Abstract
Background
Accumulating evidence demonstrates that non-coding RNAs (ncRNAs) are indispensable components of many organisms and play important roles in cellular events, regulation, and development.
Results
Here, we analysed the small non-coding RNA (ncRNA) transcriptome of Trichophyton rubrum by constructing and sequencing a cDNA library from conidia and mycelia. We identified 352 ncRNAs and their corresponding genomic loci. These ncRNA candidates included 198 entirely novel ncRNAs and 154 known ncRNAs classified as snRNAs, snoRNAs and other known ncRNAs. Further bioinformatic analysis detected 96 snoRNAs, including 56 snoRNAs that had been annotated in other organisms and 40 novel snoRNAs. All snoRNAs belonged to two major classes—C/D box snoRNAs and H/ACA snoRNAs—and their potential target sites in rRNAs and snRNAs were predicted. To analyse the evolutionary conservation of the ncRNAs in T. rubrum, we aligned all 352 ncRNAs to the genomes of six dermatophytes and to the NCBI non-redundant nucleotide database (NT). The results showed that most of the identified snRNAs were conserved in dermatophytes. Of the 352 ncRNAs, 102 also had genomic loci in other dermatophytes, and 27 were dermatophyte-specific.
Conclusions
Our systematic analysis may provide important clues to the function and evolution of ncRNAs in T. rubrum. These results also provide important information to complement the current annotation of the T. rubrum genome, which primarily comprises protein-coding genes.
Background
Numerous studies have demonstrated that non-coding RNAs (ncRNAs) are widely expressed in both prokaryotes and eukaryotes [1-4]. Furthermore, the number of ncRNAs substantially increases with the complexity of the organism, whereas the number of protein-coding genes remains relatively static. In bacteria, unicellular eukaryotes, and invertebrates, the coding sequences constitute approximately 95, 30, and 20% of the genomic DNA, respectively. In mammals, open-reading frames only account for approximately 1–2% of the genomes [5-9].
NcRNAs include highly abundant and functionally important RNAs, such as transfer RNA (tRNA) and ribosomal RNA (rRNA), as well as other small, stable RNAs, such as small nuclear RNAs (snRNAs), small nucleolar RNAs (snoRNAs), RNase P and mitochondrial RNA processing (MRP) RNA, signal recognition particle (SRP) RNA, and telomerase RNA. These RNAs have been characterised and are involved in splicing, ribosome biogenesis, translation, and chromosome replication [10,11]. Recent transcriptomic and bioinformatic studies have also identified an increasing number of new ncRNAs whose function has not been validated [12-16]. Hence, the discovery and analysis of ncRNAs has become an important step in our understanding of genomic structure and will expand our knowledge of the function and the regulatory roles of ncRNAs in the cell cycle and development.
In recent years, ncRNAs have been identified using experimental methods and computational predictions in several fungi [3,4,17-22]. A large number of non-coding RNA genes, including 33 box C/D snoRNA genes, have been predicted in the genome of Schizosaccharomyces pombe. Functional analyses of 20 Box H/ACA snoRNAs indicated that the snoRNAs evolved in coordination with rRNAs to preserve post-transcriptional modification sites among distant eukaryotes [3,4,20]. A comparative genomics analysis of seven different yeast species identified a substantial number of evolutionarily conserved, structured ncRNAs, suggesting their roles in post-transcriptional regulation [20]. NcRNAs that participate in the cleavage and processing of tRNAs were observed in Aspergillus fumigatus[21]. An extensive analysis of snoRNA genes from Neurospora crassa indicated a high diversity of post-transcriptional modification guided by snoRNAs in the fungus kingdom [22]. Thus far, the ncRNAs of dermatophytes have not been studied.
Trichophyton rubrum is the most common dermatophyte that can infect human keratinised tissue (skin, nails, and, rarely, hair) [23-25]. T. rubrum has a 22.5-Mbp haploid nuclear genome consisting of five chromosomes that range in size from 3.0–5.8 Mbp and a 27-kbp circular mitochondrial genome [26]. The Broad Institute has sequenced the T. rubrum genome and predicted more than 8,700 protein-coding genes. However, apart from rRNAs and tRNAs, no other ncRNAs have been annotated and characterised within the T. rubrum genome [26]. In the present study, we constructed an ncRNA library (ranging from 70–500 nt) and identified ncRNAs in T. rubrum using an RNA-Seq method. A total of 352 ncRNA candidates were characterised, including 198 entirely novel ncRNAs and 154 known ncRNAs. We also analysed the sequence conservation, and genomic location of these ncRNAs in six other dermatophytes. Our results may guide further studies of the important roles of ncRNA in T. rubrum and provide important complementary information to the annotation of the T. rubrum genome.
Results
Identification of ncRNA candidates in T. rubrum
To obtain a global view of ncRNAs in T. rubrum, we extracted total RNA from the conidia and mycelia phases and generated a small RNA cDNA library with size-fractionated total RNA ranging in size from 70–500 bp. After sequencing on the 454/Roche sequencing platform, a total of 87,601 reads were obtained and mapped to the T. rubrum genome. Next, the reads that mapped to the same genomic loci were clustered, resulting in 4,432 unique contigs. After removing the coding RNA and matches to tRNAs and rRNAs, the remaining 352 clusters (corresponding to 56,550 reads) were considered ncRNA candidates. Of these candidates, 154 were predicted to align with Rfam sequences and the remaining 196 were novel ncRNA candidates (Figure 1; for detailed information, see Additional file 1: Table S1).
Figure 1.
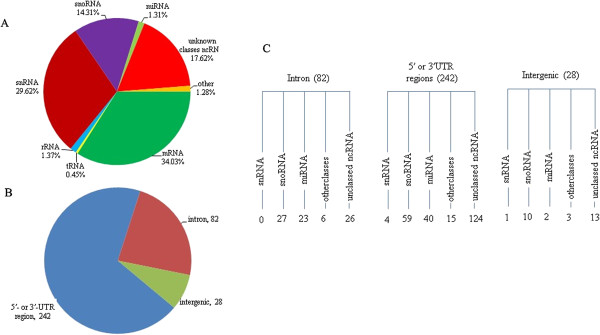
Detection of ncRNA candidates in T. rubrum by sequencing a size-fractionated cDNA library. (A) The distribution of 87,601 reads from the constructed small cDNA library of T. rubrum in different RNA classes. (B) The numbers of ncRNAs from different regions in the T. rubrum genome. (C) The number of different classes of ncRNAs are displayed in brackets.
Characteristics of ncRNA candidates
Of the 352 identified ncRNA candidates, 234 mapped to loci within 1 kb of the closest coding gene, implying a possible functional relationship. Some of the ncRNA clusters located in the immediate vicinity of a protein-coding region might be processed from the 5′- or 3′-UTR of the corresponding mRNA. Among the 352 ncRNA clusters, 82 were intronic and 29 corresponded to non-annotated intergenic regions of the T. rubrum genome (Figure 1). To verify the expression and sizes of candidate ncRNAs, we selected the spliceosomal snRNAs U1, U2, U4, U5, and U6 and 15 randomly selected novel ncRNA candidates to use in northern hybridisation. The results are shown in Figure 2.
