A plant homolog of a bacterial ribosome biogenesis factor functions in chloroplasts, associates with thylakoid membranes, and is involved in maturation of ribosomal RNA.
Abstract
Plastids (chloroplasts) possess 70S ribosomes that are very similar in structure and function to the ribosomes of their bacterial ancestors. While most components of the bacterial ribosome (ribosomal RNAs [rRNAs] and ribosomal proteins) are well conserved in the plastid ribosome, little is known about the factors mediating the biogenesis of plastid ribosomes. Here, we have investigated a putative homolog of the bacterial RbfA (for ribosome-binding factor A) protein that was identified as a cold-shock protein and an auxiliary factor acting in the 5′ maturation of the 16S rRNA. The unicellular green alga Chlamydomonas reinhardtii and the vascular plant Arabidopsis (Arabidopsis thaliana) both encode a single RbfA-like protein in their nuclear genomes. By generating specific antibodies against this protein, we show that the plant RbfA-like protein functions exclusively in the plastid, where it is associated with thylakoid membranes. Analysis of mutants for the corresponding gene (termed RBF1) reveals that the gene function is essential for photoautotrophic growth. Weak mutant alleles display reduced levels of plastid ribosomes, a specific depletion in 30S ribosomal subunits, and reduced activity of plastid protein biosynthesis. Our data suggest that, while the function in ribosome maturation and 16S rRNA 5′ end processing is conserved, the RBF1 protein has assumed an additional role in 3′ end processing. Together with the apparent absence of a homologous protein from plant mitochondria, our findings illustrate that the assembly process of the 70S ribosome is not strictly conserved and has undergone some modifications during organelle evolution.
All members of the Archaeplastida (also referred to as Plantae sensu lato) are descendants of a common ancestor that arose through endosymbiotic acquisition of a cyanobacterium by a protist-like host cell more than 1 billion years ago (Gray, 1993; Archibald, 2009). The Archaeplastida comprise the Rhodophyta, Glaucophyta, and Viridiplantae, with the latter clade including embryophyte plants and chlorophyte algae. While the cyanobacterial endosymbiont possessed thousands of genes in its genome, present-day plastid (chloroplast) genomes are much reduced and, in seed plants, harbor a relatively conserved set of only approximately 130 genes. This drastic genome shrinkage results from two evolutionary processes: (1) the loss of genetic information that became dispensable with the switch from a free-living to a symbiotic lifestyle, and (2) the massive transfer of genes from the genome of the cyanobacterial endosymbiont to the nuclear genome of its host cell (Timmis et al., 2004; Bock and Timmis, 2008; Fuentes et al., 2012).
From its bacterial ancestor, the vestigial genome of plastids has retained many typical prokaryotic features. A bacteria-type RNA polymerase (plastid-encoded RNA polymerase), whose core subunits are encoded in the plastid genome, transcribes most plastid genes. Together with nucleus-encoded sigma factors of the σ70 type, the plastid-encoded RNA polymerase enzyme recognizes promoters that contain conserved −10 (TATA) and −35 boxes and thus resemble bacterial promoters (Liere and Börner, 2007). Surprisingly, although a single RNA polymerase is sufficient to transcribe an entire bacterial genome, transcription of the plastid genome in vascular plants involves a second activity that is encoded in the nuclear genome and mediated by bacteriophage-type RNA polymerases (nucleus-encoded RNA polymerases). Similar to the operon organization of bacterial genomes, most plastid genes are arranged in operons and transcribed as polycistronic RNAs. However, the posttranscriptional processing of the polycistronic transcripts is considerably more complex in plastids than in bacteria, involving maturation of the 5′ and 3′ ends, intercistronic cleavage into monocistronic and oligocistronic units, intron splicing, and, in vascular plants, additionally RNA editing (Westhoff and Herrmann, 1988; Hayes et al., 1996; Bock, 2000; Zhou et al., 2007; Stern et al., 2010).
Translation in plastids is performed by 70S ribosomes (sometimes also referred to as chlororibosomes) that share many properties with bacterial ribosomes. The small subunit of the plastid ribosome (30S subunit) contains the 16S ribosomal RNA (rRNA) and 21 protein subunits that have clear orthologs in Escherichia coli. Twelve of these 21 subunits are encoded in the plastid genome, and the remaining nine subunits are encoded in the nuclear genome (Yamaguchi et al., 2000). The large subunit of the plastid ribosome (50S subunit) consists of three RNA subunits (23S rRNA, 5S rRNA, and 4.5S rRNA) and 31 ribosomal proteins that have orthologs in E. coli. Nine of these 31 proteins are encoded by plastid genes, whereas the remaining 22 are encoded by nuclear genes (Yamaguchi and Subramanian, 2000). Similar to ribosomal proteins in E. coli, most plastid ribosomal proteins are essential for cellular viability (Rogalski et al., 2006, 2008; Fleischmann et al., 2011). In addition to the conserved set of bacterial ribosomal proteins, plastid ribosomes also possess a small set of so-called plastid-specific ribosomal proteins that have no orthologs in E. coli (Yamaguchi et al., 2000; Yamaguchi and Subramanian, 2000; Manuell et al., 2007; Sharma et al., 2007). While some plastid-specific ribosomal proteins appear to represent bona fide ribosomal proteins (in that their knockout or knockdown results in a reduced number of plastid ribosomes and a general reduction in plastid translational activity), others are nonessential proteins of as yet unknown function (Tiller et al., 2012).
Similar to the ribosomes, the other components of the plastid translational machinery are also closely related to the counterparts in the bacterial machinery, including the set of tRNAs (Alkatib et al., 2012) and the initiation, elongation, and termination factors (Meurer et al., 2002; Albrecht et al., 2006; Motohashi et al., 2007; Shen et al., 2013). The translation of many plastid mRNAs initiates at Shine-Dalgarno (SD) sequences, a conserved purine-rich motif upstream of the start codon that acts as a ribosome-binding site by engaging in complementary base pairing with the 3′ end of the 16S rRNA. The SD consensus sequence is identical in bacteria and plastids (Ruf and Kössel, 1988; Kozak, 1999; Drechsel and Bock, 2011). However, a number of mRNAs in both bacteria and plastids do not possess SD sequences (Ruf and Kössel, 1988; Sugiura et al., 1998; Hirose and Sugiura, 2004). Translation of these mRNAs initiates by an alternative (SD-independent) mechanism that relies on start codon recognition through the local absence of RNA secondary structure (Scharff et al., 2011).
Relatively little is known about the assembly of the plastid ribosome. Interestingly, at least some plastid ribosomal proteins can faithfully assemble into E. coli ribosomes, underscoring the remarkable evolutionary conservation of the translational apparatus (Bubunenko and Subramanian, 1994; Weglöhner et al., 1997). Work in bacteria has elucidated the sequence of events in ribosome assembly and established detailed assembly maps for both ribosomal subunits (Dabbs, 1991; Talkington et al., 2005; Kaczanowska and Rydén-Aulin, 2007; Woodson, 2008; Connolly and Culver, 2009; Fournier et al., 2010). It also identified a number of auxiliary factors acting in rRNA processing and/or ribosome maturation (Davies et al., 2010; Connolly and Culver, 2013; Jacob et al., 2013). For some of these factors, homologous genes have been identified in the nuclear genomes of plants and algae, but very few of these have been studied functionally (Bang et al., 2012), and for most of the encoded proteins, their site of action (chloroplast or mitochondrion or both) and their possible role in ribosome biogenesis remain to be determined.
Surprisingly, we found a possible ribosome biogenesis factor in GreenCut, an assemblage of genes that are specifically associated with the evolution of the Viridiplantae lineage (Merchant et al., 2007; Karpowicz et al., 2011). GreenCut genes are genes that are present in the genome of the model green alga Chlamydomonas reinhardtii (Merchant et al., 2007) and have orthologs in the genomes of embryophytes (i.e. bryophytes and vascular plants) but not in the genomes of nonphotosynthetic eukaryotes and prokaryotes. Consequently, the GreenCut gene set should be enriched for genes that are specifically involved in chloroplast-related functions. The putative ribosome biogenesis factor in GreenCut was identified by its homology to RbfA (for ribosome-binding factor A), a bacterial 30S subunit-binding protein that is required for efficient 5′ processing of the 16S rRNA and whose loss of function triggers the cold-shock response in E. coli (Jones and Inouye, 1996; Xia et al., 2003; Datta et al., 2007). The presence of a putative RbfA homolog in GreenCut suggested to us that the plant protein has some unusual properties that caused its classification as potentially specific to the chloroplast-bearing lineage of eukaryotes. This prompted us to study the function of the plant gene in detail.
RESULTS
A Putative RbfA Homolog Encoded in the Nuclear Genomes of Plants and Green Algae
The surprising presence of a putative RbfA domain protein in GreenCut (Merchant et al., 2007; Karpowicz et al., 2011; see introduction) led us to investigate the evolution of this protein in prokaryotes and eukaryotes in more detail. Database searches and sequence alignments (Fig. 1; Supplemental Fig. S1) revealed that the genomes of most organisms contain a single RbfA-like protein. The high degree of sequence conservation in all prokaryotes including cyanobacteria suggests orthology to the E. coli RbfA protein (Fig. 1, A and C). Animals including humans possess an RbfA homolog that is predicted to be a mitochondrial protein (http://www.ncbi.nlm.nih.gov/gene/79863) but has not been functionally characterized so far. Interestingly, there is no obvious homolog present in the genome of Saccharomyces cerevisiae, possibly indicating that, at least in this yeast, mitochondrial ribosomes can be assembled in the absence of RbfA. The unicellular green alga C. reinhardtii (Merchant et al., 2007) and the seed plant model Arabidopsis (Arabidopsis thaliana; Arabidopsis Genome Initiative, 2000) possess RbfA-like proteins that display clear sequence similarity to bacterial proteins within the RbfA domain (Fig. 1C) but little similarity outside this domain (Fig. 1; Supplemental Fig. S1). In addition, it differs from bacterial RbfA proteins by an insertion in the RbfA domain (Fig. 1, B and C). When modeled onto the bacterial RbfA structure, this insertion appears to form a largely unstructured loop that connects two conserved α-helical domains (Fig. 1B). Considering the presence of both conserved and unique features, we tentatively named the plant proteins RBF1 (for RbfA domain-containing protein) and set out to investigate the function of RBF1 in Arabidopsis, where RBF1 is encoded by a single-copy nuclear gene harboring two introns (Fig. 2A).
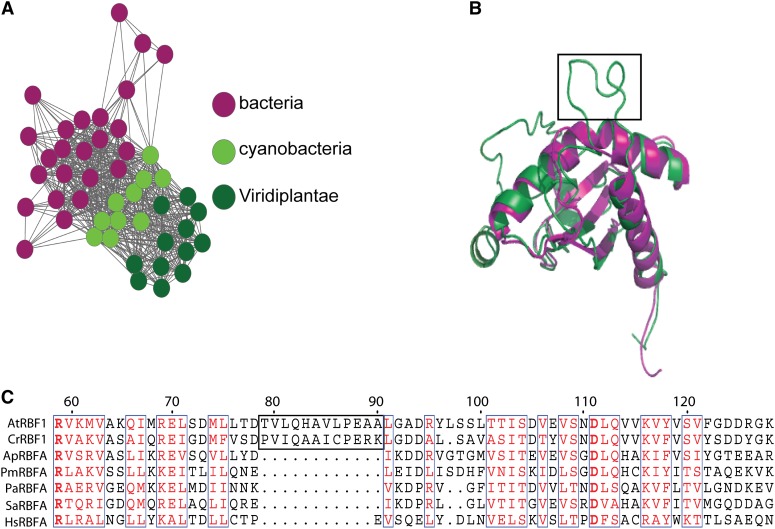
Figure 1.
