bfr to be studied was with R. sphaeroides although most work has been done on the R. capsulatus homologue (Meyer and Cusanovich, 1985). Between 900 and 1000 amorphous iron atoms along with 600 phosphate molecules are contained in each Bfr complex. It is located in the cytoplasm and its expression in “normal” growth medium is stable. Nevertheless, a moderate control by iron is observed as iron starvation induces a decrease in Bfr cellular content, while iron replete conditions promotes accumulation of Bfr in the cell (Ringeling et al., 1994; Penfold et al., 1996). The Rhodobacter Bfr structure was the second Bfr structure to be elucidated after that of E. coli Bfr (Cobessi et al., 2001).
Regarding iron storage in R. sphaeroides, some transcriptional mechanisms are reported. Transcription of Bfr encoded by orf 1546, along with its associated ferredoxin appears to be controlled by iron availability as their mRNA levels increase when iron is depleted. The transcription factor Irr is implicated in this control (Peuser et al., 2012). The other Bfr (orf 3342), which lacks a linked ferredoxin, is almost insensitive to iron levels and shows a weak Irr regulation profile (Peuser et al., 2012). Finally, the Ftn-like MbfA displays weak iron control but a strong Irr-dependence. In fact, as predicted by Rodionov et al., MbfA was shown to be under the direct regulation of Irr (Rodionov et al., 2006; Peuser et al., 2012).
VII. IRON HOMEOSTASIS REGULATORS
The regulation of iron homeostasis involves a complex overlapping set of global regulators as well as more specialized regulators dedicated to the control of specific iron homeostasis genes, such as the control of siderophore synthesis and ferrous iron uptake. Such local regulators are diverse and belong to the following families: i) AraC-type transcription factor (Pradel et al., 1998; Ducey et al., 2005; Hollander et al., 2011; Fantappiè et al., 2011); ii) two component systems (Steele et al., 2012); iii) extracytoplasmic function (ECF) σ factors (Braun, 1997; Koster et al., 1994; Braun et al., 2003); iv) LysR-type transcription factor (Litwin and Quackenbush, 2001; Vanderpool and Armstrong, 2003); v) small RNA regulators (Massé et al., 2007; Metruccio et al., 2009; Huang et al., 2012; Ducey et al., 2009; Smaldone et al., 2012). Pseudomonas aeruginosa is an example of organism that combine all the aforementioned regulators as well as several global regulators (Vasil, 2007; Cornelis et al., 2009). Computational analysis showed that R. sphaeroides possesses FecRI (orf 4274 and 4275, table 3) where FecI is a homologue of the ECF-type σ factor and FecR is periplasmic regulator of FecI (Rodionov et al., 2006). Based on its genomic environment, the fecRI operon may be co-transcribed with a siderophore uptake gene cluster that could be under its control. FecRI is involved in Fe(III)-citrate uptake in E. coli but homologues such as PupBI and HurRI have been characterized in other species. They are involved in pseudobactin siderophores and heme uptake, respectively (Koster et al., 1994; Vanderpool and Armstrong, 2003). Genome analysis shows also that R. capsulatus has four AraC-like transcription factors located next to siderophore uptake gene clusters (Rodionov et al., 2006). These AraC encoding genes are given table 3.
In many cases, transcription factors dedicated to the control of specific iron homeostasis genes are usually themselves under the control of a global transcription factor (Fantappiè et al., 2011; Pradel et al., 1998; Braun, 1997) or act as co-regulators with a global regulator (Escamilla-Hernandez and O'Brian, 2012). For a long time, the paradigm of iron homeostasis in bacteria was associated with the Ferric Uptake Regulator (Fur) that was first characterized in E. coli. Additional studies quickly showed that Fur homologues are widespread in both gram-negative (proteobacteria) and gram-positive (Firmicutes) bacteria, as well as in some cyanobacteria (Carpenter et al., 2009; Lee and Helmann, 2007). The basic mechanism of Fur regulation consists of transcriptional repression of iron uptake genes under iron replete conditions by Fur that contains a bound Fe(II) (Fe(II)-Fur). However, under conditions of iron limitation, Fur without bound iron (apo-Fur) is incapable of binding these promoters, derepressing the expression of these iron uptake genes (Rudolph et al., 2006; Lee and Helmann, 2007; Carpenter et al., 2009). The regulation by Fur is now known to be more complex since Fe(II)-Fur can also activate genes in an indirect manner, via the derepression of a small regulatory RNA (Massé et al., 2007; Metruccio et al., 2009; Huang et al., 2012; Ducey et al., 2009; Smaldone et al., 2012). Finally, direct transcriptional activation by apo-Fur on a target gene promoter was also observed (Carpenter et al., 2009; Lee and Helmann, 2007).
The Rhizobiale and Rhodobacterale phyla are also known to regulate iron homeostasis genes by the global regulator Irr (Iron response regulator) (Rodionov et al., 2006; Rudolph et al., 2006; Johnston et al., 2007). It has been proposed that upon the appearance of Irr, Fur evolved into a manganese uptake regulator. Consequently the Fur homologues in Rhizobiales and Rhodobacterales have been renamed Mur. [Another evolutionary event is the inclusion of a third iron regulator, RirA, that is present in the Rhizobiales (Rhodobacterales only have Fur/Irr) (Rodionov et al., 2006; Rudolph et al., 2006; Johnston et al., 2007)]. Fur and Irr belong to the same Fur-superfamily of metal regulators and thus share strong sequences homology. Nevertheless, their mechanisms of action are very different. Fur directly binds elemental iron which subsequently affects the ability of Fur to activate or repress gene expression. Like Fur, Irr can both activate and repress iron-dependent gene expression. However, Irr does not bind free iron but instead monitors the iron level indirectly by sensing the level of heme biosynthesis. Specifically, Irr interacts and monitors the activity of ferrochelatase, the last enzyme of the heme synthesis pathway. Irr obtains a heme from ferrochelatase which results in targeting Irr for degradation (Rudolph et al., 2006; Rodionov et al., 2006; Small et al., 2009).
Regarding Rhodobacter genomes, members of the Fur-family are represented in each of the sequenced genomes. However, a closer look reveals that: i) Fur is missing in R. capsulatus, while present in R. sphaeroides and R. ferroxidans; ii) Each genome contains a copy of Irr; iii) Each genome also has a Zur encoding gene (table 8). The latter is also a Fur-family member but specialized in zinc uptake (Lee and Helmann, 2007). It is readily identifiable as Zur is consistently located along with a znuABC gene cluster that encodes a zinc transporter. Although sharing high sequence similarity, Fur and Irr can be discriminated from their sequence analysis. A main feature distinguishing Fur and Irr is the HHDH Fe(II) binding motif that is a signature of Fur, which becomes a HHH (or HQH) in Irr. This aspartate deletion transforms this site from a Fe(II) to a heme binding site (Rudolph et al., 2006). Thus, while R. sphaeroides and R. ferroxidans display the classical Rhodobacterale iron homeostasis transcription factor features, R. capsulatus has a very unusual set of regulators. Moreover, R. capsulatus, along with Mesorhizobium loti, is among the very rare α-proteobacteria that also have a manganese MntHR uptake system, most likely acquired through horizontal gene transfer (Rodionov et al., 2006). Based on in silico analysis and studies on Rhizobiales, it has been proposed that the acquisition of Irr as global iron regulator may have pushed Fur into a more marginal role where it evolved into manganese homeostasis regulation, as Mur (Rodionov et al., 2006; Johnston et al., 2007). Following this hypothesis, R. capsulatus appears as an extreme example of this evolutionary trend.