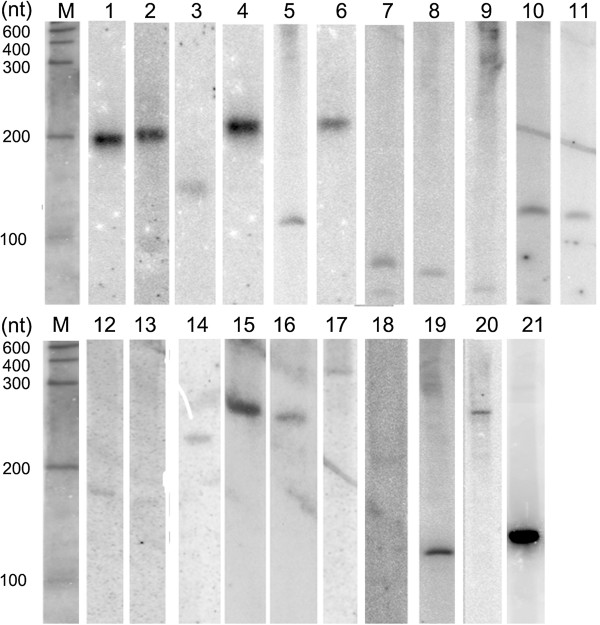
Figure 2.
Northern blotting analysis of T. rubrum ncRNA candidates. M. RiboRuler Low Range RNA Ladder (Fermentas), 1. snRNA U1, 2. snRNA U2, 3. snRNA U4, 4. snRNA U5, 5. snRNA U6, 6. Trnc_2843, 7. Trnc_3589, 8.Trnc_369, 9. Trnc_1414, 10. Trnc_293, 11. Trnc_305, 12. Trnc_1472, 13. Trnc_961, 14. Trnc_608, 15. Trnc_4262, 16. Trnc_1437, 17. Trnc_2618, 18. Trnc_3096, 19. Trnc_1686, 20. TRnc2844, and 21. 5.8S rRNA. The lengths and other information describing the ncRNAs from the northern blotting analysis are shown in Additional file 1: Table S1.
snRNA candidates
The spliceosome contains five essential small nuclear RNAs (snRNAs)—U1, U2, U4, U5, and U6—that are essential components for assembling the spliceosome and accomplishing the intricate task of intron removal from newly synthesised eukaryotic RNAs [17,18,27]. Here, we identified the genomic loci of snRNAs U1, U2, U5, and U6, each of which exhibited a unique genomic location. U5 and U6 were the most abundant snRNAs among our data, found in 15,583 and 9,034 reads, respectively. The expression of U2 and U4 was lower than the other snRNA candidates; we found only 163 reads of U2 and 146 reads of U4. These results are in agreement with those of the small ncRNA transcriptome analysis of another filamentous fungus, A. fumigatus[21,28]. U4 was not initially identified in our data. To find the U4 genomic locus in T. rubrum, we downloaded the U4 sequences of A. fumigatus, A. oryzae, and A. niger from Rfam to use as query sequences to search for homologues in the T. rubrum genome using BLASTn. One genomic locus was identified. Corresponding reads assigned to the same locus had been sequenced and clustered in our data but had been eliminated because the percentage of ORF in the cluster was greater than 80%.
We aligned the T. rubrum snRNA U1, U2, U4, U5, and U6 candidates to the genomes of six T. rubrum-related dermatophytes to predict the homologues in these genomes by BLASTn. The homologues were compared using the multiple sequence alignment software ClustalW2, revealing that all snRNAs were highly conserved in these seven dermatophytes (Table 1). High variance was observed among the sequences and lengths of these snRNAs in T. rubrum and their homologues in other fungi; however, these snRNAs were conserved at the secondary structure level, with conserved regions in the hairpin loops (Additional file 2: Figure S2). These results correspond with previous reports on A. fumigatus[21].
Table 1.
Conservation level of snRNAs in T. rubrum and related dermatophytes
| |
|
|
Genome location |
|
|
|||
|---|---|---|---|---|---|---|---|---|
| Name | Genes | Len a | Chromosome | Start | End | Position | Conserved in dermatophytes (% sequence identity) | Accession |
| Trnc_3904 |
U1 |
196 |
supercont2.8 |
159538 |
159733 |
5′UTR |
M. gypseum (98%), M. canis (98%), A. benhamiae (100%) |
KC353306 |
| Trnc_774 |
U2 |
201 |
supercont2.1 |
3545014 |
3545214 |
3′UTR |
T. tonsurans (98%), T. equinum (98%), M. gypseum (97%), T. verrucosum (99%), M. canis (96%), A. benhamiae (99%) |
KC353051 |
| Trnc_1437 |
U4 |
264 |
supercont2.17 |
13253 |
15593 |
Intergenic |
T. tonsurans (100%), T. equinum (100%), M. gypseum (99%), A. benhamiae (100%) |
KC353100 |
| Trnc_681 |
U5 |
211 |
supercont2.1 |
3061687 |
3061897 |
5′UTR |
T. tonsurans (92%), T. equinum (92%), M. gypseum (95%), T. verrucosum (93%), M. canis (91%), A. benhamiae (100%) |
KC353044 |
| Trnc_1782 | U6 | 104 | supercont2.2 | 1801544 | 1801647 | 3′UTR | T. tonsurans (100%), M. gypseum (100%), M. canis (99%), A. benhamiae (100%) | KC353131 |
Lena: the cDNA length of the snRNA. Conserved in dermatophytes (% sequence identity): the sequence identity of homologous snRNAs in other dermatophytes compared to T. rubrum; Accession is the accession number in GenBank.
snoRNAs
In eukaryotic cells, two major classes of small nucleolar ncRNA (snoRNA) have been identified: C/D box snoRNAs, which are involved in the 20-O-methylation of ribosomal, spliceosomal, and transfer RNAs (the latter in Archaea only), and H/ACA snoRNAs, which guide pseudouridylation in these RNA species [29,30].
To predict the two classes of snoRNAs and their putative targets in our data, we used the Snoscan and SnoGPS programs, defining the potential target sequences as the 5.8S, 18S, and 25S rRNAs of T. rubrum and all snRNAs identified in our data [17,18]. We identified 96 snoRNAs, including 58C/D box snoRNAs (46 had homologues in other organisms) and 38H/ACA snoRNAs (nine had homologues in other organisms). We identified 37C/D box snoRNAs as putative targets, most of which were predicted to guide methylation of 18S and 25S rRNAs. We also identified five C/D box snoRNAs (TRnc_801, TRnc_3573, TRnc_4113, TRnc_1272, and TRnc_1271) that were predicted to guide the methylation of snRNAs U1, U2, and U5. Of the 37C/D box snoRNAs, 22 had different modification sites in target rRNA or snRNA sequences. No rRNA or snRNA targets were identified in the remaining 21C/D box snoRNAs (Table 2). Additionally, the 30 identified H/ACA box snoRNAs were identified as guiding the pseudouridylation of 45 sites in rRNAs (Table 3. Detail information about potential base-paring between H/ACA box snoRNAs and rRNA shown in Additional file 3: Figure S3), whereas no pseudouridine sites were predicted on any snRNAs.
Table 2.
