Abstract
Human immunodeficiency virus regulatory protein Rev (regulator of viral expression) is translated from a monocistronic transcript produced early in the viral replication cycle. Rev binds to the cis-acting, highly structured viral RNA sequence Rev response element (RRE) and the Rev–RRE complex primarily controls nucleocytoplasmic transport of viral RNAs. Inhibition of Rev–RRE interaction therefore is an attractive target to block viral transport. We have developed a stable cell line carrying a lentiviral vector harboring a rev gene and a co-linear Rev-dependent GFP/luciferase reporter gene cassette and thus constitutively expressing the reporter proteins. Dose-dependent luciferase activity inhibition in the indicator cell line by known small molecule inhibitors Proflavin and K37 established the specificity of the assay. This novel single step assay, that involves use of very small amount of reagents/cells and addition of test material as the only manipulation, can therefore be useful for screening therapeutically potential Rev–RRE interaction inhibitors.
Electronic supplementary material
The online version of this article (doi:10.1007/s13337-013-0166-8) contains supplementary material, which is available to authorized users.
Keywords: K-37, Lentiviral vector, Luciferase assay, Proflavin, Rev–RRE interaction, Single-step assay
Introduction
Rev, a 18 kDa 116 amino acid phospoprotein, is one of the two trans-activating proteins of human immunodeficiency virus (HIV-1) which act by sequence specific interaction with their respective sites on viral mRNA [10]. As a RNA-binding protein, Rev is essential for the nucleo-cytoplasmic export of HIV-1 mRNA and subsequent availability of full-length genomic RNA for encapsidation into virus particles [3, 13]. After infecting the target cells, HIV-1 produces unspliced, incompletely and doubly spliced mRNAs. These doubly spliced mRNAs, including those coding for the regulatory proteins Tat and Rev, are small and thus exported faster to the cytoplasm. However, the unspliced and singly/incompletely spliced mRNAs require Rev for their export and very less mRNA is exported to the cytoplasm in the absence of Rev. Rev shuttles between the nucleus and cytoplasm and binds in trans to RRE, a complex 351 nucleotide sequence that is highly structured and located within the viral envelope gene and has a high-affinity binding site for Rev protein [7, 12, 27]. Any disruption of Rev–RRE interaction can therefore provide an important therapeutic modality for anti-HIV therapy [16]. The expression of HIV genes is also dependent on cis-acting inhibitory elements (INS) located within the HIV mRNAs. One such region is the p17Gag INS elements and studies have shown that INS sequences are important for Rev dependent export of RRE containing mRNA into the cytoplasm [2, 15, 20]. Several screening strategies to find potential inhibitors/novel modulators of Rev–RRE interaction have been described. These include in vitro assays based on fluorescence techniques, a Rev distribution assay in which Rev-GFP fusion proteins are used to visualize Rev nuclear export as well as other cell-based screening assays but most of these involve transient transfections [6, 18, 19, 24–26]. Earlier we described development of a HIV-2 derived multi-platform LV as well as a single step lentiviral vector (LV) based assay for rapid evaluation of inhibitors targeting HIV-1 Tat mediated LTR trans-activation [4, 5, 21]. We report here development of a LV based simple one step assay for screening of Rev–RRE interaction inhibitors. For this, a single LV platform harboring a Rev dependent EGFP/luciferase reporter cassette along with the trans-activating component rev gene was constructed. The LV construct was used to derive a stable indicator cell line, which constitutively expresses Rev protein, thereby activating the export of the Rev responsive luciferase reporter gene. Compounds or drugs which can inhibit Rev–RRE interaction leading to decreased export of the reporter gene therefore can be easily monitored with this indicator cell line. This simple assay is a promising screening method for evaluation of candidate drugs/small molecules with potential to interfere with Rev–RRE binding.
Materials and methods
Cell culture
Human embryonic kidney derived cell line HEK 293 (NCCS, Pune) and HEK 293FT (Invitrogen, USA) were cultured in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10 % fetal bovine serum (Invitrogen, USA) and 50 μg/ml Gentamicin (Nicholas-Piramal, India) in a humidified incubator at 37 °C in 5 % CO2 atmosphere.
