Abstract
Symbiotic bacteria often help their hosts acquire nutrients from their diet, showing trends of co-evolution and independent acquisition by hosts from the same trophic levels. While these trends hint at important roles for biotic factors, the effects of the abiotic environment on symbiotic community composition remain comparably understudied. In this investigation, we examined the influence of abiotic and biotic factors on the gut bacterial communities of fish from different taxa, trophic levels and habitats. Phylogenetic and statistical analyses of 25 16S rRNA libraries revealed that salinity, trophic level and possibly host phylogeny shape the composition of fish gut bacteria. When analysed alongside bacterial communities from other environments, fish gut communities typically clustered with gut communities from mammals and insects. Similar consideration of individual phylotypes (vs. communities) revealed evolutionary ties between fish gut microbes and symbionts of animals, as many of the bacteria from the guts of herbivorous fish were closely related to those from mammals. Our results indicate that fish harbour more specialized gut communities than previously recognized. They also highlight a trend of convergent acquisition of similar bacterial communities by fish and mammals, raising the possibility that fish were the first to evolve symbioses resembling those found among extant gut fermenting mammals.
Keywords: 16S rDNA, community ecology, fish, gut bacteria, microbial ecology, species interactions
Introduction
Although bacteria are ubiquitous across all life-sustaining habitats on Earth, different types of environments harbour strikingly different bacterial communities. Previous analyses have focused on factors such as salinity (Lozupone & Knight 2007), pH (Fierer & Jackson 2006; Chu et al. 2010), seasonality (Gilbert et al. 2009, 2012) and ecological interactions (Steele et al. 2011) as major factors determining the composition of free-living communities. Bacteria not only exist in such ‘external’ environments, but also associate with eukaryotic hosts. Many of these symbiotic (i.e. intimately host-associated) microbes form specific and occasionally obligate interactions with hosts including macroalgae (Lachnit et al. 2010), insects (Moran et al. 2008) and primates (Yildirim et al. 2010). Other symbiotic bacteria, such as rhizobia and Vibrio fischeri, show less dependent associations, multiplying freely in the environment while also colonizing plant or animal hosts when opportunity arises (McFall-Ngai & Ruby 1991; Fraysse et al. 2003).
Although the composition of bacterial communities is strongly determined by properties of the external environments in which they are found (Fierer & Jackson 2006), the ‘environments’ provided by eukaryotic hosts are also largely impacted by the microbes that colonize them. For example, gut bacteria often promote nutritional provisioning and nitrogen recycling for their hosts (Douglas 1998; Sabree et al. 2009). In the vertebrate gut, bacteria play important physiological roles, influencing metabolic processes such as the digestion of complex carbohydrates (Turnbaugh et al. 2006) and the regulation of fat storage (Bäckhed et al. 2004).
Most studies of vertebrate gut communities have focused on mammals, which comprise fewer than 10% of total vertebrate diversity. Far fewer have emphasized fish, which originated over 600 million years ago and encompass approximately 28 000 extant species (Nelson 2006)—nearly half the total number of vertebrate species. The antiquity, diversity and dietary variation of fish highlight the need for their inclusion as we attempt to quantify the variety and nature of symbioses exhibited across the vertebrate tree of life.
In expanding the range of studied vertebrate gut communities, of particular interest are questions about the range of environmental, ecological and evolutionary factors that shape gut microbial communities, and the functions these communities can perform. For instance, similar gut communities are found among phylogeneti-cally related mammals and among mammals with similar diets (Ley et al. 2008a,b; Muegge et al. 2011). Do the same relationships occur in fish, despite their substantial differences from mammals? Although several publications have reported on the types of bacteria associated with fish guts, the scope of their analyses has generally been narrow, with the exception of a recent effort by Roeselers et al. (2011). In this study, the authors suggest that the host gut is a selective environment based on their finding that zebrafish (Danio rerio) with different origins harboured a stable, core gut microbiome. In addition to their focus on zebrafish bacteria, the authors also analysed previously studied gut communities from other fish species, observing similarities among those found in related fish from the same order. Both findings are fascinating and suggest specialized and potentially co-evolved associations between fish and their gut bacteria. However, as was the case for this study, few statistical analyses have been performed to assess how gut communities are shaped by environmental and ecological factors. As such, the forces responsible for structuring these microbial menageries remain to be established.
To address these shortcomings, we performed a meta-analysis of 16S rRNA gene sequence data from fourteen published and two unpublished data sets on teleost gut communities, and a new data set from the Trinidadian guppy, Poecilia reticulata, generated in this study. By examining the taxonomic and phylogenetic similarity among gut bacteria of fish from different habitats, trophic levels and taxa, we investigated the factors that shape these communities. Because of the central roles microbes can play in digestive processes, we expected that fish with similar diets, especially those that are nutrient poor or difficult to digest, have been selected to harbour similar bacterial communities with similar functional capacities. We also expected, a priori, that microbes would differ notably among freshwater and marine fish based on previous findings that free-living bacterial communities from saline and nonsaline communities differ substantially (Lozupone & Knight 2007). To provide a broader perspective on fish gut bacteria, we also compared these communities to those from other vertebrate guts, nonvertebrate eukaryotes and free-living habitats. Our results, therefore, not only provide some of the first insights into the forces that structure fish gut communities, but they also illustrate their affinities to symbiotic and free-living communities from a wide range of natural environments.
Methods
Sequence acquisition
All fish gut bacterial 16S rRNA sequences were obtained from GenBank except for the guppy (Poecilia reticulata) library, which were generated through cloning and sequencing (See Supporting information for details). All nonguppy data sets were identified through Entrez queries using targeted search terms, or through a review of the published literature on fish and their gut microbes. Sequences from each of these studies had been similarly generated via Sanger sequencing from PCR products that were amplified with ‘universal’ bacterial primers as stated by the authors of the studies. Within a particular study, all sequences from one fish species were counted as a single library when acquired by the same method (i.e. culture-dependent or culture-independent techniques). However, in cases where the study specified that individuals came from different fish populations, sequences from separate populations were analysed separately as distinct libraries (exceptions are noted in Supporting information).
Sequence analysis: comparing bacterial communities
To preprocess our data set for analysis, we excluded sequences that: (i) were shorter than 200 bp, (ii) had over 10% of the sequence length composed of ambiguous nucleotides, (iii) belonged to libraries with <15 sequences or (iv) were derived from chloroplasts, as assessed by the RDP classifier tool (Wang et al. 2007). The remaining sequences, partitioned by library, were aligned using Infernal within RDP (Cole et al. 2009; Nawrocki et al. 2009).
A distance matrix was created from the RDP alignment for each library using paup with the HKY85 model of sequence evolution (Swofford 2006). The sequences within each library were grouped into OTUs based on 3% sequence divergence using the furthest neighbour algorithm in Mothur versions 1.8.0 or later (Schloss et al. 2009). Because of heterogeneity in the regions of 16S rRNA sampled for different sequence reads, we only utilized sequences greater than 750 bp from the Pomacanthus sexstriatus library, thus yielding sufficient sequence overlap for distance calculations. This led to the exclusion of 59 of the 228 eubacterial sequence reads from this library. The get.oturep command in Mothur was used to select representative sequences for each OTU. A total of 1054 representative sequences were used for all subsequent analyses. The mean length of representative sequences was 1135 bp, the median was 1340 bp, and only 1.5% of the representative sequences were shorter than 500 bp.
Because a number of authors deposited only representative sequences in NCBI, all of our remaining analyses exclusively utilized these representative 97% OTU sequences, except where noted. These analyses thus allow comparisons of OTU presence/absence across the studied libraries using qualitative metrics of community similarity but cannot be used for analyses with quantitative metrics. Based on the criteria laid out previously (Stackebrandt & Goebel 1994), comparisons between gut communities using OTUs of up to 3% divergence reveal differences in species composition, although this definition is an approximation and has many well-known exceptions.
Representative sequences were aligned in GreenG-enes NAST server (DeSantis et al. 2006), and the Lane mask (Lane 1991) was applied to exclude the hypervari-able regions of the 16S rRNA gene that can be problematic for phylogenetic analysis. Before running our phylogenetic analyses on fish-associated microbes (Fish Dataset), all sequences assigned to a bacterial phylum with ≥80% bootstrap support (using RDP Classifier) were constrained to group into monophyletic, phylum-specific clades. A de novo phylogenetic tree of representative sequences was created using RAxML 7.2.7 Black Box on the cipres web portal (Miller et al. 2010). This tree was rooted with an archaeabacterial sequence with the accession number: FJ655681. We then used Fast Uni-Frac (Hamady et al. 2009) to investigate differences among bacterial communities from different fish. This web application calculated the UniFrac distance based on the percentage of unique phylogenetic branch length among all pairs of gut communities, thus inferring the extent of phylogenetic overlap between communities from each host (Lozupone & Knight 2005; Lozupone et al. 2006).
To better understand the forces that shape fish gut communities, all fish hosts were assigned to the appropriate trophic level (herbivorous, omnivorous or carnivorous), habitat (freshwater, mixed/estuarine, marine) and taxon (at the level of order) based on published literature (Table 1). The unweighted UniFrac phylogenetic distance metric was then analysed using a principal coordinates analysis (PCoA), allowing us to assess microbial community differences across fish from these various categories. For this analysis, the anadromous salmonid species were placed with freshwater fish because they had been either reared in or collected from a freshwater environment.
Table 1.