C/D box snoRNA candidates identified in T. rubrum
| |
|
Genome position |
Homologues |
|
|
||||
|---|---|---|---|---|---|---|---|---|---|
| Name | Len a | Chromosome | Start | End | Location | Accession 1 | Genes | Putative target(s) | Accession 2 |
| TRnc_1010 |
87 |
supercont2.10 |
749220 |
749306 |
3′UTR |
RF00477 |
snosnR66 |
|
KC353070 |
| TRnc_1157 |
95 |
supercont2.11 |
539262 |
539356 |
Intron |
RF00093 |
SNORD18, U18 |
25S: Am651, Gm654; 18S: Am1159 |
KC353075 |
| TRnc_1271 |
242 |
supercont2.12 |
280437 |
280196 |
Intron |
RF01152 |
sR1 |
25S: Am2268, Am3277,Cm964,Cm961;U5: Cm103; 18S: Am1540 |
KC353083 |
| TRnc_1272 |
265 |
supercont2.12 |
280712 |
280448 |
Intron |
RF01152 |
sR1 |
25S: Cm964, Cm961;18S: Um604; U5: Cm103 |
KC353084 |
| TRnc_1299 |
109 |
supercont2.13 |
24837 |
24729 |
Intron |
RF00593 |
snoU83B |
|
KC353086 |
| TRnc_1359 |
97 |
supercont2.14 |
159345 |
159441 |
Intron |
RF00475 |
snosnR69 |
25S: Cm3322 |
KC353090 |
| TRnc_1366 |
215 |
supercont2.14 |
179253 |
179467 |
3′UTR |
RF01152 |
sR1 |
|
KC353091 |
| TRnc_1449 |
234 |
supercont2.17 |
97081 |
97314 |
5′UTR |
RF01191 |
SNORD121A |
18S: Cm673, Gm234 |
KC353101 |
| TRnc_1560 |
77 |
supercont2.2 |
546818 |
546894 |
3′UTR |
RF01139 |
sR2 |
|
KC353110 |
| TRnc_1603 |
358 |
supercont2.2 |
766347 |
766704 |
3′UTR |
RF00345 |
snoR1 |
|
KC353115 |
| TRnc_1709 |
154 |
supercont2.2 |
1400380 |
1400533 |
5′UTR |
RF01193 |
snoR20a |
|
KC353124 |
| TRnc_1825 |
309 |
supercont2.2 |
1958330 |
1958022 |
3′UTR |
|
|
25S: Um2301; Um769 |
KC353137 |
| TRnc_1841 |
143 |
supercont2.2 |
2090171 |
2090313 |
3′UTR |
RF01144 |
sR17 |
|
KC353138 |
| TRnc_2011 |
127 |
supercont2.3 |
74633 |
74759 |
3′UTR |
RF00441 |
snoZ242 |
|
KC353147 |
| TRnc_2018 |
306 |
supercont2.3 |
117035 |
116730 |
3′UTR |
|
|
18S: Um628 |
KC353149 |
| TRnc_2027 |
96 |
supercont2.3 |
166668 |
166763 |
Intron |
RF01281 |
snoR35 |
|
KC353150 |
| TRnc_2179 |
431 |
supercont2.3 |
961995 |
961565 |
Intergenic |
|
|
25S: Um413 |
KC353160 |
| TRnc_2265 |
87 |
supercont2.3 |
1276133 |
1276219 |
Intron |
RF01197 |
snR39 |
25S: Gm808 |
KC353164 |
| TRnc_2283 |
233 |
supercont2.3 |
1301587 |
1301819 |
5′UTR |
|
|
18S: Am1105; 25S: Am499, Am1453 |
KC353165 |
| TRnc_2405 |
317 |
supercont2.3 |
1975149 |
1975465 |
3′UTR |
|
|
25S: Gm1738 |
KC353175 |
| TRnc_2419 |
204 |
supercont2.3 |
2045771 |
2045974 |
5′UTR |
RF01125 |
sR4 |
18S: Am350, Gm698, Cm701;25S: Gm215, Cm3127 |
KC353177 |
| TRnc_2421 |
182 |
supercont2.3 |
2046135 |
2046316 |
5′UTR |
RF00016 |
SNORD14, U14 |
18S: Um50, Cm379;25S: Cm2352 |
KC353178 |
| TRnc_2498 |
172 |
supercont2.3 |
2451919 |
2452090 |
5′UTR |
RF00527 |
|
|
KC353188 |
| TRnc_2545 |
119 |
supercont2.3 |
2657688 |
2657806 |
3′UTR |
RF01188 |
snR56 |
18S: Gm1389,Am385 |
KC353195 |
| TRnc_2569 |
192 |
supercont2.3 |
2759920 |
2759729 |
Intron |
RF01297 |
sR40 |
|
KC353197 |
| TRnc_2594 |
143 |
supercont2.3 |
2859175 |
2859033 |
Intron |
RF01305 |
sR51 |
|
KC353199 |
| TRnc_2691 |
158 |
supercont2.4 |
233433 |
233276 |
Intergenic |
|
|
5.8S: Gm87 |
KC353216 |
| TRnc_2782 |
128 |
supercont2.4 |
669565 |
669438 |
5′UTR |
RF00630 |
P26 |
18S: Cm534; 25S: Cm1583, Cm1196, Cm3233 |
KC353223 |
| TRnc_2936 |
246 |
supercont2.4 |
1403883 |
1403638 |
Intron |
RF00312 |
snoZ206 |
25S: Gm1378 |
KC353235 |
| TRnc_3227 |
139 |
supercont2.5 |
625518 |
625380 |
Intron |
RF00594 |
SNORD86, U86 |
KC353256 |
|
| TRnc_3297 |
138 |
supercont2.5 |
896392 |
896529 |
3′UTR |
RF00610 |
SNORD110 |
KC353262 |
|
| TRnc_338 |
135 |
supercont2.1 |
1643180 |
1643314 |
Intron |
RF01223 |
snR13 |
25S: Am2267 |
KC353022 |
| TRnc_3425 |
202 |
supercont2.6 |
22581 |
22782 |
3′UTR |
|
|
25S: Gm911 |
KC353267 |
| TRnc_3426 |
98 |
supercont2.6 |
23000 |
23097 |
3′UTR |
|
|
|
KC353268 |
| TRnc_3438 |
173 |
supercont2.6 |
91295 |
91467 |
5′UTR |
RF01291 |
snoU97, SNORD97 |
KC353269 |
|
| TRnc_3573 |
95 |
supercont2.6 |
964586 |
964680 |
Intron |
RF00530 |
snoMe28S-Cm2645 |
25S: Cm2324, Um2867; U2: Um43 |
KC353276 |
| TRnc_3654 |
191 |
supercont2.7 |
14823 |
15013 |
3′UTR |
RF01140 |
sR20 |
18S: Gm832 |
KC353284 |
| TRnc_3667 |
191 |
supercont2.7 |
59063 |
59253 |
3′UTR |
RF00529 |
snoMe28S-Am2589 |
KC353285 |
|
| TRnc_3778 |
101 |
supercont2.7 |
777627 |
777727 |
5′UTR |
RF00471 |
snosnR48, snr46 |
18S: Am721; 25S: Gm2780; Am2243 |
KC353293 |
| TRnc_3833 |
109 |
supercont2.7 |
1124537 |
1124429 |
3′UTR |
RF01273 |
sR34 |
|
KC353299 |
| TRnc_3855 |
288 |
supercont2.7 |
1281447 |
1281734 |
Intron |
RF01127 |
sR42 |
|
KC353305 |
| TRnc_3911 |
80 |
supercont2.8 |
194694 |
194773 |
Intron |
RF00213 |
snoR38 |
25S: Gm2799 |
KC353308 |
| TRnc_4113 |
681 |
supercont2.8 |
1152047 |
1152727 |
Intron |
RF01274 |
sR45 |
25S: Cm1856,Cm1673; 18S: Am833; U2: Am155 |
KC353324 |
| TRnc_415 |
103 |
supercont2.1 |
1918108 |
1918210 |
Intron |
RF01121 |
Sr38 |
|
KC353027 |
| TRnc_4250 |
192 |
supercont2.9 |
658585 |
658394 |
3′UTR |
|
|
18S: Cm373 |
KC353339 |
| TRnc_4259 |
104 |
supercont2.9 |
693331 |
693434 |
5′UTR |
RF00276 |
SNORD52, U52 |
25S: Um2408 |
KC353340 |
| TRnc_4260 |
95 |
supercont2.9 |
695194 |
695288 |
Intergenic |
RF01178 |
snoR77Y,snR77 |
18S: Um565, Am564 |
KC353341 |
| TRnc_4261 |
138 |
supercont2.9 |
695445 |
695582 |
Intergenic |
RF01209 |
snR76 |
18S: Cm1674;25S: Cm2184, Am2266, Cm3294, Cm1758 |
KC353342 |
| TRnc_4262 |
273 |
supercont2.9 |
695588 |
695860 |
Intergenic |
RF01185 |
snR75, U15 |
25S: Gm2275 |
KC353343 |
| TRnc_4263 |
157 |
supercont2.9 |
695917 |
696073 |
Intergenic |
RF00086 |
SNORD27, U27, snR74 |
25S: Cm1179 |
KC353344 |