Table 8.
Iron regulators identified in the genomes of R. capsulatus SB1003, R. sphaeroides 2.4.1 and R. ferroxidans SW2.
| Regulator type | Gene name | COG | Label Rcap SB1003a | Label Rsph 2.4.1a | Label Rfer SW2a | Product |
|---|---|---|---|---|---|---|
| Fur-family | fur/mur | 0735 | N.D. | 2494 | 2373 | Ferric/manganese uptake regulator |
| irr | 0735 | 02670 | 3179 | 0819 | Iron response regulator | |
| zur | 0735 | 01134 | 3569 | 1799 | Zinc uptake regulator |
N.D.: Not detected
See table 1.
In addition to genome analysis, some experimental work has been undertaken on the role of Fur/Mur and Irr in R. sphaeroides 2.4.1. In this organism, Fur/Mur is involved in iron homeostasis and oxidative stress response upon iron scarcity, where it acts as a repressor. Interestingly, the Δfur/mur mutant shows a more hampered growth profile during manganese rather than iron limitation. Furthermore the Δfur/mur mutant also has no activation of the putative Mn(II)/Fe(II) uptake system SitABCD, highlighting this operon as a potential target for Fur (Peuser et al., 2011). The role of Irr in R. sphaeroides is also not well defined. A Δirr strain does not display much growth deficiency in an iron-depleted medium suggesting that Irr does not have a major function in iron homeostasis. Moreover, Irr in R. sphaeroides was shown to bind heme and to activate many genes beyond that of iron homeostasis such as stress response, oxidative phosphorylation, transport, and photosynthesis.
Consequently the effect of Irr on iron homeostasis may be indirect. Oxidative stress also appears to be an important part of the Irr regulon as Δirr is more resistant to this stress (Peuser et al., 2012). Such resistance could be achieved by Irr through indirect control of a catalase and possibly a small RNA involved in singlet oxygen/superoxide response. Direct Irr control was evidenced on mbfA and ccpA, encoding the membrane-bound ferritin and a cytochrome c peroxidase, respectively. Although transcription data showed weak effect, Irr was proven to directly interact with the respective promoter of these genes, confirming computational prediction of Irr regulon in R. sphaeroides (Peuser et al., 2012; Rodionov et al., 2006). This computational study of iron and manganese regulons predicted that mbfA and ccpA are Irr-regulated genes in both R. capsulatus and R. sphaeroides. Beside Irr-specific regulation, the iron regulon in R. capsulatus and R. sphaeroides were predicted to consist of 18 and 8 genes, respectively (Rodionov et al., 2006)
From the data on R. sphaeroides, neither Fur/Mur nor Irr appears as a master regulator of iron homeostasis in this species. Rodionov et al. hypothesize a potential major role of IscR, a regulator of Fe-S cluster synthesis in E. coli (Rodionov et al., 2006). While R. capsulatus and R. sphaeroides genomes have an IscR encoding gene (orf 01853 and 0443, respectively), none could be found in the R. ferroxidans genome. Nevertheless, the absence of Fur, and presence of MntR, in R. capsulatus is a good hint that it most likely regulates iron homeostasis differently from R. sphaeroides and R. ferroxidans. Sequence alignments (not shown) of Fur, Irr, Zur sequences show very high similarity and identity, in the order of 60 to 80%, respectively (table 9). But, consistently, R. sphaeroides and R. ferroxidans displays higher values with each other than with R. capsulatus.
Table 9.
Identity and similarity in Fur/Mur, Irr and Zur from R. capsulatus SB1003, R. shaeroides 2.4.1 and R. ferroxidans SW2. Similarity is indicated using parenthesis.
| Fur/Mur | Rcap | Rsph | Rfer |
|---|---|---|---|
| Rcap | No Rcap Fur | - | - |
| Rsph | No Rcap Fur | 100 (100) | - |
| Rfer | No Rcap Fur | 74.64 (84.78) | 100 (100) |
|
| |||
| Irr | Rcap | Rsph | Rfer |
|
| |||
| Rcap | 100 (100) | - | - |
| Rsph | 62.59 (76.87) | 100 (100) | - |
| Rfer | 60.00 (73.51) | 66.67 (78.23) | 100 (100) |
|
| |||
| Zur | Rcap | Rsph | Rfer |
|
| |||
| Rcap | 100 (100) | - | - |
| Rsph | 63.37 (71.51) | 100 (100) | - |
| Rfer | 68.64 (75.15) | 73.21 (80.36) | 100 (100) |
Sequences were aligned using the CLC Sequence Viewer software (CLC Bio, Denmark). Similarity and identity scores were calculated based on alignments using the “Ident and Sim” utility (http://www.bioinformatics.org/sms2/ident_sim.html) (Stothard, 2000).
VIII. CONCLUSION
Analysis of the genomes of three model Rhodobacter species, highlights the diversity of strategies used by these organisms to maintain iron homeostasis (table 10). The two extreme strategies appeared to be R. capsulatus and R. ferroxidans. On the one hand, R. capsulatus presents a whole battery of ferrous, ferric and heme iron uptake (between 10 and 12!) in addition to an iron efflux pump and several storage genes. On the other hand, R. ferroxidans displays a more minimalist arsenal with one ferrous iron uptake system, an efflux pump and one storage system. The case of R. sphaeroides seems intermediate with a balanced distribution of ferrous and ferric iron uptake systems (between 5 and 7). Also, it is notable that R. capsulatus is the only Rhodobacter with a clearly identified heme uptake system that has been mostly studied in pathogens and more recently with symbionts. Moreover, R. capsulatus is the only sequenced Rhodobacter genome that is missing the canonical iron global regulator Fur. Such a paradox suggests novel regulation of iron homeostasis in this organism. Finally, among Rhodobacter species, R. capsulatus is the only organism displaying a large portion of its genome putatively dedicated to iron uptake. This “iron island” is a ~22kb gene cluster consisting of a ferrous iron transport cassette, a heme uptake system, an ABC transporter and siderophore import system (fig. 4).
Table 10.
Summary of iron/manganese homeostasis features identified in the genomes of R. capsulatus SB1003, R. sphaeroides 2.4.1 and R. ferroxidans SW2.
| Feature type | Feature name | Rcap SB1003 | Rsph 2.4.1 | Rfer SW2 |
|---|---|---|---|---|
| Fe/Mn uptake | Feo | 2 | 1 | 1 |
| Efe | 1 | 0 | 0 | |
| MntH | 1 | 0 | 2 | |
| ABC-type | 1 | 2 | 1 | |
| Siderophore | 7 | 4 | 0 | |
| Heme | 1 | 0 | 0 | |
|
| ||||
| Total | 13 | 7 | 4 | |
|
| ||||
| Efflux pump | FieF | 1 | 1 | 1 |
|
| ||||
| Iron storage | Bfr | 1 | 2 | 0 |
| MbfA | 1 | 1 | 0 | |
|
| ||||
| Fur-type regulators | Fur | 0 | 1 | 1 |
| Irr | 1 | 1 | 1 | |
| Zur | 1 | 1 | 1 | |
|
| ||||
| Manganese regulator | MntR | 1 | 0 | 2 |
Fig 4.