A summary of the 16S rRNA sequence libraries used in this paper’s analysis
| Fish | Sampling method* |
Total sequences† |
Number of OTUs‡ |
Diet§ | Habitat¶ | Tissue type** |
Rearing†† | Order | References |
|---|---|---|---|---|---|---|---|---|---|
| Acanthurus nigricans | Uncultured | 48 | 36 | H | Saltwater | IC | W | Perciformes | Smriga et al 2010 |
| Chaenocephalus aceratus | Uncultured | 303 | 4 | C | Saltwater | IW | W | Perciformes | Ward et al 2009 |
| Chlorurus sordidus | Uncultured | 44 | 20 | H | Saltwater | FP | W | Perciformes | Smriga et al 2010 |
| Danio rerio (Eugene) | Uncultured | 1352 | 113 | O | Freshwater | DT | A | Cypriniformes | Roeselers et al 2011 |
| Danio rerio (India) | Uncultured | 403 | 74 | O | Freshwater | DT | W | Cypriniformes | Roeselers et al 2011 |
| Danio rerio (Seattle) | Uncultured | 401 | 65 | O | Freshwater | DT | A | Cypriniformes | Roeselers et al 2011 |
| Danio rerio (Chapel Hill) | Uncultured | 136 | 30 | O | Freshwater | DT | A | Cypriniformes | Roeselers et al 2011 |
| Danio rerio | Uncultured | 1456 | 145 | O | Freshwater | DT | A | Cypriniformes | Rawls et al 2006 |
| Epinephelus coioides | Cultured | 17 | 14 | C | Estuarine | DT | A | Perciformes | Sun et al 2009 |
| Gillichthys mirabilis | Uncultured | 17 | 15 | C | Estuarine | DT | W | Perciformes | Bano et al 2007 |
| Hippocampus guttulatus | Uncultured | 46 | 39 | C | Saltwater | IC | A | Gasterosteiformes | Balcázar et al 2010 |
| Lutjanus bohar | Uncultured | 44 | 18 | C | Saltwater | IC | W | Perciformes | Smriga et al 2010 |
| Naso tonganus | Uncultured | 101 | 80 | H | Saltwater | IC | W | Perciformes | Mendell, J.E., unpublished data |
| Notothenia coriiceps | Uncultured | 194 | 13 | O | Saltwater | IW | W | Perciformes | Ward et al 2009 |
| Oncorhynchus mykiss | Cultured | 15 | 13 | C | Freshwater (M) | IC | A | Salmoniformes | Huber et al 2004 |
| Oncorhynchus mykiss | Cultured | 34 | 29 | C | Freshwater (M) | IC | A | Salmoniformes | Kim et al 2007 |
| Oncorhynchus mykiss | Uncultured | 35 | 22 | C | Freshwater (M) | IC | A | Salmoniformes | Kim et al 2007 |
| Pelteobagrus fulvidraco | Uncultured | 65 | 52 | C | Freshwater | IC & IM | W | Siluriformes | Wu et al 2010 |
| Poecilia reticulata | Uncultured | 112 | 70 | O | Freshwater | DT | W | Cyprinodontiformes | This study |
| Pomacanthus sexstriatus | Uncultured | 228 | 110 | H | Saltwater | IC | W | Perciformes | Ward, R.J., unpublished data |
| Salmo salar | Cultured | 52 | 15 | C | Freshwater (M) | IC | W | Salmoniformes | Skrodenytė-Arbačiauskienė et al 2008 |
| Salmo trutta fario | Cultured | 100 | 23 | C | Freshwater | IC | W | Salmoniformes | Skrodenyte-Arbaciauskiene et al 2006 |
| Salmo trutta trutta | Cultured | 47 | 15 | C | Freshwater (M) | IC | W | Salmoniformes | Skrodenytė-Arbačiauskienė et al 2008 |
| Solea senegalensis | Cultured | 36 | 18 | C | Saltwater | DT | A | Pleuronectiformes | Martin-Antonio et al 2007 |
| Takifugu niphobles | Uncultured | 25 | 21 | C | Saltwater | IC | W | Tetraodontiformes | Shiina et al 2006 |
Sampling method refers to whether the sequences were derived from culture-independent methods (uncultured) or by culturing the bacteria before sequencing. In all cases, universal primers were used to sequence the 16S rRNA genes from the studied bacteria.
The number of all sequences published in NCBI from the listed study (except for the guppy library, which was generated as part of this study) after removing sequences that classified to chloroplasts through the RDP Classifier tool (Wang et al 2007).
The number of OTUs derived at the 3% divergence level.
Diet indicates the fish’s diet at the time of collection as H, herbivore; O, omnivore and C, carnivore.
Habitat describes the type of water in which the fish was collected. (M) indicates that the species is known to migrate between freshwater and saltwater habitats.
Tissue type describes the portion of the gut used for the analysis as DT: digestive tract including contents and wall, IC: intestinal content, IW: intestinal wall, FP: faecal pellet, IM: intestinal mucus.
Rearing indicates whether the fish was collected in an artificial environment (A), including a farm or a laboratory, or from its wild habitat (W).
We also wished to evaluate these fish gut bacterial communities within the broader context of host-associated and free-living bacterial communities. To do this, we added these samples to a previously described data set that included 99 801 nonredundant 16S rRNA sequences from 464 samples compiled from 181 published studies (Ley et al. 2008b). This data set included diverse free-living (e.g. freshwater, saltwater, soils, etc.) and host-associated (e.g. mammals, insects, etc.) bacterial assemblages (Broad Dataset). The parsimony insertion tool in ARB was used to build a phylogenetic tree (Ludwig et al. 2004), providing us with a much faster and more feasible means for phylogenetic analysis than that used for our other data sets. Unweighted UniFrac distances were used for the PCoA.
Statistics
We hypothesized that fish with similar habitats and diets would harbour similar microbial communities. Therefore, we predicted smaller UniFrac distances for comparisons between fish libraries from the same habitats and trophic levels (vs. those from different groups). We tested this prediction using t-tests that compared distances within to those between fish libraries in different groups based on the UniFrac distances derived from the RaxML de novo tree. Because the distances are not independently distributed, we used Monte Carlo simulation (5 × 105 simulated data sets) to estimate distribution under the null hypothesis for each comparison (Edgington 1990; Skroch & Nienhuis 1995). Simulations were performed using matlab version 7.11 (Mathworks Inc., Natick, MA, USA). The Monte Carlo resampling simulations were performed on all the fish libraries.
Separate multivariate analyses of variance (manovas) were run on the first three PCoA vectors across all fish libraries, and separately for the culture-independent libraries alone, to determine whether fish habitat, order, rearing environment or diet had a significant association with the PCoA axes for each data set (Table 2). If the manovas showed that fish from different groups had significantly different PCoA values, the results of each univariate analysis of variance (ANOVA) were shown for the first three PCoA vectors. This helped to illustrate which axis or axes drove the differences visible in the PCoA plots. To compare per cent composition of representative sequences within fish libraries, the data were arc sin transformed prior to the ANOVA analysis. All MANOVA and ANOVA statistics to compare PCoAs were run on SPSS 18 (SPSS Inc., Chicago, IL, USA) and were analysed according to Pillai’s Trace as the test statistic.
Table 2.
Analyses on the environmental, ecological and evolutionary factors correlated with similarity in fish gut bacterial communities
| Source of variation |
Pillai’s trace |
Hypothesis d.f. |
Error d.f. |
F | P |
|---|---|---|---|---|---|
| (a) Multivariate analysis | |||||
| All Fish | |||||
| Trophic Level | 1.246 | 6 | 42 | 12.017 | 0.000* |
| Habitat | 0.774 | 6 | 42 | 4.419 | 0.002* |
| Rearing | 0.098 | 3 | 21 | 0.763 | 0.218 |
| Order | 1.661 | 21 | 51 | 3.011 | 0.001* |
| Culture independent | |||||
| Trophic Level | 1.340 | 6 | 28 | 9.473 | 0.000* |
| Habitat | 0.783 | 6 | 28 | 3.000 | 0.027* |
| Rearing | 0.159 | 3 | 14 | 0.884 | 0.197 |
| Order | 1.707 | 18 | 33 | 2.419 | 0.201 |
| Type III SS |
d.f. | F | P | ||
| (b) Univariate analysis | |||||
| Tropic level (all fish) | |||||
| PCoA1 | 0.717 | 2 | 22.494 | 0.000* | |
| PCoA2 | 0.071 | 2 | 1.177 | 0.327 | |
| PCoA3 | 0.28 | 2 | 10.807 | 0.001* | |
| Habitat (all fish) | |||||
| PCoA1 | 0.098 | 2 | 1.115 | 0.346 | |
| PCoA2 | 0.285 | 2 | 7.018 | 0.004* | |
| PCoA3 | 0.165 | 2 | 4.546 | 0.022* | |
| Trophic Level (culture independent) | |||||
| PCoA1 | 0.288 | 2 | 9.961 | 0.176 | |
| PCoA2 | 0.063 | 2 | 0.870 | 0.000* | |
| PCoA3 | 0.265 | 2 | 10.914 | 0.065 | |
| Habitat (culture independent) | |||||
| PCoA1 | 0.036 | 2 | 0.578 | 0.254 | |
| PCoA2 | 0.216 | 2 | 4.187 | 0.008* | |
| PCoA3 | 0.199 | 2 | 6.006 | 0.398 | |
P-values from MANOVA (a) and ANOVA (b) analyses, comparing axes of principal coordinates analysis based on UniFrac distances.
P-values significant at the 0.05 level.
Sequence analysis: identifying lifestyles of related bacteria
To determine how fish gut bacteria are related to those from other environments, we performed BLASTn searches against the nr/nt database. Sequences of the top BLASTn hits for each representative OTU in our data set were downloaded and aligned with our representative OTU sequences using Infernal within RDP (Cole et al. 2009; Nawrocki et al. 2009). The aligned data set was then used for a phylogenetic analysis, performed with RAxML 7.2.7 Black Box on the cipres web portal (Miller et al. 2010). No Lane mask was performed for this analysis because its goal was to decipher relationships among close relatives—discarding hypervariable regions would have, therefore, eliminated sequence data most valuable for this aim.