Compounds
The compounds used in the study were Proflavine (PRF; Sigma, USA), K-37 (7-(3,4-dehydro-4-phenyl-1-piperidinyl)-1,4-dihydro-6-fluoro-1-methyl-8-trifluoromethyl-4-oxoquinoline-3-carboxylic acid), and 3′-Azido-3′-deoxythymidine (AZT; Sigma). A 200 μM master stock and 10 μM working stock of PRF was made in DMEM. The compound K-37 was dissolved in DMSO to make a 2 mM master stock and 10 μM working stock was prepared in DMEM. AZT was dissolved in PBS at 250 μM concentration and diluted further in DMEM to make 10 μM working stock. All the drugs were used at a final concentration of 1, 3 and 5 μM.
Plasmid construction
Rev-inducible luciferase reporter gene
HIV-1 p17Gag INS element was PCR amplified from full length HIV-I molecular clone pINDIE-C1 (from Dr. D. Mitra) and cloned in the T/A cloning vector pTZ57R (referred in the text as pTZ; MBI Fermentas, Lithuania). The fragment was released by HindIII/EcoRI digestions and sub-cloned at identical sites of pcDNA3.1+ (Invitrogen, USA) to obtain pGag. The Luciferase coding sequence (without a stop codon) was PCR amplified from pGL3 Basic plasmid (Promega, USA), cloned into pTZ and subcloned upstream of enhanced green fluorescence protein (EGFP) coding sequence at EcoRI/BamHI sites of pEGFP (Clontech, USA), to yield the Luc-EGFP fusion construct. The Luc-GFP fusion cassette was released by EcoRI/NotI digestions and cloned at identical sites downstream to p17Gag in pGag construct described above. The HIV-1 RRE sequence was PCR amplified from pINDIE-C1, cloned into pTZ and subcloned at NotI/XbaI sites downstream to EGFP in pGag-Luc-GFP, the resulting luciferase reporter plasmid, pGag-Luc-GFP-RRE, was designated as pGLG-RRE.
Rev transactivator under a constitutive cellular promoter
The HIV-1 Rev coding sequence was released from pcDNA Rev (from Dr. D. Mitra) by BamHI/XhoI digestions and cloned at identical sites of EF1α promoter bearing plasmid, pTEG (from Dr. Pierre Charneau). pcDNA was digested with BglII/NheI to release the CMV promoter and the plasmid was self ligated. The EF1α-Rev fragment was released by EcoRI/XhoI digestions and cloned at the identical sites of the (CMV) promoter less pcDNA to obtain the trans-activator construct pEFIα-Rev.
Transactivator-reporter containing LV
Both reporter as well as the activator gene cassettes was cloned next into a HIV-2 based lentiviral transfer vector, pLV-puro. First, the GLG-RRE fragment was PCR amplified from pGLG-RRE using expand long template PCR (Roche, Germany) and inserted by blunt-end ligation into pLV-puro at PmeI site to obtain pLV-GLG-RRE. Further, EF1α-Rev fragment was released from pEFIα-Rev by EcoRI/XhoI (polished) digestions and cloned into pLV-GLG-RRE at XbaI (polished) site to make a single LV transfer vector designated as pLV GLG-RRE-Rev. All PCR primers used are shown in supplementary Table 1.
Virus production and generation of stable reporter cell lines
Lentiviral particles were prepared by multiplasmid transfection of HEK293 FT cells using a modified calcium phosphate precipitation protocol, as described earlier [5, 21]. Briefly, cells were seeded at a density of 1 × 106 into 60 mm Petri plates, incubated over night and transfected in fresh medium by either CaCl2/BES method or using Lipofectamine 2000 (Invitrogen) following manufacturer’s instructions. Transfection DNA mix consisted of 12 μg transfer vector (either pLV-GLG-RRE-Rev or pLV-GLG-RRE), 8 μg pGP∆RRE, 2 μg each of pRev and pTat and 4 μg of VSV-G envelope plasmid pMD.G (from Dr. D. Trono). Cells were washed next day and cultured in fresh medium and cell free, viral supernatants were harvested after 48 h. HEK293 cells were infected using the virus preparations, cultured for 72 h and selected over 2 weeks under puromycin (500 ng/μl; Sigma) to obtain a stable indicator cell line (expressing both reporter and trans-activating rev genes) and a control cell line (expressing only reporter genes). GFP expression in the cell lines were studied by fluorescent microscopy imaging (AxioVert 200; Carl Zeiss, Germany) and flow cytometry (FACS-Calibur; Beckton-Dickinson, USA) and data analysis was performed using Cell Quest Pro software.