RBF1 is related to but distinct from bacterial RbfA proteins. A, Protein similarity network of RbfA domain proteins. RbfA domain protein sequences from bacteria, cyanobacteria, green algae, and embryophyte plants were compared in an all-versus-all BLAST with a cutoff e-value of 10−18 and visualized with Cytoscape (http://www.cytoscape.org/). Each circle represents one protein. Lines connect proteins that have e-values below the cutoff. B, Comparison of RbfA structures. The predicted structure of the RBFA protein from Arabidopsis (green) is superimposed on the crystal structure of RbfA from the thermophile bacterium Thermotoga maritima (Protein Data Bank no. 2KZF; pink). I-TASSER was used to generate the predicted structure (Roy et al., 2010). The boxed region represents the Viridiplantae-specific insertion and corresponds to the boxed sequence in C. C, Amino acid sequence alignment of RbfA-like proteins from Arabidopsis (At), C. reinhardtii (Cr), Arthrospira platensis (Ap), Prochlorococcus marinus strain MIT 9215 (Pm), Pseudomonas aeruginosa MPAO1/P2 (Pa), Staphylococcus aureus subspecies aureus N315 (Sa), and Homo sapiens (Hs), where RBFA is predicted to be a mitochondrial protein (http://www.ncbi.nlm.nih.gov/gene/79863). The region boxed in black (amino acids 120–132) represents the plant-specific insertion (see B) that is absent from all bacterial RbfA proteins. Highly conserved amino acids are boxed in blue and indicated in red font, and absolutely conserved ones are in boldface red font. Only the conserved domain of RbfA is shown in this alignment. For a complete sequence alignment, see Supplemental Figure S1.
Figure 2.
RBF1 is expressed early in plant development and is loosely associated with the thylakoid membrane. A, Domain organization and exon-intron structure of the RBF1 gene. The RBF1 genomic locus contains three exons (dark gray boxes) and two introns (black lines). Light gray boxes represent the UTRs (5′ and 3′), and the white box denotes the putative transit peptide (TP) for protein import into plastids. The region used to generate RBF1-specific antisera is indicated. B, RBF1 protein accumulation during plant development. Thylakoid membrane samples equivalent to 1 μg of chlorophyll from 1- to 6-week-old Arabidopsis plants were loaded as indicated and analyzed by immunoblotting with anti-RBF1, anti-Lhcb1.1, and anti-Lhcb1.2 antisera. A representative blot stained with Ponceau S is shown at the bottom. C, Analysis of the subcellular localization of the RBF1 protein in Arabidopsis. Total plant extract (Ex) and nuclear, mitochondrial, chloroplast, thylakoid, and chloroplast stromal fractions (Nucl, Mito, Cp, T, and S) were purified, and equivalent amounts of protein (10 μg per sample) were analyzed by immunoblotting using antibodies against RBF1 and the marker proteins D1 (a reaction center protein of PSII), cytochrome f (Cyt f; a subunit of the cytochrome b6f complex in the thylakoid membrane), the mitochondrial outer membrane protein TOM40, the nuclear protein CRYPTOCHROME2 (CRY2), and the soluble chloroplast protein RbcL (for large subunit of Rubisco). D, Weak membrane association of chloroplast RBF1. Isolated thylakoid membranes were washed with 0.4 m NaCl, and the thylakoid membranes (T) and the supernatant (S) were probed by immunoblotting with antibodies against RBF1 and the PSII reaction center protein D1.
RBF1 Localizes Exclusively to Plastids
The presence in the plant cell of two compartments with bacteria-type ribosomes raised the question of whether the Arabidopsis RBF1 protein localizes to mitochondria, to plastids, or to both organelles. While the suggested mitochondrial localization of the RbfA-like protein in humans could argue for a function of RBF1 in plant mitochondria, the closer association of the plant protein with cyanobacteria (the bacterial clade representing the ancestor of chloroplasts) than with α-proteobacteria (the clade representing the presumptive ancestor of all mitochondria) could argue for a function in plastids (Fig. 1A). Moreover, a number of conserved protein functions required in both organelles are known to be executed by dually targeted proteins (Carrie and Small, 2013). For example, some proteins involved in RNA processing (Gutmann et al., 2012) and translation (Ueda et al., 2008) are dually targeted to both plastids and mitochondria.
To address the subcellular localization of the RBF1 protein in plants, we generated a specific antibody by immunizing rabbits with a protein fragment covering most of the coding region of the Arabidopsis RBF1 locus (Fig. 2A). When total plant protein extracts were assayed with the antibody, a single protein of approximately 20 kD was recognized. This value is very similar to the calculated molecular mass of the RBF1 protein if the mass of the putative transit peptide (http://www.uniprot.org/uniprot/O65693) is subtracted.
Analysis of protein accumulation in Arabidopsis plants of different age revealed a pronounced age-dependent accumulation of the putative RBF1 protein. Expression was strongest in young (1-week old) plants and declined progressively with increasing plant age (Fig. 2B). Investigation of a developmental series of Arabidopsis leaves confirmed the strongly age-dependent accumulation pattern of RBF1. Expression was highest in very young leaves and declined progressively toward the oldest leaves at the bottom of the rosette (Supplemental Fig. S2, A and B).
To determine the cellular compartment to which the RBF1 protein localizes, nuclei, mitochondria, and chloroplasts were purified and assayed with the anti-RBF1 antibody. Immunoblot experiments including marker proteins for all compartments revealed that the RBF1 protein is only present in chloroplasts (Fig. 2C). When chloroplasts were further fractionated into thylakoids and stroma, only the thylakoid fraction showed a hybridization signal with the antibody, suggesting that RBF1 is associated with photosynthetic membranes (Fig. 2C). Treatment of isolated thylakoid membranes with the mildly chaotropic salt NaCl removed most of the RBF1 protein from the membrane fraction, indicating that RBF1 is only loosely associated with the thylakoid surface (Fig. 2D).
To explore a possible regulation of RBF1 association with photosynthetic membranes by light, plants incubated in the dark were compared with plants incubated under low-light or high-light conditions. No difference in thylakoid membrane association was seen (Supplemental Fig. S2C), indicating that the weak association of RBF1 with the thylakoid membrane is largely independent of the photosynthetic activity of the chloroplast.
Isolation of Arabidopsis Mutants for RBF1
Having established that RBF1 is a chloroplast protein, we next wanted to elucidate the function of RBF1 by reverse genetics. To this end, we searched the publicly available collections of transfer DNA (T-DNA) insertion lines and transposon mutants for the presence of rbf1 mutant alleles. Three T-DNA insertion lines potentially affecting the RBF1 locus could be identified (Fig. 3). Line rbf1-1 carries the T-DNA insertion in the 3′ portion of the RBF1 coding region (in codon 202 of the RBF1 reading frame, followed by a 5-bp deletion). Line rbf1-2 harbors a T-DNA insertion in the 5′ untranslated region (UTR; 99 bp upstream of the ATG, followed by an 8-bp deletion). In line rbf1-3, the T-DNA disrupts the intron between exons 2 and 3 (insertion 52 bp upstream of exon 3; Fig. 3A). All three lines were genotyped to confirm the T-DNA insertions, and the isolation of homozygous mutants was attempted by selfing. For all three T-DNA lines, phenotypic segregation of the progeny was observed, as expected. Homozygous rbf1-1 plants displayed the mildest phenotype of the three mutants. The plants grew autotrophically in soil, and only the young leaves were visibly different from wild-type leaves in that they show a pigmentation gradient from the leaf base to the leaf tip, with the base (corresponding to the ontogenetically youngest part of the leaf) being yellow and the tip (representing the oldest part of the leaf) being greenest (Fig. 3C; Supplemental Fig. S3). In contrast to the rbf1-1 mutant, homozygous rbf1-2 and rbf1-3 plants were incapable of autotrophic growth (Supplemental Fig. S3) and needed to be raised on Suc-containing synthetic medium in sterile culture (Fig. 3C). While rbf1-2 plants reached the young seedling stage upon growth in soil and then died (presumably of carbon starvation), rbf1-3 plants died even earlier (Supplemental Fig. S3). When rbf1-2 seedlings were raised on Suc-containing synthetic medium until they reached the early rosette stage, they could be transferred to soil and grown to maturity under autotrophic conditions, suggesting that Rbf1 function is particularly important during early plant development.
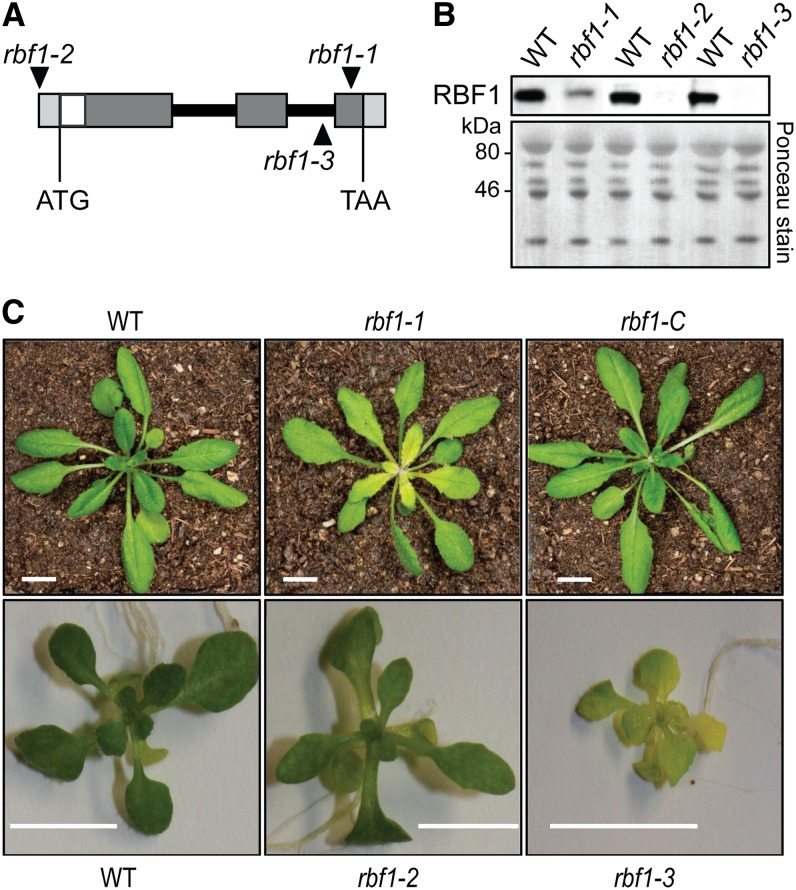
Figure 3.