| TRnc_4264 |
88 |
supercont2.9 |
696179 |
696266 |
5′UTR |
RF01207 |
snR73,U35 |
25S: Cm3333 |
KC353345 |
| TRnc_4267 |
100 |
supercont2.9 |
703004 |
703103 |
3′UTR |
|
|
18S: Um525, Gm527 |
KC353346 |
| TRnc_4316 |
97 |
supercont2.9 |
861468 |
861372 |
5′UTR |
RF01223 |
snR13 |
|
KC353347 |
| TRnc_4336 |
162 |
supercont2.9 |
996654 |
996493 |
Intron |
|
|
18S: Gm1089 |
KC353348 |
| TRnc_608 |
234 |
supercont2.1 |
2701229 |
2701462 |
3′UTR |
RF01202 |
sn2991 |
5.8S: Cm137 |
KC353041 |
| TRnc_640 |
129 |
supercont2.1 |
2869815 |
2869687 |
3′UTR |
RF00300 |
snoZ221 |
|
KC353043 |
| TRnc_801 |
488 |
supercont2.1 |
3681448 |
3681935 |
3′UTR |
RF00012 |
U3 |
18S: Um418; 25S: Cm1363, Cm1633, Cm1983, Cm3165; U1: Cm45 |
KC353053 |
| TRnc_821 |
210 |
supercont2.1 |
3768831 |
3768622 |
Intergenic |
|
|
18S: Cm1301,25S: Cm880 |
KC353055 |
| TRnc_985 | 153 | supercont2.10 | 686423 | 686575 | Intron | RF00494 | snoU2_19 | KC353066 | |
Name: the C/D box snoRNAs were numbered according to the order of identification. Lena: the cDNA length of the snoRNA. Homologues: homologues in Rfam or other organisms. Accession1 is the accession number in Rfam; Accession2 is the accession number in GenBank; Genes are homologous gene names in other organisms [19-22]. Putative target(s): the predicted modified nucleotides within rRNAs or snRNAs using the Snoscan package.
Table 3.
H/ACA box snoRNA candidates identified in T. rubrum
| |
|
Genome location |
Homologues |
|
|
||||
|---|---|---|---|---|---|---|---|---|---|
| Name | Len a | Chromosome | Start | End | Position | Accession 1 | Genes | Putative target | Accession 2 |
| Trnc_1355 |
371 |
supercont2.14 |
142837 |
142467 |
5′UTR |
|
|
18S-Ψ1434 |
KC353088 |
| Trnc_1370 |
133 |
supercont2.14 |
187697 |
187565 |
5′UTR |
RF01134 |
sR30 |
|
KC353092 |
| Trnc_203 |
308 |
supercont2.1 |
996485 |
996178 |
5′UTR |
|
|
18S-Ψ803 |
KC353013 |
| Trnc_2045 |
228 |
supercont2.3 |
296293 |
296520 |
5′UTR |
|
|
25S-Ψ2867,18S-Ψ489 |
KC353151 |
| Trnc_2579 |
349 |
supercont2.3 |
2792710 |
2793058 |
5′UTR |
|
|
18S-Ψ611 |
KC353198 |
| Trnc_2999 |
290 |
supercont2.4 |
1720998 |
1721287 |
5′UTR |
|
|
25S-Ψ2135 |
KC353240 |
| Trnc_3005 |
214 |
supercont2.4 |
1748930 |
1749143 |
5′UTR |
|
|
25S-Ψ1081 |
KC353241 |
| Trnc_3218 |
332 |
supercont2.5 |
584674 |
585005 |
5′UTR |
|
|
18S-Ψ573,25S-Ψ681,25S-Ψ2635 |
KC353255 |
| Trnc_3509 |
433 |
supercont2.6 |
608530 |
608098 |
5′UTR |
|
|
25S-Ψ2545,25S-Ψ1671 |
KC353274 |
| Trnc_5 |
289 |
supercont2.1 |
19982 |
20270 |
5′UTR |
|
|
25S-Ψ2329 |
KC352999 |
| Trnc_910 |
468 |
supercont2.10 |
343107 |
343574 |
5′UTR |
|
|
18S-Ψ12 |
KC353060 |
| Trnc_1407 |
234 |
supercont2.16 |
54707 |
54474 |
3′UTR |
|
|
25S-Ψ1155 |
KC353095 |
| Trnc_1472 |
188 |
supercont2.2 |
69663 |
69850 |
3′UTR |
RF01258 |
snR10 |
|
KC353105 |
| Trnc_1776 |
326 |
supercont2.2 |
1789188 |
1788863 |
3′UTR |
RF01231 |
snoR74 |
18S-Ψ1593,18S-Ψ412 |
KC353129 |
| Trnc_1893 |
344 |
supercont2.2 |
2393882 |
2393539 |
3′UTR |
|
|
25S-Ψ312 |
KC353142 |
| Trnc_2452 |
323 |
supercont2.3 |
2170039 |
2169717 |
3′UTR |
|
|
25S-Ψ2650 |
KC353184 |
| Trnc_2596 |
324 |
supercont2.3 |
2882125 |
2881802 |
3′UTR |
|
|
18S-Ψ1336 |
KC353200 |
| Trnc_2843 |
225 |
supercont2.4 |
976176 |
976400 |
3′UTR |
RF01251 |
snR3 |
25S-Ψ2120,25S-Ψ2251 |
KC353227 |
| Trnc_3023 |
182 |
supercont2.4 |
1839416 |
1839597 |
3′UTR |
|
|
25S-Ψ759,25S-Ψ1558,25S-Ψ520 |
KC353242 |
| Trnc_3387 |
226 |
supercont2.5 |
1472165 |
1472390 |
3′UTR |
|
|
18S-Ψ565,25S-Ψ2404 |
KC353265 |
| Trnc_3741 |
180 |
supercont2.7 |
491853 |
492032 |
3′UTR |
RF01247 |
snR32 |
|
KC353292 |
| Trnc_4007 |
239 |
supercont2.8 |
722404 |
722166 |
3′UTR |
|
|
18S-Ψ1344 |
KC353317 |
| Trnc_64 |
306 |
supercont2.1 |
267027 |
267332 |
3′UTR |
|
|
25S-Ψ2714 |
KC353002 |
| Trnc_817 |
188 |
supercont2.1 |
3719705 |
3719892 |
3′UTR |
|
|
18S-Ψ267,18S-Ψ1697 |
KC353054 |
| Trnc_920 |
310 |
supercont2.10 |
389299 |
389608 |
3′UTR |
|
|
25S-Ψ116,18S-Ψ1213 |
KC353061 |
| Trnc_1698 |
360 |
supercont2.2 |
1345609 |
1345968 |
Intron |
|
|
18S-Ψ1026 |
KC353122 |
| Trnc_2075 |
96 |
supercont2.3 |
425677 |
425772 |
Intron |
RF00405 |
SNORA44 |
|
KC353153 |
| Trnc_2172 |
126 |
supercont2.3 |
922150 |
922025 |
Intron |
RF00406 |
SNORA42 |
|
KC353159 |
| Trnc_2443 |
106 |
supercont2.3 |
2090244 |
2090349 |
Intron |
RF00428 |
SNORA38 |
|
KC353182 |
| Trnc_2531 |
75 |
supercont2.3 |
2617075 |
2617001 |
Intron |
RF00415 |
SNORA30 |
|
KC353194 |
| Trnc_2606 |
280 |
supercont2.36 |
2106 |
2385 |
Intergenic |
|
|
25S-Ψ1054 |
KC353202 |
| Trnc_2618 |
322 |
supercont2.36 |
8062 |
8383 |
Intergenic |
|
|
25S-Ψ1062 |
KC353205 |
| Trnc_2621 |
406 |
supercont2.36 |
8934 |
9339 |
Intergenic |
|
|
25S-Ψ1689 |
KC353206 |
| Trnc_2636 |
203 |
supercont2.36 |
19276 |
19478 |
Intergenic |
|
|
18S-Ψ217,25S-Ψ1890 |
KC353210 |
| Trnc_2898 |
393 |
supercont2.4 |
1199167 |
1198775 |
Intron |
|
|
25S-Ψ1718,25S-Ψ36 |
KC353231 |
| Trnc_3585 |
281 |
supercont2.6 |
1065274 |
1064994 |
Intron |
|
|
18S-Ψ867,25S-Ψ111 |
KC353278 |
| Trnc_4006 | 251 | supercont2.8 | 710950 | 711200 | Intron | RF01263 | snR191 | 18S-Ψ935,25S-Ψ1239,25-Ψ2245 | KC353316 |
Name: the H/ACA box snoRNAs were numbered according to the order of identification. Lena: the cDNA length of the snoRNA. Homologues: homologues in Rfam or other organisms. Accession1 is the accession number in Rfam; Accession2 is the accession number in GenBank; Genes are homologous gene names in other organisms [19-22]. Putative target(s): the predicted modified nucleotides within rRNAs using SnoGPS package.
