Abstract
The adoption of oligonucleotide aptamer is well on the rise, serving an ever increasing demand for versatility in biomedical field. Through the SELEX (Systematic Evolution of Ligands by EXponential enrichment), aptamer that can bind to specific target with high affinity and specificity can be obtained. Aptamers are single-stranded nucleic acid molecules that can fold into complex threedimensional structures, forming binding pockets and clefts for the specific recognition and tight binding of any given molecular target. Recently, aptamers have attracted much attention because they not only have all of the advantages of antibodies, but also have unique merits such as thermal stability, ease of synthesis, reversibility, and little immunogenicity. The advent of novel technologies is revolutionizing aptamer applications. Aptamers can be easily modified by various chemical reactions to introduce functional groups and/or nucleotide extensions. They can also be conjugated to therapeutic molecules such as drugs, drug containing carriers, toxins, or photosensitizers. Here, we discuss new SELEX strategies and stabilization methods as well as applications in drug delivery and molecular imaging.
Keywords: Aptamer, Stabilization, Modification, SELEX, Imaging, Drug delivery system
INTRODUCTION
For diagnosis, therapy, or imaging of certain disease, specific targeting is an important issue. To this end, antibodies have been widely used to target molecules of interest and made a tremendous contribution to a wide range of applications based on molecular recognition (Jayasena, 1999). The IgG2a CD3-specific transplant rejection drug OKT3 (muromonab) was the first FDA-approved therapeutic antibody (Hooks et al., 1991). Subsequently, diverse therapeutic antibodies have been applied to treat diseases (Knight et al., 1993; Mazumdar and Greenwald, 2009; Pageau, 2009; Banaszynski and Kolesar, 2013). However, many limitations have been identified, including difficulties with production, immunogenicity, expensiveness, reoptimization with each new batch, sensitivity to temperature, and irreversible denaturation. Recently, chemical antibody called aptamer has emerged as a new targeting molecule.
Aptamers consist of nucleic acids that bind to diverse targets with high affinity and selectivity. They are screened through an in vitro process and usually have higher binding affinity than traditional antibody. Aptamers are produced chemically, and no or little batch-to-batch variation is observed during aptamer production. Furthermore, aptamers can be easily modified to chemically conjugate with other molecules. Aptamer can also undergo reversible denaturation at high temperature, making it a very versatile tool for drug loading and antidote application. Moreover, aptamers elicit little or no immunogenicity in therapeutic applications (Eyetech Study Group, 2002; Foy et al., 2007; Zhu et al., 2012a). The first aptamer-based drug called Macugen (pegaptanib) was FDA-approved in 2004 (Ruckman et al., 1998), and many studies proceeded to test efficacy.
Here, the up-to-date methods for SELEX processes are introduced. Moreover, diverse strategies to increase aptamer stability in biological fluid are discussed. Also, its utility as a targeting molecule for imaging, diagnosis, therapy and targeted drug delivery are discussed.
WHAT IS AN APTAMER?
Aptamer is a single-stranded nucleic acid oligomer made of RNA or DNA which can bind to specific target molecule with high affinity and selectivity. Secondary or tertiary structures of aptamers allow target binding and determine target specificity and affinity (Ellington and Szostak, 1990; Tuerk and Gold, 1990; Schneider et al., 1995; Mosing et al., 2005; Chen et al., 2009). In the early 1990s, several research laboratories were independently experimenting with what Jack W. Szostak’s group termed “in vitro selection” (Ellington and Szostak, 1990). To describe molecular recognition properties for what were nucleic acid-based ligands, they coined the term ‘aptamer’ using the Latin word “aptus”, meaning “fitting” and the Greek word ‘‘meros’’, meaning “particle”. But naming aptamers was not nearly as interesting as discovering that their properties compete quite well with those of antibodies. Targets of aptamer may include, but are not limited to, metal ions (Kawakami et al., 2000), small organic compounds (Mann et al., 2005), biological cofactors (Lauhon and Szostak, 1995; Holland et al., 2000), metabolites (Bruno et al., 2008), proteins (Ruckman et al., 1998; White et al., 2001; Savla et al., 2011) and whole organisms such as a virus (Tang et al., 2009), bacteria (Hamula et al., 2011), yeast (Kolesnikova et al., 2010), or mammalian cell (Chen et al., 2009). Aptamers can be easily modified by various chemical reactions to introduce functional groups and/or nucleotide extensions. Aptamers can be easily conjugated to therapeutic molecules such as drugs, drug containing carriers, toxins, or photosensitizers. Thus, aptamers are promising escort molecules for drug delivery systems.
NEW METHODS FOR IN VITRO SELECTION OF APTAMERS
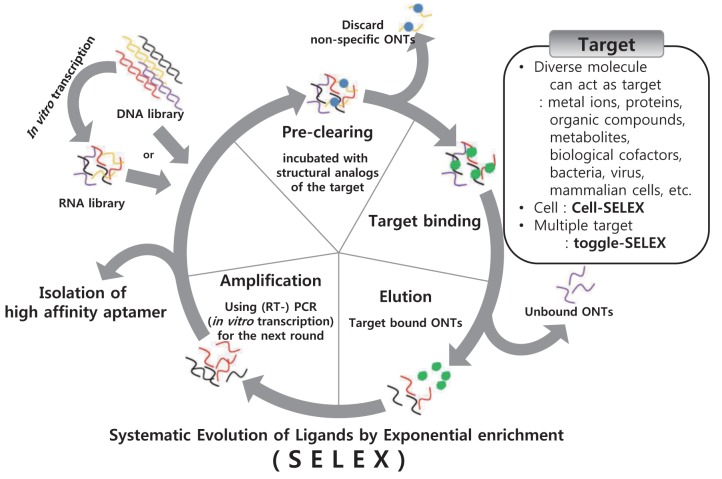
SELEX is an in vitro selection method designed to identify aptamers that are selectively bound to target molecules with high affinity. Substantive studies on aptamers have progressed since the in vitro selection process called SELEX was first reported by Gold’s and Szostak’s groups (Ellington and Szostak, 1990; Tuerk and Gold, 1990). First, the nucleic acid library, which consists of 1014-1015 random oligonucleotide strands, is incubated with a target molecule. Then, the target-bound oligonucleotide strands are separated from the unbound strands. The target-bound DNA or RNA strands are eluted from the target molecule and amplified via polymerase chain reaction to seed a new pool of nucleic acids. This selection process is continued for 6-15 rounds with increasingly stringent conditions, which ensure that the nucleic acid obtained has the highest affinity to the target molecule (Fig. 1). SELEX method can be modified in a variety of ways to increase the specificity of aptamer and efficiency of SELEX.
Fig. 1. Overview of SELEX scheme. Aptamers can be obtained through an iterative selection process known as SELEX (systematic evolution of ligands by exponential enrichment) by using single-stranded DNA or RNA. An initial pool of 1014-1015 random oligonucleotide (ONT) strands are subjected to binding with the target. Unbound ONTs are discarded and RT-PCR or PCR is performed to amplify the targetbound ONTs. This selection process is repeated 6-15 times using amplified ONTs as a new pool. This way, aptamers having high specificity and affinity are screened. Diverse molecules can be the target of the SELEX, including metal ion, protein, organic compound and cell. Toggle-SELEX performs SELEX with two different target molecules to obtain bispecific aptamers.
Counter-SELEX
The counter-SELEX method was introduced to increase the efficiency of aptamer selection by traditional SELEX (Fig. 1) (Jenison et al., 1994). Compared to traditional SELEX, counter-SELEX has a pre-clearing step using closely related structural analogs of the target to effectively discard non-specific aptamers. This allows dramatic improvement in aptamer selection and can also be applied to other modified SELEX methods.
Cell-SELEX
The SELEX target is not limited to an individual molecule. The cell-SELEX strategy has been described that utilizes living cells as the SELEX target (Fig. 1). Although traditional SELEX is typically carried out using purified target molecules, whole live cells are also employed as selection targets. Not only normal/abnormal mammalian cells such as virus-infected and cancer cells but also live pathogenic organisms such as bacteria and viruses have been utilized as cell-SELEX targets (Tang et al., 2009; Hamula et al., 2011). The aptamers generated are functional with an original conformation of the target molecule on live cells. Compared to aptamers selected using a purified target, a cell-SELEX-derived aptamer has more possibility to be used directly for in vivo and clinical applications. A screened aptamer resulting from cell-SELEX using abnormal cells can be used to detect disease or cancer. Moreover, biomarkers can be used to identify the aptamer target for a specific abnormality (Blank et al., 2001). Many strategies are available to select an anti-virus aptamer. First, SELEX can be performed using the virion directly (Wang et al., 2000). Second, aptamers can be generated that specifically bind to virus-infected cells. Through virus infection, viral proteins exist on the host cell surface due to the viral gene or enveloped protein. SELEX provides screening of virus-specific aptamers that recognize virus infected cells. Furthermore, aptamers can be selected for host cell proteins if their expression levels are upregulated upon viral infection. Lastly, stable cell lines can be generated that express viral protein as a cell-SELEX target, and the aptamer obtained can be used to detect virus infected cells (Chen et al., 2009). Furthermore, novel biomarkers can be discovered using the cell-SELEX technique. Because the target molecules of the aptamers generated by cell-SELEX may be previously unrecognized as cell-specific surface molecules, they could be novel biomarkers. Thus, cell-SELEX can be applied for de novo discovery of novel biomarkers for a desired cell by identifying the aptamer binding partner. The cell- SELEX concept can be extended for in vivo selection, which was first designed using a hepatic tumor xenograft mouse model (Mi et al., 2010). Oligonucleotides were injected intravenously, liver tumors were harvested, and the injected RNA molecules were extracted and amplified. A tissue-specific aptamer can be more easily screened through this in vivo selection process. So, a screened aptamer may be a useful target for a tissue of interest without non-specific biodistribution in the in vivo application.