The generated phylogeny was uploaded to iTOL (Letunic & Bork 2007), along with an environmental data set that was used to facilitate visualization of the tree and classification of OTUs to clades with particular characteristics. In essence, each fish gut-associated sequence and each BLASTn hit was assigned to one of the following source environments: fish gut, fish (for sequences derived from unspecified organs or body parts other than the gut), mammalian gut, mammalian (for sequences derived from unspecified organs or body parts other than the gut), other animal, plant, environmental, artificial (deriving from a man-made or heavily altered habitat) or unspecified. Branches were colour coded to reveal the source habitat category, and the tree was subsequently examined, branch-by-branch, to determine the typical source habitats for the relatives of the studied fish gut bacteria.
Common clustering of fish gut phylotypes from our meta-analysis with free-living, aquatic phylotypes would suggest that fish gut bacteria reflect communities from their surrounding environment (and perhaps exchange frequently with these communities). In contrast, if fish gut phylotypes cluster with phylotypes from other fish or other vertebrates, we would conclude that fish harbour bacteria long-adapted for a symbiotic lifestyle. To quantify these trends, we identified clades containing the fish gut bacteria, examining the lifestyles and isolation sources of other microbes within these groups (Table S1, Supporting information). Each sequence from our meta-analysis was then classified into one of fourteen categories based on the habitats colonized by relatives within their respective clades (Table 3). After classifying all fish sequences in this manner, a bar graph was constructed to illustrate the category assignments for representative OTU sequences from each library. In doing so, we illustrated the typical lifestyles of the closest relatives to fish gut bacteria, shedding light on the origins of the microbial species that colonize the guts of the studied fish.
Table 3.
Lifestyles exhibited by the closest relatives of fish gut microbes. These categories are referred to in Fig. 4
| Category | Description |
|---|---|
| 1 | Vertebrate gut generalists: fish gut associates cluster within clades consisting of all or mostly (≥85% with n ≥ 10 clade members) bacteria from vertebrate guts |
| 2 | Animal and vertebrate gut generalists: fish gut associates belonged to clades containing some bacteria, but <85%, from mammalian or bird guts. All other bacteria within these clades were derived from nonvertebrate animals or nongut tissues from vertebrates |
| 3 | Fish gut generalists: fish gut associates belonged to lineages consisting only of microbes from the guts of fish. Each clade harboured bacteria from multiple host species |
| 4 | Fish gut specialists—multi-host population: fish gut associates belonged to lineages consisting only of microbes from the guts of fish. Each clade harboured bacteria from multiple host populations of a single species |
| 5 | Fish gut specialists—single-host population: fish gut associates belonged to lineages consisting only of microbes from the guts of fish. Each clade harboured bacteria from just one host population from one species |
| 6 | Animal generalists: fish gut associates grouped into lineages for which animals (e.g. invertebrates such as insects and corals; nongut habitats from fish and mammals) and fish guts were the only source habitats |
| 7 | Terrestrial environmental microbes: fish gut associates were most closely related to bacteria from terrestrial environments. |
| 8 | Terrestrial freshwater microbes: fish gut associates were most closely related to bacteria from freshwater environments |
| 9 | Marine environmental microbes: fish gut associates were most closely related to bacteria from marine or estuarine environments |
| 10 | Miscellaneous environmental microbes: fish gut associates were most closely related to bacteria from other types of environments (e.g. snow, ice, air) |
| 11 | Plant-associated microbes: fish gut associates were most closely related to bacteria isolated from plants |
| 12 | Microbes from artificial habitats: fish gut associates were most closely related to bacteria from a human-made or human-altered environment |
| 13 | Undefined, basal clade members: fish gut associates were basal to clades with mixed habitat sources |
| 14 | Microbes with unspecified habitats: fish gut associates were most closely related to bacteria with unspecified isolation sources |
Results
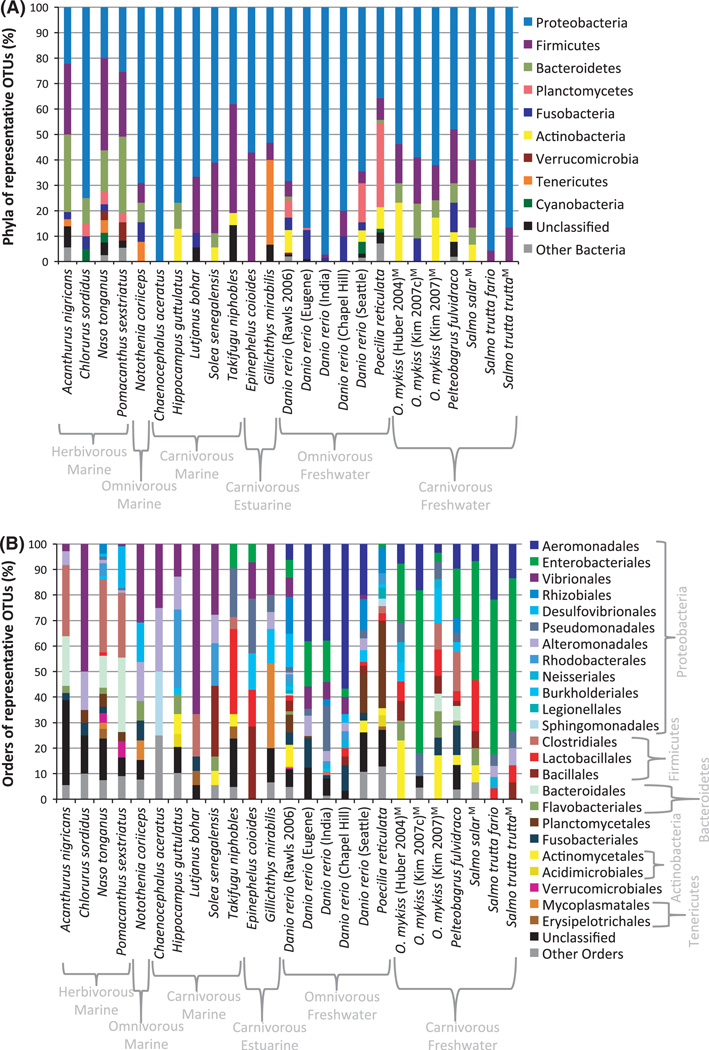
We obtained a total of 5199 sequences from 25 GenBank libraries and generated one guppy library of 112 sequences for this study. A total of 1054 representative sequences were selected from all of these libraries, with each serving as a unique representative of each 97% OTU group per host population (Table 1). Of the representative sequences, the dominant phylum was the Proteobacteria (Fig. 1), revealing that most species of fish gut bacteria are from this particular taxon as previously demonstrated (Rawls et al. 2006; see Nayak 2010 for review). The phyla found in fish guts and their mean prevalence (i.e. number of representative species for 97% OTU groups) within each library were as follows: Proteobacteria (62.51%), Firmicutes (15.2%), Bacteroidetes (6.04%), Actinobacteria (3.70%), Fusobacteria (2.88%), Planctomycetes (2.68%), Tenericutes (1.88%), Cyanobacteria (0.57%), Verrucomicrobia (0.45%), Spiro-chaetes (0.25%), TM7 (0.25%), Lentisphaerae (0.15%), Synergistetes (0.12%), Acidobacteria (0.11%), OP10 (0.07%), Chloroflexi (0.03%) and Nitrospira (0.03%).
Fig. 1.
(A) Bacterial species from 25 fish gut communities classified into the phylum level. Bar graphs for each library represent the percentage of species assigned to each phylum with ≥80% bootstrap confidence. Names of the fish hosts are listed along the horizontal axis. To differentiate between libraries of the same species, the authors of the relevant studies are listed in parentheses, while ‘CI’ indicates the sequences culture independently derived from the Kim et al. (2007) Oncorhynchus mykiss samples. The D. rerio libraries from the Roeselers et al. (2011) study are differentiated by location name in parentheses. (B) Bacterial OTUs from 25 fish gut communities classified to the order level. Bar graphs for each library represent the percentage of species assigned to each order with ≥80% bootstrap confidence.
When representative bacteria were classified into orders, a strong distinction emerged among fish from saltwater vs. freshwater habitats (Fig. 1). Specifically, freshwater fish harboured a greater proportion of Aeromonadales (two-tailed t-test, P = 0.000) and Enterobacteriales (two-tailed t-test, P = 0.000) species, while marine fish communities consisted of higher proportions of species from the Vibrionales (two-tailed t-test, P = 0.018). Among trophic levels, the herbivorous fish were enriched with Clostridiales (ANOVA, P = 0.005), Bacteroidales (ANOVA, P = 0.000) and Verrucomicrobiales (ANOVA, P = 0.003) while omnivorous fish were enriched with species from the Rhizobiales (ANOVA, P = 0.019), Fusobacteriales (ANOVA, P = 0.042) and Planctomycetales (ANOVA, P = 0.008). Both carnivores and omnivores tended to have more Desulfovibrionales (ANOVA, P = 0.036) and Aeromonadales (ANOVA, P = 0.020).