Isolation of Arabidopsis rbf1 mutants, and characterization of their phenotypes. A, T-DNA insertion sites in tagged Arabidopsis rbf1 mutants. T-DNA insertion sites (black triangles) are shown in relation to the RBF1 gene structure. The three rbf1 alleles analyzed in this study are denoted as rbf1-1, rbf1-2, and rbf1-3. The RBF1 coding region is indicated by the translational start (ATG) and stop (TAA) codons. The exon-intron structure of the RBF1 genomic locus is represented as in Figure 2. B, Immunoblot detection of RBF1 in leaf extracts of wild-type (WT) and rbf1 mutant plants using anti-RBF1 antibodies. Leaf total protein extracts from 3-week-old wild-type and rbf1 seedlings (for growth conditions, see “Materials and Methods”; 10 μg of protein loaded) were separated by SDS-PAGE and probed with anti-RBF1 antibodies. A representative blot stained with Ponceau S is shown at the bottom. C, Phenotypes of rbf1 mutants. The top row shows, from left to right, a wild-type plant, an rbf1-1 mutant plant (exhibiting pale young leaves), and a complemented rbf1-1 mutant plant (rbf1-C). The bottom row shows, from left to right, a wild-type plant, an rbf1-2 mutant plant, and an rbf1-3 mutant plant grown on Suc-containing synthetic medium. Plants were grown under long-day conditions (16 h of light, 8 h of dark) at a photon flux density of 120 μmol m−2 s−1. Bars = 1 cm.
Comparative analysis of the phenotypes of the three mutant alleles tentatively suggested that rbf1-3 represents a null allele, presumably because the T-DNA insertion blocks intron splicing (Fig. 3A). rbf1-2 may be a slightly leaky allele that allows for some residual expression due to the location of the T-DNA insertion outside of the coding region (in the 5′ UTR). This conclusion is further supported by the significantly better growth of rbf1-2 mutant plants on Suc-containing medium compared with rbf1-3 mutant plants (which are severely pigment deficient also under in vitro conditions; Fig. 3C). rbf1-1 is likely to represent a hypomorphic allele that entails a partial loss of gene function, most likely due to insertion into the (lowly conserved) 3′ portion of the coding region (Fig. 3A; Supplemental Fig. S1).
To confirm this interpretation of the rbf1 phenotypes, we assessed RBF1 protein accumulation in the three mutants. While the protein could be readily detected in rbf1-1 mutant plants (albeit at significantly reduced levels compared with the wild type), no protein was detectable in rbf1-2 and rbf1-3 plants (Fig. 3B). Although the slightly milder phenotype of the rbf1-2 mutant (Supplemental Fig. S3) had suggested that there is some residual RBF1 expression, this expression level appears to be below the detection limit of our antibodies, which is approximately 10% of wild-type levels (Supplemental Fig. S4).
To determine the RBF1 expression level in the hypomorphic rbf1-1 mutant, protein accumulation was assessed by comparison with a dilution series of wild-type protein extract. The results of these analyses suggest that the RBF1 accumulation level in the rbf1-1 mutant is between 20% and 40% of the wild-type level (Supplemental Fig. S4, A and B).
Assuming that translation of the rbf1-1 mRNA terminates at the first in-frame stop codon within the T-DNA sequence, a chimeric protein would be produced in the mutant that carries 21 amino acids derived from translation of the T-DNA sequence at its C terminus. The molecular mass of this fusion protein is very similar to that of the wild-type RBF1 protein (Fig. 3B; Supplemental Fig. S4).
Genetic Complementation of the rbf1-1 Mutant
To ultimately confirm that the mutant phenotypes are caused by T-DNA insertion into the RBF1 locus, we sought to complement a mutant line with an allele derived from the wild-type RBF1 complementary DNA (cDNA) sequence. Due to the severe mutant phenotypes of the homozygous rbf1-2 and rbf1-3 plants, this experiment was performed with homozygous rbf1-1 mutant plants, which flowered normally, thus allowing transformation by the floral dip method (Clough and Bent, 1998). To be able to distinguish the RBF1-1 protein from the protein made from the transgenic construct, the RBF1 cDNA was translationally fused to the coding region of firefly luciferase. Transgenic expression of the chimeric RBF1-luciferase fusion protein in stably transformed rbf1-1 plants fully rescued the mutant phenotype (Fig. 3C), confirming that the T-DNA insertion into the RBF1 locus is causally linked to the observed pale-green phenotype.
In immunoblot analyses, the anti-RBF1 antibody detected three bands in the complemented lines: an approximately 20-kD band representing the RBF1-1 protein (calculated molecular mass of the mature protein, 18.1 kD), an approximately 80-kD band corresponding to the expected molecular mass of the RBF1-luciferase fusion protein, and an approximately 55-kD band presumably being a degradation product of the fusion protein (Supplemental Fig. S4C). Faithful recognition of the RBF1-luciferase fusion protein in three independently generated complemented lines also provided an ultimate confirmation of the specificity of our anti-RBF1 antibody.
Altered Chloroplast Ultrastructure, Reduced Thylakoid Membrane Protein Accumulation, and Impaired Photosynthesis in rbf1 Mutant Plants
Together with the chloroplast localization of the RBF1 protein, the pale phenotype of young leaves of rbf1-1 mutant plants suggested an RBF1 function related to chloroplast biogenesis. To test this assumption, we examined chloroplast ultrastructure in wild-type and mutant plants by transmission electron microscopy. Consistent with the reduced leaf pigmentation, chloroplasts in young leaves of rbf1-1 mutant plants showed an underdeveloped thylakoid membrane system, from which grana stacks were largely absent (Fig. 4). Taken together with the observed weak association of the RBF1 protein with thylakoid membranes, this finding provided preliminary evidence for a (direct or indirect) involvement of RBF1 in thylakoid membrane biogenesis.
Figure 4.
Comparison of chloroplast ultrastructure in wild-type (WT) and rbf1-1 mutant plants. Young leaves from wild-type and rbf1-1 mutant plants were analyzed by electron microscopy. A, Chloroplast ultrastructure. Note the strongly reduced grana stacking in the rbf1-1 mutant. B, Enlargement of the region boxed in A showing the structure of thylakoid membranes in the wild type and the mutant.
As thylakoid membranes are highly protein rich, altered thylakoid membrane morphology is often accompanied by changes in the accumulation of the four major thylakoidal protein complexes: PSII, the cytochrome b6f complex, PSI, and ATP synthase. When the accumulation of these complexes was investigated using antibodies against diagnostic (i.e. essential) protein subunits, all four complexes were found to be reduced in the rbf1-1 mutant (Fig. 5A). This reduction was confirmed at the level of the fully assembled complexes by blue-native gel electrophoresis. All detectable complexes and supercomplexes containing PSII and its light-harvesting antenna (light-harvesting complex II) are present in visibly lower amounts in the rbf1-1 mutant (Fig. 5B).
Figure 5.
Thylakoid protein accumulation and photosynthesis in the rbf1-1 mutant. A, Immunoblot analysis of thylakoid proteins diagnostic for PSII, PSI, and ATP synthase. For quantitative comparison, a dilution series for the wild type (WT; 25%, 50%, and 100%) was loaded. Protein extracts were separated by SDS-PAGE and probed with specific antibodies directed against D1 (PSII reaction center subunit), Lhcb1 (PSII antenna protein), PsaA (PSI reaction center subunit), Lhca1 (PSI antenna protein), CF1αβ (ATP synthase subunits), and the large subunit of Rubisco (RbcL), a soluble protein in the stroma. A representative gel stained with Coomassie blue is shown below the blots to confirm even loading of the gel. B, Blue-native gel electrophoretic analysis of thylakoidal protein complexes from young (YL) and mature (ML) leaves. C, False-color display of Fv/Fm of rbf1-1 and wild-type plants. The color scale representing Fv/Fm is given at the right.
Finally, we wanted to test whether altered thylakoid morphology and reduced accumulation of thylakoidal protein complexes translate into a lower efficiency of photosynthetic electron transport in young leaves of the rbf1-1 mutant. Determination of the maximum quantum efficiency of PSII (Fv/Fm) by fluorescence imaging revealed strongly reduced Fv/Fm values in young mutant leaves, whereas mature (fully expanded) leaves had similar values to the corresponding leaves of wild-type plants (Fig. 5C). Consistent with their wild-type-like phenotype, the complemented lines had Fv/Fm values that were similarly high to those of wild-type plants (Supplemental Fig. S4D).
Reduced Accumulation of 30S Subunits of the Chloroplast Ribosome in rbf1 Mutants
In E. coli, RbfA is involved in ribosome biogenesis through mediating efficient processing of the 5′ end of the 16S rRNA during assembly of the 30S ribosomal subunit (Xia et al., 2003; Datta et al., 2007). While the leaf age-dependent phenotype observed in the rbf1-1 mutant could be compatible with a similar function of the RBF1 protein in chloroplasts (Fleischmann et al., 2011), the unusual structural properties of RBF1 (Fig. 1; Supplemental Fig. S1), its presence in GreenCut, and its association with the thylakoid membrane (Fig. 2; Supplemental Fig. S2) had cast some doubt on a function of RBF1 in ribosome biogenesis. To address a possible ribosome-related function of RBF1 in chloroplasts, we employed a microfluidics-based method to quantify the rRNA species of cytosolic 80S ribosomes and plastid 70S ribosomes (Walter et al., 2010; Tiller et al., 2012). Since rRNAs do not accumulate if not incorporated into ribosomal subunits, the abundance of individual rRNA species serves as a proxy for the accumulation of the respective ribosomal subunits. Moreover, as the large and small subunits of the 70S and 80S ribosomes associate with each other only during the translation process and otherwise are present in the dissociated state, the two ribosomal subunits accumulate largely independently of each other. Consequently, the comparative quantification of rRNA species also provides a suitable method to distinguish between defects in the biogenesis of the large versus the small subunits of the ribosome (Walter et al., 2010).
To identify possible defects in chloroplast ribosome biogenesis in rbf1 mutant plants, the ratios of plastid to cytosolic ribosomal subunits and the ratio of small (30S) to large (50S) subunits of the chloroplast ribosome were measured for wild-type plants and the rbf1-1 and rbf1-2 mutants (Fig. 6). Young leaves of rbf1-1 mutant plants showed severely reduced levels of 30S plastid ribosomal subunits (as evidenced by low ratios of plastid 16S rRNA to cytosolic 18S rRNA and plastid 16S rRNA to plastid 23S rRNA; Fig. 6A), whereas the 50S subunit accumulated to wild-type-like levels. In contrast, chloroplast ribosomal subunit accumulation in mature rbf1-1 leaves was not significantly different from that of wild-type leaves (Fig. 6B), in agreement with the wild-type-like phenotype of fully expanded rbf1-1 leaves.
Figure 6.