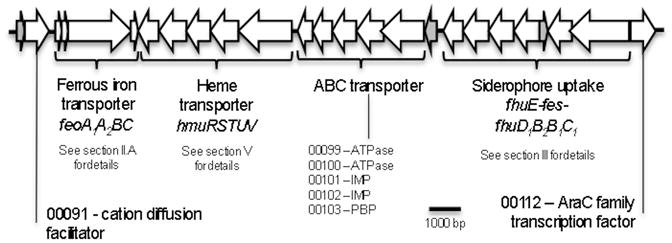
Putative “Iron island” in R. capsulatus SB1003. Genes encoding “hypothetical proteins” are shown in grey. IMP: Inner-membrane protein; PBP: Periplasmic binding protein. ORF number is given with the 5 digits only of the nomenclature of R. capsulatus SB1003 (for example, 12345 correspond to locus RCAP_rcc12345 for a chromosomal gene and p12345 to RCAP_rcp12345 for a plasmid gene).
Among the question that remains to be solved is the transport of iron through the periplasmic membrane. Indeed, unlike siderophores and heme, which have specific outer-membrane receptors, the access of elemental iron to the inner-membrane acquisition systems (Feo, Efe or ABC-type) has not been characterized. The paradigm for such a small compound consists in passive diffusion but recent research indicates that porin-like channels in the outer-membrane can transfer specific divalent metal cations to the periplasm (Hohle et al., 2011).
The field of metal homeostasis, and the regulation of iron uptake in particular, is a complex one. Indeed, after decades of research on iron regulation in a well-known organism such as E. coli, iron uptake systems are still being discovered (Koch et al., 2011). Decades of studies of Fur are also not enough to suppress debates about its mechanism of action as well as discriminating related representatives such as Mur, Zur, Nur (Lee and Helmann, 2007). In addition, a new layer of complexity is emerging in the field concerning the overlap between iron and manganese homeostasis which are two metals that have crucial and yet opposite effects on oxidative stress (Puri et al., 2010; Jakubovics and Jenkinson, 2001; Horsburgh et al., 2002). Depending on whether cells are challenged with an oxidative stress response or DNA synthesis under iron scarcity, organisms have been shown to be able to switch from an Fe-based enzyme activity to its Mn-based equivalent (Andrews et al., 2003; Andrews, 2011; Lee and Helmann, 2007).The various strategies to maintain metal homeostasis are printed in the genomes of these Rhodobacter strains and clearly highlight the diversity of this genus.
References
- Amarelle V, Koziol U, Rosconi F, Noya F, O'Brian MR, Fabiano E. A new small regulatory protein, HmuP, modulates haemin acquisition in Sinorhizobium meliloti. Microbiology. 2010;156:1873–1882. doi: 10.1099/mic.0.037713-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Amarelle V, O'Brian MR, Fabiano E. ShmR is essential for utilization of heme as a nutritional iron source in Sinorhizobium meliloti. Applied and environmental microbiology. 2008;74:6473–6475. doi: 10.1128/AEM.01590-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Amin SA, Green DH, Waheeb AlD, Gärdes A, Carrano CJ. Iron transport in the genus Marinobacter. Biometals : an international journal on the role of metal ions in biology, biochemistry, and medicine. 2012;25:135–147. doi: 10.1007/s10534-011-9491-9. [DOI] [PubMed] [Google Scholar]
- Andrews SC. Making DNA without iron - induction of a manganese-dependent ribonucleotide reductase in response to iron starvation. FEMS microbiology letters. 2011;80:286–289. doi: 10.1111/j.1365-2958.2011.07594.x. [DOI] [PubMed] [Google Scholar]
- Andrews SC. The Ferritin-like superfamily: Evolution of the biological iron storeman from a rubrerythrin-like ancestor. Biochimica et biophysica acta. 2010;1800:691–705. doi: 10.1016/j.bbagen.2010.05.010. [DOI] [PubMed] [Google Scholar]
- Andrews SC, Robinson AK, Rodríguez-Quiñones F. Bacterial iron homeostasis. FEMS microbiology reviews. 2003;27:215–237. doi: 10.1016/S0168-6445(03)00055-X. [DOI] [PubMed] [Google Scholar]
- Angerer A, Klupp B, Braun V. Iron transport systems of Serratia marcescens. The Journal of Bacteriology. 1992;174:1378–1387. doi: 10.1128/jb.174.4.1378-1387.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anzaldi LL, Skaar EP. Overcoming the heme paradox: heme toxicity and tolerance in bacterial pathogens. Infection and Immunity. 2010;78:4977–4989. doi: 10.1128/IAI.00613-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Archibald F. Lactobacillus plantarum, an organism not requiring iron. FEMS microbiology letters. 1983;19:29–32. [Google Scholar]
- Badarau A, Firbank SJ, Waldron KJ, Yanagisawa S, Robinson NJ, Banfield MJ, Dennison C. FutA2 is a ferric binding protein from Synechocystis PCC 6803. The Journal of biological chemistry. 2008;283:12520–12527. doi: 10.1074/jbc.M709907200. [DOI] [PubMed] [Google Scholar]
- Baichoo N, Wang T, Ye R, Helmann JD. Global analysis of the Bacillus subtilis Fur regulon and the iron starvation stimulon. Molecular microbiology. 2002;45:1613–1629. doi: 10.1046/j.1365-2958.2002.03113.x. [DOI] [PubMed] [Google Scholar]
- Barker KD, Barkovits K, Wilks A. Metabolic flux of extracellular heme uptake in Pseudomonas aeruginosa is driven by the iron-regulated heme oxygenase (HemO) The Journal of biological chemistry. 2012;287:18342–18350. doi: 10.1074/jbc.M112.359265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bearden SW, Perry RD. The Yfe system of Yersinia pestis transports iron and manganese and is required for full virulence of plague. Molecular microbiology. 1999;32:403–414. doi: 10.1046/j.1365-2958.1999.01360.x. [DOI] [PubMed] [Google Scholar]
- Bendtsen JD, Nielsen H, Widdick D, Palmer T, Brunak S. Prediction of twin-arginine signal peptides. BMC bioinformatics. 2005;6:167. doi: 10.1186/1471-2105-6-167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Biel SW, Biel AJ. Isolation of a Rhodobacter capsulatus mutant that lacks c-type cytochromes and excretes porphyrins. Journal of bacteriology. 1990;172:1321–1326. doi: 10.1128/jb.172.3.1321-1326.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bou-Abdallah F. The iron redox and hydrolysis chemistry of the ferritins. Biochimica et biophysica acta. 2010;1800:719–731. doi: 10.1016/j.bbagen.2010.03.021. [DOI] [PubMed] [Google Scholar]
- Braun V. Surface signaling: novel transcription initiation mechanism starting from the cell surface. Archives of microbiology. 1997;167:325–331. doi: 10.1007/s002030050451. [DOI] [PubMed] [Google Scholar]
- Braun V, Hantke K. Recent insights into iron import by bacteria. Current Opinion in Chemical Biology. 2011;15:328–334. doi: 10.1016/j.cbpa.2011.01.005. [DOI] [PubMed] [Google Scholar]
- Braun V, Mahren S, Ogierman M. Regulation of the FecI-type ECF sigma factor by transmembrane signalling. Current Opinion in Microbiology. 2003;6:173–180. doi: 10.1016/s1369-5274(03)00022-5. [DOI] [PubMed] [Google Scholar]