Capillary Electrophoresis-SELEX
The SELEX process has disadvantages in that it is time consuming to repeat the rounds. Some molecular biological methods have been introduced to SELEX to overcome these disadvantages. Capillary electrophoresis-SELEX (CE-SELEX) was designed for selecting aptamers to reduce repeating rounds with low dissociation constants (Mosing et al., 2005). In this method, the nucleic acids that bind the target migrate with different mobilities than those of unbound sequences, allowing them to be collected as separate fractions. Although traditional SELEX requires 6-12 rounds, CE-SELEX significantly reduces the number of rounds and increases binding affinity (Schneider et al., 1995). CE-SELEX decreases the time, effort, and cost to screen a higher affinity aptamer compared to those of traditional SELEX. Based on CE technology, a non- SELEX method that selects an aptamer without amplification was demonstrated (Berezovski et al., 2006). Non-equilibrium capillary electrophoresis of equilibrium mixtures (NECEEM), a highly efficient affinity method, has been used to partition the DNA-target complex from the free DNA. An aptamer with nanomolar affinity for the protein farnesyltransferase was selected in a single round of partitioning using NECEEM (Berezovski et al., 2005). The advantage of non-SELEX is its speed and simplicity. NECEEM-based non-SELEX selection took only 1 h in contrast to several days or several weeks required for a traditional SELEX procedure by conventional partitioning methods. This procedure can be automated using a single commercially available capillary electrophoresis instrument.
One step MonoLEX
The MonoLEX approach combines a single affinity chromatography step with subsequent physical segmentation of the affinity resin and one single final exponential amplification step of the bound aptamers (Nitsche et al., 2007). Using the Biomek 2000, an automatic SELEX device (Cox et al., 1998), Cox et al. demonstrated that 10 rounds of selection against eight targets in parallel can be performed in 4-5 days.
Microfluidic SELEX
Microfluidic SELEX (mSELEX) was designed that combines traditional SELEX with a microfluidic system. Protein immobilization is the most important aspect of mSELEX. A mSELEX prototype device was developed and biotinylated lysozymes were immobilized to silica microline previously coated with streptavidin (Hybarger et al., 2006). In another study, a magnetic bead-based SELEX process with microfluidics technology and a continuous-flow magnetic activated chipbased separation device was developed (Lou et al., 2009). Using this mSELEX method, an enriched aptamer pool was obtained that tightly bound to the light chain of recombinant Botulinum neurotoxin type A after a single round of selection with a 33 nM Kd value. Recently, a novel formulation of solgel protein microarray material was developed, which elicited physical properties suitable for protein immobilization, proteinprotein interactions, and immunoassays (Kim et al., 2006). Based on sol-gel, the authors developed a microfluidic device and method for binding nucleic acids to immobilized proteins and efficiently eluting and recovering the intact nucleic acids (Park et al., 2009). The mSELEX system enhanced selection efficiency and reduced time and effort compared to those of traditional SELEX.
Toggle-SELEX
Furthermore, toggle-SELEX has been established for in vitro animal model assays using a single aptamer (Fig. 1). The “toggle” selection process was repeated during SELEX rounds using a target applied to human thrombin for even rounds and porcine thrombin for odd rounds to select a species cross-reactive aptamer (White et al., 2001). A species cross-reactive aptamer from the toggle-SELEX strategy may contribute directly to animal models and clinical applications. Thus, toggle-SELEX makes the aptamer technique closer to a “bench-to-clinic” concept.
Based on these novel methods, high-throughput screening of aptamers and improved efficiency of the SELEX process is possible. Moreover, consilience technology such as microfluidic technology and engineering could reduce effort, time, and cost of aptamer selection and analysis, while increasing its efficacy. Taken together, aptamers may play a key role in future theragnosis because of higher binding affinity, lower cost, and ease of synthesis compared to antibodies.
STABILIZATION OF THE OLIGONUCLEOTIDE APTAMER
One of the ultimate goals during aptamer selection is a clinical application or an escort molecule. However, unmodified nucleic acids per se are unstable in biological fluids due to enzymatic degradation or a short half-life. Various strategies have been established to increase serum stability and overcome the degradation of oligonucleotides by nuclease (Fig. 2, 3).
Fig. 2. Method for obtaining Spiegelmer. SELEX is performed with mirror-imaged target and D-form RNA. Based on the selected D-form RNA sequence, L-form RNA aptamer can be synthesized. Now L-RNA, the chiral form of the acquired D-form RNA, can bind to natural target. This L-form RNA is nuclease-resistant and suitable for in vivo application. The world ‘Spiegelmer’ is derived from German ‘Spiegel’ meaning ‘mirror’.
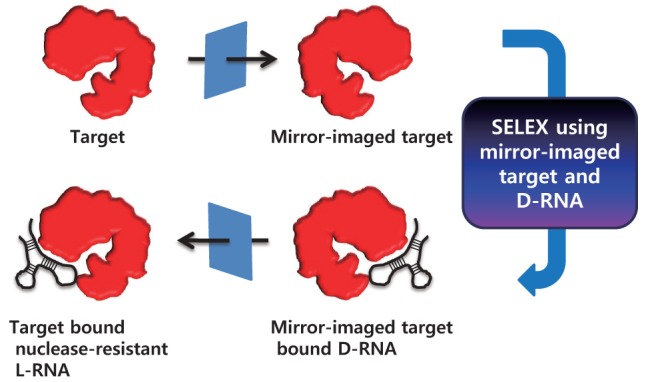
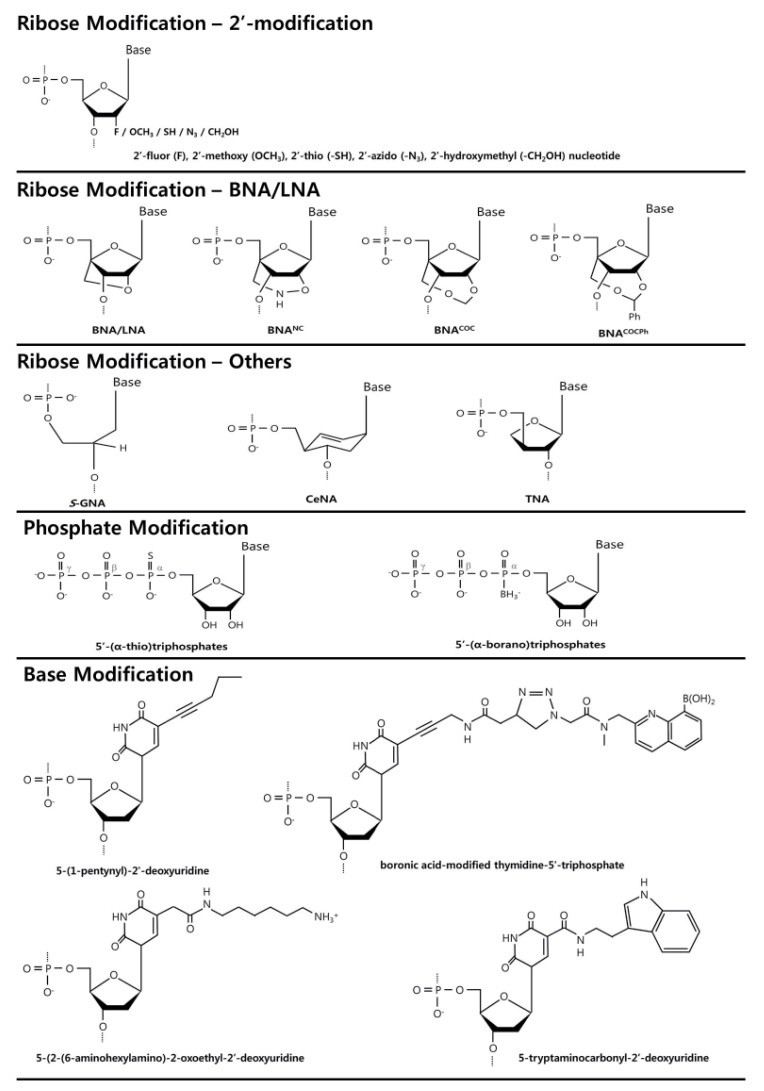
Fig. 3. Structure of chemically modified nucleotides (Modified form Kasahara and Kuwahara, 2012). The simplest ribose 2’ modification is widely used to increase aptamer stability in vivo. Phosphate and base modification are also used for this purpose. Substitution of F, OCH3, SH or CH2OH for 2’-OH (H) is widely used. BNA/LNA was designed to structurally protect 2’ site. In addition, thiol (S) or borane (BH3) group is introduced to α phosphate to strengthen oligonucleotide backbone. Functional groups can also be introduced into the base.