Other types of ncRNA in T. rubrum
We also identified 51 other ncRNA genomic loci, such as pri-miRNAs or pre-miRNAs, RNAse MRP, and telomerase RNA. miRNAs related transcriptional loci were the most widely distributed ncRNAs in the T. rubrum genome; for example, the mir-598 miRNA family had 13 transcriptional regions and mir-533 had eight. In our data, these miRNA homologies of ncRNAs, which varied from 70–270 bp, were much longer than the lengths of mature miRNAs (18–25 bp), they may be pri- or pre-miRNAs candidates.
Evolutionary conservation of the ncRNAs in T. rubrum
To analyse the evolutionary conservation of ncRNAs in T. rubrum, we used BLASTn to align the sequences of all 352 ncRNAs to the genomes of six related dermatophytes: T. equinum, T. tonsurans, T. verrucosum, A. benhamiae, M. gypseum, and M. canis. The loci of 102 of these sncRNAs were also identified in all six genomes (Additional file 4: Table S4). We found that the sequences of these sncRNAs were highly conserved, with sequence identities above 85%. Of the 352 ncRNAs, ten had no hits in other genomes and might be specifically expressed in T. rubrum (Table 4). To further analyse the conserved ncRNAs in dermatophytes, we employed BLASTn to align all of the sncRNAs with the NCBI non-redundant nucleotide database (NT) after excluding Arthrodermataceae. These BLASTn results were processed by MEGAN4, which placed each ncRNA sequence in a node in the NCBI taxonomy [31].
Table 4.
The ncRNA candidates specifically expressed in T. rubrum
| |
|
|
|
Genome location |
|
|||
|---|---|---|---|---|---|---|---|---|
| Name | Class | Reads | Len a | Supercontig | Start | End | Position | Accession |
| Trnc_20 |
|
1 |
94 |
supercont2.1 |
48466 |
48559 |
3′UTR |
KC353103 |
| Trnc_1456 |
|
1 |
94 |
supercont2.18 |
53193 |
53100 |
3′UTR |
KC353000 |
| Trnc_2606 |
snoRNA;H/ACA-box |
2 |
280 |
supercont2.36 |
2106 |
2385 |
Intergenic |
KC353202 |
| Trnc_2609 |
|
4 |
255 |
supercont2.36 |
4048 |
4302 |
Intergenic |
KC353203 |
| Trnc_2621 |
snoRNA;H/ACA-box |
97 |
406 |
supercont2.36 |
8934 |
9339 |
Intergenic |
KC353206 |
| Trnc_2633 |
|
297 |
597 |
supercont2.36 |
17132 |
17728 |
Intergenic |
KC353209 |
| Trnc_2636 |
snoRNA;H/ACA-box |
1 |
203 |
supercont2.36 |
19276 |
19478 |
Intergenic |
KC353210 |
| Trnc_2640 |
|
2 |
71 |
supercont2.36 |
21309 |
21379 |
Intergenic |
KC353211 |
| Trnc_2649 |
|
2 |
79 |
supercont2.36 |
23976 |
24054 |
Intergenic |
KC353212 |
| Trnc_3096 | 1 | 201 | supercont2.4 | 2153644 | 2153444 | 3′UTR | KC353244 | |
Lena: the cDNA length of the ncRNAs; Accession is the accession number in GenBank. This table shows the lengths and genomic loci of ten ncRNAs that might be specifically expressed in T. rubrum. These ncRNAs have no hits assigned to the NCBI NT database using BLASTn.
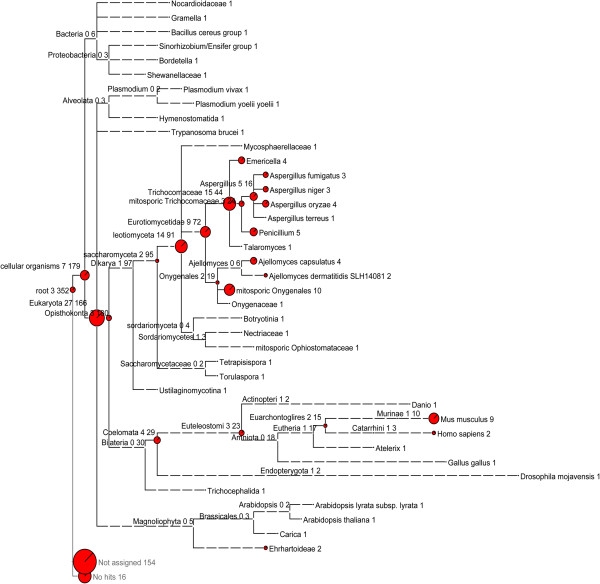
As shown in Figure 3, a total of 179 ncRNA sequences were classified under cellular organisms, with 166 clustered to the Eukaryota node (approximately 47.2% of the total 352 ncRNAs). Of these ncRNAs, 97 were assigned to Fungi, indicating that these ncRNAs were conserved in fungi; all snRNAs were assigned to this node. Of the ncRNAs under the Fungi taxonomic level, 16 and 44 were assigned to Onygenales and Trichocomaceae, respectively, supporting the close relationship between the dermatophytes and the fungi in these families. Seventy-three ncRNAs were assigned to phyla distantly related to fungi, including three assigned to the root, seven to cellular organisms, 27 to the Eukaryota node, 30 under Bilateria, and six under Bacteria. These results suggest that some ancient ncRNAs are preserved in T. rubrum.
Figure 3.
MEGAN phylogenetic analysis of T. rubrum ncRNA candidates. A MEGAN tree with the taxonomic affiliation of 352 ncRNAs that were identified by BLASTN of all sequences in NT after excluding Arthrodermataceae according to NCBI taxonomy. Each circle of the MEGAN tree represents a taxon in the NCBI taxonomy database and is labelled by its name and the number of snRNAs that were assigned to the taxon and not to a subtaxon. The size of the circles represents the number of ncRNAs.