- Brickman TJ, Armstrong SK. Iron and pH-responsive FtrABCD ferrous iron utilization system of Bordetella species. Molecular microbiology. 2012;86:580–593. doi: 10.1111/mmi.12003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brickman TJ, Cummings CA, Liew SY, Relman DA, Armstrong SK. Transcriptional profiling of the iron starvation response in Bordetella pertussis provides new insights into siderophore utilization and virulence gene expression. Journal of bacteriology. 2011;193:4798–4812. doi: 10.1128/JB.05136-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Caiazza NC, Lies DP, Newman DK. Phototrophic Fe(II) oxidation promotes organic carbon acquisition by Rhodobacter capsulatus SB1003. Applied and environmental microbiology. 2007;73:6150–6158. doi: 10.1128/AEM.02830-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cao J, Woodhall MR, Alvarez J, Cartron ML, Andrews SC. EfeUOB (YcdNOB) is a tripartite, acid-induced and CpxAR-regulated, low-pH Fe2+ transporter that is cryptic in Escherichia coli K-12 but functional in E. coli O157:H7. Molecular microbiology. 2007;65:857–875. doi: 10.1111/j.1365-2958.2007.05802.x. [DOI] [PubMed] [Google Scholar]
- Carpenter BM, Whitmire JM, Merrell DS. This is not your mother's repressor: the complex role of fur in pathogenesis. Infection and Immunity. 2009;77:2590–2601. doi: 10.1128/IAI.00116-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carrondo MA. Ferritins, iron uptake and storage from the bacterioferritin viewpoint. The EMBO journal. 2003;22:1959–1968. doi: 10.1093/emboj/cdg215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cartron ML, Maddocks S, Gillingham P, Craven CJ, Andrews SC. Feo--transport of ferrous iron into bacteria. Biometals : an international journal on the role of metal ions in biology, biochemistry, and medicine. 2006;19:143–157. doi: 10.1007/s10534-006-0003-2. [DOI] [PubMed] [Google Scholar]
- Chao TC, Becker A, Buhrmester J, Pühler A, Weidner S. The Sinorhizobium meliloti fur gene regulates, with dependence on Mn(II), transcription of the sitABCD operon, encoding a metal-type transporter. The Journal of Bacteriology. 2004;186:3609–3620. doi: 10.1128/JB.186.11.3609-3620.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chiancone E, Ceci P, Ilari A, Ribacchi F, Stefanini S. Iron and proteins for iron storage and detoxification. Biometals : an international journal on the role of metal ions in biology, biochemistry, and medicine. 2004;17:197–202. doi: 10.1023/b:biom.0000027692.24395.76. [DOI] [PubMed] [Google Scholar]
- Chin N, Frey J, Chang CF, Chang YF. Identification of a locus involved in the utilization of iron by Actinobacillus pleuropneumoniae. FEMS microbiology letters. 1996;143:1–6. doi: 10.1111/j.1574-6968.1996.tb08452.x. [DOI] [PubMed] [Google Scholar]
- Chu BC, Garcia-Herrero A, Johanson TH, Krewulak KD, Lau CK, Peacock RS, Slavinskaya Z, Vogel HJ. Siderophore uptake in bacteria and the battle for iron with the host; a bird's eye view. Biometals : an international journal on the role of metal ions in biology, biochemistry, and medicine. 2010;23:601–611. doi: 10.1007/s10534-010-9361-x. [DOI] [PubMed] [Google Scholar]
- Cobessi D, Huang LS, Ban M, Pon NG, Daldal F, Berry EA. The 2.6 A resolution structure of Rhodobacter capsulatus bacterioferritin with metal-free dinuclear site and heme iron in a crystallographic ‘special position’. Acta Crystallographica Section D Biological Crystallography. 2001;58:29–38. doi: 10.1107/s0907444901017267. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cornelis P, Matthijs S, Van Oeffelen L. Iron uptake regulation in Pseudomonas aeruginosa. Biometals : an international journal on the role of metal ions in biology, biochemistry, and medicine. 2009;22:15–22. doi: 10.1007/s10534-008-9193-0. [DOI] [PubMed] [Google Scholar]
- Croal LR, Jiao Y, Newman DK. The fox operon from Rhodobacter strain SW2 promotes phototrophic Fe(II) oxidation in Rhodobacter capsulatus SB1003. The Journal of Bacteriology. 2007;189:1774–1782. doi: 10.1128/JB.01395-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- D'Onofrio A, Crawford JM, Stewart EJ, Witt K, Gavrish E, Epstein S, Clardy J, Lewis K. Siderophores from neighboring organisms promote the growth of uncultured bacteria. Chemistry and biology. 2010;17:254–264. doi: 10.1016/j.chembiol.2010.02.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dailey HA, Septer AN, Daugherty L, Thames D, Gerdes S, Stabb EV, Dunn AK, Dailey TA, Phillips JD. The Escherichia coli protein YfeX functions as a porphyrinogen oxidase, not a heme dechelatase. mBio. 2011;2:e00248–11. doi: 10.1128/mBio.00248-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dashper SG, Butler CA, Lissel JP, Paolini RA, Hoffmann B, Veith PD, O'Brien-Simpson NM, Snelgrove SL, Tsiros JT, Reynolds EC. A novel Porphyromonas gingivalis FeoB plays a role in manganese accumulation. The Journal of biological chemistry. 2005;280:28095–28102. doi: 10.1074/jbc.M503896200. [DOI] [PubMed] [Google Scholar]
- Dubbels BL, DiSpirito AA, Morton JD, Semrau JD, Neto JNE, Bazylinski DA. Evidence for a copper-dependent iron transport system in the marine, magnetotactic bacterium strain MV-1. Microbiology (Reading, England) 2004;150:2931–2945. doi: 10.1099/mic.0.27233-0. [DOI] [PubMed] [Google Scholar]
- Ducey TF, Carson MB, Orvis J, Stintzi AP, Dyer DW. Identification of the iron-responsive genes of Neisseria gonorrhoeae by microarray analysis in defined medium. The Journal of Bacteriology. 2005;187:4865–4874. doi: 10.1128/JB.187.14.4865-4874.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ducey TF, Jackson L, Orvis J, Dyer DW. Transcript analysis of nrrF, a Fur repressed sRNA of Neisseria gonorrhoeae. Microbial Pathogenesis. 2009;46:166–170. doi: 10.1016/j.micpath.2008.12.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ehrenreich A, Widdel F. Anaerobic oxidation of ferrous iron by purple bacteria, a new type of phototrophic metabolism. Applied and environmental microbiology. 1994;60:4517–4526. doi: 10.1128/aem.60.12.4517-4526.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Escamilla-Hernandez R, O'Brian MR. HmuP is a coactivator of Irr-dependent expression of heme utilization genes in Bradyrhizobium japonicum. Journal of bacteriology. 2012;194:3137–3143. doi: 10.1128/JB.00071-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fantappiè L, Scarlato V, Delany I. Identification of the in vitro target of an iron-responsive AraC-like protein from Neisseria meningitidis that is in a regulatory cascade with Fur. Microbiology. 2011;157:2235–2247. doi: 10.1099/mic.0.048033-0. [DOI] [PubMed] [Google Scholar]