For example, the first FDA-approved aptamer drug Macugen, which specifically binds to VEGF165, is a modified oligonucleotide prepared by SELEX processes (Ruckman et al., 1998). The RNA aptamer was screened from a library of modified RNAs involving 2'-fluoropyrimidine nucleotides (U, C) and natural purine nucleotides (A, G) in which the 2′-position of ribose is a nuclease attack site. Then, the natural A and G are properly replaced with 2'-methoxy nucleotides to enhance nuclease resistance and avoid a decrease in affinity. Like Macugen, 2’-fluor or methoxy or amine substituted nucleotides was widely used to protect from nuclease (Zhou et al., 2008, 2011a, 2011b, 2013) (Fig. 3). Additionally, 2’-thio (-SH), 2’-azido (-N3), 2’-hydroxymethyl (-CH2OH) modified nucleotides was developed for protection of oligonucleotide. Moreover, bridged/locked, S-glycerol, cyclohexenyl, and threose nucleic acid (BNA/LNA, S-GNA, CeNA, and TNA, respectively) were applied to aptamer building block (Horhota et al., 2005; Ichida et al., 2005; Kempeneers et al., 2005; Tsai et al., 2007; Kuwahara et al., 2008; Barciszewski et al., 2009; Kasahara et al., 2013) (Fig. 3). For example, Schmidt group reported that LNA modification increased plasma stability more than 2’-OMe modification using TTA1 aptamer which bind to Tenascin-C (Schmidt et al., 2004). Furthermore, for the substitute phosphodiester bond in backbone, 5’-(α-thio)triphosphates and 5’-(α-borano) triphosphates were developed (Andreola et al., 2000; Lato et al., 2002) , with increased stability in the biological milieu (Yang et al., 1999). Moreover, peptide nucleic acid (PNA) whose sugar-phosphate backbone is changed to repeating N-(2-aminoethyl)-glycine units linked by peptide bonds, increases biostability (Wittung et al., 1994). In other aspect, base modification could increase aptamer stability in novel thrombin aptamer containing 5-(1-pentynyl)-2'-deoxyuridine (Latham et al., 1994). In the thalidomide aptamer, a modified deoxyuridine bearing a cationic functional group via a hydrophobic methylene linker at the C5 position (Shoji et al., 2007) could improve stability against nucleases and the binding affinity to target. Fibrinogen aptamer using boronic acidmodified thymidine-5'-triphosphate (Li et al., 2008) , TNFRSF9 aptamer using 5-tryptaminocarbonyl-dU (Vaught et al., 2010) were also reported (Fig. 3). Certain thermophilic and mutant polymerases such as Pwo, Vent (exo-), Deep Vent (exo-), and KOD Dash have been actively developed for incorporation of modified nucleotides during polymerization (Gudima et al., 1998; Kuwahara et al., 2003; Leal et al., 2006; Kuwahara et al., 2009; Veedu et al., 2009; Kuwahara et al., 2010).
A chiral form of an aptamer is an alternative strategy to prevent enzymatic degradation in biological fluids by altering nuclease recognition. NOXXON Pharma established Spiegelmer (from German Spiegel “mirror”, Fig. 2) to screen nucleaseresistant aptamers, which are RNA-like molecules built from L-ribose units. This Spiegelmer technology circumvents nuclease activity in biological fluids. In short, a chiral form of the target molecule and D-form oligonucleotides are used during the SELEX process. Subsequently, an L-form of the aptamer is synthesized that has the same sequence as the selected Dform aptamer. Thus, an enzymatic degradation-insensitive aptamer, which also binds to the original target molecule, can be obtained (Fig. 2) (Klussmann et al., 1996; Nolte et al., 1996; Williams et al., 1997; Leva et al., 2002).
Several authors have inhibited in vivo clearance to increase half-life of aptamers. An anti-platelet derived growth factor aptamer was conjugated with 40 kDa polyethylene glycol (PEG) to slow down renal clearance (Floege et al., 1999). In another study, Healy et al. reported an aptamer conjugated with 20 and 40 kDa PEG to prolong aptamer half-life in circulation (Healy et al., 2004). Furthermore, the DNA aptamer ARCC1779, which binds to the A1 domain of the von Willebrand factor, was also 5′-conjugated with a 20 kDa PEG to reduce renal filtration (Diener et al., 2009). Many studies are underway to increase the biostability and half-life of aptamers in different settings. However, post-SELEX modification can influence target binding affinity or secondary and tertiary structure formation, which, in turn, can alter aptamer function.
APTAMERS AS AGONISTS/ANTAGONISTS
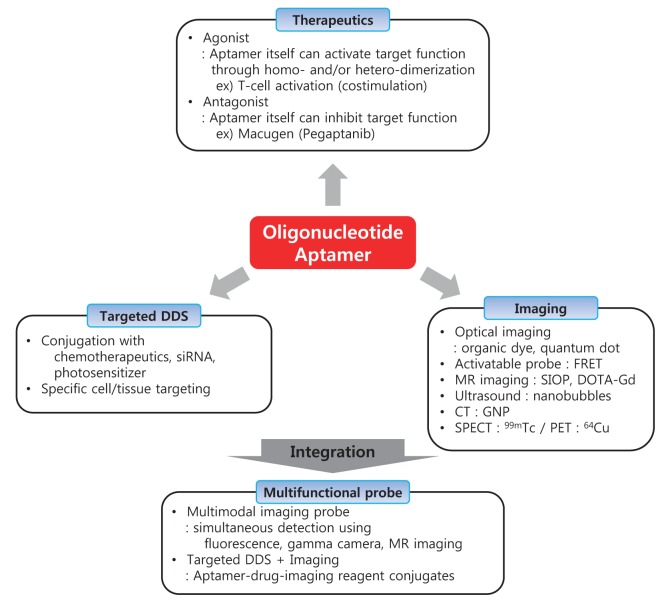
Individual aptamers can be equipped in a modular manner with additional functions and can be specifically tailored for many potential applications in biotechnology and molecular medicine (Fig. 4). In most cases, aptamers not only bind their cognate protein but also efficiently inhibit the function of target as an antagonist. Many aptamer therapeutic studies have been conducted to treat certain diseases by inhibiting therapeutic target activity and decreasing the partner binding property (Holahan et al., 2011; Gissel et al., 2012; Siller-Matula et al., 2012; Vater et al., 2013). Macugen, the first FDAapproved aptamer drug for age-related macular degeneration, is a typical antagonistic aptamer (Zhou and Wang, 2006). Macugen (pegaptanib) is a 28-base RNA oligonucleotide with two branched 20 kDa PEG moieties (Ruckman et al., 1998). It selectively binds to the vascular endothelial growth factor (VEGF)165 isoform when introduced intravitreally. Thus, the intracellular signal cascade is inhibited by the pegaptanib- VEGF165 complex, which cannot bind to the VEGF receptor. Thus, pegaptanib has the potential to inhibit growth of blood vessels and vascular leakage by inhibiting the intracellular signal cascade induced by the pegaptanib-VEGF165 complex. Diverse therapeutics based on antagonistic functions of aptamers are undergoing clinical trials (Table 1). They inhibit the signaling cascade by decreasing activity of the target protein or altering binding with the physiological binding partner.
Fig. 4. Applications of aptamer. Aptamer can be used as therapeutics, targeted drug delivery system (DDS) and imaging. These three parts can be put together using aptamer and conjugation in multifunctional probe.
Table 1.
Undergoing aptamer clinical trials.
| Name | Target | Condition | Phase |
|---|---|---|---|
|
| |||
| Pegaptanib sodium (Macugen) | VEGF | Age-Related Macular Degeneration | Approved |
| EYE001 | VEGF | Macular Degeneration / Choroidal Neovascularization | Phase 2/3 |
| E10030 | PDGF | Age-Related Macular Degeneration | Phase 2 |
| Pegaptanib sodium (Macugen) | VEGF | Diabetic Macular Edema | Phase 2 |
| AS1411 | Nucleolin | Acute myeloid leukemia | Phase 2 |
| ARC1779 | A1 domain of von Willebrand Factor | von Willebrand Disease / Purpura / Thrombotic Thrombocytopenic | Phase 2 |
| REG1 (RB006 and RB007) | Coagulation factor IXa | Coronary Artery Disease | Phase 2 |
| NOX-E36 | CCL2 | Chronic Inflammatory Diseases / Type 2 Diabetes Mellitus / Systemic Lupus Erythematous | Phase 1 |
| NOX-A12 | CXCL12 | Autologous or Hematopoietic Stem Cell Transplantation | Phase 1 |
| ARC19499 | tissue factor pathway inhibitor (TFPI) | Hemophilia | Phase 1 |
| ARC1905 | Complement Component 5 (C5) | Age-Related Macular Degeneration | Phase 1 |
| EYE001 | VEGF | von Hippel-Lindau Disease | Phase 1 |
The first report about aptamers as agonists came from the study that reported agonistic use of an aptamer with 4-1BB, which is a major costimulatory receptor (McNamara et al., 2008). 4-1BB increased expansion and survival of activated T cells. Dimerization of the anti-4-1BB aptamer treated with the CD3 antibody not only increased T-cell activation and interferon-γ in vitro but also tumor regression in vivo. Moreover, Dollins et al. designed an aptamer-based agonist by dimerizing the OX4 RNA aptamer, which targets one of the tumor necrosis factor receptor families (Dollins et al., 2008). Two 2′-fluoropyrimidine-modified RNA aptamers were selected that bind to CD28 (CD28Apt2 and CD28Apt7) (Pastor et al., 2013). That study suggested that an agonistic dimerized anti- CD28 aptamer increases costimulatory efficacy and reduces therapeutic dose and toxicity compared to conventional agonistic anti-CD28 antibodies. Similar research on aptamers for use as costimulatory agonistic molecules is being conducted (Suntharalingam et al., 2006; Römer et al., 2011). Moreover, not only homodimerization of an aptamer for agonistic use but also heterodimerization of two individual aptamers that recognize different targets for dual purposes, specific targeting and stimulating, have been reported (Gilboa et al., 2013). Two individual aptamers were heterodimerized including a 4-1BB binding aptamer to stimulate the immune response and a prostate-specific membrane antigen (PSMA) binding aptamer for prostate cancer targeting. This bispecific aptamer was developed to regress prostate tumors through the host immune response. A bispecific aptamer is more effective than and synergizes with vaccination and exhibits a superior therapeutic index compared with nontargeted costimulation with 4-1BB antibodies or 4-1BB aptamers. Agonistic/antagonistic aptamers offer significant advantages over antibodies in terms of synthesis, cost, and immunogenicity as well as conjugation chemistry.