Luciferase Assay
For transient transfection experiments, HEK293 (2 × 103/100 μl) cells were seeded in antibiotic-free media in 96-well flat bottom plates and transfected next day with either pGLG-RRE alone or co-transfected together with pGLG-RRE and pEFIα-Rev. Reporter activity was determined after 48 h using Steady-Glo Luciferase assay following manufacturer’s instructions (Promega, USA) followed by detection of luminescent signal using a microplate reader (Mithras LB-940; Berthold, Germany). For drug assay, cells from both the control and indicator cell lines were cultured (5 × 103 cells/well/100 μl medium) in 96 well flat bottom plate (Nunc) for 16 h followed by addition of putative drug compounds and reporter activity determination identically after 48 h. Percent inhibition of luciferase activity was calculated after normalization with the basal level of reporter expression in the control cell line.
MTT Assay
1 × 104 cells from the indicator cell line were cultured in a 96-well microtiter plate, incubated for 16 h, followed by addition of different concentrations of drug candidates and further incubation for 48 h. 20 μl MTT ([3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide]; 5 mg/ml in PBS; USB Corporation, USA) was added to each well, incubated for 4 h, followed by the addition of 50 μl DMSO (Sigma, USA) per well and 10 min incubation on a shaker. Absorbance was measured at 550/650 nm (Spectra Max 190; Molecular Devices, USA).
Immunoblotting
1 × 105 cells from the indicator cell line were cultured in a 6-well plate for 48 h under the indicated doses of PRF and K-37. Cells were harvested after 48 h and lysed using Proteojet (MBI Fermentas). Proteins were separated on a 15 % SDS-PAGE gel, transferred to PVDF membrane (Immobilon-P; Millipore, USA). The membranes were blocked (5 % non fat dry milk in Tris-buffered saline with 0.1 % Tween 20) for 1 h followed by overnight incubation at 4 °C with HIV-1 Rev antibody [8, 28]. Membrane was washed and incubated with HRPO conjugated secondary antibody for 1 h and signal was detected as autoradiograph using ECL+ chemiluminescence detection system (GE Healthcare, USA). Densitometry of the blots was carried out using ImageJ 1.43 (NIH) software.
Results
Molecular clones and stable cell lines
Two basic plasmids on pcDNA backbone were derived to act as the Rev dependent reporter construct that harbors EGFP and luciferase as reporters and the Rev (transactivator) construct, respectively (Fig. 1a, b). Subsequently the two cassettes, i.e., reporter and transactivator, were cloned in cis on a LV platform to derive a transactivator-reporter configuration with each expression cassette under control of different promoters, CMV and EF1α, respectively (Fig. 1c). A control LV was also derived that harbors only the reporter cassette (Fig. 1d). The two cell lines obtained by transduction with viruses made from these two LV platforms were designated accordingly as indicator and control cell lines, respectively and both the stable cell clones were EGFP positive after puromycin selection (Suppl. Fig. 1). The relative constitutive luciferase activity of the indicator cell line showed a significant difference (~2.6-fold) in comparison to that of the control cell line (Suppl. Fig. 2).
Fig. 1.