Accumulation of rRNAs as a proxy for the corresponding ribosomal subunits of the cytosolic and plastid ribosomes. The ratios of plastid 16S rRNA (component of the 30S subunit) to plastid 23S rRNA (component of the 50S subunit), plastid rRNAs to the cytosolic 18S rRNA, and cytosolic 18S rRNA to cytosolic 25S rRNA are shown for the wild type (WT) and the rbf1-1 and rbf1-2 mutants. “23SH1” is the largest “hidden break” product of the 23S rRNA (Delp and Kössel, 1991; Tiller et al., 2012), and “23S” is the 2.8-kb precursor. Significant differences (P < 0.05) are marked with asterisks. Data are from three technical replicates for each plant line, and error bars indicate sd. A, rbf1-1 young leaves. B, rbf1-1 mature (fully expanded) leaves. C, rbf1-2 mature (fully expanded) leaves.
To test if mature leaves are affected in the rbf1 mutants with more severe phenotypes, we also analyzed plastid ribosome accumulation in mature leaves of the rbf1-2 mutant. Indeed, rbf1-2 mutant plants showed strongly reduced levels of 30S ribosomal subunits (as assessed by the ratio of 16S to 23S or 18S rRNA) also in mature leaves (Fig. 6C), in line with their much more severe phenotype than the rbf1-1 mutant plants (Supplemental Fig. S3).
Impaired rRNA Processing in rbf1 Mutants
The ribosome quantification established that rbf1 mutant plants display a specific deficiency in the small ribosomal subunit, suggesting a role for the RBF1 protein in the biogenesis of the 30S subunit. To determine if reduced accumulation of the small subunit of the chloroplast ribosome is correlated with defects in rRNA processing, RNA gel-blot experiments with specific probes against the 16S and 23S rRNAs were performed. These analyses revealed inefficient processing of the 16S rRNA in homozygous rbf1-1 and rbf1-2 mutants (Fig. 7). Whereas in wild-type plants, the 16S rRNA was nearly exclusively present in its mature form (1.5-kb species; Fig. 7, A and B), a significant amount of unprocessed precursor (1.7-kb species) accumulated in the mutants. In contrast, the processing of the 23S rRNA did not differ much between wild-type and mutant plants of similar age and developmental stage (Fig. 7C). However, a slight overaccumulation of the 3-kb 23S-4.5S rRNA precursor was seen in the mutant plants. This is likely a secondary consequence of disturbed 30S subunit function and has been observed previously in other mutants with defects in the plastid translational machinery (Yu et al., 2008; Tiller et al., 2012). It is noteworthy in this respect that several mutants with defects in chloroplast rRNA maturation have been described in the literature (Barkan, 1993; Bisanz et al., 2003; Park et al., 2011; Asakura et al., 2012; Stoppel et al., 2012), but for only a few of them has a direct role in rRNA processing been established (Bollenbach et al., 2005; Stoppel and Meurer, 2012). Many mild disturbances in plastid rRNA maturation are likely secondary effects of defects elsewhere in plastid gene expression (Koussevitzky et al., 2007; Yu et al., 2008).
Figure 7.
Analysis of rRNA processing in rbf1 mutant plants. A, Physical map and transcript pattern of the plastid rRNA operon. The 7.3-kb primary transcripts, the various processing intermediates, and the mature forms of the 16S and 23S rRNAs are shown. Note that the 23S rRNA is cleaved into three pieces, a phenomenon known as hidden break processing (Delp and Kössel, 1991). The positions of the hybridization probes for both genes are also indicated. B, Analysis of 16S rRNA accumulation in wild-type plants (WT) and the rbf1-1 and rbf1-2 mutants. Young and mature leaves from soil-grown plants were analyzed for the rbf1-1 mutant (which grows autotrophically; Fig. 3), whereas plant material grown on Suc-containing synthetic medium (in vitro) was analyzed for the rbf1-2 mutant (because of its seedling-lethal phenotype; Fig. 3; Supplemental Fig. S3). As a loading control, the ethidium bromide-stained gel region containing the cytosolic 25S rRNA is shown. C, Analysis of 23S rRNA accumulation by RNA gel-blot hybridization. The same samples used in B were analyzed.
To investigate the processing defect in the 16S rRNA of rbf1 mutants in more detail and test whether it exclusively affects the 5′ end, as reported for rbfA mutants in E. coli (Xia et al., 2003; Datta et al., 2007), RNA gel-blot experiments with probes specifically recognizing 5′ unprocessed and 3′ unprocessed 16S rRNA precursors were conducted. Surprisingly, these analyses revealed that both 5′ end maturation and 3′ end maturation of the 16S rRNA are impaired in the rbf1 mutants (Fig. 8). This result suggests a striking difference between the bacterial RbfA protein and the plastid RBF1 protein.
Figure 8.
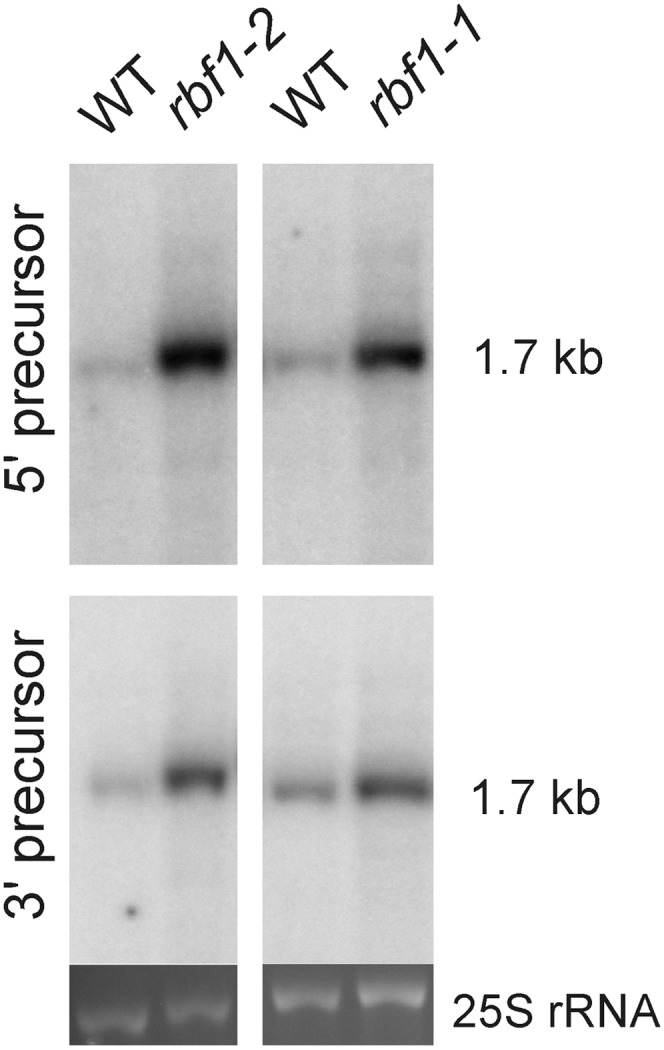
Detection of 5′ unprocessed and 3′ unprocessed precursors of the 16S rRNA. RNA blots were hybridized to probes specifically recognizing 16S precursors with 5′ extensions (5′ precursor; top) or 3′ extensions (3′ precursor; bottom; Supplemental Table S1). The rbf1-2 mutant and the corresponding wild type (WT) were grown on synthetic medium (due to the severe phenotype of the rbf1-2 mutant; compare with Supplemental Fig. S3), whereas the rbf1-1 mutant and its wild-type control were grown in soil. As a loading control, the ethidium bromide-stained gel region containing the cytosolic 25S rRNA is shown.
Chloroplast Translation in rbf1 Mutants
Together with the retarded growth and the reduced photosynthetic performance of rbf1 mutants, the observed defects in 16S rRNA processing and 30S ribosomal subunit accumulation suggest that the rbf1 mutant phenotypes are due to impaired chloroplast translation. To test for translational activity in the mutants, polysome profiles were comparatively analyzed in wild-type plants and rbf1-1 mutant plants. Polysomes are complexes of mRNAs and translating ribosomes, and their migration into continuous Suc gradients upon ultracentrifugation correlates with the number of ribosomes associated with a particular mRNA. Intensely translated mRNAs are loaded with many ribosomes and migrate deeply into the Suc gradient, whereas untranslated and inefficiently translated mRNAs accumulate in the upper fractions of the gradient.
When electrophoretically separated RNA samples extracted from individual fractions of the polysome gradients were hybridized to specific probes for the 16S rRNA and its 1.7-kb precursor as well as the 23S rRNA and its 3.0-kb precursor, it became apparent that at least some of the partially processed rRNA precursor molecules are associated with translating ribosomes. The rRNA precursors are more abundant in polysomes of the rbf1-1 mutant than in polysomes of the wild type (Fig. 9). Proportionally more mature rRNAs were detected in the bottom fraction of the wild-type gradient than in that of the mutant (Fig. 9, A and C; compare fractions 2 and 3 with fraction 6). A similar slight shift to lighter fractions in the mutant was seen when selected mRNAs were assayed (Fig. 10), suggesting that mRNAs in the rbf1-1 mutant are, on average, loaded with fewer ribosomes. It should be noted that, since polysome isolation requires substantial amounts of leaf material, the mutant with the mildest phenotype (rbf1-1) had to be used in these experiments. Based on the severity of the phenotype and the ribosome quantitation data, the reduction in translational activity is likely to be significantly stronger in the rbf1-2 and rbf1-3 mutants.
Figure 9.
Distribution of plastid rRNAs and rRNA precursors between free ribosomes and polysomes in the rbf1-1 mutant and the wild type (WT). Free ribosomes and polysomes were separated using Suc gradient centrifugation, followed by RNA isolation and gel-blot analysis with probes specific for plastid rRNAs and their precursors. The wedges at the bottom indicate the increasing Suc density in the gradients. The light gradient fractions (left lanes) are enriched in free ribosomes, whereas the heavy fractions (right lanes) are enriched in polysomes. The methylene blue-stained cytosolic 18S rRNA is shown as a loading control below each blot. A, Hybridization to a 16S rRNA-specific probe (compare with Fig. 7A). Note the stronger presence of precursor molecules in the rbf1-1 mutant (best visible in fractions 1–3) and the underrepresentation of the 16S rRNA in the heaviest fraction (fraction 6). B, Hybridization to a 16S rRNA precursor-specific probe. The probe hybridizes downstream of the mature 3′ end (nucleotide positions +39 to +116). C, Hybridization to a 23S rRNA-specific probe (compare with Fig. 7A). D, Hybridization to a 23S rRNA precursor-specific probe. The probe binds upstream of the mature 5′ end (nucleotide positions −100 to −26). E, Identification of polysome-containing gradient fractions by centrifugation in the presence of the polysome-dissociating agent puromycin. The blot was hybridized to the 16S rRNA-specific probe.