These patterns were also generally consistent with the composition of culture-independent libraries from a subset of the analysed fish for which OTU-abundance information was known (i.e. the sequence data were not de-replicated by picking OTUs before the analysis; Fig. S1, Supporting information). First, consistent with the enrichment of species-level OTUs from the Aeromonadales in freshwater fish guts, bacteria from this group were the most dominant in the complete freshwater gut libraries, making up a median of 40.1% of the sequences in these samples. No Aeromonadales were detected among the smaller subset of analysed marine hosts (i.e. those with full sequence libraries available in NCBI). Second, Vibrionales bacteria comprised a median of 3.6% of sequence reads from the analysed freshwater fish gut communities, compared to 69.8% for the analysed marine fish communities. Third, the most abundant orders in the marine herbivore gut communities were Bacteroidales and Clostridiales, which comprised a median of 12.7% and 22.6% of sequence reads per library. Bacteroidales were not found among omnivorous or carnivorous fish, while Clostridiales had a maximum representation of 9.1% with a median of 0% among these potential hosts. Combined, these trends highlight the robustness of our patterns across different analytical approaches.
Relationships between fish gut bacterial communities: Fish Dataset
The phylogeny generated from our maximum likelihood analysis (Fig. S2, Supporting information) was used for UniFrac analyses examining phylogenetic overlap between fish gut bacterial communities. The PCoA, performed using unweighted UniFrac distances, showed clustering based on trophic level, habitat salinity and method of sampling (i.e. culture-dependent vs. culture-independent) (Fig 2 and Fig S3, Supporting information).
Fig. 2.
Results of principal coordinates analysis based on pairwise unweighted UniFrac distances. Symbols representing individual communities are coloured by trophic level (green: herbivores, red: carnivores and yellow: omnivores). Shapes represent the habitats from which the samples were derived (circle: saltwater, triangle: estuarine, square: freshwater, diamond: migratory fish species between fresh and saltwater, all of which were sampled from freshwater). Open symbols represent cultured bacterial communities, while solid symbols represent communities studied through culture-independent means. Symbols with ‘D’ on the top left illustrate individual zebrafish libraries, while those with ‘O’ represent rainbow trout libraries.
Differences along PCoA axes were found to be significant according to trophic level and habitat (Table 2). More specifically, when considering the full data set, bacterial communities of fish from different trophic levels were significantly different across PCoA1 and PCoA3, while bacterial communities from the culture-independent subset were significantly different across PCoA2 according to trophic level. Bacterial communities of fish from different habitats were significantly different across PCoA2 and PCoA3 for the full data set and just PCoA2 for the culture-independent communities. For comparisons involving either the full data set or culture-independent data set only, bacterial communities did not differ based on the rearing environment (i.e. whether the fish were from artificial environments, such as a laboratory or aquaculture setting, or their natural habitats). The MANOVA comparing fish order was significant for the full data set but not for the culture-independent subset. There were marginal concerns about the sphericity of the data for the MANOVA analyses, which were used to explore whether fish having the same trophic level, habitat, rearing condition and order had more similar PCoA values, but because they were highly significant, the results were not likely to change very much because of the violation of the sphericity assumption.
When pairwise comparisons of UniFrac distances for all 25 fish libraries were analysed with Monte Carlo resampling, distances between different trophic levels were found to be significantly greater than those within trophic levels (P < 0.001, by Monte Carlo simulation). While the UniFrac distances did not differ significantly between vs. within habitats (P > 0.05, by Monte Carlo simulation), the trophic level × habitat interaction effect was significant (P < 0.001, by Monte Carlo simulation).
Relationships between fish gut communities and other free-living and host-associated bacterial communities: Broad Dataset
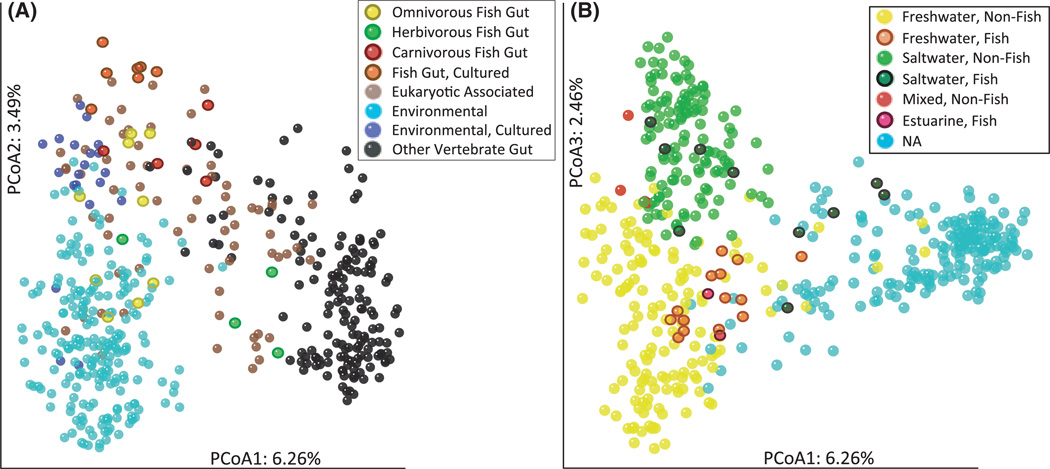
As seen for analyses on the Fish Dataset, PCoA analyses on the Broad Dataset phylogeny (not shown) revealed clustering of bacterial communities based on trophic level, salinity and method of sampling (i.e. whether the bacteria were cultured prior to sequencing) (Fig. 3). Yet, when the fish libraries were compared to those from other habitats, it was clear that they did not cluster together. Instead, they were dispersed among communities from vertebrate guts, other eukaryotes and various free-living environments (Fig. 3A).
Fig. 3.
(A) Principal coordinates analysis (PCoA) analysis illustrating similarity of bacterial communities from fish guts and other communities (Axes 1 and 2) across trophic levels and sampling methodologies. Fish are distributed among other bacterial communities, but they cluster with each other along trophic levels. (B) PCoA analysis illustrating similarity of bacterial communities from fish guts and other communities (Axes 1 and 3) based on habitat salinity. In this figure, PCoA 3 is shown because the difference in habitat salinity separates along this axis. The fish from estuarine habitats appear to be more similar to fish from freshwater, but the environmental communities from mixed salinity habitats cluster more with saltwater bacterial communities.
As has been previously described (Ley et al. 2008b), the first PCoA vector in this analysis separated free-living bacterial communities and those associated with most invertebrates, from those that associated with the gut of most vertebrates, particularly herbivorous and omnivorous mammals. Intermediate along this axis were bacterial communities associated with the termite gut, carnivorous mammals, and the human mouth and vagina. Interestingly, the fish gut samples were spread across this axis, with three of the four herbivorous fish gut communities (including Pomacanthus sexstriatus, Acanthurus nigricans and Naso tonganus) clustering closest to those from mammalian guts. The carnivorous fish mostly clustered near carnivorous mammals, in an intermediate position along this axis. In contrast, the omnivorous fish all clustered near the free-living/invertebrate-associated communities rather than with omnivorous mammals.
While PCoA2 separated cultured bacterial communities from those that were acquired by culture-independent techniques (Fig. 3A), the variation across PCoA3 correlated with differences in habitat salinity—microflora from saltwater environments tended to have higher PCoA3 values (green points, Fig. 3B) than those from freshwater habitats (yellow/orange points, Fig. 3B). Along PCoA3, the fish from estuarine environments tended to group with the freshwater fish even though the mixed salinity, nonfish environmental samples (from estuaries and an intertidal hotspring) grouped more with the saltwater samples. The herbivorous fish communities were all from marine hosts and were generally found on an intermediate position along PCoA3. Future sampling designs that include fish with different diets from all salinity classes would therefore be very useful.
Cluster analysis (Fig. S4, Supporting information) performed using the UniFrac distance matrix helped to further reveal the diversity of fish communities—which were dispersed across the generated dendrogram. All herbivorous fish communities grouped together on this tree, along with a community from one marine carnivore.
This cluster was, in turn, related to communities from the guts of mammals and invertebrates (earthworms, herbivorous beetles and termites) and from anoxic rice paddy soil. Other clusters containing fish communities tended to group with invertebrate and vertebrate gut communities, with two clusters showing closer affinities to environmental bacteria (i.e. one cluster containing the guppy and two zebrafish communities, and one consisting of communities from the seahorse, Hippocampus gu-tulatus, and the Senegalese sole, Solea senegalensis).
