Abstract
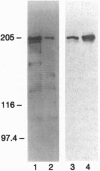
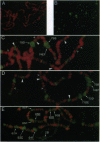
Previously, we reported on the discovery and characterization of a mammalian chromatin-associated protein, CHD1 (chromo-ATPase/helicase-DNA-binding domain), with features that led us to suspect that it might have an important role in the modification of chromatin structure. We now report on the characterization of the Drosophila melanogaster CHD1 homologue (dCHD1) and its localization on polytene chromosomes. A set of overlapping cDNAs encodes an 1883-aa open reading frame that is 50% identical and 68% similar to the mouse CHD1 sequence, including conservation of the three signature domains for which the protein was named. When the chromo and ATPase/helicase domain sequences in various CHD1 homologues were compared with the corresponding sequences in other proteins, certain distinctive features of the CHD1 chromo and ATPase/helicase domains were revealed. The dCHD1 gene was mapped to position 23C-24A on chromosome 2L. Western blot analyses with antibodies raised against a dCHD1 fusion protein specifically recognized an approximately 210-kDa protein in nuclear extracts from Drosophila embryos and cultured cells. Most interestingly, these antibodies revealed that dCHD1 localizes to sites of extended chromatin (interbands) and regions associated with high transcriptional activity (puffs) on polytene chromosomes from salivary glands of third instar larvae. These observations strongly support the idea that CHD1 functions to alter chromatin structure in a way that facilitates gene expression.
Full text
PDF




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Andres A. J., Thummel C. S. Hormones, puffs and flies: the molecular control of metamorphosis by ecdysone. Trends Genet. 1992 Apr;8(4):132–138. doi: 10.1016/0168-9525(92)90371-A. [DOI] [PubMed] [Google Scholar]
- Ashburner M. Sequential gene activation by ecdysone in polytene chromosomes of Drosophila melanogaster. II. The effects of inhibitors of protein synthesis. Dev Biol. 1974 Jul;39(1):141–157. doi: 10.1016/s0012-1606(74)80016-3. [DOI] [PubMed] [Google Scholar]
- Bork P., Koonin E. V. An expanding family of helicases within the 'DEAD/H' superfamily. Nucleic Acids Res. 1993 Feb 11;21(3):751–752. doi: 10.1093/nar/21.3.751. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown N. H., Kafatos F. C. Functional cDNA libraries from Drosophila embryos. J Mol Biol. 1988 Sep 20;203(2):425–437. doi: 10.1016/0022-2836(88)90010-1. [DOI] [PubMed] [Google Scholar]
- Burtis K. C., Thummel C. S., Jones C. W., Karim F. D., Hogness D. S. The Drosophila 74EF early puff contains E74, a complex ecdysone-inducible gene that encodes two ets-related proteins. Cell. 1990 Apr 6;61(1):85–99. doi: 10.1016/0092-8674(90)90217-3. [DOI] [PubMed] [Google Scholar]
- Cairns B. R., Kim Y. J., Sayre M. H., Laurent B. C., Kornberg R. D. A multisubunit complex containing the SWI1/ADR6, SWI2/SNF2, SWI3, SNF5, and SNF6 gene products isolated from yeast. Proc Natl Acad Sci U S A. 1994 Mar 1;91(5):1950–1954. doi: 10.1073/pnas.91.5.1950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cavener D. R., Ray S. C. Eukaryotic start and stop translation sites. Nucleic Acids Res. 1991 Jun 25;19(12):3185–3192. doi: 10.1093/nar/19.12.3185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Champlin D. T., Lis J. T. Distribution of B52 within a chromosomal locus depends on the level of transcription. Mol Biol Cell. 1994 Jan;5(1):71–79. doi: 10.1091/mbc.5.1.71. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Churchill M. E., Travers A. A. Protein motifs that recognize structural features of DNA. Trends Biochem Sci. 1991 Mar;16(3):92–97. doi: 10.1016/0968-0004(91)90040-3. [DOI] [PubMed] [Google Scholar]