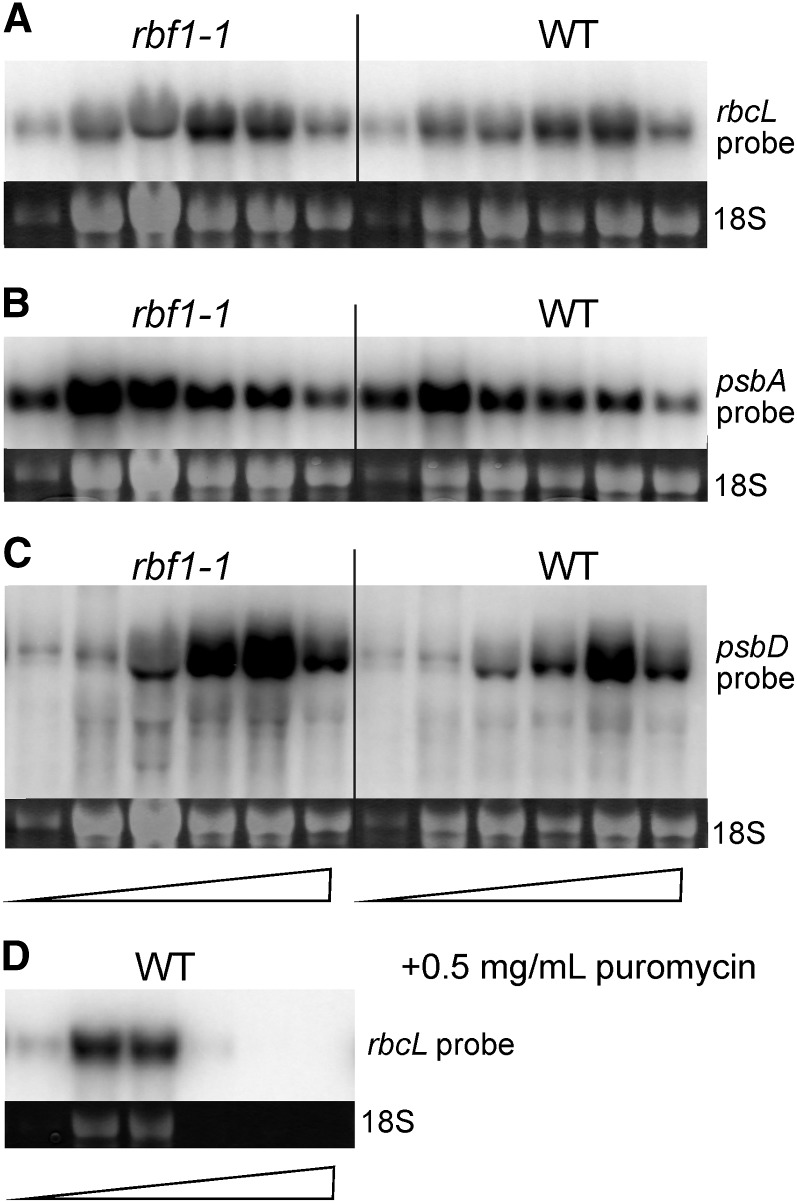
Figure 10.
Polysome loading of selected plastid mRNAs in the wild type (WT) and the rbf1-1 mutant. Blot hybridization analyses of RNAs extracted from fractionated polysome gradients are shown. The methylene blue-stained cytosolic 18S rRNA is shown as a loading control below each blot. The wedges at the bottom indicate the increasing Suc density in the gradient. A, Hybridization to an rbcL-specific probe. Note that the rbcL mRNA peaks in fraction 5 in the wild type but peaks already in fraction 4 in the mutant. B, Hybridization to a psbA-specific probe. C, Hybridization to a psbD-specific probe. Note that the psbD mRNA peaks in fraction 5 in the wild type but in fractions 4 and 5 in the mutant. D, Identification of polysome-containing gradient fractions by centrifugation in the presence of the polysome-dissociating agent puromycin. The blot was hybridized to the rbcL-specific probe.
DISCUSSION
In this work, we have identified the plant RBF1 protein as a factor involved in the biogenesis of the 30S subunit of the chloroplast ribosome. We have shown that impaired RBF1 gene function results in delayed chloroplast development, presumably due to reduced plastid translational capacity. Chloroplast development is known to entail an extraordinarily high demand for de novo protein biosynthesis, because the entire photosynthetic apparatus needs to be built up in a relatively short time. Therefore, even mild disturbances of the translational machinery in the plastid can result in delayed greening of developing leaves. As the leaf ages, the demand for plastid translational capacity decreases and the translational machinery in these mutants gets time to catch up and restore wild-type-like levels of chloroplast proteins (Fleischmann et al., 2011). Thus, the delayed greening observed in the rbf1 mutants is likely to result from plastid translational capacity limiting the speed of thylakoid membrane biogenesis.
Our localization data strongly suggest that RBF1 specifically functions in chloroplast ribosome biogenesis and is not involved in the biogenesis of mitochondrial ribosomes. Searches of sequenced plant genomes revealed no other rbfA or RBF1 homolog that potentially could be targeted to mitochondria. Together with the absence of an rbfA gene from the yeast genome, this finding suggests that the biogenesis of the small subunit of the mitochondrial ribosome can proceed independently of RBF1 function.
Surprisingly, RBF1 was found in GreenCut, a set of genes identified as specifically associated with the evolution of the Viridiplantae lineage (Merchant et al., 2007; Karpowicz et al., 2011). The reason for this is that the plant sequences are very highly related to each other but rather diverged from bacterial RbfA sequences outside the RbfA domain. Since GreenCut required an expect score of e−10 for the identification of a homolog, the proteins in various bacteria and the mitochondrial forms in animals were not viewed as orthologs.
Quantitation of rRNA accumulation and analysis of chloroplast rRNA processing indicate that RBF1 is an ortholog of the bacterial RbfA protein. Its absence causes specific processing defects in the chloroplast 16S rRNA. However, while for bacterial rbfA mutants only defective 5′ end processing of the 16S rRNA has been described (Xia et al., 2003; Datta et al., 2007), analysis of the chloroplast 16S rRNA precursor molecules accumulating in Arabidopsis rbf1 mutants indicates that both 5′ and 3′ end processing are impaired (Figs. 7–9). It is important to note that the effect of the bacterial RbfA protein on 16S rRNA processing is unlikely to be enzymatic in nature. The protein does not appear to possess endoribonucleolytic or exoribonucleolytic activity, nor has any other enzymatic activity been identified (Datta et al., 2007; Connolly and Culver, 2013). Instead, during maturation of the 30S subunit, RbfA displaces 16S rRNA helices that are associated with the decoding process in the ribosome (especially helix 44; Datta et al., 2007). It thus appears that the role of RbfA in the biogenesis of the 30S subunit of the bacterial ribosome is more a structural one in that it may facilitate conformational changes and/or structural transitions during 30S maturation. Whether the additional effect on 3′ end processing in chloroplasts is due to structural differences between RBF1 and RbfA (Fig. 1; Supplemental Fig. S1) or differences in 16S rRNA structure (or both) remains to be elucidated.
Our localization analyses have indicated that RBF1 is associated with the thylakoid membrane. Given that the protein is required for biogenesis of the 30S ribosomal subunit, its association with photosynthetic membranes is somewhat surprising. Although the insertion of some nascent polypeptide chains into the thylakoid membrane occurs cotranslationally (Gray, 1992; Houben et al., 1999; Nilsson et al., 1999) and a significant fraction of chloroplast ribosomes can be associated with the thylakoid membrane (for review, see Peled-Zehavi and Danon, 2007), chloroplast ribosome biogenesis is not known to take place at the thylakoid membrane. However, in the light of our findings on RBF1 localization and the earlier notion that some fusion proteins of GFP with plastid ribosomal proteins are not homogenously distributed within the chloroplast (Tiller et al., 2012), a thorough investigation of the site of plastid ribosome biogenesis is certainly warranted. A possible connection between ribosome biogenesis and thylakoids could be provided by the plastid nucleoid that recently has been implicated in ribosome biogenesis (Majeran et al., 2012). The nucleoid, in turn, appears to be associated with the thylakoid membrane (Liu and Rose, 1992; Jeong et al., 2003), raising the possibility that the plastid nucleoid provides a molecular bridge that tethers ribosome biogenesis to thylakoids. Consistent with this hypothesis, a proteomic analysis has detected RBF1 in megadalton-size complexes that also contain RNA polymerase subunits and various other DNA-binding proteins (Olinares et al., 2010).
In summary, our work here has identified a thylakoid-associated protein that is involved in the biogenesis of the 30S subunit of the chloroplast ribosome. Although the protein exhibits several unusual features compared with its bacterial homologs, the basic function in ribosome maturation and 16S rRNA 5′ processing appears to be conserved. However, the chloroplast protein is additionally required for efficient 3′ end processing. This as well as the apparent absence of a homologous protein from plant (and yeast) mitochondria illustrates that, even for the assembly of a complex macromolecular machine that is as conserved as the prokaryotic ribosome, evolution can develop different solutions.
MATERIALS AND METHODS
Plant Growth Conditions and Mutant Characterization
Arabidopsis (Arabidopsis thaliana) Columbia and Landsberg erecta wild-type plants were obtained from the Arabidopsis Biological Resource Center. Three T-DNA insertion mutants for RBF1 (gene identifier At4g34730) were obtained: 008178 (SALK; Columbia background), 058490 (SALK; Columbia background), and ET7188 (Cold Spring Harbor Laboratory; Landsberg erecta background). Plants in soil were grown in a Conviron growth chamber (model MTR26) under long-day conditions (16 h of light, 8 h of dark) at 120 µmol photons m−2 s−1. For exposure to high-light conditions, plants were grown in a controlled-climate chamber (Percival; model E30BHOC8) at 800 µmol photons m−2 s−1. For growth on Suc-containing synthetic medium, seeds were surface sterilized and plated on Murashige and Skoog agar medium containing 0.8% Suc (Murashige and Skoog, 1962). Mutant plants were screened by PCR for heterozygosity or homozygosity of the T-DNA insertion using primers 008178_F and 008178_R for SALK_008178, primers 058490_F and 058490_R for SALK_058490, primers ET7188_F and ET7188_R for ET7188, and T-DNA left border primers LBd1 and Ds3-4 for SALK and Cold Spring Harbor Laboratory lines, respectively. DNA fragments were amplified by 35 cycles of denaturation at 94°C for 1 min, annealing at 54°C for 55 s, and polymerization at 72°C for 1 min followed by a final extension for 10 min at 72°C.
Organelle Isolation and Biochemical Fractionation
Four-week-old Arabidopsis plants were used for the preparation of chloroplasts (Spetea et al., 2004), thylakoids (Spetea et al., 2004), and nuclei (Folta and Kaufman, 2006) by biochemical fractionation on ice in dim light using ice-cold buffers. Forty grams of Arabidopsis leaves collected after light acclimation (3 h post illumination, 120 μmol photons m−2 s−1) was homogenized in 30 mL of preparation buffer (25 mm Tricine-NaOH, pH 7.8, 330 mm sorbitol, 1 mm EDTA, 10 mm KCl, 0.15% [w/v] bovine serum albumin, 4 mm sodium ascorbate, and 7 mm l-Cys) in a precooled Waring blender for five periods of 1 s at high speed. The homogenate was immediately filtered through four layers of Miracloth (20-μm pore size), and the pellet was collected from the filtrate by centrifugation for 3 min at 1,000g in the cold (4°C). The pellet was resuspended in the same buffer and centrifuged again for 5 min at 1,000g. Intact chloroplasts were purified on Percoll gradients. The resuspended chloroplast pellet was carefully layered on top of a 35%/80% (v/v) Percoll step gradient and separated by centrifugation at 2,000g for 15 min in a swinging-bucket rotor. The lower green band contains intact chloroplasts and was carefully collected from the gradient.