TARGETED DRUG DELIVERY
Targeted drug delivery is one of the most important areas in therapeutic research. Aptamers exhibit many desirable properties for targeted drug delivery such as ease of selection and synthesis, high binding affinity and specificity, low immunogenicity, and versatile synthetic accessibility. The therapeutic agents that have been delivered using aptamers as the targeting ligands can be categorized into three major classes of drugs, toxins, and small interfering RNAs (siRNAs) (Fig. 4) (Zhang et al., 2011).
Doxorubicin (Dox) used in chemotherapy for hematological malignancies, carcinomas, and soft tissue sarcomas can intercalate within the double-stranded CG sequences of DNA and RNA. Based on the intercalation property of Dox to the aptamer, targeted Dox delivery for targeting cancer and Doxmediated cancer therapy has been actively studied (Liu et al., 2012; Meng et al., 2012; Subramanian et al., 2012). A Dox intercalated aptamer that binds to PSMA, which is abundantly expressed on the cell surface of a human prostatic adenocarcinoma (LNCaP) metastatic lesion has been studied. PSMA expressing LNCaP-specific cell death was observed by treatment of Dox-intercalated PSMA-specific 2′-fluoropyrimidine aptamer, but not in PSMA-negative PC3 cells (Bagalkot et al., 2006). Covalent conjugation of aptamer harboring the drug is relatively stable such that drug release during transport is prevented before the drug localizes at its cellular target. Drugs can be chemically modified to form stable ester, amide, and disulfide bonds that serve to conjugate the drug to the aptamer. The sgc8c DNA aptamer which specifically binds to protein tyrosine kinase 7 (PTK7) was covalently conjugated with Dox using hydrazone (Huang et al., 2009). As a result, PTK7 expressing CCRF-CEM cell-specific Dox delivery and cell death were observed. Other chemotherapy drugs such as daunorubicin, thalidomide, and dactinoimycin can also intercalate to double-stranded oligonucleotides. They are expected to show a good therapeutic effect in specific cancers, and their application in various aptamers is actively sought.
The small size and lipid bilayer structure of liposomes make them an excellent drug delivery carrier. Aptamer-tagged liposomes were established that contained the anticancer drug paclitaxel and a fluorescent dye (Nile Red) for tumor-specific drug delivery and imaging (Aravind et al., 2012). In another study, Cy3-labeled, carboxylated, thiolated oligonucleotide aptamer (thioaptamer) against E-selectin (ESTA, E-Selectin ThioAptamer)was designed. This modified ESTA was coupled to an amino PEGylated stealth liposome to make the vasculature targeting ESTA-conjugated liposome (Mann et al., 2011), which achieved tumor vasculature-specific drug delivery. Similarly, this aptamer can be covalently coupled via NHS chemistry to toxic proteins. The targeting properties of the anti-PSMA aptamer can also be combined with the toxic characteristics of gelonin, a ribosomal toxin (Chu et al., 2006a). Moreover, a photosensitizer-aptamer conjugate was developed for cancerspecific delivery to be used in photodynamic cancer therapy (Yang et al., 2011; Han et al., 2013).
siRNA delivery to specific cells using an aptamer has been actively investigated. In these studies, receptor-mediated endocytosis of the aptamer is necessary. In general, non-covalent conjugation through a connector and covalent linkage through formation of an aptamer-siRNA chimera are two major strategies for aptamer and siRNA conjugation. In 2006, two biotinylated anti-PSMA aptamers were linked to two biotinylated anti-lamin A/C siRNAs using streptavidin as the connector (Chu et al., 2006b). As a result, PSMA-positive prostate cancer cell-specific siRNA was internalized and siRNA-specific gene expression was inhibited in vitro. However, the siRNA release mechanism and an in vivo experiment have not been demonstrated. The “sticky bridge” concept was used to noncovalently conjugate the anti-gp120 aptamer with siRNAs to suppress HIV replication in vitro (Horhota et al., 2005). An aptamer-siRNA chimera was developed using an aptamer consisting of single-stranded oligonucleotides. An aptamer was covalently linked to the passenger strands of siRNAs followed by annealing of the siRNA guide strands to the passenger strands to create a functional siRNA duplex (McNamara et al., 2006). That study found that aptamer-siRNA chimera-mediated gene silencing was dependent on Dicer and occurred via the RNAi pathway. Moreover, anti-PSMA aptamer-Plk1 did not trigger interferon responses in vitro, and promoted tumor regression in a xenograft model of prostate cancer. Structural modifications were introduced that increased the valency of the chimera aptamer to enhance therapeutic efficacy of the aptamer-siRNA targeting PSMA-expressing system (Wullner et al., 2008). In their design, the siRNA sequence served as a linker to join the two aptamers, or to join the 3′ end of one of the aptamer dimers as a shRNA. Such modifications did not affect the critical sequence of the targeting regions and facilitated formation of the active conformation. Targeted siRNA delivery using an aptamer-siRNA chimera is being actively studied (Neff et al., 2011; Zhu et al., 2012b; Hussain et al., 2013).
APTAMER-BASED IMAGING
Another application of aptamers is diagnosis via in vivo and in vitro imaging, using an aptamer that is conjugated to a fluorophore, quantum dots, or other material such as gadolinium, which is useful for magnetic resonance imaging (MRI) (Fig. 4). Optical imaging is a cost- effective imaging method that typically uses fluorescent or bioluminescent molecules. Aptamerbased optical imaging can be divided into direct targeting and activatable probes (Hong et al., 2011). Direct conjugation with an aptamer and a fluorescent molecule is widely studied (Shi et al., 2010; Cui et al., 2011; Talbot et al., 2011; Zhang et al., 2012; Song et al., 2013; Zhang et al., 2013). It is the simplest way to image via a visualized aptamer by fluorescence. Activatable probes are based on conformational changes in aptamer. It is designed such that fluorescence will “turn-on” when the aptamer binds to its target. Typically, a fluorescently labeled substrate is designed to be maximally quenched by a quencher in close proximity because of FRET (McIntyre and Matrisian et al., 2003; Tsien, 2005). For example, Zhang et al., designed a bifunctional fluorescent oligonucleotide probe for the detection of adenosine triphosphate (ATP) and thrombin (Tmb). The molecular beacon contains two hairpin loops that serve as the sensing elements, and a fluorophore at one end and a quencher at the opposite end, as the reporter (Li et al., 2013). “Turn-on” of fluorescence mechanism by quenching is widely used in aptamer-based biosensor. Comparing to a conventional “always-on” probe, the activatable probe could substantially minimize the background signal originating from non-target tissues, thereby giving significantly enhanced image contrast (Hong et al., 2011). Not only optical imaging, but superparamagnetic iron oxide nanoparticle-aptamer conjugate was used to MR imaging (Yigit et al., 2007). T1-weighted MR imaging was designed that uses aptamer-gadolinium-tetraazacyclododecanetetraacetic acid (DOTA-Gd) (Xu and Lu, 2011). It consists of anti-adenosine aptamer and its complementary sequence which conjugated with DOTA-Gd. Formation of adenosine-aptamer complex leads to release of DOTAGd- complementary sequence, which allows MR imaging.
Aptamer can also be applied for targeted ultrasound imaging using covalent conjugation of nanobubbles (Wang et al., 2011). These aptamer-conjugated nanobubbles can be harnessed to release certain amount of drugs upon targeted ultrasound-trigger (Maul et al., 2010). Another application of aptamer is radioisotope labeling and its use in SPECT and PET. The TTA1 aptamer that can bind to tenascin-C, which is overexpressed in many solid tumors, was labeled with 99mTc for SPECT imaging (Hicke et al., 2006). 99mTc-labeled TTA1 aptamer exhibited rapid blood clearance with a circulation half-life of <2 min but tumor penetration in several xenograft models. The durable tumor retention in combination with fast blood clearance yielded an excellent tumor-to-blood ratio, and various tumors were clearly visualized by planar scintigraphy. Furthermore, PET imaging application was studied using 64Cu-labeled RNA aptamer (Rockey et al., 2011). In this study, the choice of chelators and radiolabeling parameters such as pH and temperature were investigated for the development of 64Cu-labeled RNA aptamers for potential PET imaging. In other aspect, aptamer can be coupled to iodinated nanoparticles (NPs) or polymer-coated Bi2S3 NPs which can be used as computed tomography (CT) contrast agents (Cai and Chen, 2007). Because each imaging method has its own advantages and disadvantages, a multimodality imaging probe based on AS1411 aptamer was evaluated (Hwang et al., 2010). For this purpose, a cobalt-ferrite NP coated with rhodamine was modified with the AS1411 aptamer, which was then conjugated to a chelating agent and further labeled with 67Ga. This multimodal imaging probe could be simultaneously used for fluorescence, gamma camera and MR imaging in vivo.