- Côté J., Quinn J., Workman J. L., Peterson C. L. Stimulation of GAL4 derivative binding to nucleosomal DNA by the yeast SWI/SNF complex. Science. 1994 Jul 1;265(5168):53–60. doi: 10.1126/science.8016655. [DOI] [PubMed] [Google Scholar]
- Delmas V., Stokes D. G., Perry R. P. A mammalian DNA-binding protein that contains a chromodomain and an SNF2/SWI2-like helicase domain. Proc Natl Acad Sci U S A. 1993 Mar 15;90(6):2414–2418. doi: 10.1073/pnas.90.6.2414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dingwall A. K., Beek S. J., McCallum C. M., Tamkun J. W., Kalpana G. V., Goff S. P., Scott M. P. The Drosophila snr1 and brm proteins are related to yeast SWI/SNF proteins and are components of a large protein complex. Mol Biol Cell. 1995 Jul;6(7):777–791. doi: 10.1091/mbc.6.7.777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eisen J. A., Sweder K. S., Hanawalt P. C. Evolution of the SNF2 family of proteins: subfamilies with distinct sequences and functions. Nucleic Acids Res. 1995 Jul 25;23(14):2715–2723. doi: 10.1093/nar/23.14.2715. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fleischmann G., Pflugfelder G., Steiner E. K., Javaherian K., Howard G. C., Wang J. C., Elgin S. C. Drosophila DNA topoisomerase I is associated with transcriptionally active regions of the genome. Proc Natl Acad Sci U S A. 1984 Nov;81(22):6958–6962. doi: 10.1073/pnas.81.22.6958. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Franke A., DeCamillis M., Zink D., Cheng N., Brock H. W., Paro R. Polycomb and polyhomeotic are constituents of a multimeric protein complex in chromatin of Drosophila melanogaster. EMBO J. 1992 Aug;11(8):2941–2950. doi: 10.1002/j.1460-2075.1992.tb05364.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henikoff S. Transcriptional activator components and poxvirus DNA-dependent ATPases comprise a single family. Trends Biochem Sci. 1993 Aug;18(8):291–292. doi: 10.1016/0968-0004(93)90037-n. [DOI] [PubMed] [Google Scholar]
- Imbalzano A. N., Kwon H., Green M. R., Kingston R. E. Facilitated binding of TATA-binding protein to nucleosomal DNA. Nature. 1994 Aug 11;370(6489):481–485. doi: 10.1038/370481a0. [DOI] [PubMed] [Google Scholar]
- Jamrich M., Greenleaf A. L., Bautz E. K. Localization of RNA polymerase in polytene chromosomes of Drosophila melanogaster. Proc Natl Acad Sci U S A. 1977 May;74(5):2079–2083. doi: 10.1073/pnas.74.5.2079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kozak M. An analysis of 5'-noncoding sequences from 699 vertebrate messenger RNAs. Nucleic Acids Res. 1987 Oct 26;15(20):8125–8148. doi: 10.1093/nar/15.20.8125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kwon H., Imbalzano A. N., Khavari P. A., Kingston R. E., Green M. R. Nucleosome disruption and enhancement of activator binding by a human SW1/SNF complex. Nature. 1994 Aug 11;370(6489):477–481. doi: 10.1038/370477a0. [DOI] [PubMed] [Google Scholar]
- Muchardt C., Yaniv M. A human homologue of Saccharomyces cerevisiae SNF2/SWI2 and Drosophila brm genes potentiates transcriptional activation by the glucocorticoid receptor. EMBO J. 1993 Nov;12(11):4279–4290. doi: 10.1002/j.1460-2075.1993.tb06112.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Orlando V., Paro R. Mapping Polycomb-repressed domains in the bithorax complex using in vivo formaldehyde cross-linked chromatin. Cell. 1993 Dec 17;75(6):1187–1198. doi: 10.1016/0092-8674(93)90328-n. [DOI] [PubMed] [Google Scholar]
- Paro R. Imprinting a determined state into the chromatin of Drosophila. Trends Genet. 1990 Dec;6(12):416–421. doi: 10.1016/0168-9525(90)90303-n. [DOI] [PubMed] [Google Scholar]
- Pearce J. J., Singh P. B., Gaunt S. J. The mouse has a Polycomb-like chromobox gene. Development. 1992 Apr;114(4):921–929. doi: 10.1242/dev.114.4.921. [DOI] [PubMed] [Google Scholar]