For fractionation into stromal and thylakoid protein fractions, 30 mL of preparation buffer was gently mixed with the chloroplasts collected from the gradient. Subsequently, the chloroplasts were recovered by centrifugation at 2,500g for 4 min, resuspended in 3 mL of chloroplast lysis buffer (10 mm Tricine-NaOH, pH 7.8, and 5 mm MgCl2), and incubated on ice for 15 min. A Pyrex Potter-Elvehjem tissue grinder (homogenizer) was used to mediate complete lysis of the chloroplasts (four strokes). Afterward, the thylakoid membranes were collected by centrifugation for 5 min at 8,000g, resuspended in buffer (100 mm sorbitol, 25 mm Tricine-NaOH, pH 7.8, 5 mm MgCl2, and 10 mm KCl), and purified on a Suc gradient (40%–80%) by centrifugation at 30,000g for 1 h. The thylakoid membranes sediment at the bottom of the tube as a green pellet. The supernatant from the chloroplast lysis step contains the soluble stromal proteins. The stromal fraction was centrifuged at 8,000g for 4 min to remove any membrane particles and concentrated in a SpeedVac to a 500-µL volume.
Nuclei were isolated by a slightly modified published procedure (Folta and Kaufman, 2006). Briefly, 10 g of Arabidopsis leaves was washed by adding 30 mL of cold ethanol (−20°C), and the plant material was chopped continuously for 5 min. Subsequently, the ethanol was removed and the tissue was washed three times for 5 min each with extraction buffer (2 m hexylene glycol, 20 mm PIPES-KOH, pH 7, 10 mm MgCl2, and 5 mm β-mercaptoethanol) followed by homogenization in a Waring blender six times for 1 s at high speed. The homogenate was filtered through four layers of Miracloth into a glass beaker under magnetic stirring. Triton X-100 (25%) was added dropwise to a final concentration of 1% (v/v), and the solution was incubated at 4°C for 10 min to disintegrate all cellular membranes and organelles except the inner nuclear envelope. Following centrifugation for 30 min at 2,000g, the supernatant was removed, and 5 mL of gradient buffer (0.5 m hexylene glycol, 5 mm PIPES-KOH, pH 7.0, 10 mm MgCl2, 5 mm β-mercaptoethanol, and 1% Triton X-100) was added to the pellet. The nuclei were carefully resuspended (taking care to not resuspend contaminating green debris) and then washed five times by resuspension in gradient buffer and centrifugation for 10 min at 2,000g.
Plant mitochondria were isolated from fresh organic spinach (Spinacia oleracea; 480 g) essentially as described (Keech et al., 2005). Before isolation, the leaves were incubated in the dark for 12 h at 4°C to reduce their starch content. The leaves were then blended in grinding buffer (0.3 m Suc, 60 mm Tricine-NaOH, pH 8.0, 2 mm EDTA, 10 mm KH2PO4, 25 mm tetrasodium pyrophosphate, 1 mm Gly, 1% [w/v] polyvinylpyrrolidone 40, 50 mm sodium ascorbate, 1% [w/v] bovine serum albumin, and 20 mm Cys) and filtered through one layer of Miracloth. The filtrate was centrifuged at 2,500g for 5 min, and the supernatant containing the mitochondria was recovered and centrifuged again at 15,000g for 20 min. The mitochondrial pellet was resuspended in 1 mL of wash buffer (0.3 m Suc, 10 mm Tricine-NaOH, pH 7.5, and 10 mm KH2PO4), gently layered onto a Percoll gradient, and centrifuged at 15,000g for 20 min. Gradients were produced by mixing 25 mL of 2× gradient buffer with 25 mL of Percoll followed by centrifugation at 39,000g for 40 min at 4°C in a fixed-angle rotor. The mitochondrial fraction (representing a white band near the bottom of the tube) was collected and washed with 20 volumes of wash buffer.
Protein Isolation, Gel Electrophoretic Separation, and Immunodetection
The various subcellular fractions were separated by SDS-PAGE (6% stacking gel, 14% separation gel, with 6 m urea), and the proteins were subsequently transferred to polyvinylidene difluoride membranes (Immobilone; Millipore). Membranes were blocked with 10% skim milk for 1 h at room temperature with gentle shaking and then incubated with the primary antibody overnight at 4°C. An antibody specific for RBF1 was produced by immunization of rabbits with a recombinant protein corresponding to amino acids 52 to 215 of RBF1 (Covance). Antibodies against the DE loop in the D1 protein (AS10 704), Lhcb1 (AS01 004), and Lhcb2 (AS01 003) were obtained from Agrisera. The CF1αβ antibody was generated by immunization with the purified CF1 complex from spinach (Selman-Reimer et al., 1981). A PsaA-specific antibody was kindly provided by Dr. Jean-David Rochaix (University of Geneva; Rivier et al., 2001). The antibody against Cry2 was described previously (Lin et al., 1998). A maize cytochrome f antibody was kindly provided by Dr. Alice Barkan (University of Oregon; Voelker and Barkan, 1995), the TOM40 antibody was kindly provided by Dr. Jim Whelan (University of Western Australia; Carrie et al., 2009), and the Rubisco large subunit antibody was a gift from Dr. Steve Rodermel (Iowa State University; Rodermel et al., 1988). Blotted membranes were washed six times for 10 min using Tris-buffered saline with 1% (v/v) Tween 20 (TTBS) buffer and incubated at room temperature with horseradish peroxidase-conjugated secondary antibody for 1 h. The membranes were then washed six times for 10 min with TTBS buffer, and the immunoreactive proteins were visualized following a 5-min incubation with detection reagents from the SuperSignal WestPico horseradish peroxidase detection kit (Thermo Scientific). Quantification of the immunoblots was done using the Fujifilm LAS-1000 software.
For blue-native gel electrophoresis, purified thylakoid membranes were resuspended in 20% (w/v) glycerol and 25 mm BisTris-HCl, pH 7.0, to a final chlorophyll concentration of 2 mg mL−1. An equal volume of 1.5% (w/v) n-dodecyl β-d-maltoside dissolved in resuspension buffer was added, and the mixture was incubated on ice for 10 min. After centrifugation at 14,000g for 30 min, the supernatant was supplemented with 0.1 volume of sample buffer (100 mm BisTris-HCl, pH 7.0, 0.5 m ε-amino-n-caproic acid, 30% [w/v] Suc, and 50 mg mL−1 Serva blue G) and subjected to blue-native gel electrophoresis with a gradient of 5% to 13.5% acrylamide on the separation gel. Electrophoresis was performed at 4°C and 100 to 200 V (with an increment of 10 V every 30 min) for 5 h.
Complementation of the rbf1-1 Mutant with the At4g34730 Gene
For transgenic complementation analysis, an RBF1 cDNA fragment was amplified by PCR from a cDNA clone using a high-fidelity polymerase (Phusion; New England Biolabs) and primers RBF1_cloning_1-F(start) and RBF1_cloning_1-R(stop) (Supplemental Table S2). The PCR product was subcloned into a modified pEGAD vector, generating a translational fusion with firefly luciferase. The resulting plasmid pEGAD-RBF1 was transformed into Agrobacterium tumefaciens strain AGL0 by electroporation using standard protocols. Transformation of homozygous rbf1-1 plants was carried out using the floral dip method (Clough and Bent, 1998). Transformants were selected in soil with the herbicide Basta (120 mg L−1; Bayer Scientific; Chemical Abstract Service, CAS no. 77182-82-2). The T1 and T2 generations of selected transformants were assayed by phenotypic analysis, immunoblotting, and determination of chlorophyll fluorescence.
Electron Microscopy
To obtain leaf material for electron microscopy, plants were grown in soil under a 16-h-light/8-h-dark cycle at a light intensity of 120 μmol m−2 s−1. After 3 h into the light period, pieces of leaves from 4-week-old wild-type and mutant seedlings were directly fixed in a solution containing 2% (v/v) glutaraldehyde and 2% (v/v) paraformaldehyde in 0.1 m phosphate-buffered saline (PBS) buffer (pH 7.4) for 2 h at room temperature and then incubated at 4°C overnight. Subsequently, 0.5% (w/v) tannic acid was added, and the samples were incubated for 1 h at room temperature. The leaf pieces were then washed five times in 0.1 m PBS buffer and postfixed in a solution of 1% (w/v) OsO4 (in PBS; pH 7.2–7.4). The combined treatment with tannic acid, glutaraldehyde, and paraformaldehyde followed by osmification enhanced the staining of membranes. The samples were washed four times in 0.1 m sodium acetate buffer (pH 5.5) and then block stained in 0.5% (w/v) uranyl acetate (in 0.1 m sodium acetate buffer, pH 5.5) for 12 h at 4°C. Subsequently, the samples were dehydrated in graded ethanol (50%, 75%, 95%, 100%, 100%, and 100%) for 10 min per step, rinsed with propylene oxide, and infiltrated in mixtures of Epon 812 and propylene oxide (1:1 and then 2:1 for 2 h each), followed by infiltration in pure Epon 812 overnight. Embedding was performed in pure Epon 812, and curing was done in an oven at 60°C for 48 h. Sections of 60 nm thickness (gray interference color) were cut with an ultramicrotome (RMC MTX) using a diamond knife. The sections were deposited on single-hole grids coated with formvar and carbon and double-stained in aqueous solutions of 8% (w/v) uranyl acetate for 25 min at 60°C and lead citrate for 3 min at room temperature. Thin sections were examined with a JEOL 100CX electron microscope.
Chlorophyll Fluorescence Analyses
Imaging of Fv/Fm was performed on whole Arabidopsis plants with a FluorCam 700 MF system (Photon Systems Instruments) using the quenching analysis effect settings. Fluorcam version 1.6 software was used to control the imaging system and to process the images. Pulses of actinic light and continuous illumination were generated by two arrays, each made up of 345 light-emitting diodes (620 nm). Plants were dark adapted for 20 min prior to the measurements.
rRNA and Polysome Analysis
rbf1-1 (SALK_008178) mutant plants were germinated on Murashige and Skoog medium (Murashige and Skoog, 1962) with 2% (w/v) Suc, transferred to soil after 2 weeks, and then cultivated for 2 weeks under an 8-h-light (approximately 140 μmol m−2 s−1, 20°C)/16-h-dark (16°C) cycle. Material from young leaves was compared with mature (i.e. fully expanded) leaves. rbf1-2 (SALK_058490) mutant plants were germinated under identical conditions and, after 3 weeks, transferred to boxes with 0.5× Murashige and Skoog medium supplemented with 1% (w/v) Suc and grown for 3 weeks at a light intensity of approximately 50 μmol m−2 s−1. Material from mature leaves was analyzed. Total RNA was isolated with the TriFast reagent (Peqlab) following the manufacturer’s protocol. rRNAs were quantified by analyzing total RNA with an Agilent 2100 Bioanalyzer using the Agilent RNA 6000 nano kit according to the instructions of the manufacturer (Agilent Technologies). rRNA ratios were determined as described previously (Walter et al., 2010; Tiller et al., 2012). For RNA gel-blot analyses, total RNA was separated on formaldehyde-containing 1.25% (w/v) agarose gels and blotted onto Hybond XL membranes (GE Healthcare). Polysome isolation and analysis were done as described previously (Barkan et al., 1994; Kahlau and Bock, 2008). Control gradients for the identification of fractions containing free mRNAs contained 0.5 mg mL−1 puromycin. Hybridization probes (Supplemental Table S1) were generated by PCR amplification, purified by agarose gel electrophoresis, and then labeled with [α-32P]dCTP by random priming (Multiprime DNA labeling system; GE Healthcare). Hybridizations were performed at 65°C according to standard protocols.