Genomic configuration of the Rev-dependent reporter and trans-activator Rev cassettes in the plasmids. a pGLG-RRE, b pEFIα-Rev and reporter-activator and reporter in lentiviral transfer vector. c pLV-GLG-RRE-Rev, d pLV-GLGRRE, respectively
Reporter assay of inhibitors
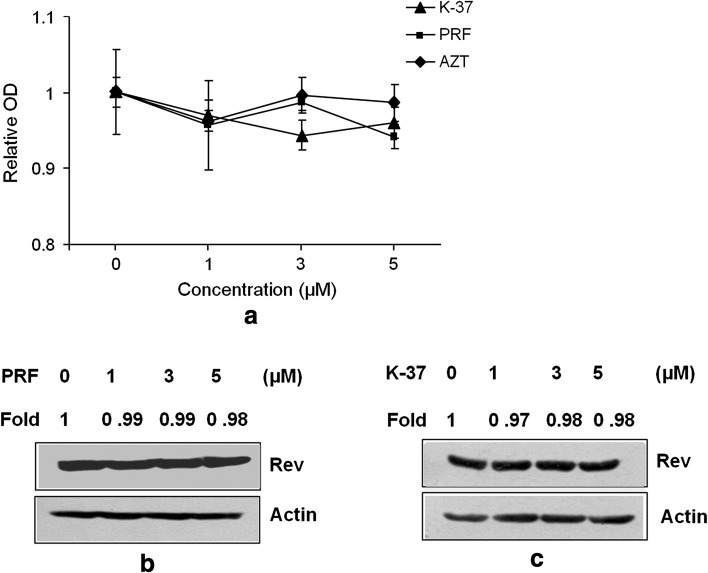
Luciferase reporter expression was evaluated using three compounds PRF, K-37, and AZT at three different concentrations (1, 3 and 5 μM). A characteristic dose-dependent inhibition profile was obtained at increasing concentrations of K-37 and PRF, reaching up to a maximum of 68 and 70 % respectively at 5 μM dose. AZT did not show any appreciable inhibition even at 5 μM concentration (Fig. 2a, b). Effect of the three compounds were tested on the indicator cell line at the test doses by MTT assay and no significant cellular cytotoxicity was observed in the presence of the drugs at any concentration (Fig. 3a).The compounds were also tested identically on the control cell line to yield similar observations (data not shown). Effect of the compounds was studied, at same doses, on the expression profile of the transactivator Rev in the indicator cell line. Expression of Rev protein was not influenced at any dose of the drugs (Fig. 3b, c).
Fig. 2.
Inhibition of Rev-mediated luciferase reporter trans-activation under different doses of a Proflavin and b K-37 AZT served as a negative control. Columns and error bars are mean ± SD (n = 3)
Fig. 3.
Effect of drugs on indicator cell viability and Rev expression. a Cell viability at indicated doses of K37, Proflavin and AZT by MTT assay; Rev protein expression in presence of b Proflavin and c K-37. Densitometric analysis of Rev expression levels represented as fold changes, respectively, and beta-actin served as loading control. Columns and error bars are mean ± SD (n = 3)
Discussion
While most currently used drugs for treatment of AIDS are targeted to HIV enzymes such as reverse transcriptase, protease and integrase (and recently the entry blockers), frequent emergence of drug resistance is a persistent concern and logical alternative molecular targets are the crucial HIV regulatory proteins. Rev, like Tat, is an essential regulatory protein for HIV replication and in its absence viral genomic RNA and other sub-genomic mRNAs cannot exit nucleus efficiently. Thus inhibiting Rev function offers an attractive alternative to block viral replication. The classical approach towards drug discovery has been the screening of a vast number of compounds/drugs and this approach also has been utilized to discover lead compounds capable of inhibiting Rev–RRE system. A number of small-molecule compounds, aminoglycoside antibiotics such as neomycin, RRE decoys, transdominant-negative version of the Rev protein and diphenylfuran cations have been screened for inhibiton of Rev–RRE interaction [9, 11, 14, 22, 23, 28]. Most of the screening assays described so far are based on transfections, require a lot of experimental manipulations and are time consuming. The indicator cell line we developed for this quick assay contains a composite reporter gene construct under control of CMV promoter and the rev activator gene on the same LV backbone. The promoter for rev was therefore changed to EF1α (by replacing CMV promoter) to avoid promoter competition. Two compounds, PRF and K-37 were used to prove efficacy of the assay. PRF interferes with Rev–RRE interaction by competing directly with the Rev binding site on RRE [8] whereas K-37 is a fluoroquinoline derivative, a class of small RNA binding molecule that inhibits Tat and other RNA dependent trans-activations [1, 17]. Since, the drugs used in the study did not show any cellular cytotoxicity or alteration in the expression of the transactivator but significantly inhibited reporter gene expression, it was confirmed that the reporter expression down regulation was specifically due to interference of Rev–RRE interaction. A graphical abstract shows the simple operational principle of the assay (Suppl. Fig. 3). Luciferase has a short half-life, high sensitivity and high signal to background ratio and thus reporter assay using luciferase activity as end point offers a robust, versatile, cost-effective, and technically simple reporter system to screen in a high throughput mode. Expression of GFP provides an optional cell selection method by sorting. The host stabilized GFP expressing cell lines were (antibiotic) selected and the two cell lines being non-clonal origin with substantially less highly fluorescent population, MFI values did not show significant change (Suppl. Fig. 1). GFP here was used only for monitoring the vector status in the cell line and is indicative of active transcription under the CMV promoter. Also, the availability of a LV derived stable indicator cell line ensures antibiotic selection free maintenance. A basal level of constitutive transport of some small HIV mRNAs bearing RRE is well documented [6, 24, 25] .