- Fetherston JD, Mier I, Truszczynska H, Perry RD. The Yfe and Feo Transporters Are Involved in Microaerobic Growth and the Virulence of Yersinia pestis in Bubonic Plague. Infection and Immunity. 2012;80:3880–3891. doi: 10.1128/IAI.00086-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fisher CR, Davies NMLL, Wyckoff EE, Feng Z, Oaks EV, Payne SM. Genetics and virulence association of the Shigella flexneri sit iron transport system. Infection and Immunity. 2009;77:1992–1999. doi: 10.1128/IAI.00064-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Frankenberg-Dinkel N. Bacterial heme oxygenases. Antioxidants and redox signaling. 2004;6:825–834. doi: 10.1089/ars.2004.6.825. [DOI] [PubMed] [Google Scholar]
- Grass G, Otto M, Fricke B, Haney CJ, Rensing C, Nies DH, Munkelt D. FieF (YiiP) from Escherichia coli mediates decreased cellular accumulation of iron and relieves iron stress. Archives of microbiology. 2005;183:9–18. doi: 10.1007/s00203-004-0739-4. [DOI] [PubMed] [Google Scholar]
- Grifantini R, Sebastian S, Frigimelica E, Draghi M, Bartolini E, Muzzi A, Rappuoli R, Grandi G, Genco CA. Identification of iron-activated and -repressed Fur-dependent genes by transcriptome analysis of Neisseria meningitidis group B. Proceedings of the National Academy of Sciences of the United States of America. 2003;100:9542–9547. doi: 10.1073/pnas.1033001100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grosse C, Scherer J, Koch D, Otto M, Taudte N, Grass G. A new ferrous iron-uptake transporter, EfeU (YcdN), from Escherichia coli. Molecular microbiology. 2006;62:120–131. doi: 10.1111/j.1365-2958.2006.05326.x. [DOI] [PubMed] [Google Scholar]
- Guo J, Nair MKM, Galván EM, Liu SL, Schifferli DM. Tn5AraOut mutagenesis for the identification of Yersinia pestis genes involved in resistance towards cationic antimicrobial peptides. Microbial Pathogenesis. 2011;51:121–132. doi: 10.1016/j.micpath.2011.04.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hantke K. Ferrous iron transport mutants in Escherichia coli K12. FEMS microbiology letters. 1987;44:53–57. doi: 10.1111/j.1574-6968.1993.tb06082.x. [DOI] [PubMed] [Google Scholar]
- Hantke K. Is the bacterial ferrous iron transporter FeoB a living fossil? Trends in microbiology. 2003;11:192–195. doi: 10.1016/s0966-842x(03)00100-8. [DOI] [PubMed] [Google Scholar]
- He J, Miyazaki H, Anaya C, Yu F, Yeudall WA, Lewis JP. Role of Porphyromonas gingivalis FeoB2 in metal uptake and oxidative stress protection. Infection and Immunity. 2006;74:4214–4223. doi: 10.1128/IAI.00014-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hider RC, Kong X. Chemistry and biology of siderophores. Natural product reports. 2010;27:637–657. doi: 10.1039/b906679a. [DOI] [PubMed] [Google Scholar]
- Hohle TH, Franck WL, Stacey G, O'Brian MR. Bacterial outer membrane channel for divalent metal ion acquisition. Proceedings of the National Academy of Sciences of the United States of America. 2011;108:15390–15395. doi: 10.1073/pnas.1110137108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hollander A, Mercante AD, Shafer WM, Cornelissen CN. The iron-repressed, AraC-like regulator MpeR activates expression of fetA in Neisseria gonorrhoeae. Infection and Immunity. 2011;79:4764–4776. doi: 10.1128/IAI.05806-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horsburgh MJ, Wharton SJ, Karavolos M, Foster SJ. Manganese: elemental defence for a life with oxygen. Trends in microbiology. 2002;10:496–501. doi: 10.1016/s0966-842x(02)02462-9. [DOI] [PubMed] [Google Scholar]
- Huang SH, Wang CK, Peng HL, Wu CC, Chen YT, Hong YM, Lin CT. Role of the small RNA RyhB in the Fur regulon in mediating the capsular polysaccharide biosynthesis and iron acquisition systems in Klebsiella pneumoniae. BMC microbiology. 2012;12:148. doi: 10.1186/1471-2180-12-148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jakubovics NS, Jenkinson HF. Out of the iron age: new insights into the critical role of manganese homeostasis in bacteria. Microbiology (Reading, England) 2001;147:1709–1718. doi: 10.1099/00221287-147-7-1709. [DOI] [PubMed] [Google Scholar]
- Janakiraman A, Slauch JM. The putative iron transport system SitABCD encoded on SPI1 is required for full virulence of Salmonella typhimurium. Molecular microbiology. 2000;35:1146–1155. doi: 10.1046/j.1365-2958.2000.01783.x. [DOI] [PubMed] [Google Scholar]
- Johnston AWB, Todd JD, Curson AR, Lei S, Nikolaidou-Katsaridou N, Gelfand MS, Rodionov DA. Living without Fur: the subtlety and complexity of iron-responsive gene regulation in the symbiotic bacterium Rhizobium and other alpha-proteobacteria. Biometals : an international journal on the role of metal ions in biology, biochemistry, and medicine. 2007;20:501–511. doi: 10.1007/s10534-007-9085-8. [DOI] [PubMed] [Google Scholar]
- Kammler M, Schön C, Hantke K. Characterization of the ferrous iron uptake system of Escherichia coli. 175:6212–6219. doi: 10.1128/jb.175.19.6212-6219.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Katoh H, Hagino N, Ogawa T. Iron-binding activity of FutA1 subunit of an ABC-type iron transporter in the cyanobacterium Synechocystis sp. Strain PCC 6803. Plant and cell physiology. 2001a;42:823–827. doi: 10.1093/pcp/pce106. [DOI] [PubMed] [Google Scholar]
- Katoh H, Hagino N, Grossman AR, Ogawa T. Genes essential to iron transport in the cyanobacterium Synechocystis sp. strain PCC 6803. The Journal of Bacteriology. 2001b;183:2779–2784. doi: 10.1128/JB.183.9.2779-2784.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kehres DG, Janakiraman A, Slauch JM, Maguire ME. SitABCD is the alkaline Mn(2+) transporter of Salmonella enterica serovar Typhimurium. The Journal of Bacteriology. 2002;184:3159–3166. doi: 10.1128/JB.184.12.3159-3166.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khun HH, Kirby SD, Lee BC. A Neisseria meningitidis fbpABC mutant is incapable of using nonheme iron for growth. Infection and Immunity. 1998;66:2330–2336. doi: 10.1128/iai.66.5.2330-2336.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim H, Lee H, Shin D. The FeoA protein is necessary for the FeoB transporter to import ferrous iron. Biochemical and biophysical research communications. 2012;423:733–738. doi: 10.1016/j.bbrc.2012.06.027. [DOI] [PubMed] [Google Scholar]