Describing the lifestyles of related bacteria
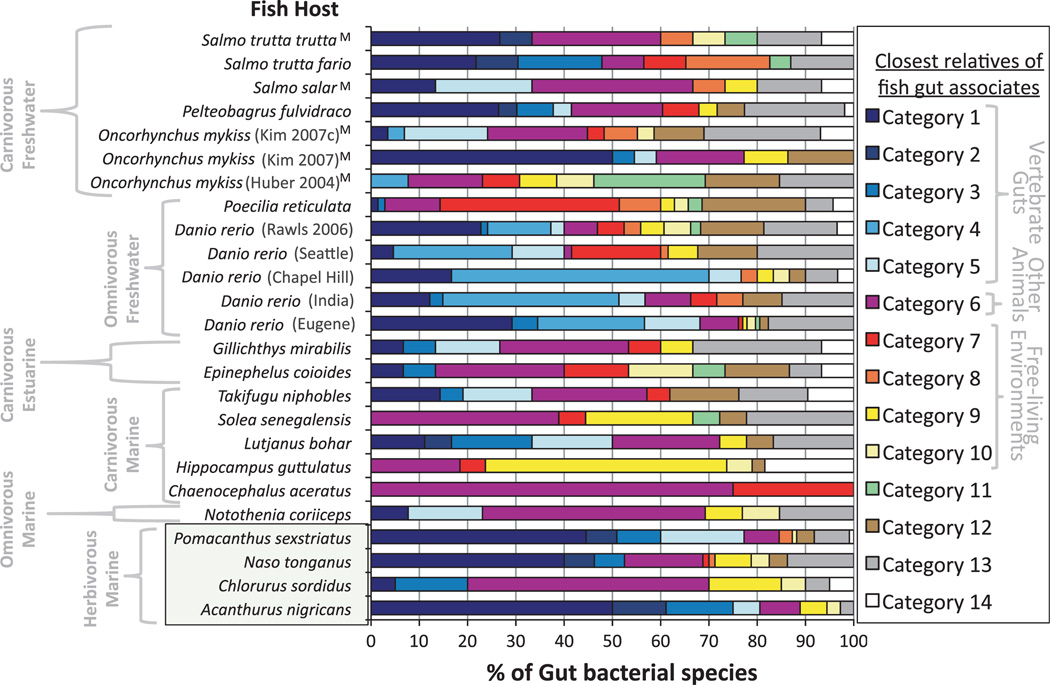
To place the gut bacteria of fish into a broader evolutionary context, we constructed a maximum likelihood phylogeny of all representative sequences along with their top BLASTn hits. Consideration of clade membership in the light of habitat source (detailed fully in Table S1, Supporting information) revealed that over half of the species from 17/25 fish libraries had closest phylogenetic affinity to bacteria from vertebrate guts (blue bars in Fig. 4 corresponding to categories 1–5). Three of four herbivorous gut communities were unique in that most of their representative bacterial species belonged to lineages consisting of microbes from the guts of other vertebrates—namely birds and mammals (categories 1 and 2). Bacteria from these three fish hosts grouped into a diverse range of gut-associated clades (17, 21 and 27 different vertebrate-associated clades across A. nigricans, P. sexstriatus and N. tong-anus, respectively), most commonly within the Bacteroidia, Clostridia and Verrucomicrobia. Many of these consisted only of gut bacteria from one to three of these fish plus microbes from various herbivorous and omnivorous mammalian counterparts. Another common class of fish gut associates (category 6) showed closest phylogenetic affinity to microbes isolated from other animals (mostly insects and corals) and nongut tissues of vertebrates. Free-living bacteria with environmental lifestyles (red, orange and yellow bars in Fig. 4), however, were comparatively rare among the closest relatives of fish gut bacteria. In fact, only 2/25 studied fish communities harboured a majority species with affinities to environmental microbes (categories 7–10).
Fig. 4.
Lifestyles of bacteria related to fish gut microbes. After constructing a phylogeny with 16S rRNA sequences from bacteria of fish guts and their top BLASTn hits, all representative sequences (one per 97% OTU) were categorized based on their relatedness to microbes from vertebrate guts, other animals (or vertebrate tissues), plants and both natural environmental and artificial habitats. The percentage of species (i.e. 97% OTUs) from each library falling into one of the 14 designated categories (see Table 3 for classification criteria) is illustrated using colour-coded bar graphs. For further clarity, trophic level and habitat type for studied fish hosts are indicated to the left of the Y-axis (herbivorous fish are highlighted at the bottom of the graph), while general groups of bacterial lifestyles are revealed to the right of the category legend. ‘M‘ symbols after host species names indicate that these species are known to migrate between fresh and saltwater. Note that herbivores show enrichment for microbes from categories 1 and 2, indicating that they are closely related to gut associates of other vertebrates—mostly mammals.
While we identified many clades of vertebrate gut associates, the two largest contained 42 and 51 species and were distributed across 12 and 10 fish host libraries, respectively (Table S1, Supporting information). The first (clade 44 from the Fusobacteria) corresponded to the Cetobacterium group identified by Roeselers et al. 2011. Members of this group were predominantly found in freshwater fish. Bacteria from the second of these lineages (clade 36 from the Gammapro-teobacteria) were confined to freshwater fish and polar bear faeces. BLASTn searches of selected representatives revealed that these were related to Plesiomonas shingello-ides (Gammaproteobacteria: Enterobacteriales), a bacterium that has previously been found in freshwater fish, mammals and bodies of freshwater.
Discussion
It has previously been shown that fish gut communities vary within species because of factors such as dietary input, season, developmental stage and the surrounding habitat (Luczkovich & Stellwag 1993; Hansen & Olafsen 1999; Nayak 2010). This work has also shown that microbes differ on broader scales between freshwater and saltwater fish, with bacteria such as Aeromonas and Plesiomonas (Gammaproteobacteria) enriched in freshwater specimens and anadromous fish collected from freshwater habitats, and Vibrio exhibiting greater prevalence in marine species (see Nayak 2010 for review). But until now, most research on fish gut communities has relied on cultivation, while most culture-independent studies have not looked beyond a small number of sampled hosts (but see Roeselers et al. 2011). As such, our consideration of the ecological and evolutionary factors that correlate with community composition has shed novel and quantitative insights into the factors that structure bacterial gut communities across the most ancient and diverse group of vertebrates.
Overall, our findings indicate that host trophic level, habitat and possibly host phylogeny play a role in shaping gut microbial communities of fish. Our results also conform to the expectation that cultivation-based studies yield a distorted picture of the actual community composition (Fig 2 and Fig 3). The assertion that salinity and host relatedness shape fish microbial communities was also suggested by the Roeselers et al. (2011) study, which included a subset of the fish bacterial communities analysed here. Although there is some agreement between our findings, the results here did show weaker clustering of the zebrafish microbial communities (Fig 2 and Fig S3, Supporting information). In spite of these differences, the tendency for conspecific fish (i.e. zebrafish and rainbow trout) to harbour similar gut bacteria suggests that core gut communities may be common across a broader range of fish.
Although our analyses focused on representative species from each host library, considerations of whole libraries yielded trends consistent with effects of habitat salinity and trophic level (Fig. S1, Supporting information). Analyses alongside communities from other hosts and habitats similarly revealed that these patterns were robust (Fig. 3), as did statistics performed exclusively on culture-independent libraries (Table 2). Furthermore, potential biases stemming from differences in primers or sampling effort across studies are not likely explanations for the patterns observed in our analyses (see Supporting information for discussion).
While our study is among the first to quantify the effects of ecological and environmental factors on fish gut communities, another attribute of this work is that it provides the first systematic investigation into the types of microbes and communities showing closest affinities to the gut flora of fish. UniFrac PCoA and cluster analyses (Fig 3A and Fig S4, Supporting information) indicated that most fish gut communities bear resemblance to those from vertebrate and invertebrate guts, with few showing greatest similarity to environmental microbial assemblages. Phylogenetic analysis of representative OTUs and their closest relatives similarly uncovered frequent clustering of fish gut associates with symbionts of animals, including vertebrate gut colonists (Fig. 4). We must, therefore, consider the possibility that a large percentage of the bacterial species in fish guts may make their living as more specialized members of symbiotic communities rather than as free-living, environmental bacteria.
While the above findings hint at symbiotic origins for many of the microbes found in fish guts, it should be noted that most freshwater and saltwater gut communities clustered, respectively, with those from other freshwater and saltwater environments. This resembles previous findings on the importance of salinity in structuring communities from free-living habitats (Lozupone & Knight 2007). It is also consistent with the colonization of the gut habitat by at least some environmental bacteria, including common gut associates such as Vibrio and Aeromonas. This trend indicates that bacteria adapted to associate with the guts of marine fish species must also be adapted to tolerate the saline environment of their hosts. Marine herbivores provided an exception to this trend (Fig. 1B), harbouring few of the typical environmental marine bacteria yet many close relatives of bacteria from mammalian guts.
Gut communities of fish and mammals
Comparisons of fish gut bacterial communities with those from other environments, including lakes, soils, oceans and eukaryotic hosts, reveal that fish gut bacteria can be quite distinct between different hosts (Fig 3A and Fig S4, Supporting information). It will, thus, be interesting to determine whether gut communities of other non-mammalian vertebrates with closer related-ness to mammals—amphibians, birds and reptiles—show similar dispersion patterns or whether their communities appear derived and unique like those of the mammals.
In spite of the diverse gut communities found across fish, most were united by the dominance of microbes from the Proteobacteria (Fig 1 and S1, Supporting information), in keeping with previous findings (Huber et al. 2004; Rawls et al. 2006; Wu et al. 2010; Roeselers et al. 2011). In stark contrast, Proteobacteria are outnumbered within healthy adult mammalian guts by species from the Bacteroidetes and Firmicutes (Ley et al. 2008a). This distinction between dominant phyla present within fish and mammals is likely driven by evolved differences in the selectivity of the gut environment (Rawls et al. 2006).
An exception to this pattern was, again, found for three of the four studied marine herbivores—P. sexstriatus, N. tonganus and A. nigricans (see Supporting information for a description of the exception). Proteo-bacterial species comprised <30% of the OTUs from their gut communities, which were instead unique in their combined enrichment of species from the Bacteroidetes and Firmicutes (>50% of the OTUs in each; Fig. 1). A consideration of full libraries (vs. only representative species) for these fish confirmed the numerical abundance of these species (Fig. S1, Supporting information), indicating that the trend was not simply because of the presence of many rare varieties from these phyla. As this resemblance to mammalian guts was also reflected in our UniFrac analyses (Fig 3A and Fig S4, Supporting information), our findings hint that the microbial communities of these fish may function similarly to those in their mammalian counterparts. Indeed, the presence of short-chain fatty acids in the guts of marine herbivores (Clements & Choat 1995) suggests that herbivorous fish and mammals are united through the process of gut fermentation.
A need for greater sampling of symbiotic gut communities across the fish
The fish phylogeny exhibits great species diversity, but is also diverse in the varieties of exhibited morphologies, physiologies and ecologies. Fish are found in drastically varied environments, including deep ocean habitats with extreme pressure (Carney 2005; Zintzen et al. 2011), hypersaline salt ponds (Carpelan 1957; Lenanton 1977) and many habitats in between these extremes. Aside from their varied habitats, fish can occupy diverse trophic niches, ranging from specialists on plankton or fish scales to omnivorous generalists and piscivorous carnivores (Gerking 1994). Fish can also have vastly distinct morphological features within their intestinal tracts—some species have additional chambers and structures, while others have lost intestinal features such as stomachs (Jobling 1995).