Rev binds RRE and docks it to the export protein CRM1 and the whole complex including the viral transcript is then transported together from the nucleus into the cytoplasm. Besides the RRE-sequence, unspliced and partially spliced HIV-1 RNAs contain several short INS. While RRE is a binding site for positive RNA-transport factors (Rev and CRM1), the INS elements are thought to be sites for negative nuclear retention factors for sequestration of INS-containing RNAs in the nucleus. Thus, RRE/INS containing RNAs only become constitutively competent for nuclear-cytoplasmic export in the presence of Rev. Additionally, independent of its activity in exporting RRE-containing RNA from the nucleus, Rev enhances the translation of RRE-containing RNAs such as Gag by several folds. Notably, the observed increase in translation may be due to Rev-mediated stabilization of RRE/INS-RNAs. Hence we generated a reporter construct that harbors essential elements required for the “Rev-dependent” reporter RNA transport to mimic the HIV genome transcription dynamics in a minimalistic fashion. Reporter genes in tandem without a stop codon and IRES (coding for a Luc-GFP fusion protein with unaltered reporter properties) was made to reduce the packaging size constraints and were placed downstream to the CMV promoter flanked by INS (5′) and RRE element (3′) to block nuclear export of the transcript. However, the basal level activity is still seen in absence of Rev suggesting leaky expression, which is also found during HIV replication. The idea of placing the rev gene under EF1α promoter is to express the protein in a constitutive manner, thereby activating the export of the Rev responsive luciferase reporter transcript. The two cell lines obtained by transduction with viruses made from these two LV platforms were made (to measure Rev-dependent and independent transports) and designated accordingly as indicator and control cell lines, respectively. Using the specific small molecules targeting Rev–RRE interaction it was confirmed that the reporter expression down regulation was specifically due to interference of Rev–RRE interaction.
Nonetheless, though two cell lines are required for the assay, addition of drug is the only experimental manipulation here. The key factors determining the utility of an assay are time, cost, sensitivity, specificity and reproducibility. The assay described here, that does not require infectious virus input, is to our knowledge the simplest user friendly and rapid single step assay for screening of Rev–RRE interaction inhibitors.
Electronic supplementary material
Fluorescent microscopy imaging (Upper panel) and corresponding Flow cytometry profile (Lower panel) of EGFP expression in (a) HEK293 cells, (b) control cell line and (c) indicator cell line (TIF 597 kb)
Rev-dependent reporter (luciferase) expression augmentation in the indicator cell line compared to the control cell line. Columns and error bars are mean + sd (n = 3) (TIF 76 kb)
A graphical abstract of the assay principle (TIF 245 kb)
Acknowledgments
We thank Dr. Debashis Mitra, National Cell Science Centre, Pune, for providing pcDNA-Rev and pINDIE-C1, Dr. Didier Trono, Swiss Institute of Technology, Lausanne, Switzerland for the pMD.G, and Dr. Pierre Charneau, Pasteur Institute, Paris, France for pTEG. The HIV-1 Rev antibody (Cat #7376) was obtained through the NIH AIDS Reagent Program, Division of AIDS, NIAID, NIH: HIV-1 Rev Monoclonal Antibody (1G7) from Dr. Anne Marie Szilvay. The study was supported by institutional core grant. SR and AGC were supported by research fellowships from ACTREC, Tata Memorial Centre.