- Koch D, Chan ACK, Murphy MEP, Lilie H, Grass G, Nies DH. Characterization of a dipartite iron uptake system from uropathogenic Escherichia coli strain F11. The Journal of biological chemistry. 2011;286:25317–25330. doi: 10.1074/jbc.M111.222745. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kosman DJ. Molecular mechanisms of iron uptake in fungi. Molecular microbiology. 2003;47:1185–1197. doi: 10.1046/j.1365-2958.2003.03368.x. [DOI] [PubMed] [Google Scholar]
- Koster M, van Klompenburg W, Bitter W, Leong J, Weisbeek P. Role for the outer membrane ferric siderophore receptor PupB in signal transduction across the bacterial cell envelope. The EMBO journal. 1994;13:2805–2813. doi: 10.1002/j.1460-2075.1994.tb06574.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Köster W. ABC transporter-mediated uptake of iron, siderophores, heme and vitamin B12. Research in microbiology. 2001;152:291–301. doi: 10.1016/s0923-2508(01)01200-1. [DOI] [PubMed] [Google Scholar]
- Krewulak KD, Vogel HJ. Structural biology of bacterial iron uptake. Biochimica et biophysica acta. 2008;1778:1781–1804. doi: 10.1016/j.bbamem.2007.07.026. [DOI] [PubMed] [Google Scholar]
- Krewulak KD, Vogel HJ. TonB or not TonB: is that the question? Biochemistry and cell biology = Biochimie et biologie cellulaire. 2011;89:87–97. doi: 10.1139/o10-141. [DOI] [PubMed] [Google Scholar]
- LASCELLES J. The synthesis of porphyrins and bacteriochlorophyll by cell suspensions of Rhodopseudomonas spheroides. The Biochemical journal. 1956;62:78–93. doi: 10.1042/bj0620078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee JW, Helmann JD. Functional specialization within the Fur family of metalloregulators. Biometals : an international journal on the role of metal ions in biology, biochemistry, and medicine. 2007;20:485–499. doi: 10.1007/s10534-006-9070-7. [DOI] [PubMed] [Google Scholar]
- Létoffé S, Heuck G, Delepelaire P, Lange N, Wandersman C. Bacteria capture iron from heme by keeping tetrapyrrol skeleton intact. Proceedings of the National Academy of Sciences of the United States of America. 2009;106:11719–11724. doi: 10.1073/pnas.0903842106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Litwin CM, Quackenbush J. Characterization of a Vibrio vulnificus LysR homologue, HupR, which regulates expression of the haem uptake outer membrane protein, HupA. Microbial Pathogenesis. 2001;31:295–307. doi: 10.1006/mpat.2001.0472. [DOI] [PubMed] [Google Scholar]
- Liu M, Boulouis H-J, Biville F. Heme degrading protein HemS is involved in oxidative stress response of Bartonella henselae. PloS one. 2012;7:e37630. doi: 10.1371/journal.pone.0037630. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu X, Du Q, Wang Z, Zhu D, Huang Y, Li N, Wei T, Xu S, Gu L. Crystal structure and biochemical features of EfeB/YcdB from Escherichia coli O157: ASP235 plays divergent roles in different enzyme-catalyzed processes. The Journal of biological chemistry. 2011;286:14922–14931. doi: 10.1074/jbc.M110.197780. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu M, Fu D. Structure of the Zinc Transporter YiiP. Science (New York, NY) 2007;317:1746–1748. doi: 10.1126/science.1143748. [DOI] [PubMed] [Google Scholar]
- Massé E, Salvail H, Desnoyers G, Arguin M. Small RNAs controlling iron metabolism. Current Opinion in Microbiology. 2007;10:140–145. doi: 10.1016/j.mib.2007.03.013. [DOI] [PubMed] [Google Scholar]
- Metruccio MME, Fantappiè L, Serruto D, Muzzi A, Roncarati D, Donati C, Scarlato V, Delany I. The Hfq-dependent small noncoding RNA NrrF directly mediates Fur-dependent positive regulation of succinate dehydrogenase in Neisseria meningitidis. Journal of bacteriology. 2009;191:1330–1342. doi: 10.1128/JB.00849-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meyer TE, Cusanovich MA. Soluble cytochrome composition of the purple phototrophic bacterium, Rhodopseudomonas sphaeroides ATCC 17023. Biochimica et biophysica acta. 1985;807:308–319. doi: 10.1016/0005-2728(85)90263-4. [DOI] [PubMed] [Google Scholar]
- Montanini B, Blaudez D, Jeandroz S, Sanders D, Chalot M. Phylogenetic and functional analysis of the Cation Diffusion Facilitator (CDF) family: improved signature and prediction of substrate specificity. BMC genomics. 2007;8:107. doi: 10.1186/1471-2164-8-107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Munkelt D, Grass G, Nies DH. The chromosomally encoded cation diffusion facilitator proteins DmeF and FieF from Wautersia metallidurans CH34 are transporters of broad metal specificity. The Journal of Bacteriology. 2004;186:8036–8043. doi: 10.1128/JB.186.23.8036-8043.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nienaber A, Hennecke H, Fischer HM. Discovery of a haem uptake system in the soil bacterium Bradyrhizobium japonicum. Molecular microbiology. 2001;41:787–800. doi: 10.1046/j.1365-2958.2001.02555.x. [DOI] [PubMed] [Google Scholar]
- Nies DH. Biochemistry. How cells control zinc homeostasis. Science (New York, NY) 2007;317:1695–1696. doi: 10.1126/science.1149048. [DOI] [PubMed] [Google Scholar]
- Nies DH. How iron is transported into magnetosomes. Molecular microbiology. 2011;82:792–796. doi: 10.1111/j.1365-2958.2011.07864.x. [DOI] [PubMed] [Google Scholar]
- O'Neill MJ, Bhakta MN, Fleming KG, Wilks A. Induced fit on heme binding to the Pseudomonas aeruginosa cytoplasmic protein (PhuS) drives interaction with heme oxygenase (HemO) Proceedings of the National Academy of Sciences of the United States of America. 2012;109:5639–5644. doi: 10.1073/pnas.1121549109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ollinger J, Song KB, Antelmann H, Hecker M, Helmann JD. Role of the Fur regulon in iron transport in Bacillus subtilis. The Journal of Bacteriology. 2006;188:3664–3673. doi: 10.1128/JB.188.10.3664-3673.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paustian ML, May BJ, Kapur V. Pasteurella multocida gene expression in response to iron limitation. Infection and Immunity. 2001;69:4109–4115. doi: 10.1128/IAI.69.6.4109-4115.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Penfold CN, Ringeling PL, Davy SL, Moore GR, McEwan AG, Spiro S. Isolation, characterisation and expression of the bacterioferritin gene of Rhodobacter capsulatus. FEMS microbiology letters. 1996;139:143–148. doi: 10.1111/j.1574-6968.1996.tb08194.x. [DOI] [PubMed] [Google Scholar]