Given the great variation exhibited across ichthyofauna, it is important to note that a relatively narrow range of habitats and ecologies are represented in our meta-analysis because of the limited availability of sequence-based data sets. Furthermore, of the 62 extant orders and 515 extant fish families, just 8 orders and 15 families are represented in this study, and all come from the Actinopterygii (ray-finned fish), one of two classes of the bony fish. When we consider that fish comprise slightly more than one-half of all recognized living vertebrate species (Nelson 2006), while representing the first organisms to evolve adaptive immune systems, their vast ecological, physiological and morphological diversities combine to make them an attractive system for the exploration of symbiosis. We advocate that such sampling begin by tackling: (i) more basal groups of fish, including jawless and cartilaginous forms, and (ii) members of the Sarcopterygii, including the lungfishes that represent the closest relatives of tet-rapods. A special emphasis should also be made to study close relatives from different trophic levels or habitats. This combined approach will help to better understand how these communities vary, the forces that drive this variation and how they have changed over time. Targeted sampling will also help to disentangle the effects of habitat, trophic and genetic similarities on the composition of gut communities.
Explorations on the consequences of symbiosis
In further explorations of fish and their gut bacterial communities, it will also be essential to paint a clearer picture of the significance of symbiosis and its roles in fish ecology and evolution. Indeed, certain microbial gut associates may confer fitness advantages, affecting the evolutionary trajectories of the creatures that harbour them. Interestingly, prior studies have shown that fish gut bacteria play roles in nutrition, immunity and defence (see Nayak 2010 for review). For instance, gut bacteria shape fish nutrition through modulating cholesterol metabolism (Rawls et al. 2004), the provisioning of vitamins (Sugita et al. 1991) and the synthesis of digestive enzymes that degrade a variety of substrates (MacDonald et al. 1986; Bairagi et al. 2002; Sugita & Ito 2006). The detection of short-chain fatty acids (SCFAs) in the intestines of marine herbivores also indicates digestive and nutritional roles for their gut bacteria (Clements & Choat 1995). Furthermore, it is known that the gut microbiome is crucial for the development of both the gut and immune system within some fish hosts (Perez et al. 2010). With the increasing availability and accessibility of molecular technologies, novel and high-throughput approaches can be utilized to gain an increased understanding of the functional roles of these fish microbes. A metagenomic approach, for instance, would provide insight into the consequences of symbioses with gut bacteria, revealing whether the functional differences among gut communities from mammals at different trophic levels (Muegge et al. 2011) are also found within fish.
Explorations of the mechanisms of symbiosis and community assembly
Another unexplored avenue relating to fish and their gut communities relates to the mechanisms of symbiosis and, in particular, the means by which symbiotic communities assemble within the gut. The gut habitats of various organisms have complex factors that determine which microbes flourish and which are extirpated. Factors affecting environmental bacteria, such as pH (Fierer & Jackson 2006; Chu et al. 2010), can also affect microbial colonization in animal alimentary tracts (Duncan et al. 2009). Immune systems can also discriminate between symbiotic and pathogenic bacteria (Cerf-Bensussan & Gaboriau-Routhiau 2010; Atarashi et al. 2011), providing a means of selecting for or against particular microbial constituents. Like mammals, fish do show a similar capacity to retain characteristic microbes, as illustrated by Rawls et al. (2006) who seeded germ-free zebrafish guts with gut flora from mice. Over time, the resulting gut communities shifted to look more like those from a typical zebrafish gut, with Proteobacteria proliferating at the expense of Bacteroidetes and Firmicutes. This implies that the autochthonous microbes are not a passive reflection of their seeding communities and agrees with our finding that most bacterial species in the guts of fish are not closest to environmental bacteria, but to those found as symbionts of other animals.
So what forces shape the colonization of fish guts? The prevailing view of the bacterial colonization of a fish gut is that it depends on the bacterial composition of rearing water and diet, which directly seeds the gut of fry (Nayak 2010). As fish become active feeders, the bacterial load in their guts increases, and autochthonous bacteria become attached to the intestinal mucus or epithelial surface forming the resident gut inhabitants, while allochthonous or transient bacteria fail to become established (Nayak 2010).
The belief that fish gut bacterial communities are entirely seeded from the ambient bacterioplankton, the bacteria in their food or those in the environment (Nayak 2010), however, neglects the fact that a number of bacteria have only been isolated from fish guts. An example of such a fish-specific symbiont includes one of the largest bacteria known, Epulopiscium fishelsoni. This microbe can reach sizes greater than 600 × 80 µm and has only been found in surgeonfish (Angert et al. 1993). Because no free-living forms of E. fishelsoni have been found, alternative means of bacterial acquisition aside from the environment should be considered. Additionally, not all fish develop from externally deposited eggs. Two of the nineteen fish species studied in this meta-analysis have alternative reproductive strategies, including the guppy, which is oviparous, and H. guttulatus, which relies on male brooding of offspring (Breden et al. 1999; Foster & Vincent 2004). Cichlid species also exhibit parental care, including a number of species that brood their offspring in their mouths (Goodwin et al. 1998). At least three cichlid species, including Etroplus maculatus, Symphysodon discus and Cichlasoma citrinellum, have been observed exhibiting a specific behaviour in which young fish contact a parent ‘as though feeding from the surface of their bodies’, which Noakes & Barlow (1973) speculated is a way for the fish to acquire mucus and/or other essential material. Although unproven, this type of behaviour could clearly serve as a mechanism for bacterial transmission, which may be vital for growth and development, as few young survive when isolated from their parents (Hildemann 1959; Ward & Barlow 1967). Such behaviours and reproductive strategies introduce the possibility for more direct modes of transmission resembling those seen in mammals. Furthermore, within insects, gut bacteria have been isolated from surface-sterilized eggs, suggesting that vertical transmission may occur even in egg-laying animals (Pinto-Tomás et al. 2011). Combined with the enrichment of symbiotic (and mammalian gut) bacteria within the guts of fish, these findings suggest a need for further studies on nontraditional means of microbial acquisition.
Fish may also acquire gut bacteria through cyclic transmission, where hosts obtain their symbiotic community from the environment. This type of transmission can promote prolonged associations between bacteria and hosts, especially when the adult population seeds the environment with their bacteria (Mcfall-Ngai 1998). McFall-Ngai suggests also that over evolutionary time, selection could occur within a host’s development that leads it to gain appropriate environmental bacteria through recognition and adherence mechanisms within its cell surfaces (Mcfall-Ngai 1998). Such a phenomenon could provide an additional mechanism to promote the differentiation of gut communities among fish from similar environments (e.g. marine herbivores vs. omnivores and carnivores).
As one can see, our current understanding of the mechanisms of gut community assembly stems from a small number of insights, often derived from studies in other systems. This wide-open topic should, thus, provide many opportunities for research on the roles of host immunity and physiology, host genotype (e.g. Whitham et al. 2008) and microbial interactions (e.g. Keller & Surette 2006) in shaping the types of communities in fish guts (see Supporting information for further discussion).
Conclusions
In conclusion, fish appear to have more complex relationships with their gut bacteria than previously described, and their gut microbiome may not be a simple reflection of the microbes from their environment. This seems especially true for species of marine herbivorous fish, whose inhabitants show close ties to microbes from mammalian guts. Given their antiquity and the overlap of their gut communities with those from mammals, we propose that fish may have served as the first vertebrate hosts for many of the microbes that have come to proliferate in the guts of mammals. This would imply that because fish were the first hosts with adaptive immune systems encountered by these bacteria, the guts of several fish lineages served as training grounds for microbes that would eventually evolve relationships with mammals. Of course, it is alternatively possible that the sharing of similar gut bacteria between mammals and fish could reflect the outcomes of convergent evolution with fish and mammals separately domesticating related free-living lineages. Discriminating between these hypotheses will require more thorough sampling of gut communities across the fish in search of a tendency for their gut microbes to fall out as basal branches within larger clades of vertebrate gut symbionts. Should this prediction be met, humans can count their gut bacteria among the many other attributes that constitute their ‘inner fish’ (Shubin 2008).
Supplementary Material
Acknowledgements
We thank Yi Hu and Drew McQuade for assistance with data collection, the National Science Foundation funded Frontiers in Integrative Biological Research (FIBR) Guppy Project (DEB-0623632EF), and in particular Eugenia Zandona` and Rana El-Sabaawi for providing the guppy samples used in this study, Katriina Ilves for useful consultation on the fish phylogeny and Esther Angert for helpful discussion. We would also like to thank the L.D. Betz Chair of Environmental Science for the support of K.E.S. during this research.
Biography
K.E.S. is a doctoral candidate investigating gut microbial communities, trophic ecology and digestive physiology of fish. S.D.E. is a doctoral candidate focusing on bioinformatics tool development for metagenomic applications. M.O.C. studies biophysical constraints on the ecology of vertebrates and quantitative theoretical ecology. G.L.R. investigates improvements to ecological ordination methods and taxonomic classification from next-generation sequencers. C.A.L. and R.K. are bioinfor-matic scientists and microbiologists with an interest in the ecology and evolution of human gut bacteria and global patterns in bacterial diversity. S.S.K. studies trophic ecology and food web dynamics in aquatic ecosystems. J.A.R. studies the ecology and evolution of symbioses between animals and microbes.