Apart from the classified ncRNAs, the remaining 170 ncRNA candidates had no significant similarity to any nucleotide sequence in NT, including 154 unassigned ncRNAs and 16 ncRNAs with no hits. Of these unclassified ncRNAs, 27 existed in and were conserved in all six dermatophytes, indicating that these 27 ncRNAs were dermatophyte-specific ncRNAs (Table 5).
Table 5.
The ncRNA candidates specifically expressed in dermatophytes
| |
|
|
Genome location |
|
|||
|---|---|---|---|---|---|---|---|
| Name | Len a | Reads | Chromosome | Start | End | Position | Accession |
| Trnc_817 |
188 |
323 |
supercont2.1 |
3719705 |
3719892 |
3′UTR |
KC353054 |
| Trnc_733 |
174 |
1 |
supercont2.1 |
3371115 |
3371288 |
3′UTR |
KC353049 |
| Trnc_2676 |
156 |
2 |
supercont2.4 |
110438 |
110593 |
3′UTR |
KC353213 |
| Trnc_3999 |
178 |
5 |
supercont2.8 |
672734 |
672557 |
3′UTR |
KC353314 |
| Trnc_1167 |
177 |
1 |
supercont2.11 |
544895 |
545071 |
3′UTR |
KC353076 |
| Trnc_2448 |
161 |
1 |
supercont2.3 |
2123075 |
2122915 |
5′UTR |
KC353183 |
| Trnc_4219 |
104 |
1 |
supercont2.9 |
449429 |
449532 |
5′UTR |
KC353335 |
| Trnc_956 |
241 |
2 |
supercont2.10 |
559285 |
559525 |
5′UTR |
KC353063 |
| Trnc_305 |
97 |
579 |
supercont2.1 |
1515685 |
1515781 |
Intron |
KC353018 |
| Trnc_500 |
203 |
1 |
supercont2.1 |
2298649 |
2298447 |
Intron |
KC353035 |
| Trnc_1792 | 251 | 1 | supercont2.2 | 1856556 | 1856806 | Intron | KC353132 |
Lena: the cDNA length of the ncRNAs; Accession is the accession number in GenBank. This table shows the lengths and genomic loci of ten sncRNAs that might be specifically expressed in dermatophytes. These ncRNAs were conserved in all six dermatophytes but have no homologues in NT.
Discussion
RNA is emerging as a central player in cellular regulation, with active roles in multiple regulatory layers, including transcription, RNA maturation, RNA modification, and translational regulation [32]. Recent studies have revealed an unexpected complexity of regulatory RNAs, even in bacteria [2,33]. In the present study, we first used an RNA-Seq method to analyse the ncRNAs in the genome of the dermatophyte fungus T. rubrum. We identified 352 sncRNA candidates, including snRNAs, snoRNAs, miRNAs, and other types of ncRNAs; 196 novel ncRNAs were predicted. We further confirmed the genomic loci of these ncRNAs in T. rubrum. This work provides an important complement to the current annotation of the T. rubrum genome, which is currently comprised primarily of protein-coding genes.
Five types of snRNAs (U1, U2, U4, U5, and U6) were identified, and their secondary structures were predicted by RNAfold [27]. We found these snRNAs to be highly conserved among dermatophytes. We also detected 96 snoRNAs, including 55 that were annotated in other organisms and 41 that were novel snoRNAs. Using the Snoscan and snoGPS programs, we bioinformatically identified their potential target sites on rRNAs and snRNAs. miRNAs have been previously reported in some fungi, such as S. pombe, but have not been found in A. fumigatus[21,34]. In our data, we detected 68 genomic loci corresponding to 12 miRNA families; the lengths of these ncRNAs varied from 80–270 bp, suggesting that they were pri-miRNAs or pre-miRNAs [35]. To analyse the evolutionary conservation of ncRNAs, we aligned the 352 snRNAs to six other dermatophyte genomes and the NT database; we found 27 dermatophyte-specific ncRNAs and 11 T. rubrum-specific ncRNAs.
Conclusions
In this study, sequences for ncRNAs were obtained in T.rubrum and characterized by sequence comparison to know ncRNAs in other organisms, some of which were presumably functionally characterized in other work. This will prove to be a valuable resource but real understanding of regulatory mechanisms will come from followon work from this strong beginning.
Methods
Strain and culture conditions
The T. rubrum strain BMU01672 was grown on potato glucose agar (Difco) at 28°C for ten days to produce conidia. The conidia were isolated as previously reported, introduced into YPD medium (2% dextrose, 2% Bacto-Peptone, and 1% yeast extract), and incubated at 28°C with constant shaking at 200 rpm (Innova 4230 Refrigerated Incubator Shaker; New Brunswick Scientific, Edison NJ) [36]. After culture, the mycelia were harvested and ground to a powder in liquid nitrogen for RNA extraction.
RNA extraction and cDNA library construction
Total RNA was extracted from conidia and mycelia using the RNeasy Plant Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. Same amount of total RNA from conidia and mycelia was mixed and pooled on a denaturing 8% polyacrylamide gel [7 M urea and 1× TBE buffer (90 mM Tris, 64.6 mM boric acid, 2.5 mM EDTA, pH 8.3)]. We collected gel bands containing RNAs of 70–500 bp, excluding the 5.8S rRNA band. RNAs were passively eluted and then ethanol-precipitated. RNA size and concentration were quantified with the Agilent 2100 Bioanalyser and the Agilent RNA 6000 Pico Kit according to the manufacturer’s protocols. The fractionated RNA was dephosphorylated with FastAP (Fermentas) and ligated to the 3′-adaptor oligonucleotide (UUUUGACCACGGTACCCAG, RNA is underlined) by T4 RNA ligase (Promega). Subsequently, the RNA was reverse transcribed using oligo 3RT (CTGGGTACCGTGGTCAAA) and converted into double-stranded cDNA with a SuperScript Double-Stranded cDNA Synthesis Kit (Invitrogen). The ds-cDNA was purified using the MinElute Reaction Cleanup Kit (Qiagen) according to the manufacturer’s protocol.
454/Roche sequencing and data bioinformatic analysis
For 454/Roche sequencing, approximately 5 μg of the size-fractionated cDNA sample (70–500 bp) was blunted. The pieces were then ligated with short adaptors prior to amplification and sequencing. The sequencing run was performed using the method of Margulies et al.[37].
After 454 sequencing, the 5′ and 3′ adaptors were removed from the reads. Genome data for T. rubrum and six related dermatophytes (Trichophyton equinum, Trichophyton tonsurans, Trichophyton verrucosum, Arthroderma benhamiae, Microsporum gypseum, and Microsporum canis) were downloaded from the Broad Institute web site (http://www.broadinstitute.org/annotation/genome/dermatophyte_comparative/MultiDownloads.html).
The high-quality reads were mapped to the genome using BLAST (version 2.2.22) (Eval < 1e − 5). Then, reads that were 80% mapped to the genome were clustered according to their genomic position and assembled into contigs according to the genomic sequence at the corresponding loci. The ORFs in the contigs were predicted using getorf in the EMBOSS program (version 6.3.1). Contigs with less than 80% ORF were aligned to TrED EST sequences and the NCBI non-redundant protein sequence database (NR) [38,39]. The clusters with no hits in the TrED EST sequences and NR were used for the following steps: (1) alignment to non-coding RNA sequences with rRNA sequences downloaded from Rfam and GenBank [40], (2) identification of tRNAs with tRNAscan-SE (version 1.1) [41], and (3) alignment of clusters to Rfam sequences using HMMER (version 3.0) [42] and INFERNAL (version 1.0.2). The criteria for identification of known ncRNAs were as follows: (1) percentage of ORF less than 80%, (2) no hits in NR, (3) not mRNA, and (4) with homologues in Rfam [Eval (HMMER and INFERNAL) < 0.01]. For new ncRNA identification, the criteria were as follows: (1) percentage of ORF less than 80%, (2) no hits in NR, (3) not mRNA, (4) not rRNA, (5) not tRNA, and (6) no hits in Rfam (Eval > 0.01).