- Peterson C. L., Dingwall A., Scott M. P. Five SWI/SNF gene products are components of a large multisubunit complex required for transcriptional enhancement. Proc Natl Acad Sci U S A. 1994 Apr 12;91(8):2905–2908. doi: 10.1073/pnas.91.8.2905. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peterson C. L., Tamkun J. W. The SWI-SNF complex: a chromatin remodeling machine? Trends Biochem Sci. 1995 Apr;20(4):143–146. doi: 10.1016/s0968-0004(00)88990-2. [DOI] [PubMed] [Google Scholar]
- Reeves R., Nissen M. S. The A.T-DNA-binding domain of mammalian high mobility group I chromosomal proteins. A novel peptide motif for recognizing DNA structure. J Biol Chem. 1990 May 25;265(15):8573–8582. [PubMed] [Google Scholar]
- Shaffer C. D., Wallrath L. L., Elgin S. C. Regulating genes by packaging domains: bits of heterochromatin in euchromatin? Trends Genet. 1993 Feb;9(2):35–37. doi: 10.1016/0168-9525(93)90171-D. [DOI] [PubMed] [Google Scholar]
- Singh P. B., Miller J. R., Pearce J., Kothary R., Burton R. D., Paro R., James T. C., Gaunt S. J. A sequence motif found in a Drosophila heterochromatin protein is conserved in animals and plants. Nucleic Acids Res. 1991 Feb 25;19(4):789–794. doi: 10.1093/nar/19.4.789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stokes D. G., Perry R. P. DNA-binding and chromatin localization properties of CHD1. Mol Cell Biol. 1995 May;15(5):2745–2753. doi: 10.1128/mcb.15.5.2745. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thummel C. S., Burtis K. C., Hogness D. S. Spatial and temporal patterns of E74 transcription during Drosophila development. Cell. 1990 Apr 6;61(1):101–111. doi: 10.1016/0092-8674(90)90218-4. [DOI] [PubMed] [Google Scholar]
- Tsukiyama T., Daniel C., Tamkun J., Wu C. ISWI, a member of the SWI2/SNF2 ATPase family, encodes the 140 kDa subunit of the nucleosome remodeling factor. Cell. 1995 Dec 15;83(6):1021–1026. doi: 10.1016/0092-8674(95)90217-1. [DOI] [PubMed] [Google Scholar]
- Weeks J. R., Hardin S. E., Shen J., Lee J. M., Greenleaf A. L. Locus-specific variation in phosphorylation state of RNA polymerase II in vivo: correlations with gene activity and transcript processing. Genes Dev. 1993 Dec;7(12A):2329–2344. doi: 10.1101/gad.7.12a.2329. [DOI] [PubMed] [Google Scholar]
- Wilson C. J., Chao D. M., Imbalzano A. N., Schnitzler G. R., Kingston R. E., Young R. A. RNA polymerase II holoenzyme contains SWI/SNF regulators involved in chromatin remodeling. Cell. 1996 Jan 26;84(2):235–244. doi: 10.1016/s0092-8674(00)80978-2. [DOI] [PubMed] [Google Scholar]
- Winston F., Carlson M. Yeast SNF/SWI transcriptional activators and the SPT/SIN chromatin connection. Trends Genet. 1992 Nov;8(11):387–391. doi: 10.1016/0168-9525(92)90300-s. [DOI] [PubMed] [Google Scholar]
- Wreggett K. A., Hill F., James P. S., Hutchings A., Butcher G. W., Singh P. B. A mammalian homologue of Drosophila heterochromatin protein 1 (HP1) is a component of constitutive heterochromatin. Cytogenet Cell Genet. 1994;66(2):99–103. doi: 10.1159/000133676. [DOI] [PubMed] [Google Scholar]
- Yoshinaga S. K., Peterson C. L., Herskowitz I., Yamamoto K. R. Roles of SWI1, SWI2, and SWI3 proteins for transcriptional enhancement by steroid receptors. Science. 1992 Dec 4;258(5088):1598–1604. doi: 10.1126/science.1360703. [DOI] [PubMed] [Google Scholar]
- Zhao K., Hart C. M., Laemmli U. K. Visualization of chromosomal domains with boundary element-associated factor BEAF-32. Cell. 1995 Jun 16;81(6):879–889. doi: 10.1016/0092-8674(95)90008-x. [DOI] [PubMed] [Google Scholar]
- Zink B., Engström Y., Gehring W. J., Paro R. Direct interaction of the Polycomb protein with Antennapedia regulatory sequences in polytene chromosomes of Drosophila melanogaster. EMBO J. 1991 Jan;10(1):153–162. doi: 10.1002/j.1460-2075.1991.tb07931.x. [DOI] [PMC free article] [PubMed] [Google Scholar]