References
- 1.Baba M, Okamoto M, Kawamura M, Makino M, Higashida T, Takashi T, Kimura Y, Ikeuchi T, Tetsuka T, Okamoto T. Inhibition of human immunodeficiency virus type 1 replication and cytokine production by fluoroquinoline derivatives. Mol Pharmacol. 1998;53:1097–1103. [PubMed] [Google Scholar]
- 2.Berthold E, Maldarelli F. Cis-acting elements in human immunodeficiency virus type 1 RNAs direct viral transcripts to distinct intranuclear locations. J Virol. 1996;70:4667–4682. doi: 10.1128/jvi.70.7.4667-4682.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Brandt S, Blissenbach M, Grewe B, Konietzny R, Grunwald T, Uberla K. Rev proteins of human and simian immunodeficiency virus enhance RNA encapsidation. PLoS Pathog. 2007;3:e54. doi: 10.1371/journal.ppat.0030054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Chande AG, Baba M, Mukhopadhyaya R. A single step assay for rapid evaluation of inhibitors targeting HIV type 1 Tat-mediated long terminal repeat transactivation. AIDS Res Hum Retrovir. 2012;28:902–906. doi: 10.1089/aid.2011.0228. [DOI] [PubMed] [Google Scholar]
- 5.Chande AG, Raina S, Dhamne H, Kamat RH, Mukhopadhyaya R. Multiple platforms of a HIV-2 derived lentiviral vector for expanded utility. Plasmid. 2013;69:90–95. doi: 10.1016/j.plasmid.2012.09.006. [DOI] [PubMed] [Google Scholar]
- 6.Daelemans D, Afonina E, Nilsson J, Werner G, Kjems J, De Clercq E, Pavlakis GN, Vandamme AM. A synthetic HIV-1 Rev inhibitor interfering with the CRM1-mediated nuclear export. Proc Natl Acad Sci USA. 2002;99:14440–14445. doi: 10.1073/pnas.212285299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Daly TJ, Cook KS, Gray GS, Maione TE, Rusche JR. Specific binding of HIV-1 recombinant Rev protein to the Rev-responsive element in vitro. Nature. 1989;342:816–819. doi: 10.1038/342816a0. [DOI] [PubMed] [Google Scholar]
- 8.DeJong ES, Chang CE, Gilson MK, Marino JP. Proflavine acts as a Rev inhibitor by targeting the high-affinity Rev binding site of the Rev responsive element of HIV-1. Biochemistry. 2003;42:8035–8046. doi: 10.1021/bi034252z. [DOI] [PubMed] [Google Scholar]
- 9.Fang J, Kubota S, Pomerantz RJ. A trans-dominant negative HIV type 1 Rev with intact domains of NLS/NOS and NES. AIDS Res Hum Retrovir. 2002;18:705–709. doi: 10.1089/088922202760072320. [DOI] [PubMed] [Google Scholar]
- 10.Frankel AD, Young JA. HIV-1: fifteen proteins and an RNA. Annu Rev Biochem. 1998;67:1–25. doi: 10.1146/annurev.biochem.67.1.1. [DOI] [PubMed] [Google Scholar]
- 11.Hamasaki K, Ueno A. Aminoglycoside antibiotics, neamine and its derivatives as potent inhibitors for the RNA–protein interactions derived from HIV-1 activators. Bioorg Med Chem Lett. 2001;11:591–594. doi: 10.1016/S0960-894X(01)00005-1. [DOI] [PubMed] [Google Scholar]
- 12.Heaphy S, Dingwall C, Ernberg I, Gait MJ, Green SM, Karn J, Lowe AD, Singh M, Skinner MA. HIV-1 regulator of virion expression (Rev) protein binds to an RNA stem-loop structure located within the Rev response element region. Cell. 1990;60:685–693. doi: 10.1016/0092-8674(90)90671-Z. [DOI] [PubMed] [Google Scholar]
- 13.Malim MH, Hauber J, Le SY, Maizel JV, Cullen BR. The HIV-1 rev trans-activator acts through a structured target sequence to activate nuclear export of unspliced viral mRNA. Nature. 1989;338:254–257. doi: 10.1038/338254a0. [DOI] [PubMed] [Google Scholar]
- 14.Malim MH, Freimuth WW, Liu J, Boyle TJ, Lyerly HK, Cullen BR, Nabel GJ. Stable expression of transdominant Rev protein in human T cells inhibits human immunodeficiency virus replication. J Exp Med. 1992;176:1197–1201. doi: 10.1084/jem.176.4.1197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Mikaelian I, Krieg M, Gait MJ, Karn J. Interactions of INS (CRS) elements and the splicing machinery regulate the production of Rev-responsive mRNAs. J Mol Biol. 1996;257:246–264. doi: 10.1006/jmbi.1996.0160. [DOI] [PubMed] [Google Scholar]