Analysis of snRNAs folding and predication of snoRNAs putative targets
T. rubrum snRNAs are compared with the homologs in other fungi using the multiple sequence alignment software ClustalW2. The secondary structures of aligned sequences are predicted by RNAalifold [28]. The putative targets of snoRNAs were predicted by Snoscan and SnoGPS programs [17,18]. The potential target sequences as the 5.8S, 18S, and 25S rRNAs of T. rubrum were downloaded from GenBank under the accession number JX431933.
To predict the two classes of snoRNAs and their putative targets in our data, we used the Snoscan and SnoGPS programs, defining the potential target sequences as the 5.8S, 18S, and 25S rRNAs of T. rubrum and all snRNAs identified in our data [17,18].
Northern blot analysis
For the northern blot analysis, 10 μg of total RNA was separated by electrophoresis on an 8% polyacrylamide gel containing 7 M urea and then electrotransferred onto a nylon membrane (Hybond-N+; Amersham) using a semi-dry blotting apparatus (BioRad). A total of 24–30 mer DNA oligonucleotides antisense to snRNAs and 15 randomly selected ncRNA candidates were end-labelled with (γ32P)-ATP and hybridised at 45°C for 16 hr. After stringency washes, the blots were exposed to phosphor storage screens, which were then scanned with a Typhoon 9200 imager (GE Healthcare).
Nucleotide sequence accession numbers
The 352 ncRNAs sequences of T. rubrum were submitted to GenBank under the following accession numbers: KC352999 – KC353350.
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
TL, TX and XX performed experiments. XR, TX and JY analyzed and interpreted data. JD and LS carried out the ncRNA cDNA sequencing. TL and XR wrote the paper. RC and QJ proposed the research goal, supervised the whole studies and provided a critical review of the manuscript. All authors read and approved the final manuscript.
Supplementary Material
Detailed information on ncRNAs identified in T. rubrum.
Secondary structure predictions of aligned snRNAs.
Potential base-paring between H/ACA box snoRNAs and rRNAs predicted by snoGPS.
Conversed sncRNAs in all seven dermatophytes.
Contributor Information
Tao Liu, Email: liutao@ipbcams.ac.cn.
Xianwen Ren, Email: renxwise@ipbcams.ac.cn.
Tengfei Xiao, Email: xtf25@163.com.
Jian Yang, Email: yangj@ipbcams.ac.cn.
Xingye Xu, Email: xuxingye@ipbcams.ac.cn.
Jie Dong, Email: jiedong@ipbcams.ac.cn.
Lilian Sun, Email: sunlilian@ipbcams.ac.cn.
Runsheng Chen, Email: crs@sun5.ibp.ac.cn.
Qi Jin, Email: zdsys@vip.sina.com.
Acknowledgements
This work was supported by the National Nature Science Foundation of China (Grant No. 30870104), the National High Technology Research and Development Program of China (Grant No. 2012AA020303), the National Science and Technology Major Project of China (Grant No. 2013ZX10004-601), and an intramural grant from the Institute of Pathogen Biology, Chinese Academy of Medical Sciences (Grant No. 2006IPB008).
References
- Szymanski M, Erdmann VA, Barciszewski J. Database issue. Noncoding RNAs database (ncRNAdb) Nucleic Acids Res. 2007;14:D162–D164. doi: 10.1093/nar/gkl994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rivas E, Klein RJ, Jones TA, Eddy SR. Computational identification of noncoding RNAs in E. coli by comparative genomics. Curr Biol. 2001;14(17):1369–1373. doi: 10.1016/S0960-9822(01)00401-8. [DOI] [PubMed] [Google Scholar]
- Li SG, Zhou H, Luo YP, Zhang P, Qu LH. Identification and functional analysis of 20 Box H/ACA small nucleolar RNAs (snoRNAs) from Schizosaccharomyces pombe. J Biol Chem. 2005;14(16):16446–16455. doi: 10.1074/jbc.M500326200. [DOI] [PubMed] [Google Scholar]
- Wood V, Gwilliam R, Rajandream MA, Lyne M, Lyne R, Stewart A, Sgouros J, Peat N, Hayles J, Baker S. et al. The genome sequence of Schizosaccharomyces pombe. Nature. 2002;14(6874):871–880. doi: 10.1038/nature724. [DOI] [PubMed] [Google Scholar]
- Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J, Devon K, Dewar K, Doyle M, FitzHugh W. et al. Initial sequencing and analysis of the human genome. Nature. 2001;14(6822):860–921. doi: 10.1038/35057062. [DOI] [PubMed] [Google Scholar]
- Waterston RH, Lindblad-Toh K, Birney E, Rogers J, Abril JF, Agarwal P, Agarwala R, Ainscough R, Alexandersson M, An P. et al. Initial sequencing and comparative analysis of the mouse genome. Nature. 2002;14(6915):520–562. doi: 10.1038/nature01262. [DOI] [PubMed] [Google Scholar]
- Taft RJ, Pheasant M, Mattick JS. The relationship between non-protein-coding DNA and eukaryotic complexity. Bioessays. 2007;14(3):288–299. doi: 10.1002/bies.20544. [DOI] [PubMed] [Google Scholar]
- Ponting CP, Belgard TG. Transcribed dark matter: meaning or myth? Hum Mol Genet. 2010;14(R2):R162–R168. doi: 10.1093/hmg/ddq362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Amaral PP, Mattick JS. Noncoding RNA in development. Mamm Genome. 2008;14(7–8):454–492. doi: 10.1007/s00335-008-9136-7. [DOI] [PubMed] [Google Scholar]
- Ponting CP, Oliver PL, Reik W. Evolution and functions of long noncoding RNAs. Cell. 2009;14(4):629–641. doi: 10.1016/j.cell.2009.02.006. [DOI] [PubMed] [Google Scholar]
- Brown JW, Marshall DF, Echeverria M. Intronic noncoding RNAs and splicing. Trends Plant Sci. 2008;14(7):335–342. doi: 10.1016/j.tplants.2008.04.010. [DOI] [PubMed] [Google Scholar]
- Cheng J, Kapranov P, Drenkow J, Dike S, Brubaker S, Patel S, Long J, Stern D, Tammana H, Helt G. et al. Transcriptional maps of 10 human chromosomes at 5-nucleotide resolution. Science. 2005;14(5725):1149–1154. doi: 10.1126/science.1108625. [DOI] [PubMed] [Google Scholar]
- Birney E, Stamatoyannopoulos JA, Dutta A, Guigo R, Gingeras TR, Margulies EH, Weng Z, Snyder M, Dermitzakis ET, Thurman RE. et al. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature. 2007;14(7146):799–816. doi: 10.1038/nature05874. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Washietl S, Pedersen JS, Korbel JO, Stocsits C, Gruber AR, Hackermuller J, Hertel J, Lindemeyer M, Reiche K, Tanzer A. et al. Structured RNAs in the ENCODE selected regions of the human genome. Genome Res. 2007;14(6):852–864. doi: 10.1101/gr.5650707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kapranov P, Willingham AT, Gingeras TR. Genome-wide transcription and the implications for genomic organization. Nat Rev Genet. 2007;14(6):413–423. doi: 10.1038/nrg2083. [DOI] [PubMed] [Google Scholar]
- Lu ZJ, Yip KY, Wang G, Shou C, Hillier LW, Khurana E, Agarwal A, Auerbach R, Rozowsky J, Cheng C. et al. Prediction and characterization of noncoding RNAs in C. elegans by integrating conservation, secondary structure, and high-throughput sequencing and array data. Genome Res. 2011;14(2):276–285. doi: 10.1101/gr.110189.110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lowe TM, Eddy SR. A computational screen for methylation guide snoRNAs in yeast. Science. 1999;14(5405):1168–1171. doi: 10.1126/science.283.5405.1168. [DOI] [PubMed] [Google Scholar]