Footnotes
Data accessibility
DNA sequences of Poecilia reticulata gut bacteria: GenBank accession numbers JQ253406–JQ253517. Additional information regarding the metadata included in this paper is available in Tables S1 and S2 (Supporting information).
Supporting information
Additional supporting information may be found in the online version of this article.
Table S1 Habitats, lifestyles, and phylogenetic affiliations of fish gut bacteria and their closest GenBank relatives.
Table S2 List of primers used in all studies included in this meta-analysis.
Fig. S1 Ordinal classifications of sequences from whole libraries of fish that were derived from culture independent methods.
Fig. S2 Phylogenetic tree of all representative fish gut bacterial sequences used in the study.
Fig. S3 Principal coordinates analysis based on pairwise unweighted UniFrac distances for bacterial communities from culture-independent data only.
Fig. S4 UniFrac cluster analysis dendrogram illustrating similarity of fish gut microbiota to communities from other hosts and habitats.
Fig. S5 Principal coordinates analysis as shown in figure 2 with each sample labeled by the initials of each fish species and genus.
Please note: Wiley-Blackwell are not responsible for the content or functionality of any supporting information supplied by the authors. Any queries (other than missing material) should be directed to the corresponding author for the article.
References
- Angert ER, Clements KD, Pace NR. The largest bacterium. Nature. 1993;362:239–241. doi: 10.1038/362239a0. [DOI] [PubMed] [Google Scholar]
- Atarashi K, Tanoue T, Shima T, et al. Induction of colonic regulatory T cells by indigenous Clostridium species. Science. 2011;331:337–341. doi: 10.1126/science.1198469. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bäckhed F, Ding H, Wang T, et al. The gut microbiota as an environmental factor that regulates fat storage. Proceedings of the National Academy of Sciences. 2004;101:15718–15723. doi: 10.1073/pnas.0407076101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bairagi A, Ghosh KS, Sen SK, Ray AK. Enzyme producing bacterial flora isolated from fish digestive tracts. Aquaculture International. 2002;10:109–121. [Google Scholar]
- Balcázar JL, Lee NM, Pintado J, Planas M. Phylogenetic characterization and in situ detection of bacterial communities associated with seahorses (Hippocampus guttulatus) in captivity. Systematic and Applied Microbiology. 2010;33:71–77. doi: 10.1016/j.syapm.2009.11.005. [DOI] [PubMed] [Google Scholar]
- Bano N, DeRae Smith A, Bennett W, Vasquez L, Hollibaugh JT. Dominance of Mycoplasma in the guts of the Long-Jawed Mudsucker, Gillichthys mirabilis, from five California salt marshes. Environmental Microbiology. 2007;9:2636–2641. doi: 10.1111/j.1462-2920.2007.01381.x. [DOI] [PubMed] [Google Scholar]
- Breden F, Ptacek MB, Rashed M, Taphorn D, Figueiredo CA. Molecular phylogeny of the live-bearing fish genus Poecilia (Cyprinodontiformes: Poeciliidae) Molecular Phylogenetics and Evolution. 1999;12:95–104. doi: 10.1006/mpev.1998.0600. [DOI] [PubMed] [Google Scholar]
- Carney RS. Zonation of deep biota on continental margins. Oceanography and Marine Biology: An Annual Review. 2005;43:211–278. [Google Scholar]
- Carpelan LH. Hydrobiology of the Alviso salt ponds. Ecology. 1957;38:376–390. [Google Scholar]
- Cerf-Bensussan N, Gaboriau-Routhiau V. The immune system and the gut microbiota: friends or foes? Nature Reviews Immunology. 2010;10:735–744. doi: 10.1038/nri2850. [DOI] [PubMed] [Google Scholar]
- Chu H, Fierer N, Lauber CL, et al. Soil bacterial diversity in the Arctic is not fundamentally different from that found in other biomes. Environmental Microbiology. 2010;12:2998–3006. doi: 10.1111/j.1462-2920.2010.02277.x. [DOI] [PubMed] [Google Scholar]
- Clements KD, Choat JH. Fermentation in tropical marine herbivorous fishes. Physiological Zoology. 1995;68:355–378. [Google Scholar]
- Cole JR, Wang Q, Cardenas E, et al. The Ribosomal Database Project: improved alignments and new tools for rRNA analysis. Nucleic Acids Research. 2009;37:D141–D145. doi: 10.1093/nar/gkn879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DeSantis TZ, Hugenholtz P, Keller K, et al. NAST: a multiple sequence alignment server for comparative analysis of 16S rRNA genes. Nucleic Acids Research. 2006;34:W394–W399. doi: 10.1093/nar/gkl244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Douglas AE. Nutritional interactions in insect-microbial symbioses: aphids and their symbiotic bacteria Buchnera. Annual Review of Entomology. 1998;43:17–37. doi: 10.1146/annurev.ento.43.1.17. [DOI] [PubMed] [Google Scholar]
- Duncan SH, Louis P, Thomson JM, Flint HJ. The role of pH in determining the species composition of the human colonic microbiota. Environmental Microbiology. 2009;11:2112–2122. doi: 10.1111/j.1462-2920.2009.01931.x. [DOI] [PubMed] [Google Scholar]
- Edgington E. Randomization Tests. Marcel Dekker, Inc., New York City; New York: 1990. [Google Scholar]
- Fierer N, Jackson RB. The diversity and biogeography of soil bacterial communities. Proceedings of the National Academy of Sciences. 2006;103:626–631. doi: 10.1073/pnas.0507535103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Foster SJ, Vincent ACJ. Life history and ecology of seahorses: implications for conservation and management. Journal of Fish Biology. 2004;65:1–61. [Google Scholar]
- Fraysse N, Couderc F, Poinsot V. Surface polysaccharide involvement in establishing the rhizobium-legume symbiosis. European Journal of Biochemistry. 2003;270:1365–1380. doi: 10.1046/j.1432-1033.2003.03492.x. [DOI] [PubMed] [Google Scholar]
- Gerking SD. Feeding Ecology of Fish. Academic Press: San Diego; 1994. [Google Scholar]
- Gilbert JA, Field D, Swift P, et al. The seasonal structure of microbial communities in the Western English Channel. Environmental Microbiology. 2009;11:3132–3139. doi: 10.1111/j.1462-2920.2009.02017.x. [DOI] [PubMed] [Google Scholar]
- Gilbert JA, Steele JA, Caporaso JG, et al. Defining seasonal marine microbial community dynamics. ISME J. 2012;6:298–308. doi: 10.1038/ismej.2011.107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goodwin NB, Balshine-Earn S, Reynolds JD. Evolutionary transitions in parental care in cichlid fish. Proceedings of the Royal Society of London Series B: Biological Sciences. 1998;265:2265–2272. doi: 10.1098/rstb.2001.0930. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hamady M, Lozupone C, Knight R. Fast UniFrac: facilitating high-throughput phylogenetic analyses of microbial communities including analysis of pyrosequencing and PhyloChip data. ISME Journal. 2009;4:17–27. doi: 10.1038/ismej.2009.97. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hansen GH, Olafsen JA. Bacterial interactions in early life stages of marine cold water fish. Microbial Ecology. 1999;38:1–26. doi: 10.1007/s002489900158. [DOI] [PubMed] [Google Scholar]
- Hildemann WH. A cichlid fish, Symphysodon discus, with unique nurture habits. The American Naturalist. 1959;93:27–34. [Google Scholar]
- Huber I, Spanggaard B, Appel KF, et al. Phylogenetic analysis and in situ identification of the intestinal microbial community of rainbow trout (Oncorhynchus mykiss, Walbaum) Journal of Applied Microbiology. 2004;96:117–132. doi: 10.1046/j.1365-2672.2003.02109.x. [DOI] [PubMed] [Google Scholar]
- Jobling M. Environmental Biology of Fishes. London: Chapman & Hall; 1995. [Google Scholar]
- Keller L, Surette MG. Communication in bacteria: an ecological and evolutionary perspective. Nature Reviews Microbiology. 2006;4:249–258. doi: 10.1038/nrmicro1383. [DOI] [PubMed] [Google Scholar]
- Kim DH, Brunt J, Austin B. Microbial diversity of intestinal contents and mucus in rainbow trout (Oncorhynchus mykiss) Journal of Applied Microbiology. 2007;102:1654–1664. doi: 10.1111/j.1365-2672.2006.03185.x. [DOI] [PubMed] [Google Scholar]
- Lachnit T, Meske D, Wahl M, Harder T, Schmitz R. Epibacterial community patterns on marine macroalgae are host-specific but temporally variable. Environmental Microbiology. 2010;13:655–665. doi: 10.1111/j.1462-2920.2010.02371.x. [DOI] [PubMed] [Google Scholar]
- Lane DJ. 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M, editors. Nucleic Acid Techniques in Bacterial Systematic. New York: Wiley, New York City; 1991. [Google Scholar]
- Lenanton RCJ. Fishes from the hypersaline waters of the stromatolite zone of Shark Bay, Western Australia. Copeia. 1977;1977:387–390. [Google Scholar]
- Letunic I, Bork P. Interactive Tree Of Life (iTOL): an online tool for phylogenetic tree display and annotation. Bioinformatics. 2007;23:127–128. doi: 10.1093/bioinformatics/btl529. [DOI] [PubMed] [Google Scholar]