- 16.Nakaya T, Iwai S, Fujinaga K, Otsuka E, Ikuta K. Inhibition of HIV-1 replication by targeting the Rev protein. Leukemia. 1997;11(Suppl 3):134–137. [PubMed] [Google Scholar]
- 17.Okamoto H, Cujec TP, Okamoto M, Peterlin BM, Baba M, Okamoto T. Inhibition of the RNA-dependent transactivation and replication of human immunodeficiency virus type 1 by a fluoroquinoline derivative K-37. Virology. 2000;272:402–408. doi: 10.1006/viro.2000.0396. [DOI] [PubMed] [Google Scholar]
- 18.Qian-Cutrone J, Huang S, Chang LP, Pirnik DM, Klohr SE, Dalterio RA, Hugill R, Lowe S, Alam M, Kadow KF. Harziphilone and fleephilone, two new HIV REV/RRE binding inhibitors produced by Trichoderma harzianum. J antibiot. 1996;49:990–997. doi: 10.7164/antibiotics.49.990. [DOI] [PubMed] [Google Scholar]
- 19.Qian-Cutrone J, Huang S, Trimble J, Li H, Lin PF, Alam M, Klohr SE, Kadow KF. Niruriside, a new HIV REV/RRE binding inhibitor from Phyllanthus niruri. J Nat Prod. 1996;59:196–199. doi: 10.1021/np9600560. [DOI] [PubMed] [Google Scholar]
- 20.Rosen CA, Terwilliger E, Dayton A, Sodroski JG, Haseltine WA. Intragenic cis-acting art gene-responsive sequences of the human immunodeficiency virus. Proc Natl Acad Sci USA. 1988;85:2071–2075. doi: 10.1073/pnas.85.7.2071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Santhosh CV, Tamhane MC, Kamat RH, Patel VV, Mukhopadhyaya R. A lentiviral vector with novel multiple cloning sites: stable transgene expression in vitro and in vivo. Biochem Biophys Res Commun. 2008;371:546–550. doi: 10.1016/j.bbrc.2008.04.106. [DOI] [PubMed] [Google Scholar]
- 22.Symensma TL, Baskerville S, Yan A, Ellington AD. Polyvalent Rev decoys act as artificial Rev-responsive elements. J Virol. 1999;73:4341–4349. doi: 10.1128/jvi.73.5.4341-4349.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Takahashi T, Hamasaki K, Ueno A, Mihara H. Construction of peptides with nucleobase amino acids: design and synthesis of the nucleobase-conjugated peptides derived from HIV-1 Rev and their binding properties to HIV-1 RRE RNA. Bioorg Med Chem. 2001;9:991–1000. doi: 10.1016/S0968-0896(00)00324-2. [DOI] [PubMed] [Google Scholar]
- 24.Tang RY, Su Y. Construction of a cell-based high-flux assay for the rev protein of HIV-1. J Virol Methods. 1997;65:153–158. doi: 10.1016/S0166-0934(97)02176-9. [DOI] [PubMed] [Google Scholar]
- 25.Wolff B, Sanglier JJ, Wang Y. Leptomycin B is an inhibitor of nuclear export: inhibition of nucleo-cytoplasmic translocation of the human immunodeficiency virus type 1 (HIV-1) Rev protein and Rev-dependent mRNA. Chem Biol. 1997;4:139–147. doi: 10.1016/S1074-5521(97)90257-X. [DOI] [PubMed] [Google Scholar]
- 26.Xiao G, Kumar A, Li K, Rigl CT, Bajic M, Davis TM, Boykin DW, Wilson WD. Inhibition of the HIV-1 Rev–RRE complex formation by unfused aromatic cations. Bioorg Med Chem. 2001;9:1097–1113. doi: 10.1016/S0968-0896(00)00344-8. [DOI] [PubMed] [Google Scholar]
- 27.Zapp ML, Green MR. Sequence-specific RNA binding by the HIV-1 Rev protein. Nature. 1989;342:714–716. doi: 10.1038/342714a0. [DOI] [PubMed] [Google Scholar]
- 28.Zapp ML, Stern S, Green MR. Small molecules that selectively block RNA binding of HIV-1 Rev protein inhibit Rev function and viral production. Cell. 1993;74:969–978. doi: 10.1016/0092-8674(93)90720-B. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Fluorescent microscopy imaging (Upper panel) and corresponding Flow cytometry profile (Lower panel) of EGFP expression in (a) HEK293 cells, (b) control cell line and (c) indicator cell line (TIF 597 kb)
Rev-dependent reporter (luciferase) expression augmentation in the indicator cell line compared to the control cell line. Columns and error bars are mean + sd (n = 3) (TIF 76 kb)
A graphical abstract of the assay principle (TIF 245 kb)