- Schattner P, Decatur WA, Davis CA, Ares M Jr, Fournier MJ, Lowe TM. Genome-wide searching for pseudouridylation guide snoRNAs: analysis of the Saccharomyces cerevisiae genome. Nucleic Acids Res. 2004;14(14):4281–4296. doi: 10.1093/nar/gkh768. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Torchet C, Badis G, Devaux F, Costanzo G, Werner M, Jacquier A. The complete set of H/ACA snoRNAs that guide rRNA pseudouridylations in Saccharomyces cerevisiae. RNA. 2005;14(6):928–938. doi: 10.1261/rna.2100905. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steigele S, Huber W, Stocsits C, Stadler PF, Nieselt K. Comparative analysis of structured RNAs in S. cerevisiae indicates a multitude of different functions. BMC Biol. 2007;14:25. doi: 10.1186/1741-7007-5-25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jochl C, Rederstorff M, Hertel J, Stadler PF, Hofacker IL, Schrettl M, Haas H, Huttenhofer A. Small ncRNA transcriptome analysis from Aspergillus fumigatus suggests a novel mechanism for regulation of protein synthesis. Nucleic Acids Res. 2008;14(8):2677–2689. doi: 10.1093/nar/gkn123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu N, Xiao ZD, Yu CH, Shao P, Liang YT, Guan DG, Yang JH, Chen CL, Qu LH, Zhou H. SnoRNAs from the filamentous fungus Neurospora crassa: structural, functional and evolutionary insights. BMC Genomics. 2009;14:515. doi: 10.1186/1471-2164-10-515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Costa M, Passos XS, Hasimoto e Souza LK, Miranda AT, Lemos Jde A, Oliveira JG Jr, Silva Mdo R Jr. [Epidemiology and etiology of dermatophytosis in Goiania, GO, Brazil] Rev Soc Bras Med Trop. 2002;14(1):19–22. doi: 10.1590/S0037-86822002000100004. [DOI] [PubMed] [Google Scholar]
- Jennings MB, Weinberg JM, Koestenblatt EK, Lesczczynski C. Study of clinica suspected onychomycosis in a podiatric population. J Am Podiatr Med Assoc. 2002;14(6):327–330. doi: 10.7547/87507315-92-6-327. [DOI] [PubMed] [Google Scholar]
- Monod M, Jaccoud S, Zaugg C, Lechenne B, Baudraz F, Panizzon R. Survey of dermatophyte infections in the Lausanne area Switzerland. Dermatology. 2002;14(2):201–203. doi: 10.1159/000063913. [DOI] [PubMed] [Google Scholar]
- Martinez DA, Oliver BG, Graser Y, Goldberg JM, Li W, Martinez-Rossi NM, Monod M, Shelest E, Barton RC, Birch E. et al. Comparative genome analysis of Trichophyton rubrum and related dermatophytes reveals candidate genes involved in infection. mBio. 2012;14(5):e00212–e00259. doi: 10.1128/mBio.00259-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gruber AR, Lorenz R, Bernhart SH, Neubock R, Hofacker IL. The Vienna RNA websuite. Nucleic Acids Res. 2008;14(Web Server issue):W70–W74. doi: 10.1093/nar/gkn188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bernhart SH, Hofacker IL, Will S, Gruber AR, Stadler PF. RNAalifold: improved consensus structure prediction for RNA alignments. BMC Bioinformatics. 2008;14:474. doi: 10.1186/1471-2105-9-474. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bachellerie JP, Cavaille J, Huttenhofer A. The expanding snoRNA world. Biochimie. 2002;14(8):775–790. doi: 10.1016/S0300-9084(02)01402-5. [DOI] [PubMed] [Google Scholar]
- Maden BE. The numerous modified nucleotides in eukaryotic ribosomal RNA. Prog Nucleic Acid Res Mol Biol. 1990;14:241–303. doi: 10.1016/s0079-6603(08)60629-7. [DOI] [PubMed] [Google Scholar]
- Huson DH, Mitra S, Ruscheweyh HJ, Weber N, Schuster SC. Integrative analysis of environmental sequences using MEGAN4. Genome Res. 2011;14(9):1552–1560. doi: 10.1101/gr.120618.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bompfunewerer AF, Flamm C, Fried C, Fritzsch G, Hofacker IL, Lehmann J, Missal K, Mosig A, Muller B, Prohaska SJ. et al. Evolutionary patterns of non-coding RNAs. Theory Biosci. 2005;14(4):301–369. doi: 10.1016/j.thbio.2005.01.002. [DOI] [PubMed] [Google Scholar]
- Storz G, Vogel J, Wassarman KM. Regulation by small RNAs in bacteria: expanding frontiers. Mol Cell. 2011;14(6):880–891. doi: 10.1016/j.molcel.2011.08.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Emmerth S, Schober H, Gaidatzis D, Roloff T, Jacobeit K, Buhler M. Nuclear retention of fission yeast dicer is a prerequisite for RNAi-mediated heterochromatin assembly. Dev Cell. 2010;14(1):102–113. doi: 10.1016/j.devcel.2009.11.011. [DOI] [PubMed] [Google Scholar]
- Lee Y, Jeon K, Lee JT, Kim S, Kim VN. MicroRNA maturation: stepwise processing and subcellular localization. EMBO J. 2002;14(17):4663–4670. doi: 10.1093/emboj/cdf476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu T, Zhang Q, Wang L, Yu L, Leng W, Yang J, Chen L, Peng J, Ma L, Dong J. et al. The use of global transcriptional analysis to reveal the biological and cellular events involved in distinct development phases of Trichophyton rubrum conidial germination. BMC Genomics. 2007;14:100. doi: 10.1186/1471-2164-8-100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Margulies M, Egholm M, Altman WE, Attiya S, Bader JS, Bemben LA, Berka J, Braverman MS, Chen YJ, Chen Z. et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature. 2005;14(7057):376–380. doi: 10.1038/nature03959. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang L, Ma L, Leng W, Liu T, Yu L, Yang J, Yang L, Zhang W, Zhang Q, Dong J. et al. Analysis of the dermatophyte Trichophyton rubrum expressed sequence tags. BMC Genomics. 2006;14:255. doi: 10.1186/1471-2164-7-255. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang J, Chen L, Wang L, Zhang W, Liu T, Jin Q. TrED: the Trichophyton rubrum Expression Database. BMC Genomics. 2007;14:250. doi: 10.1186/1471-2164-8-250. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burge SW, Daub J, Eberhardt R, Tate J, Barquist L, Nawrocki EP, Eddy SR, Gardner PP, Bateman A. Rfam 11.0: 10 years of RNA families. Nucleic Acids Res. 2013;14(Database issue)):D226–D232. doi: 10.1093/nar/gks1005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lowe TM, Eddy SR. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 1997;14(5):955–964. doi: 10.1093/nar/25.5.955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eddy SR. Accelerated profile HMM searches. PLoS Comput Biol. 2011;14(10):e1002195. doi: 10.1371/journal.pcbi.1002195. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Detailed information on ncRNAs identified in T. rubrum.
Secondary structure predictions of aligned snRNAs.
Potential base-paring between H/ACA box snoRNAs and rRNAs predicted by snoGPS.
Conversed sncRNAs in all seven dermatophytes.