- Perry RD, Mier I, Fetherston JD. Roles of the Yfe and Feo transporters of Yersinia pestis in iron uptake and intracellular growth. Biometals : an international journal on the role of metal ions in biology, biochemistry, and medicine. 2007;20:699–703. doi: 10.1007/s10534-006-9051-x. [DOI] [PubMed] [Google Scholar]
- Petersen TN, Brunak S, von Heijne G, Nielsen H. SignalP 4.0: discriminating signal peptides from transmembrane regions. Nature methods. 2011;8:785–786. doi: 10.1038/nmeth.1701. [DOI] [PubMed] [Google Scholar]
- Peuser V, Metz S, Klug G. Response of the photosynthetic bacterium Rhodobacter sphaeroides to iron limitation and the role of a Fur orthologue in this response. Environ Microbiol Rep. 2011;3:397–404. doi: 10.1111/j.1758-2229.2011.00245.x. [DOI] [PubMed] [Google Scholar]
- Peuser V, Remes B, Klug G. Role of the Irr protein in the regulation of iron metabolism in Rhodobacter sphaeroides. PloS one. 2012;7:e42231. doi: 10.1371/journal.pone.0042231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Planavsky NJ, McGoldrick P, Scott CT, Li C, Reinhard CT, Kelly AE, Chu X, Bekker A, Love GD, Lyons TW. Widespread iron-rich conditions in the mid-Proterozoic ocean. Nature. 2011;477:448–451. doi: 10.1038/nature10327. [DOI] [PubMed] [Google Scholar]
- Posey JE. Lack of a Role for Iron in the Lyme Disease Pathogen. Science (New York, NY) 2000;288:1651–1653. doi: 10.1126/science.288.5471.1651. [DOI] [PubMed] [Google Scholar]
- Poulain AJ, Newman DK. Rhodobacter capsulatus catalyzes light-dependent Fe(II) oxidation under anaerobic conditions as a potential detoxification mechanism. Applied and environmental microbiology. 2009;75:6639–6646. doi: 10.1128/AEM.00054-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pradel E, Guiso N, Locht C. Identification of AlcR, an AraC-type regulator of alcaligin siderophore synthesis in Bordetella bronchiseptica and Bordetella pertussis. The Journal of Bacteriology. 1998;180:871–880. doi: 10.1128/jb.180.4.871-880.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Puri S, O'Brian MR. The hmuQ and hmuD genes from Bradyrhizobium japonicum encode heme-degrading enzymes. The Journal of Bacteriology. 2006;188:6476–6482. doi: 10.1128/JB.00737-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Puri S, Hohle TH, O'Brian MR. Control of bacterial iron homeostasis by manganese. Proceedings of the National Academy of Sciences of the United States of America. 2010;107:10691–10695. doi: 10.1073/pnas.1002342107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rajasekaran MB, Mitchell SA, Gibson TM, Hussain R, Siligardi G, Andrews SC, Watson KA. Isolation and characterisation of EfeM, a periplasmic component of the putative EfeUOBM iron transporter of Pseudomonas syringae pv. syringae. Biochemical and biophysical research communications. 2010a;398:366–371. doi: 10.1016/j.bbrc.2010.06.072. [DOI] [PubMed] [Google Scholar]
- Rajasekaran MB, Nilapwar S, Andrews SC, Watson KA. EfeO-cupredoxins: major new members of the cupredoxin superfamily with roles in bacterial iron transport. Biometals : an international journal on the role of metal ions in biology, biochemistry, and medicine. 2010b;23:1–17. doi: 10.1007/s10534-009-9262-z. [DOI] [PubMed] [Google Scholar]
- Reissbrodt R, Kingsley R, Rabsch W, Beer W, Roberts M, Williams PH. Iron-regulated excretion of alpha-keto acids by Salmonella typhimurium. The Journal of Bacteriology. 1997;179:4538–4544. doi: 10.1128/jb.179.14.4538-4544.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ringeling PL, Davy SL, Monkara FA, Hunt C, Dickson DP, McEwan AG, Moore GR. Iron metabolism in Rhodobacter capsulatus. Characterisation of bacterioferritin and formation of non-haem iron particles in intact cells. European journal of biochemistry / FEBS. 1994;223:847–855. doi: 10.1111/j.1432-1033.1994.tb19061.x. [DOI] [PubMed] [Google Scholar]
- Rodionov DA, Gelfand MS, Todd JD, Curson ARJ, Johnston AWB. Computational reconstruction of iron- and manganese-responsive transcriptional networks in alpha-proteobacteria. PLoS computational biology. 2006;2:e163. doi: 10.1371/journal.pcbi.0020163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rong C, Zhang C, Zhang Y, Qi L, Yang J, Guan G, Li Y, Li J. FeoB2 Functions in Magnetosome Formation and Oxidative Stress Protection in Magnetospirillum gryphiswaldense Strain MSR-1. Journal of bacteriology. 2012;194:3972–3976. doi: 10.1128/JB.00382-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rudolph G, Hennecke H, Fischer HM. Beyond the Fur paradigm: iron-controlled gene expression in rhizobia. FEMS microbiology reviews. 2006;30:631–648. doi: 10.1111/j.1574-6976.2006.00030.x. [DOI] [PubMed] [Google Scholar]
- Runyen-Janecky LJ, Brown AN, Ott B, Tujuba HG, Rio RVM. Regulation of high-affinity iron acquisition homologues in the tsetse fly symbiont Sodalis glossinidius. Journal of bacteriology. 2010;192:3780–3787. doi: 10.1128/JB.00161-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sabri M, Léveillé S, Dozois CM. A SitABCD homologue from an avian pathogenic Escherichia coli strain mediates transport of iron and manganese and resistance to hydrogen peroxide. Microbiology (Reading, England) 2006;152:745–758. doi: 10.1099/mic.0.28682-0. [DOI] [PubMed] [Google Scholar]
- Sanders JD, Cope LD, Hansen EJ. Identification of a locus involved in the utilization of iron by Haemophilus influenzae. Infection and Immunity. 1994;62:4515–4525. doi: 10.1128/iai.62.10.4515-4525.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sandy M, Butler A. Microbial Iron Acquisition: Marine and Terrestrial Siderophores. Chemical reviews. 2009;109:4580–4595. doi: 10.1021/cr9002787. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saraiva IH, Newman DK, Louro RO. Functional characterization of the FoxE iron oxidoreductase from the photoferrotroph Rhodobacter ferrooxidans SW2. The Journal of biological chemistry. 2012;287:25541–25548. doi: 10.1074/jbc.M112.360636. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sarma R, Barney BM, Hamilton TL, Jones A, Seefeldt LC, Peters JW. Crystal structure of the L protein of Rhodobacter sphaeroides light-independent protochlorophyllide reductase with MgADP bound: a homologue of the nitrogenase Fe protein. Biochemistry. 2008;47:13004–13015. doi: 10.1021/bi801058r. [DOI] [PubMed] [Google Scholar]
- Septer AN, Wang Y, Ruby EG, Stabb EV, Dunn AK. The haem-uptake gene cluster in Vibrio fischeri is regulated by Fur and contributes to symbiotic colonization. Environmental microbiology. 2011;13:2855–2864. doi: 10.1111/j.1462-2920.2011.02558.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sirijovski N, Mamedov F, Olsson U, Styring S, Hansson M. Rhodobacter capsulatus magnesium chelatase subunit BchH contains an oxygen sensitive iron-sulfur cluster. Archives of microbiology. 2007;188:599–608. doi: 10.1007/s00203-007-0282-1. [DOI] [PubMed] [Google Scholar]