- Ley RE, Hamady M, Lozupone C, et al. Evolution of mammals and their gut microbes. Science. 2008a;320:1647–1651. doi: 10.1126/science.1155725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ley RE, Lozupone CA, Hamedy M, Knight R, Gordon JI. Worlds within worlds: evolution of the vertebrate gut microbiota. Nature Reviews Microbiology. 2008b;6:776–788. doi: 10.1038/nrmicro1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lozupone C, Knight R. UniFrac: a new phylogenetic method for comparing microbial communities. Applied and Environmental Microbiology. 2005;71:8228–8235. doi: 10.1128/AEM.71.12.8228-8235.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lozupone CA, Knight R. Global patterns in bacterial diversity. Proceedings of the National Academy of Sciences. 2007;104:11436–11440. doi: 10.1073/pnas.0611525104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lozupone C, Hamady M, Knight R. UniFrac – an online tool for comparing microbial community diversity in a phylogenetic context. BMC Bioinformatics. 2006;7:371. doi: 10.1186/1471-2105-7-371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luczkovich JJ, Stellwag EJ. Isolation of cellulolytic microbes from the intestinal tract of the pinfish, Lagodon rhomboides: size-related changes in diet and microbial abundance. Marine Biology. 1993;116:381–388. [Google Scholar]
- Ludwig W, Strunk O, Westram R, et al. ARB: a software environment for sequence data. Nucleic Acids Research. 2004;32:1363–1371. doi: 10.1093/nar/gkh293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- MacDonald NL, Stark JR, Austin B. Bacterial microflora in the gastro-intestinal tract of Dover sole (Solea solea L.), with emphasis on the possible role of bacteria in the nutrition of the host. FEMS Microbiology Letters. 1986;35:107–111. [Google Scholar]
- Martin-Antonio B, Manchado M, Infante C, et al. Intestinal microbiota variation in Senegalese sole (Solea senegalensis) under different feeding regimes. Aquaculture Research. 2007;38:1213–1222. [Google Scholar]
- Mcfall-Ngai MJ. The development of cooperative associations between animals and bacteria: establishing detente among domains. American Zoologist. 1998;38:593–608. [Google Scholar]
- McFall-Ngai M, Ruby E. Symbiont recognition and subsequent morphogenesis as early events in an animal-bacterial mutualism. Science. 1991;254:1491–1494. doi: 10.1126/science.1962208. [DOI] [PubMed] [Google Scholar]
- Miller M, Holder M, Vos R, et al. The CIPRES Portals. 2010 [Google Scholar]
- Moran NA, McCutcheon JP, Nakabachi A. Genomics and evolution of heritable bacterial symbionts. Annual Review of Genetics. 2008;42:165–190. doi: 10.1146/annurev.genet.41.110306.130119. [DOI] [PubMed] [Google Scholar]
- Muegge BD, Kuczynski J, Knights D, et al. Diet drives convergence in gut microbiome functions across mammalian phylogeny and within humans. Science. 2011;332:970–974. doi: 10.1126/science.1198719. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nawrocki EP, Kolbe DL, Eddy SR. Infernal 1.0: inference of RNA alignments. Bioinformatics. 2009;25:1335–1337. doi: 10.1093/bioinformatics/btp157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nayak SK. Role of gastrointestinal microbiota in fish. Aquaculture Research. 2010;41:1553–1573. [Google Scholar]
- Nelson JS. Fishes of the World. 4th edn. Hoboken, New Jersey: John Wiley & Sons, Inc.; 2006. [Google Scholar]
- Noakes D, Barlow G. Ontogeny of parent-contacting behavior in young Cichlasoma citrinellum (Pisces, Cichlidae) Behaviour. 1973;46:221–255. [Google Scholar]
- Perez T, Balcazar JL, Ruiz-Zarzuela I, et al. Host-microbiota interactions within the fish intestinal ecosystem. Mucosal Immunology. 2010;3:355–360. doi: 10.1038/mi.2010.12. [DOI] [PubMed] [Google Scholar]
- Pinto-Tomás AA, Sittenfeld A, Uribe-Lorío L, et al. Comparison of midgut bacterial diversity in tropical caterpillars (Lepidoptera: Saturniidae) fed on different diets. Environmental Entomology. 2011;40:1111–1122. doi: 10.1603/EN11083. [DOI] [PubMed] [Google Scholar]
- Rawls JF, Samuel BS, Gordon JI. Gnotobiotic zebrafish reveal evolutionarily conserved responses to the gut microbiota. Proceedings of the National Academy of Sciences. 2004;101:4596–4601. doi: 10.1073/pnas.0400706101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rawls JF, Mahowald MA, Ley RE, Gordon JI. Reciprocal gut microbiota transplants from zebrafish and mice to germ-free recipients reveal host habitat selection. Cell. 2006;127:423–433. doi: 10.1016/j.cell.2006.08.043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roeselers G, Mittge EK, Stephens WZ, et al. Evidence for a core gut microbiota in the zebrafish. ISME Journal. 2011;5:1595–1608. doi: 10.1038/ismej.2011.38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sabree ZL, Kambhampati S, Moran NA. Nitrogen recycling and nutritional provisioning by Blattabacterium, the cockroach endosymbiont. Proceedings of the National Academy of Sciences. 2009;106:19521–19526. doi: 10.1073/pnas.0907504106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schloss PD, Westcott SL, Ryabin T, et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Applied and Environment Microbiology. 2009;75:7537–7541. doi: 10.1128/AEM.01541-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shiina A, Itoi S, Washio S, Sugita H. Molecular identification of intestinal microflora in Takifugu niphobles . Comparative Biochemistry and Physiology Part D: Genomics and Proteomics. 2006;1:128–132. doi: 10.1016/j.cbd.2005.10.001. [DOI] [PubMed] [Google Scholar]
- Shubin N. Your Inner Fish: A Journey into the 3.5-Billion-Year History of the Human Body. New York City, New York: Random House, Inc; 2008. [Google Scholar]
- Skroch P, Nienhuis J. Impact of scoring error and reproducibility RAPD data on RAPD based estimates of genetic distance. TAG Theoretical and Applied Genetics. 1995;91:1086–1091. doi: 10.1007/BF00223923. [DOI] [PubMed] [Google Scholar]
- Skrodenyte-Arbaciauskiene V, Sruoga A, Butkauskas D. Assessment of microbial diversity in the river trout Salmo trutta fario L. intestinal tract identified by partial 16S rRNA gene sequence analysis. Fisheries Science. 2006;72:597–602. [Google Scholar]
- Skrodenytė-Arbačiauskienė V, Sruoga A, Butkauskas D, Skrupskelis K. Phylogenetic analysis of intestinal bacteria of freshwater salmon Salmo salar and sea trout Salmo trutta trutta and diet. Fisheries Science. 2008;74:1307–1314. [Google Scholar]
- Smriga S, Sandin SA, Azam F. Abundance, diversity, and activity of microbial assemblages associated with coral reef fish guts and feces. FEMS Microbiology Ecology. 2010;73:31–42. doi: 10.1111/j.1574-6941.2010.00879.x. [DOI] [PubMed] [Google Scholar]
- Stackebrandt E, Goebel BM. A place for DNA-DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. International Journal of Systematic Bacteriology. 1994;44:846–849. [Google Scholar]
- Steele JA, Countway PD, Xia L, et al. Marine bacterial, archaeal and protistan association networks reveal ecological linkages. ISME Journal. 2011;5:1414–1425. doi: 10.1038/ismej.2011.24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sugita H, Ito Y. Identification of intestinal bacteria from Japanese flounder (Paralichthys olivaceus) and their ability to digest chitin. Letters in Applied Microbiology. 2006;43:336–342. doi: 10.1111/j.1472-765X.2006.01943.x. [DOI] [PubMed] [Google Scholar]
- Sugita H, Miyajima C, Deguchi Y. The vitamin B12-producing ability of the intestinal microflora of freshwater fish. Aquaculture. 1991;92:267–276. [Google Scholar]
- Sun Y, Yang H, Ling Z, Chang J, Ye J. Gut microbiota of fast and slow growing grouper Epinephelus coioides . African Journal of Microbiology Research. 2009;3:637–640. [Google Scholar]
- Swofford D. pPAUP* 4.0, Version 4.0b10. Sinauer, Sunderland: Massachusetts; 2006. [Google Scholar]
- Turnbaugh PJ, Ley RE, Mahowald MA, et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature. 2006;444:1027–1131. doi: 10.1038/nature05414. [DOI] [PubMed] [Google Scholar]
- Wang Q, Garrity GM, Tiedje JM, Cole JR. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Applied and Environmental Microbiology. 2007;73:5261–5267. doi: 10.1128/AEM.00062-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ward JA, Barlow GW. The maturation and regulation of glancing off the parents by young orange chromides (Etroplus maculatus: Pisces, Cichlidae) Behaviour. 1967;29:1–56. [Google Scholar]
- Ward NL, Steven B, Penn K, Methé BA, Detrich WH. Characterization of the intestinal microbiota of two Antarctic notothenioid fish species. Extremophiles. 2009;13:679–685. doi: 10.1007/s00792-009-0252-4. [DOI] [PubMed] [Google Scholar]
- Whitham TG, DiFazio SP, Schweitzer JA, et al. Extending genomics to natural communities and ecosystems. Science. 2008;320:492–495. doi: 10.1126/science.1153918. [DOI] [PubMed] [Google Scholar]
- Wu S, Gao T, Zheng Y, et al. Microbial diversity of intestinal contents and mucus in yellow catfish (Pelteobagrus fulvidraco) Aquaculture. 2010;303:1–7. [Google Scholar]
- Yildirim S, Yeoman CJ, Sipos M, et al. Characterization of the fecal microbiome from non-human wild primates reveals species specific microbial communities. PLoS One. 2010;5:e13963. doi: 10.1371/journal.pone.0013963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zintzen V, Roberts CD, Clark MR, et al. Composition, distribution and regional affinities of the deepwater ichthyofauna of the Lord Howe Rise and Norfolk Ridge, south-west Pacific Ocean. Deep Sea Research Part II: Topical Studies in Oceanography. 2011;58:933–947. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.