There are also attempts to not only image but simultaneously treat the disease. Diagnosis and therapy or theragnosis can be much simpler and easier when the disadvantages of each part are complemented through conjugation with aptamers. Recently, drug-loaded aptamer-gold nanoparticle (GNP) conjugate was designed for combined CT imaging and chemotherapy (Kim et al., 2010). The PSMA aptamer was conjugated with GNP for CT imaging, and Dox was loaded as a chemotherapeutic drug for prostate cancer targeting. This conjugate has been successfully used for targeting, imaging, and therapy of PMSA-expressing prostate cancer cells. Similarly, anti-mucin1 aptamer-QD-Dox conjugate was designed for optical imaging and chemotherapy of ovarian cancer (Salva et al., 2011). The key to effective and successful imaging would be the development of delivery platforms having high efficiency and ultrasensitive molecular probes for specific targets of interest
SUMMARY AND CONCLUSIONS
Aptamers have many advantages compared to traditional targeting molecules and antibodies including high affinity, low cost and immunogenicity, thermal stability, ease of production and high reproducibility. Diverse molecules such as chemotherapy drugs, toxins, siRNAs, imaging probes, and nanomaterials can be conjugated to aptamers, which serves to meet the increasing demand for versatile applications. Many new SELEX methods have been developed to easily and rapidly select for an aptamer. Compared to the traditional SELEX method, microfluidic, CE-SELEX saves time and effort in addition to providing better affinity and selectivity. In the near future, high-throughput selection of aptamers will allow various rapid aptamer selections and this will help to replace the role of antibodies in many applications from research to the clinic. Aptamers help to overcome the problem of immunogenicity when using antibodies in clinical applications and have provided a safe therapy and/or diagnosis without an immune response. Furthermore, aptamers are easily obtained by an in vitro chemical reaction and can overcome ethical issues of antibodies obtained from animals at a high cost. Natural oligonucleotides are unstable in biological fluids because of endogenous nuclease. This problem can be addressed by using modified oligonucleotides during SELEX or by a post- SELEX modification. Although a modified aptamer is obtained, careful consideration will be needed for post-SELEX modifications because the modification can affect properties of the aptamer such as binding affinity and/or function. Aptamers can be easily conjugated with other molecules in a chemical reaction. Aptamer-imaging probe conjugates can be used in various imaging modalities including optical imaging, MRI, PET, SPECT, CT, and ultrasound. Moreover, aptamers can also be conjugated with anti-cancer drugs. An aptamer-based multimodal imaging probe loaded with drug(s) will allow more precise real-time monitoring and therapy. The theragnostic potential of aptamer is gaining new momentum and will see a most auspicious development.
Acknowledgments
The present research was conducted by the research fund of Dankook University in 2011.
References
- 1.Andreola M. L., Calmels C., Michel J., Toulmé J. J., Litvak S. Towards the selection of phosphorothioate aptamers:optimizing in vitro selection steps with phosphorothioate nucleotides. Eur. J. Biochem. (2000);267:5032–5040. doi: 10.1046/j.1432-1327.2000.01557.x. [DOI] [PubMed] [Google Scholar]
- 2.Aravind A., Jeyamohan P., Nair R., Veeranarayanan S., Nagaoka Y., Yoshida Y., Maekawa T., Kumar D. S. AS1411 aptamer tagged PLGA-lecithin-PEG nanoparticles for tumor cell targeting and drug delivery. Biotechnol. Bioeng. (2012);109:2920–2931. doi: 10.1002/bit.24558. [DOI] [PubMed] [Google Scholar]
- 3.Bagalkot V., Farokhzad O. C., Langer R., Jon S. An aptamer-doxorubicin physical conjugate as a novel targeted drugdelivery platform. Angew. Chem. Int. Ed. Engl. (2006);45:8149–8152. doi: 10.1002/anie.200602251. [DOI] [PubMed] [Google Scholar]
- 4.Banaszynski M., Kolesar J. M. Vemurafenib and ipilimumab: New agents for metastatic melanoma. Am. J. Health Syst. Pharm. (2013);70:1205–1210. doi: 10.2146/ajhp120260. [DOI] [PubMed] [Google Scholar]
- 5.Barciszewski J., Medgaard M., Koch T., Kurreck J., Ermann V. A. Locked nucleic acid aptamers. Methods Mol. Biol. (2009);535:165–186. doi: 10.1007/978-1-59745-557-2_10. [DOI] [PubMed] [Google Scholar]
- 6.Berezovski M., Drabovich A., Krylova S. M., Musheev M., Okhonin V., Petrov A., Krylov S. N. Nonequilibrium capillary electrophoresis of equilibrium mixtures: a universal tool for development of aptamers. J. Am. Chem. Soc. (2005);127:3165–3171. doi: 10.1021/ja042394q. [DOI] [PubMed] [Google Scholar]
- 7.Berezovski M., Musheev M., Drabovich A., Krylov S. N. Non-SELEX selection of aptamers. J. Am. Chem. Soc. (2006);128:1410–1411. doi: 10.1021/ja056943j. [DOI] [PubMed] [Google Scholar]
- 8.Blank M., Weinschenk T., Priemer M., Schluesener H. Systematic evolution of a DNA aptamer binding to rat brain tumor microvessels. selective targeting of endothelial regulatory protein pigpen. J. Biol. Chem. (2001);276:16464–16468. doi: 10.1074/jbc.M100347200. [DOI] [PubMed] [Google Scholar]
- 9.Bruno J. G., Carrillo M. P., Phillips T., Vail N. K., Hanson D. Competitive FRET-aptamer-based detection of methylphosphonic acid, a common nerve agent metabolite. J. Fluoresc. (2008);18:867–876. doi: 10.1007/s10895-008-0316-3. [DOI] [PubMed] [Google Scholar]
- 10.Cai W., Chen X. Nanoplatforms for targeted molecular imaging in living subjects. Small. (2007);3:1840–1854. doi: 10.1002/smll.200700351. [DOI] [PubMed] [Google Scholar]
- 11.Chen F., Hu Y., Li D., Chen H., Zhang X. L. CS-SELEX generates high-affinity ssDNA aptamers as molecular probes for hepatitis C virus envelope glycoprotein E2. PLoS One. (2009);4:e8142. doi: 10.1371/journal.pone.0008142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Chu T. C., Marks J. W. 3rd, Lavery L. A., Faulkner S., Rosenblum M. G., Ellington A. D., Levy M. Aptamer:toxin conjugates that specifically target prostate tumor cells. Cancer Res. (2006a);66:5989–5992. doi: 10.1158/0008-5472.CAN-05-4583. [DOI] [PubMed] [Google Scholar]
- 13.Chu T. C., Twu K. Y., Ellington A. D., Levy M. Aptamer mediated siRNA delivery. Nucleic Acids Res. (2006b);34:e73. doi: 10.1093/nar/gkl388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Cox J. C., Rudolph P., Ellington A. D. Automated RNA selection. Biotechnol. Prog. . (1998);14:845–850. doi: 10.1021/bp980097h. [DOI] [PubMed] [Google Scholar]
- 15.Cui Q., Ren Q., Wei H. P., Chen Z., Deng J. Y., Zhang Z. P., Zhang X. E. Quantum dot-aptamer nanoprobes for recognizing and labeling influenza A virus particles. Nanoscale. (2011);3:2454–2457. doi: 10.1039/c1nr10218d. [DOI] [PubMed] [Google Scholar]
- 16.Diener J. L., Daniel Lagassé H. A., Duerschmied D., Merhi Y., Tanguay J. F., Hutabarat R., Gilbert J., Wagner D. D., Schaub R. Inhibition of von Willebrand factor-mediated platelet activation and thrombosis by the anti-von Willebrand factor A1-domain aptamer ARC1779. J. Thromb. Haemost. (2009);7:1155–1162. doi: 10.1111/j.1538-7836.2009.03459.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Dollins C. M., Nair S., Boczkowski D., Lee J., Layzer J. M., Gilboa E., Sullenger B. A. Assembling OX40 aptamers on a molecular scaffold to create a receptor-activating aptamer. Chem. Biol. (2008);15:675–682. doi: 10.1016/j.chembiol.2008.05.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Ellington A. D., Szostak J. W. In vitro selection of RNA molecules that bind specific ligands. Nature. (1990);346:818–822. doi: 10.1038/346818a0. [DOI] [PubMed] [Google Scholar]
- 19.Eyetech Study Group. Preclinical and phase 1A clinical evaluation of an anti-vegf pegylated aptamer (Eye001) for the treatment of exudative age-related macular degeneration. Retina. (2002);22:143–152. doi: 10.1097/00006982-200204000-00002. [DOI] [PubMed] [Google Scholar]
- 20.Floege J., Ostendorf T., Janssen U., Burg M., Radeke H. H., Vargeese C., Gill S. C., Green L. S., Janjić N. Novel approach to specific growth factor inhibition in vivo: antagonism of platelet-derived growth factor in glomerulonephritis by aptamers. Am. J. Pathol. (1999);154:169–179. doi: 10.1016/S0002-9440(10)65263-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Foy J. W., Rittenhouse K., Modi M., Patel M. Local tolerance and systemic safety of pegaptanib sodium in the dog and rabbit. J. Ocul. Pharmacol. Ther. (2007);23:452–466. doi: 10.1089/jop.2006.0149. [DOI] [PubMed] [Google Scholar]