- Skaar EP, Gaspar AH, Schneewind O. Bacillus anthracis IsdG, a heme-degrading monooxygenase. The Journal of Bacteriology. 2006;188:1071–1080. doi: 10.1128/JB.188.3.1071-1080.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smaldone GT, Revelles O, Gaballa A, Sauer U, Antelmann H, Helmann JD. A global investigation of the Bacillus subtilis iron-sparing response identifies major changes in metabolism. Journal of bacteriology. 2012;194:2594–2605. doi: 10.1128/JB.05990-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Small SK, Puri S, O'Brian MR. Heme-dependent metalloregulation by the iron response regulator (Irr) protein in Rhizobium and other Alpha-proteobacteria. Biometals : an international journal on the role of metal ions in biology, biochemistry, and medicine. 2009;22:89–97. doi: 10.1007/s10534-008-9192-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stearman R, Yuan DS, Yamaguchi-Iwai Y, Klausner RD, Dancis A. A permease-oxidase complex involved in high-affinity iron uptake in yeast. Science (New York, NY) 1996;271:1552–1557. doi: 10.1126/science.271.5255.1552. [DOI] [PubMed] [Google Scholar]
- Steele KH, O'Connor LH, Burpo N, Kohler K, Johnston JW. Characterization of a ferrous iron-responsive two-component system in nontypeable Haemophilus influenzae. Journal of bacteriology. 2012;194:6162–6173. doi: 10.1128/JB.01465-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stothard P. The sequence manipulation suite: JavaScript programs for analyzing and formatting protein and DNA sequences. BioTechniques. 2000;28:1102–1104. doi: 10.2144/00286ir01. [DOI] [PubMed] [Google Scholar]
- Strange HR, Zola TA, Cornelissen CN. The fbpABC operon is required for Ton-independent utilization of xenosiderophores by Neisseria gonorrhoeae strain FA19. Infection and Immunity. 2011;79:267–278. doi: 10.1128/IAI.00807-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sturm A, Schierhorn A, Lindenstrauss U, Lilie H, Brüser T. YcdB from Escherichia coli reveals a novel class of Tat-dependently translocated hemoproteins. The Journal of biological chemistry. 2006;281:13972–13978. doi: 10.1074/jbc.M511891200. [DOI] [PubMed] [Google Scholar]
- Sugano Y. DyP-type peroxidases comprise a novel heme peroxidase family. Cellular and molecular life sciences : CMLS. 2009;66:1387–1403. doi: 10.1007/s00018-008-8651-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suzuki T, Okamura Y, Calugay RJ, Takeyama H, Matsunaga T. Global gene expression analysis of iron-inducible genes in Magnetospirillum magneticum AMB-1. The Journal of Bacteriology. 2006;188:2275–2279. doi: 10.1128/JB.188.6.2275-2279.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Theil EC, Goss DJ. Living with iron (and oxygen): questions and answers about iron homeostasis. Chemical reviews. 2009;109:4568–4579. doi: 10.1021/cr900052g. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tom-Yew SAL, Cui DT, Bekker EG, Murphy MEP. Anion-independent iron coordination by the Campylobacter jejuni ferric binding protein. The Journal of biological chemistry. 2005;280:9283–9290. doi: 10.1074/jbc.M412479200. [DOI] [PubMed] [Google Scholar]
- Touati D. Iron and oxidative stress in bacteria. Archives of biochemistry and biophysics. 2000;373:1–6. doi: 10.1006/abbi.1999.1518. [DOI] [PubMed] [Google Scholar]
- Van Der Giezen M, Lenton TM. The rise of oxygen and complex life. The Journal of eukaryotic microbiology. 2012;59:111–113. doi: 10.1111/j.1550-7408.2011.00605.x. [DOI] [PubMed] [Google Scholar]
- van Vliet AH, Wooldridge KG, Ketley JM. Iron-responsive gene regulation in a Campylobacter jejuni fur mutant. The Journal of Bacteriology. 1998;180:5291–5298. doi: 10.1128/jb.180.20.5291-5298.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vanderpool CK, Armstrong SK. Heme-responsive transcriptional activation of Bordetella bhu genes. The Journal of Bacteriology. 2003;185:909–917. doi: 10.1128/JB.185.3.909-917.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vasil ML. How we learnt about iron acquisition in Pseudomonas aeruginosa: a series of very fortunate events. Biometals : an international journal on the role of metal ions in biology, biochemistry, and medicine. 2007;20 (3–4):587–601. doi: 10.1007/s10534-006-9067-2. [DOI] [PubMed] [Google Scholar]
- Wandersman C, Delepelaire P. Bacterial iron sources: from siderophores to hemophores. Annual review of microbiology. 2004;58:611–647. doi: 10.1146/annurev.micro.58.030603.123811. [DOI] [PubMed] [Google Scholar]
- Watson RJ, Millichap P, Joyce SA, Reynolds S, Clarke DJ. The role of iron uptake in pathogenicity and symbiosis in Photorhabdus luminescens TT01. BMC microbiology. 2010;10:177. doi: 10.1186/1471-2180-10-177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weinberg ED. The Lactobacillus anomaly: total iron abstinence. Perspectives in biology and medicine. 1997;40:578–583. doi: 10.1353/pbm.1997.0072. [DOI] [PubMed] [Google Scholar]
- Wen Y, Kim IH, Son JS, Lee BH, Kim KS. Iron and quorum sensing coordinately regulate the expression of vulnibactin biosynthesis in Vibrio vulnificus. The Journal of biological chemistry. 2012;287:26727–26739. doi: 10.1074/jbc.M112.374165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wexler M, Yeoman KH, Stevens JB, de Luca NG, Sawers G, Johnston AW. The Rhizobium leguminosarum tonB gene is required for the uptake of siderophore and haem as sources of iron. Molecular microbiology. 2001;41:801–816. doi: 10.1046/j.1365-2958.2001.02556.x. [DOI] [PubMed] [Google Scholar]
- Widdel F, Schnell S, Heising S, Ehrenreich A, Assmus B, Schink B. Ferrous iron oxidation by anoxygenic phototrophic bacteria. Nature. 1993;362:834–836. [Google Scholar]
- Wyckoff EE, Mey AR, Leimbach A, Fisher CF, Payne SM. Characterization of ferric and ferrous iron transport systems in Vibrio cholerae. The Journal of Bacteriology. 2006;188:6515–6523. doi: 10.1128/JB.00626-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yasmin S, Andrews SC, Moore GR, Le Brun NE. A new role for heme, facilitating release of iron from the bacterioferritin iron biomineral. The Journal of biological chemistry. 2011;286:3473–3483. doi: 10.1074/jbc.M110.175034. [DOI] [PMC free article] [PubMed] [Google Scholar]