Protein Similarity Networks, Structural Predictions, and Sequence Alignments
A protein similarity network was constructed from bacterial and Viridiplantae RbfA-like sequences (Supplemental Table S3) as described previously (Carrie et al., 2009). The three-dimensional structure of the Arabidopsis RBF1 protein was predicted by I-TASSER from the amino acid sequence without the transit peptide (Roy et al., 2010). Alignments of amino acid sequences were produced with MUSCLE (Edgar, 2004).
Supplemental Data
The following materials are available in the online version of this article.
Supplemental Figure S1. Amino acid sequence alignment of RbfA-like proteins.
Supplemental Figure S2. RBF1 protein abundance and thylakoid association in Arabidopsis leaves.
Supplemental Figure S3. Phenotypes of the rbf-1-1 mutant, the complemented rbf-1-1 mutant (rbf1-C), the rbf1-3 mutant, and the rbf1-2 mutant.
Supplemental Figure S4. RBF1 abundance in the rbf1-1 mutant and characterization of complemented rbf1-1 lines (rbf1-C).
Supplemental Table S1. Probes used for RNA gel-blot analyses.
Supplemental Table S2. Oligonucleotides used as primers for PCR.
Supplemental Table S3. Sequences used to construct the protein similarity network.
Acknowledgments
Electron microscopy was performed at the Electron Microscopy Services Center of the University of California, Los Angeles, Brain Research Institute. We thank Steven Karpowicz (Department of Biology, University of Central Oklahoma) for useful scientific discussion and Mark Arbing and Annie Shin (University of California, Los Angeles-Department of Energy Protein Expression Technology Center) for assistance with the expression of recombinant RBF1.
Glossary
- rRNA
ribosomal RNA
- SD
Shine-Dalgarno
- T-DNA
transfer DNA
- UTR
untranslated region
- cDNA
complementary DNA
- Fv/Fm
maximum quantum efficiency of PSII
- PBS
phosphate-buffered saline
References
- Albrecht V, Ingenfeld A, Apel K. (2006) Characterization of the snowy cotyledon 1 mutant of Arabidopsis thaliana: the impact of chloroplast elongation factor G on chloroplast development and plant vitality. Plant Mol Biol 60: 507–518 [DOI] [PubMed] [Google Scholar]
- Alkatib S, Scharff LB, Rogalski M, Fleischmann TT, Matthes A, Seeger S, Schöttler MA, Ruf S, Bock R. (2012) The contributions of wobbling and superwobbling to the reading of the genetic code. PLoS Genet 8: e1003076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arabidopsis Genome Initiative (2000) Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408: 796–815 [DOI] [PubMed] [Google Scholar]
- Archibald JM. (2009) The puzzle of plastid evolution. Curr Biol 19: R81–R88 [DOI] [PubMed] [Google Scholar]
- Asakura Y, Galarneau E, Watkins KP, Barkan A, van Wijk KJ. (2012) Chloroplast RH3 DEAD box RNA helicases in maize and Arabidopsis function in splicing of specific group II introns and affect chloroplast ribosome biogenesis. Plant Physiol 159: 961–974 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bang WY, Chen J, Jeong IS, Kim SW, Kim CW, Jung HS, Lee KH, Kweon HS, Yoko I, Shiina T, et al (2012) Functional characterization of ObgC in ribosome biogenesis during chloroplast development. Plant J 71: 122–134 [DOI] [PubMed] [Google Scholar]
- Barkan A. (1993) Nuclear mutants of maize with defects in chloroplast polysome assembly have altered chloroplast RNA metabolism. Plant Cell 5: 389–402 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barkan A, Walker M, Nolasco M, Johnson D. (1994) A nuclear mutation in maize blocks the processing and translation of several chloroplast mRNAs and provides evidence for the differential translation of alternative mRNA forms. EMBO J 13: 3170–3181 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bisanz C, Bégot L, Carol P, Perez P, Bligny M, Pesey H, Gallois JL, Lerbs-Mache S, Mache R. (2003) The Arabidopsis nuclear DAL gene encodes a chloroplast protein which is required for the maturation of the plastid ribosomal RNAs and is essential for chloroplast differentiation. Plant Mol Biol 51: 651–663 [DOI] [PubMed] [Google Scholar]
- Bock R. (2000) Sense from nonsense: how the genetic information of chloroplasts is altered by RNA editing. Biochimie 82: 549–557 [DOI] [PubMed] [Google Scholar]
- Bock R, Timmis JN. (2008) Reconstructing evolution: gene transfer from plastids to the nucleus. Bioessays 30: 556–566 [DOI] [PubMed] [Google Scholar]
- Bollenbach TJ, Lange H, Gutierrez R, Erhardt M, Stern DB, Gagliardi D. (2005) RNR1, a 3′-5′ exoribonuclease belonging to the RNR superfamily, catalyzes 3′ maturation of chloroplast ribosomal RNAs in Arabidopsis thaliana. Nucleic Acids Res 33: 2751–2763 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bubunenko MG, Subramanian AR. (1994) Recognition of novel and divergent higher plant chloroplast ribosomal proteins by Escherichia coli ribosome during in vivo assembly. J Biol Chem 269: 18223–18231 [PubMed] [Google Scholar]
- Carrie C, Kühn K, Murcha MW, Duncan O, Small ID, O’Toole N, Whelan J. (2009) Approaches to defining dual-targeted proteins in Arabidopsis. Plant J 57: 1128–1139 [DOI] [PubMed] [Google Scholar]
- Carrie C, Small I. (2013) A reevaluation of dual-targeting of proteins to mitochondria and chloroplasts. Biochim Biophys Acta 1833: 253–259 [DOI] [PubMed] [Google Scholar]
- Clough SJ, Bent AF. (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16: 735–743 [DOI] [PubMed] [Google Scholar]
- Connolly K, Culver G. (2009) Deconstructing ribosome construction. Trends Biochem Sci 34: 256–263 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Connolly K, Culver G. (2013) Overexpression of RbfA in the absence of the KsgA checkpoint results in impaired translation initiation. Mol Microbiol 87: 968–981 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dabbs ER. (1991) Mutants lacking individual ribosomal proteins as a tool to investigate ribosomal properties. Biochimie 73: 639–645 [DOI] [PubMed] [Google Scholar]
- Datta PP, Wilson DN, Kawazoe M, Swami NK, Kaminishi T, Sharma MR, Booth TM, Takemoto C, Fucini P, Yokoyama S, et al (2007) Structural aspects of RbfA action during small ribosomal subunit assembly. Mol Cell 28: 434–445 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davies BW, Köhrer C, Jacob AI, Simmons LA, Zhu J, Aleman LM, Rajbhandary UL, Walker GC. (2010) Role of Escherichia coli YbeY, a highly conserved protein, in rRNA processing. Mol Microbiol 78: 506–518 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Delp G, Kössel H (1991) rRNAs and rRNA genes of plastids. In L Bogorad, IK Vasil, eds, Cell Culture and Somatic Cell Genetics of Plants. Academic Press, New York, pp 139–167 [Google Scholar]
- Drechsel O, Bock R. (2011) Selection of Shine-Dalgarno sequences in plastids. Nucleic Acids Res 39: 1427–1438 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Edgar RC. (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32: 1792–1797 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fleischmann TT, Scharff LB, Alkatib S, Hasdorf S, Schöttler MA, Bock R. (2011) Nonessential plastid-encoded ribosomal proteins in tobacco: a developmental role for plastid translation and implications for reductive genome evolution. Plant Cell 23: 3137–3155 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Folta KM, Kaufman LS. (2006) Isolation of Arabidopsis nuclei and measurement of gene transcription rates using nuclear run-on assays. Nat Protoc 1: 3094–3100 [DOI] [PubMed] [Google Scholar]
- Fournier GP, Neumann JE, Gogarten JP. (2010) Inferring the ancient history of the translation machinery and genetic code via recapitulation of ribosomal subunit assembly orders. PLoS ONE 5: e9437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fuentes I, Karcher D, Bock R. (2012) Experimental reconstruction of the functional transfer of intron-containing plastid genes to the nucleus. Curr Biol 22: 763–771 [DOI] [PubMed] [Google Scholar]
- Gray JC. (1992) Cytochrome f: structure, function and biosynthesis. Photosynth Res 34: 359–374 [DOI] [PubMed] [Google Scholar]
- Gray MW. (1993) Origin and evolution of organelle genomes. Curr Opin Genet Dev 3: 884–890 [DOI] [PubMed] [Google Scholar]
- Gutmann B, Gobert A, Giegé P. (2012) PRORP proteins support RNase P activity in both organelles and the nucleus in Arabidopsis. Genes Dev 26: 1022–1027 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayes R, Kudla J, Schuster G, Gabay L, Maliga P, Gruissem W. (1996) Chloroplast mRNA 3′-end processing by a high molecular weight protein complex is regulated by nuclear encoded RNA binding proteins. EMBO J 15: 1132–1141 [PMC free article] [PubMed] [Google Scholar]
- Hirose T, Sugiura M. (2004) Multiple elements required for translation of plastid atpB mRNA lacking the Shine-Dalgarno sequence. Nucleic Acids Res 32: 3503–3510 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Houben E, de Gier JW, van Wijk KJ. (1999) Insertion of leader peptidase into the thylakoid membrane during synthesis in a chloroplast translation system. Plant Cell 11: 1553–1564 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacob AI, Köhrer C, Davies BW, RajBhandary UL, Walker GC. (2013) Conserved bacterial RNase YbeY plays key roles in 70S ribosome quality control and 16S rRNA maturation. Mol Cell 49: 427–438 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jeong SY, Rose A, Meier I. (2003) MFP1 is a thylakoid-associated, nucleoid-binding protein with a coiled-coil structure. Nucleic Acids Res 31: 5175–5185 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones PG, Inouye M. (1996) RbfA, a 30S ribosomal binding factor, is a cold-shock protein whose absence triggers the cold-shock response. Mol Microbiol 21: 1207–1218 [DOI] [PubMed] [Google Scholar]
- Kaczanowska M, Rydén-Aulin M. (2007) Ribosome biogenesis and the translation process in Escherichia coli. Microbiol Mol Biol Rev 71: 477–494 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kahlau S, Bock R. (2008) Plastid transcriptomics and translatomics of tomato fruit development and chloroplast-to-chromoplast differentiation: chromoplast gene expression largely serves the production of a single protein. Plant Cell 20: 856–874 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karpowicz SJ, Prochnik SE, Grossman AR, Merchant SS. (2011) The GreenCut2 resource, a phylogenomically derived inventory of proteins specific to the plant lineage. J Biol Chem 286: 21427–21439 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keech O, Dizengremel P, Gardeström P. (2005) Preparation of leaf mitochondria from Arabidopsis thaliana. Physiol Plant 124: 403–409 [Google Scholar]