- 22.Gilboa E., McNamara J. 2nd, Pastor F. Use of oligonucleotide aptamer ligands to modulate the function of immune receptors. Clin. Cancer Res. (2013);19:1054–1062. doi: 10.1158/1078-0432.CCR-12-2067. [DOI] [PubMed] [Google Scholar]
- 23.Gissel M., Orfeo T., Foley J. H., Butenas S. Effect of BAX499 aptamer on tissue factor pathway inhibitor function and thrombin generation in models of hemophilia. Thromb. Res. (2012);130:948–955. doi: 10.1016/j.thromres.2012.08.299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Gudima S.O., Kostyuk D. A., Grishchenko O. I., Tunitskaya V. L., Memelova L. V., Kochetkov S. N. Synthesis of mixed ribo/deoxyribopolynucleotides by mutant T7 RNA polymerase. FEBS Lett. (1998);439:302–306. doi: 10.1016/S0014-5793(98)01393-3. [DOI] [PubMed] [Google Scholar]
- 25.Hamula C. L., Le X. C., Li X. F. DNA aptamers binding to multiple prevalent M-types of Streptococcus pyogenes. Anal. Chem. (2011);83:3640–3647. doi: 10.1021/ac200575e. [DOI] [PubMed] [Google Scholar]
- 26.Han D., Zhu G., Wu C., Zhu Z., Chen T., Zhang X., Tan W. Engineering a cell-surface aptamer circuit for targeted and amplified photodynamic cancer therapy. ACS Nano. (2013);7:2312–2319. doi: 10.1021/nn305484p. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Healy J. M., Lewis S. D., Kurz M., Boomer R. M., Thompson K. M., Wilson C., McCauley T. G. Pharmacokinetics and biodistribution of novel aptamer compositions. Pharm. Res. (2004);21:2234–2246. doi: 10.1007/s11095-004-7676-4. [DOI] [PubMed] [Google Scholar]
- 28.Hicke B. J., Stephens A. W., Gould T., Chang Y. F., Lynott C. K., Heil J., Borkowski S., Hilger C. S., Cook G., Warren S., Schmidt P. G. Tumor targeting by an aptamer. J. Nucl. Med. (2006);47:668–678. [PubMed] [Google Scholar]
- 29.Holahan M. R., Madularu D., McConnell E. M., Walsh R., DeRosa M. C. Intra-accumbens injection of a dopamine aptamer abates MK-801-induced cognitive dysfunction in a model of schizophrenia. PLoS One. (2011);6:e22239. doi: 10.1371/journal.pone.0022239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Holland C. A., Henry A. T., Whinna H. C., Church F. C. Effect of oligodeoxynucleotide thrombin aptamer on thrombin inhibition by heparin cofactor II and antithrombin. FEBS Lett. (2000);484:87–91. doi: 10.1016/S0014-5793(00)02131-1. [DOI] [PubMed] [Google Scholar]
- 31.Hong H., Goel S., Zhang Y., Cai W. Molecular imaging with nucleic acid aptamers. Curr. Med. Chem. (2011);18:4195–4205. doi: 10.2174/092986711797189691. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Hooks M. A., Wade C. S., Millikan W. J. Muromonab CD-3: a review of its pharmacology, pharmacokinetics, and clinical use in transplantation. Pharmacotherapy. (1991);11:26–37. [PubMed] [Google Scholar]
- 33.Horhota A., Zou K., Ichida J. K., Yu B., McLaughlin L. W., Szostak J. W., Chaput J. C. Kinetic analysis of an efficient DNAdependent TNA polymerase. J. Am. Chem. Soc. (2005);127:7427–7434. doi: 10.1021/ja0428255. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Huang Y. F., Shangguan D., Liu H., Phillips J. A., Zhang X., Chen Y., Tan W. Molecular assembly of an aptamer-drug conjugate for targeted drug delivery to tumor cells. Chembiochem. (2009);10:862–868. doi: 10.1002/cbic.200800805. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Hussain A. F., Tur M. K., Barth S. An aptamer-siRNA chimera silences the eukaryotic elongation factor 2 gene and induces apoptosis in cancers expressing αvβ3 integrin. Nucleic Acid Ther. (2013);23:203–212. doi: 10.1089/nat.2012.0408. [DOI] [PubMed] [Google Scholar]
- 36.Hwang do W., Ko H. Y., Lee J. H., Kang H., Ryu S. H., Song I. C., Lee D. S., Kim S. A nucleolin-targeted multimodal nanoparticle imaging probe for tracking cancer cells using an aptamer. J. Nucl. Med. (2010);51:98–105. doi: 10.2967/jnumed.109.069880. [DOI] [PubMed] [Google Scholar]
- 37.Hybarger G., Bynum J., Williams R. F., Valdes J. J., Chambers J. P. A microfluidic SELEX prototype. Anal. Bioanal. Chem. (2006);384:191–198. doi: 10.1007/s00216-005-0089-3. [DOI] [PubMed] [Google Scholar]
- 38.Ichida J. K., Zou K., Horhota A., Yu B., McLaughlin L. W., Szostak J. W. An in vitro selection system for TNA. J. Am. Chem. Soc. (2005);127:2802–2803. doi: 10.1021/ja045364w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Jayasena S. D. Aptamers: an emerging class of molecules that rival antibodies in diagnostics. Clin. Chem. (1999);45:1628–1650. [PubMed] [Google Scholar]
- 40.Jenison R. D., Gill S. C., Pardi A., Polisky B. High-resolution molecular discrimination by RNA. Science. (1994);263:1425–1429. doi: 10.1126/science.7510417. [DOI] [PubMed] [Google Scholar]
- 41.Kasahara Y., Kuwahara M. Artificial specific binders directly recovered from chemically modified nucleic Acid libraries. J. Nucleic Acids. (2012);2012:156482. doi: 10.1155/2012/156482. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Kasahara Y., Irisawa Y., Ozaki H., Obika S., Kuwahara M. 2',4'-BNA/LNA aptamers: CE-SELEX using a DNA-based library of full-length 2'-O, 4'-C-methylene-bridged/linked bicyclic ribonucleotides. Bioorg. Med. Chem. Lett. (2013);23:1288–1292. doi: 10.1016/j.bmcl.2012.12.093. [DOI] [PubMed] [Google Scholar]
- 43.Kawakami J., Imanaka H., Yokota Y., Sugimoto N. in vitro selection of aptamers that act with Zn2+. J. Inorg. Biochem. (2000);82:197–206. doi: 10.1016/S0162-0134(00)00158-6. [DOI] [PubMed] [Google Scholar]
- 44.Kempeneers V., Renders M., Froeyen M., Herdewijn P. Investigation of the DNA-dependent cyclohexenyl nucleic acid polymerization and the cyclohexenyl nucleic acid-dependent DNA polymerization. Nucleic Acids Res. (2005);33:3828–3836. doi: 10.1093/nar/gki695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Kim D., Jeong Y. Y., Jon S. A drug-loaded aptamer-gold nanoparticle bioconjugate for combined CT imaging and therapy of prostate cancer. ACS Nano. (2010);4:3689–3696. doi: 10.1021/nn901877h. [DOI] [PubMed] [Google Scholar]
- 46.Kim S., Kim Y., Kim P., Ha J., Kim K., Sohn M., Yoo J. S., Lee J., Kwon J. A., Lee K. N. Improved sensitivity and physical properties of sol-gel protein chips using large-scale material screening and selection. Anal. Chem. (2006);78:7392–7396. doi: 10.1021/ac0520487. [DOI] [PubMed] [Google Scholar]
- 47.Klussmann S., Nolte A., Bald R., Erdmann V. A., Fürste J. P. Mirror-image RNA that binds D-adenosine. Nat. Biotechnol. (1996);14:1112–1115. doi: 10.1038/nbt0996-1112. [DOI] [PubMed] [Google Scholar]
- 48.Knight D. M., Trinh H., Le J., Siegel S., Shealy D., McDonough M., Scallon B., Moore M. A., Vilcek J., Daddona P. Construction and initial characterization of a mouse-human chimeric anti-TNF antibody. Mol. Immunol. (1993);30:1443–1453. doi: 10.1016/0161-5890(93)90106-L. [DOI] [PubMed] [Google Scholar]
- 49.Kolesnikova O., Kazakova H., Comte C., Steinberg S., Kamenski P., Martin R. P., Tarassov I., Entelis N. Selection of RNA aptamers imported into yeast and human mitochondria. RNA. (2010);16:926–941. doi: 10.1261/rna.1914110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Kuwahara M., Obika S., Nagashima J., Ohta Y., Suto Y., Ozaki H., Sawai H., Imanishi T. Systematic analysis of enzymatic DNA polymerization using oligo-DNA templates and triphosphate analogs involving 2',4'-bridged nucleosides. Nucleic Acids Res. (2008);36:4257–4265. doi: 10.1093/nar/gkn404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Kuwahar M., Takahata Y., Shoji A., Ozaki A. N., Ozaki H., Sawai H. Substrate properties of C5-substituted pyrimidine 2’-deoxynucleoside 5’-triphosphates for thermostable DNA polymerases during PCR. Bioorg. Med. Chem. Lett. (2003);13:3735–3738. doi: 10.1016/j.bmcl.2003.08.001. [DOI] [PubMed] [Google Scholar]
- 52.Kuwahara M., Takano Y., Kasahara Y., Nara H., Ozaki H., Sawai H., Sugiyama A., Obika S. Study on suitability of KOD dna polymerase for enzymatic production of artificial nucleic acids using base/sugar modified nucleoside triphosphates. Molecules. (2010);15:8229–8240. doi: 10.3390/molecules15118229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Kuwahara M., Takeshima H., Nagashima J., Minezaki S., Ozaki H., Sawai H. Transcription and reverse transcription of artificial nucleic acids involving backbone modification by template directed DNA polymerase reactions. Bioorg. Med. Chem. (2009);17:3782–3788. doi: 10.1016/j.bmc.2009.04.045. [DOI] [PubMed] [Google Scholar]
- 54.Latham J. A., Johnson R., Toole J. J. The application of a modified nucleotide in aptamer selection: novel thrombin aptamers containing 5-(1-pentynyl)-2’-deoxyuridine. Nucleic Acids Res. (1994);22:2817–2822. doi: 10.1093/nar/22.14.2817. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Lato S. M., Ozerova N. D., He K., Sergueeva Z., Shaw B. R., Burke D. H. Boron-containing aptamers to ATP. Nucleic Acids Res. (2002);30:1401–1407. doi: 10.1093/nar/30.6.1401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Lauhon C. T., Szostak J. W. RNA aptamers that bind flavin and nicotinamide redox cofactors. J. Am. Chem. Soc. (1995);117:1246–1257. doi: 10.1021/ja00109a008. [DOI] [PubMed] [Google Scholar]
- 57.Leal N. A., Sukeda M., Benner S. A. Dynamic assembly of primers on nucleic acid templates. Nucleic Acids Res. (2006);34:4702–4710. doi: 10.1093/nar/gkl625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Leva S., Lichte A., Burmeister J., Muhn P., Jahnke B., Fesser D., Erfurth J., Burgstaller P., Klussmann S. GnRH binding RNA and DNA Spiegelmers: a novel approach toward GnRH antagonism. Chem. Biol. (2002);9:351–359. doi: 10.1016/S1074-5521(02)00111-4. [DOI] [PubMed] [Google Scholar]
- 59.Li F., Du Z., Yang L., Tang B. Selective and sensitive turnon detection of adenosine triphosphate and thrombin based on bifunctional fluorescent oligonucleotide probe. Biosens. Bioelectron. (2013);41:907–910. doi: 10.1016/j.bios.2012.10.007. [DOI] [PubMed] [Google Scholar]
- 60.Li M., Lin N., Huang Z., Du L., Altier C., Fang H., Wang B. Selecting aptamers for a glycoprotein through the incorporation of the boronic acid moiety. J. Am. Chem. Soc. (2008);130:12636–12638. doi: 10.1021/ja801510d. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Liu Z., Duan J. H., Song Y. M., Ma J., Wang F. D., Lu X., Yang X. D. Novel HER2 aptamer selectively delivers cytotoxic drug to HER2-positive breast cancer cells in vitro. J. Transl. Med. (2012);10:148. doi: 10.1186/1479-5876-10-148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Lou X., Qian J., Xiao Y., Viel L., Gerdon A. E., Lagally E. T., Atzberger P., Tarasow T. M., Heeger A. J., Soh H. T. Micromagnetic selection of aptamers in microfluidic channels. Proc. Natl. Acad. Sci. U.S.A. (2009);106:2989–2994. doi: 10.1073/pnas.0813135106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Mann A. P., Bhavane R. C., Somasunderam A., Liz Montalvo-Ortiz B., Ghaghada K. B., Volk D., Nieves-Alicea R., Suh K. S., Ferrari M., Annapragada A., Gorenstein D. G., Tanaka T. Thioaptamer conjugated liposomes for tumor vasculature targeting. Oncotarget. (2011);2:298–304. doi: 10.18632/oncotarget.261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Mann D., Reinemann C., Stoltenburg R., Strehlitz B. in vitro selection of DNA aptamers binding ethanolamine. Biochem. Biophys. Res. Commun. (2005);338:1928–1934. doi: 10.1016/j.bbrc.2005.10.172. [DOI] [PubMed] [Google Scholar]
- 65.Maul T. M., Dudgeon D. D., Beste M. T., Hammer D. A., Lazo J. S., Villanueva F. S., Wagner W. R. Optimization of ultrasound contrast agents with computational models to improve selection of ligands and binding strength. Biotechnol. Bioeng. (2010);107:854–864. doi: 10.1002/bit.22857. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Mazumdar S., Greenwald D. Golimumab. MAbs. (2009);1:422–431. doi: 10.4161/mabs.1.5.9286. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.McIntyre J. O., Matrisian L. M. Molecular imaging of proteolytic activity in cancer. J. Cell. Biochem. (2003);90:1087–1097. doi: 10.1002/jcb.10713. [DOI] [PubMed] [Google Scholar]
- 68.McNamara J. O. 2nd, Andrechek E. R., Wang Y., Viles K. D., Rempel R. E., Gilboa E., Sullenger B. A., Giangrande P. H. Cell type-specific delivery of siRNAs with aptamer-siRNA chimeras. Nat. Biotechnol. (2006);24:1005–1015. doi: 10.1038/nbt1223. [DOI] [PubMed] [Google Scholar]
- 69.McNamara J. O., Kolonias D., Pastor F., Mittler R. S., Chen L., Giangrande P. H., Sullenger B., Gilboa E. Multivalent 4-1BB binding aptamers costimulate CD8+ T cells and inhibit tumor growth in mice. J. Clin. Invest. (2008);118:376–386. doi: 10.1172/JCI33365. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Meng L., Yang L., Zhao X., Zhang L., Zhu H., Liu C., Tan W. Targeted delivery of chemotherapy agents using a liver cancer-specific aptamer. PLoS One. (2012);7:e33434. doi: 10.1371/journal.pone.0033434. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Mi J., Liu Y., Rabbani Z. N., Yang Z., Urban J. H., Sullenger B. A., Clary B. M. in vivo selection of tumor-targeting RNA motifs. Nat. Chem. Biol. (2010);6:22–24. doi: 10.1038/nchembio.277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Mosing R. K., Mendonsa S. D., Bowser M. T. Capillary electrophoresis-SELEX selection of aptamers with affinity for HIV-1 reverse transcriptase. Anal. Chem. (2005);77:6107–6112. doi: 10.1021/ac050836q. [DOI] [PubMed] [Google Scholar]
- 73.Neff C. P., Zhou J., Remling L., Kuruvilla J., Zhang J., Li H., Smith D. D., Swiderski P., Rossi J. J., Akkina R. An aptamersiRNA chimera suppresses HIV-1 viral loads and protects from helper CD4(+) T cell decline in humanized mice. Sci. Transl. Med. (2011);3:66ra6. doi: 10.1126/scitranslmed.3001581. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Nitsche A., Kurth A., Dunkhorst A., Pänke O., Sielaff H., Junge W., Muth D., Scheller F., Stöcklein W., Dahmen C., Pauli G., Kage A. One-step selection of Vaccinia virus-binding DNA aptamers by MonoLEX. BMC Biotechnol. (2007);7:48. doi: 10.1186/1472-6750-7-48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Nolte A., Klussmann S., Bald R., Erdmann V. A., Furste J. P. Mirror-design of L-oligonucleotide ligands binding to L-arginine. Nat. Biotechnol. (1996);14:1116–1119. doi: 10.1038/nbt0996-1116. [DOI] [PubMed] [Google Scholar]
- 76.Pageau S. C. Denosumab. MAbs. (2009);1:210–5. doi: 10.4161/mabs.1.3.8592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Park S. M., Ahn J. Y., Jo M., Lee D. K., Lis J. T., Craighead H. G., Kim S. Selection and elution of aptamers using nanoporous sol-gel arrays with integrated microheaters. Lab Chip. (2009);9:1206–1212. doi: 10.1039/b814993c. [DOI] [PubMed] [Google Scholar]
- 78.Pastor F., Soldevilla M. M., Villanueva H., Kolonias D., Inoges S., de Cerio A. L., Kandzia R., Klimyuk V., Gleba Y., Gilboa E., Bendandi M. CD28 aptamers as powerful immune response modulators. Mol. Ther. Nucleic Acids. (2013);2:e98. doi: 10.1038/mtna.2013.26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Rockey W. M., Huang L., Kloepping K. C., Baumhover N. J., Giangrande P. H., Schultz M. K. Synthesis and radiolabeling of chelator-RNA aptamer bioconjugates with copper-64 for targeted molecular imaging. Bioorg. Med. Chem. (2011);19:4080–4090. doi: 10.1016/j.bmc.2011.05.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Römer P. S., Berr S., Avota E., Na S. Y., Battaglia M., ten Berge I., Einsele H., Hünig T. Preculture of PBMCs at high cell density increases sensitivity of T-cell responses, revealing cytokine release by CD28 superagonist TGN1412. Blood. (2011);118:6772–6782. doi: 10.1182/blood-2010-12-319780. [DOI] [PubMed] [Google Scholar]
- 81.Ruckman J., Green L. S., Beeson J., Waugh S., Gillette W. L., Henninger D. D., Claesson-Welsh L., Janjić N. 2'-Fluoropyrimidine RNA-based aptamers to the 165-amino acid form of vascular endothelial growth factor (VEGF165). Inhibition of receptor binding and VEGF-induced vascular permeability through interactions requiring the exon 7-encoded domain. J. Biol. Chem. (1998);273:20556–20567. doi: 10.1074/jbc.273.32.20556. [DOI] [PubMed] [Google Scholar]
- 82.Savla R., Taratula O., Garbuzenko O., Minko T. Tumor targeted quantum dot-mucin 1 aptamer-doxorubicin conjugate for imaging and treatment of cancer. J. Control. Release. (2011);153:16–22. doi: 10.1016/j.jconrel.2011.02.015. [DOI] [PubMed] [Google Scholar]
- 83.Schmidt K. S., Borkowski S., Kurreck J., Stephens A. W., Bald R., Hecht M., Friebe M., Dinkelborg L., Erdmann V. A. Application of locked nucleic acids to improve aptamer in vivo stability and targeting function. Nucleic Acids Res. (2004);32:5757–5765. doi: 10.1093/nar/gkh862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Schneider D. J., Feigon J., Hostomsky Z., Gold L. Highaffinity ssDNA inhibitors of the reverse transcriptase of type 1 human immunodeficiency virus. Biochemistry. (1995);34:9599–9610. doi: 10.1021/bi00029a037. [DOI] [PubMed] [Google Scholar]
- 85.Shi H., Tang Z., Kim Y., Nie H., Huang Y. F., He X., Deng K., Wang K., Tan W. in vivo fluorescence imaging of tumors using molecular aptamers generated by cell-SELEX. Chem. Asian J. (2010);5:2209–2213. doi: 10.1002/asia.201000242. [DOI] [PubMed] [Google Scholar]
- 86.Shoji A., Kuwahara M., Ozaki H., Sawai H. Modified DNA aptamer that binds the (R)-isomer of a thalidomide derivative with high enantioselectivity. J. Am. Chem. Soc. (2007);129:1456–1464. doi: 10.1021/ja067098n. [DOI] [PubMed] [Google Scholar]