- Koussevitzky S, Stanne TM, Peto CA, Giap T, Sjögren LLE, Zhao Y, Clarke AK, Chory J. (2007) An Arabidopsis thaliana virescent mutant reveals a role for ClpR1 in plastid development. Plant Mol Biol 63: 85–96 [DOI] [PubMed] [Google Scholar]
- Kozak M. (1999) Initiation of translation in prokaryotes and eukaryotes. Gene 234: 187–208 [DOI] [PubMed] [Google Scholar]
- Liere K, Börner T. (2007) Transcription and transcriptional regulation in plastids. Top Curr Genet 19: 121–174 [Google Scholar]
- Lin C, Yang H, Guo H, Mockler T, Chen J, Cashmore AR. (1998) Enhancement of blue-light sensitivity of Arabidopsis seedlings by a blue light receptor cryptochrome 2. Proc Natl Acad Sci USA 95: 2686–2690 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu JW, Rose RJ. (1992) The spinach chloroplast chromosome is bound to the thylakoid membrane in the region of the inverted repeat. Biochem Biophys Res Commun 184: 993–1000 [DOI] [PubMed] [Google Scholar]
- Majeran W, Friso G, Asakura Y, Qu X, Huang M, Ponnala L, Watkins KP, Barkan A, van Wijk KJ. (2012) Nucleoid-enriched proteomes in developing plastids and chloroplasts from maize leaves: a new conceptual framework for nucleoid functions. Plant Physiol 158: 156–189 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manuell AL, Quispe J, Mayfield SP. (2007) Structure of the chloroplast ribosome: novel domains for translation regulation. PLoS Biol 5: e209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Merchant SS, Prochnik SE, Vallon O, Harris EH, Karpowicz SJ, Witman GB, Terry A, Salamov A, Fritz-Laylin LK, Maréchal-Drouard L, et al. (2007) The Chlamydomonas genome reveals the evolution of key animal and plant functions. Science 318: 245–250 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meurer J, Lezhneva L, Amann K, Gödel M, Bezhani S, Sherameti I, Oelmüller R. (2002) A peptide chain release factor 2 affects the stability of UGA-containing transcripts in Arabidopsis chloroplasts. Plant Cell 14: 3255–3269 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Motohashi R, Yamazaki T, Myouga F, Ito T, Ito K, Satou M, Kobayashi M, Nagata N, Yoshida S, Nagashima A, et al. (2007) Chloroplast ribosome release factor 1 (AtcpRF1) is essential for chloroplast development. Plant Mol Biol 64: 481–497 [DOI] [PubMed] [Google Scholar]
- Murashige T, Skoog F. (1962) A revised medium for rapid growth and bioassays with tobacco tissue culture. Physiol Plant 15: 473–497 [Google Scholar]
- Nilsson R, Brunner J, Hoffman NE, van Wijk KJ. (1999) Interactions of ribosome nascent chain complexes of the chloroplast-encoded D1 thylakoid membrane protein with cpSRP54. EMBO J 18: 733–742 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olinares PD, Ponnala L, van Wijk KJ. (2010) Megadalton complexes in the chloroplast stroma of Arabidopsis thaliana characterized by size exclusion chromatography, mass spectrometry, and hierarchical clustering. Mol Cell Proteomics 9: 1594–1615 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Park YJ, Cho HK, Jung HJ, Ahn CS, Kang H, Pai HS. (2011) PRBP plays a role in plastid ribosomal RNA maturation and chloroplast biogenesis in Nicotiana benthamiana. Planta 233: 1073–1085 [DOI] [PubMed] [Google Scholar]
- Peled-Zehavi H, Danon A. (2007) Translation and translational regulation in chloroplasts. Top Curr Genet 19: 249–281 [Google Scholar]
- Rivier C, Goldschmidt-Clermont M, Rochaix JD. (2001) Identification of an RNA-protein complex involved in chloroplast group II intron trans-splicing in Chlamydomonas reinhardtii. EMBO J 20: 1765–1773 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rodermel SR, Abbott MS, Bogorad L. (1988) Nuclear-organelle interactions: nuclear antisense gene inhibits ribulose bisphosphate carboxylase enzyme levels in transformed tobacco plants. Cell 55: 673–681 [DOI] [PubMed] [Google Scholar]
- Rogalski M, Ruf S, Bock R. (2006) Tobacco plastid ribosomal protein S18 is essential for cell survival. Nucleic Acids Res 34: 4537–4545 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rogalski M, Schöttler MA, Thiele W, Schulze WX, Bock R. (2008) Rpl33, a nonessential plastid-encoded ribosomal protein in tobacco, is required under cold stress conditions. Plant Cell 20: 2221–2237 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roy A, Kucukural A, Zhang Y. (2010) I-TASSER: a unified platform for automated protein structure and function prediction. Nat Protoc 5: 725–738 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruf M, Kössel H. (1988) Occurrence and spacing of ribosome recognition sites in mRNAs of chloroplasts from higher plants. FEBS Lett 240: 41–44 [Google Scholar]
- Scharff LB, Childs L, Walther D, Bock R. (2011) Local absence of secondary structure permits translation of mRNAs that lack ribosome-binding sites. PLoS Genet 7: e1002155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Selman-Reimer S, Merchant S, Selman BR. (1981) Isolation, purification, and characterization of coupling factor 1 from Chlamydomonas reinhardtii. Biochemistry 20: 5476–5482 [DOI] [PubMed] [Google Scholar]
- Sharma MR, Wilson DN, Datta PP, Barat C, Schluenzen F, Fucini P, Agrawal RK. (2007) Cryo-EM study of the spinach chloroplast ribosome reveals the structural and functional roles of plastid-specific ribosomal proteins. Proc Natl Acad Sci USA 104: 19315–19320 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shen Y, Li C, McCarty DR, Meeley R, Tan BC. (2013) Embryo defective12 encodes the plastid initiation factor 3 and is essential for embryogenesis in maize. Plant J 74: 792–804 [DOI] [PubMed] [Google Scholar]
- Spetea C, Hundal T, Lundin B, Heddad M, Adamska I, Andersson B. (2004) Multiple evidence for nucleotide metabolism in the chloroplast thylakoid lumen. Proc Natl Acad Sci USA 101: 1409–1414 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stern DB, Goldschmidt-Clermont M, Hanson MR. (2010) Chloroplast RNA metabolism. Annu Rev Plant Biol 61: 125–155 [DOI] [PubMed] [Google Scholar]
- Stoppel R, Manavski N, Schein A, Schuster G, Teubner M, Schmitz-Linneweber C, Meurer J. (2012) RHON1 is a novel ribonucleic acid-binding protein that supports RNase E function in the Arabidopsis chloroplast. Nucleic Acids Res 40: 8593–8606 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stoppel R, Meurer J. (2012) The cutting crew: ribonucleases are key players in the control of plastid gene expression. J Exp Bot 63: 1663–1673 [DOI] [PubMed] [Google Scholar]
- Sugiura M, Hirose T, Sugita M. (1998) Evolution and mechanism of translation in chloroplasts. Annu Rev Genet 32: 437–459 [DOI] [PubMed] [Google Scholar]
- Talkington MWT, Siuzdak G, Williamson JR. (2005) An assembly landscape for the 30S ribosomal subunit. Nature 438: 628–632 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tiller N, Weingartner M, Thiele W, Maximova E, Schöttler MA, Bock R. (2012) The plastid-specific ribosomal proteins of Arabidopsis thaliana can be divided into non-essential proteins and genuine ribosomal proteins. Plant J 69: 302–316 [DOI] [PubMed] [Google Scholar]
- Timmis JN, Ayliffe MA, Huang CY, Martin W. (2004) Endosymbiotic gene transfer: organelle genomes forge eukaryotic chromosomes. Nat Rev Genet 5: 123–135 [DOI] [PubMed] [Google Scholar]
- Ueda M, Nishikawa T, Fujimoto M, Takanashi H, Arimura Si, Tsutsumi N, Kadowaki K-i. (2008) Substitution of the gene for chloroplast RPS16 was assisted by generation of a dual targeting signal. Mol Biol Evol 25: 1566–1575 [DOI] [PubMed] [Google Scholar]
- Voelker R, Barkan A. (1995) Nuclear genes required for post-translational steps in the biogenesis of the chloroplast cytochrome b6f complex in maize. Mol Gen Genet 249: 507–514 [DOI] [PubMed] [Google Scholar]
- Walter M, Piepenburg K, Schöttler MA, Petersen K, Kahlau S, Tiller N, Drechsel O, Weingartner M, Kudla J, Bock R. (2010) Knockout of the plastid RNase E leads to defective RNA processing and chloroplast ribosome deficiency. Plant J 64: 851–863 [DOI] [PubMed] [Google Scholar]
- Weglöhner W, Jünemann R, von Knoblauch K, Subramanian AR. (1997) Different consequences of incorporating chloroplast ribosomal proteins L12 and S18 into the bacterial ribosomes of Escherichia coli. Eur J Biochem 249: 383–392 [DOI] [PubMed] [Google Scholar]
- Westhoff P, Herrmann RG. (1988) Complex RNA maturation in chloroplasts: the psbB operon from spinach. Eur J Biochem 171: 551–564 [DOI] [PubMed] [Google Scholar]
- Woodson SA. (2008) RNA folding and ribosome assembly. Curr Opin Chem Biol 12: 667–673 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xia B, Ke H, Shinde U, Inouye M. (2003) The role of RbfA in 16S rRNA processing and cell growth at low temperature in Escherichia coli. J Mol Biol 332: 575–584 [DOI] [PubMed] [Google Scholar]
- Yamaguchi K, Subramanian AR. (2000) The plastid ribosomal proteins: identification of all the proteins in the 50 S subunit of an organelle ribosome (chloroplast). J Biol Chem 275: 28466–28482 [DOI] [PubMed] [Google Scholar]
- Yamaguchi K, von Knoblauch K, Subramanian AR. (2000) The plastid ribosomal proteins: identification of all the proteins in the 30 S subunit of an organelle ribosome (chloroplast). J Biol Chem 275: 28455–28465 [DOI] [PubMed] [Google Scholar]
- Yu F, Liu X, Alsheikh M, Park S, Rodermel S. (2008) Mutations in SUPPRESSOR OF VARIEGATION1, a factor required for normal chloroplast translation, suppress var2-mediated leaf variegation in Arabidopsis. Plant Cell 20: 1786–1804 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou F, Karcher D, Bock R. (2007) Identification of a plastid intercistronic expression element (IEE) facilitating the expression of stable translatable monocistronic mRNAs from operons. Plant J 52: 961–972 [DOI] [PMC free article] [PubMed] [Google Scholar]