- 87.Siller-Matula J. M., Merhi Y., Tanguay J. F., Duerschmied D., Wagner D. D., McGinness K. E., Pendergrast P. S., Chung J. K., Tian X., Schaub R. G., Jilma B. ARC15105 is a potent antagonist of von Willebrand factor mediated platelet activation and adhesion. Arterioscler. Thromb. Vasc. Biol. (2012);32:902–909. doi: 10.1161/ATVBAHA.111.237529. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Song Y., Zhu Z., An Y., Zhang W., Zhang H., Liu D., Yu C., Duan W., Yang C. J. Selection of DNA aptamers against epithelial cell adhesion molecule for cancer cell imaging and circulating tumor cell capture. Anal. Chem. (2013);85:4141–4149. doi: 10.1021/ac400366b. [DOI] [PubMed] [Google Scholar]
- 89.Subramanian N., Raghunathan V., Kanwar J. R., Kanwar R. K., Elchuri S. V., Khetan V., Krishnakumar S. Target-specific delivery of doxorubicin to retinoblastoma using epithelial cell adhesion molecule aptamer. Mol. Vis. (2012);18:2783–2795. [PMC free article] [PubMed] [Google Scholar]
- 90.Suntharalingam G., Perry M. R., Ward S., Brett S. J., Castello-Cortes A., Brunner M. D., Panoskaltsis N. Cytokine storm in a phase 1 trial of the anti-CD28 monoclonal antibody TGN1412. N. Engl. J. Med. (2006);355:1018–1028. doi: 10.1056/NEJMoa063842. [DOI] [PubMed] [Google Scholar]
- 91.Talbot L. J., Mi Z., Bhattacharya S. D., Kim V., Guo H., Kuo P. C. Pharmacokinetic characterization of an RNA aptamer against osteopontin and demonstration of in vivo efficacy in reversing growth of human breast cancer cells. Surgery. (2011);150:224–230. doi: 10.1016/j.surg.2011.05.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Tang Z., Parekh P., Turner P., Moyer R. W., Tan W. Generating aptamers for recognition of virus-infected cells. Clin. Chem. (2009);55:813–822. doi: 10.1373/clinchem.2008.113514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Tsai C. H., Chen J., Szostak J. W. Enzymatic synthesis of DNA on glycerol nucleic acid templates without stable duplex formation between product and template. Proc. Natl. Acad. Sci. USA. (2007);104:14598–14603. doi: 10.1073/pnas.0704211104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Tsien R. Y. Building and breeding molecules to spy on cells and tumors. FEBS Lett. (2005);579:927–932. doi: 10.1016/j.febslet.2004.11.025. [DOI] [PubMed] [Google Scholar]
- 95.Tuerk C., Gold L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science. (1990);249:505–510. doi: 10.1126/science.2200121. [DOI] [PubMed] [Google Scholar]
- 96.Vater A., Sell S., Kaczmarek P., Maasch C., Buchner K., Pruszynska-Oszmalek E., Kolodziejski P., Purschke W. G., Nowak K. W., Strowski M. Z., Klussmann S. A mixed mirror-image DNA/RNA aptamer inhibits glucagon and acutely improves glucose tolerance in models of type 1 and type 2 diabetes. J. Biol. Chem. (2013);288:21136–21147. doi: 10.1074/jbc.M112.444414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Vaught J. D., Bock C., Carter J., Fitzwater T., Otis M., Schneider D., Rolando J., Waugh S., Wilcox S. K., Eaton B. E. Expanding the chemistry of DNA for in vitro selection. J. Am. Chem. Soc. (2010);132:4141–4151. doi: 10.1021/ja908035g. [DOI] [PubMed] [Google Scholar]
- 98.Veedu R. N., Vester B., Wengel J. Efficient enzymatic synthesis of LNA-modified DNA duplexes using KOD DNA polymerase. Org. Biomol. Chem. (2009);7:1404–1409. doi: 10.1039/b819946a. [DOI] [PubMed] [Google Scholar]
- 99.Wang C. H., Huang Y. F., Yeh C. K. Aptamer-conjugated nanobubbles for targeted ultrasound molecular imaging. Langmuir. (2011);27:6971–6976. doi: 10.1021/la2011259. [DOI] [PubMed] [Google Scholar]
- 100.Wang J., Jiang H., Liu F. in vitro selection of novel RNA ligands that bind human cytomegalovirus and block viral infection. RNA. (2000);6:571–583. doi: 10.1017/S1355838200992215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.White R., Rusconi C., Scardino E., Wolberg A., Lawson J., Hoffman M., Sullenger B. Generation of species crossreactive aptamers using "toggle" SELEX. Mol. Ther. (2001);4:567–573. doi: 10.1006/mthe.2001.0495. [DOI] [PubMed] [Google Scholar]
- 102.Williams K. P., Liu X. H., Schumacher T. N., Lin H. Y., Ausiello D. A., Kim P. S., Bartel D. P. Bioactive and nuclease-resistant L-DNA ligand of vasopressin. Proc. Natl. Acad. Sci. U.S.A. (1997);94:11285–11290. doi: 10.1073/pnas.94.21.11285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Wittung P., Nielsen P. E., Buchardt O., Egholm M., Nordén B. DNA-like double helix formed by peptide nucleic acid. Nature. (1994);368:561–563. doi: 10.1038/368561a0. [DOI] [PubMed] [Google Scholar]
- 104.Wullner U., Neef I., Eller A., Kleines M., Tur M. K., Barth S. Cell-specific induction of apoptosis by rationally designed bivalent aptamer-siRNA transcripts silencing eukaryotic elongation factor 2. Curr. Cancer Drug Targets. (2008);8:554–565. doi: 10.2174/156800908786241078. [DOI] [PubMed] [Google Scholar]
- 105.Xu W., Lu Y. A smart magnetic resonance imaging contrast agent responsive to adenosine based on a DNA aptamer-conjugated gadolinium complex. Chem. Commun. (2011);47:4998–5000. doi: 10.1039/c1cc10161g. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Yang X., Fennewald S., Luxon B. A., Aronson J., Herzog N. K., Gorenstein D. G. Aptamers containing thymidine 3’-Ophosphorodithioates: synthesis and binding to nuclear factor-κB. Bioorg. Med. Chem. Lett. (1999);9:3357–3362. doi: 10.1016/S0960-894X(99)00600-9. [DOI] [PubMed] [Google Scholar]
- 107.Yang X., Huang J., Wang K., Li W., Cui L., Li X. Angiogenin-mediated photosensitizer-aptamer conjugate for photodynamic therapy. ChemMedChem. (2011);6:1788–1780. doi: 10.1002/cmdc.201100226. [DOI] [PubMed] [Google Scholar]
- 108.Yigit M. V., Mazumdar D., Kim H. K., Lee J. H., Odintsov B., Lu Y. Smart "turn-on" magnetic resonance contrast agents based on aptamer-functionalized superparamagnetic iron oxide nanoparticles. Chembiochem. (2007);8:1675–1678. doi: 10.1002/cbic.200700323. [DOI] [PubMed] [Google Scholar]
- 109.Zhang C., Ji X., Zhang Y., Zhou G., Ke X., Wang H., Tinnefeld P., He Z. One-pot synthesized aptamer-functionalized CdTe:Zn2+ quantum dots for tumor-targeted fluorescence imaging in vitro and in vivo. Anal. Chem. (2013);85:5843–5849. doi: 10.1021/ac400606e. [DOI] [PubMed] [Google Scholar]
- 110.Zhang M. Z., Yu R. N., Chen J., Ma Z. Y., Zhao Y. D. Targeted quantum dots fluorescence probes functionalized with aptamer and peptide for transferrin receptor on tumor cells. Nanotechnology. (2012);23:485104. doi: 10.1088/0957-4484/23/48/485104. [DOI] [PubMed] [Google Scholar]
- 111.Zhang Y., Hong H., Cai W. Tumor-targeted drug delivery with aptamers. Curr. Med. Chem. (2011);18:4185–4194. doi: 10.2174/092986711797189547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Zhou B., Wang B. Pegaptanib for the treatment of agerelated macular degeneration. Exp. Eye Res. (2006);83:615–619. doi: 10.1016/j.exer.2006.02.010. [DOI] [PubMed] [Google Scholar]
- 113.Zhou J., Li H., Li S., Zaia J., Rossi J. J. Novel dual inhibitory function aptamer-siRNA delivery system for HIV-1 therapy. Mol. Ther. (2008);16:1481–1489. doi: 10.1038/mt.2008.92. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Zhou J., Li H., Zhang J., Piotr S., Rossi J. Development of cell-type specific anti-HIV gp120 aptamers for siRNA delivery. J. Vis. Exp. (2011a);23:2954. doi: 10.3791/2954. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Zhou J., Neff C. P., Swiderski P., Li H., Smith D. D., Aboellail T., Remling-Mulder L., Akkina R., Rossi J. J. Functional in vivo delivery of multiplexed anti-HIV-1 siRNAs via a chemically synthesized aptamer with a sticky bridge. Mol. Ther. (2013);21:192–200. doi: 10.1038/mt.2012.226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Zhou J., Shu Y., Guo P., Smith D. D., Rossi J. J. Dual functional RNA nanoparticles containing phi29 motor pRNA and anti-gp120 aptamer for cell-type specific delivery and HIV-1 inhibition. Methods. (2011b);54:284–294. doi: 10.1016/j.ymeth.2010.12.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Zhu G., Ye M., Donovan M. J., Song E., Zhao Z., Tan W. Nucleic acid aptamers: an emerging frontier in cancer therapy. Chem. Commun. (Camb) (2012A);48:10472–10480. doi: 10.1039/c2cc35042d. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Zhu Q., Shibata T., Kabashima T., Kai M. Inhibition of HIV-1 protease expression in T cells owing to DNA aptamer-mediated specific delivery of siRNA. Eur. J. Med. Chem. (2012B);56:396–399. doi: 10.1016/j.ejmech.2012.07.045. [DOI] [PubMed] [Google Scholar]