Abstract
The genome of the facultative anaerobic γ-proteobacterium Shewanella oneidensis MR-1 encodes for three terminal oxidases: a bd-type quinol oxidase and two heme-copper oxidases, a A-type cytochrome c oxidase and a cbb 3-type oxidase. In this study, we used a biochemical approach and directly measured oxidase activities coupled to mass-spectrometry analysis to investigate the physiological role of the three terminal oxidases under aerobic and microaerobic conditions. Our data revealed that the cbb 3-type oxidase is the major terminal oxidase under aerobic conditions while both cbb 3-type and bd-type oxidases are involved in respiration at low-O2 tensions. On the contrary, the low O2-affinity A-type cytochrome c oxidase was not detected in our experimental conditions even under aerobic conditions and would therefore not be required for aerobic respiration in S. oneidensis MR-1. In addition, the deduced amino acid sequence suggests that the A-type cytochrome c oxidase is a ccaa 3-type oxidase since an uncommon extra-C terminal domain contains two c-type heme binding motifs. The particularity of the aerobic respiratory pathway and the physiological implication of the presence of a ccaa 3-type oxidase in S. oneidensis MR-1 are discussed.
Introduction
In the oxygen respiratory systems, electrons of low-redox potential electron donors are transferred through a series of membrane-bound proteins or complexes and finally, the reduction of molecular oxygen to water is catalyzed by enzymes called terminal oxidases. These oxygen reductases are complicated integral membrane multi-subunit complexes grouped into two major superfamilies. Most of them belong to the well-characterized heme-copper oxidases (HCO) superfamily [1]. HCO have been named cytochrome c oxidases or quinol oxidases, depending on the nature of their electron donor and are able to pump protons across membrane. Additionally, based on biochemical and structural differences in their catalytic subunits and on phylogenetic analysis, a classification of HCO into three families was proposed [2]: i) type A (mitochondrial-like oxidases or aa 3-type), ii) type B (ba 3-type oxidases) and iii) type C (cbb 3-type oxidases) only detected in bacteria [3]. Cytochrome bd-type oxidases, phylogenetically unrelated to the HCO [4], represent the second major oxidases superfamily [5]. Widely distributed among prokaryotes, bd-type oxygen reductases function as quinol-oxidases and are bioenergetically less efficient than HCO since they generate a proton motive force by transmembrane charge separation rather than by pumping protons [6]–[8]. In addition to its role in cell bioenergetics, many studies suggest that cytochrome bd oxidase may be implicated in other important physiological functions. In particular, the enzyme seems to be involved in the bacterial response to a wide variety of stress conditions such as alkalinization of the medium, high temperatures, hydrogen peroxide and nitrosative (NO) stresses [5], [9]–[11]. Cytochrome bd oxidase could also play a determinant role in bacterial pathogenicity by protecting bacteria against the NO-mediated host immune response [12].
In contrast to the mitochondrial respiratory systems, most bacteria have branched-respiratory chains terminating in multiple oxidases or use alternative electron acceptors. This enables them to respond to changes in the environment and contributes to their ability to colonize many microoxic and anoxic environments [13]–[15]. Each oxidase type in bacteria is expected to have a specific affinity for O2 [16]. Under O2 limitation, many bacteria induce high O2-affinity oxidases to respire traces of molecular oxygen. The cbb 3-type cytochrome c oxidase and the bd-type quinol oxidase are generally considered to have a high affinity for oxygen [17], [18]. On the other hand, the A-type oxidases have low affinity for oxygen and are predicted to provide the capacity to grow in high-O2 environments only [16].
Shewanella species are facultative anaerobic gram-negative γ-proteobacteria that exhibit extensive respiratory versatility using a broad spectrum of terminal electron acceptors such as fumarate, dimethyl sulfoxide, NO2 −, NO3 −, Fe(III), As(V), U(VI), Mn(IV), Cr(VI), Tc(VII) elemental sulfur and azo dyes [19]–[22]. Genome analysis of Shewanella oneidensis MR-1 revealed the presence of genes coding for terminal oxidases: a bd-type quinol oxidase and two HCO, a A-type cytochrome c oxidase and a cbb 3-type oxidase [19]. Latterly, it has been reported that cytochrome bd oxidase confers nitrite resistance to S. oneidensis MR-1 and plays a significant role in oxygen respiration under microaerobic but not aerobic conditions [23]. Marritt et al. [24] hypothesized that, in accordance with their predicted affinity for molecular oxygen, S. oneidensis MR-1 A-type and C-type cytochrome c oxidases should operate under aerobic and microaerobic conditions respectively. However, the physiological role of the specific terminal oxidases in S. oneidensis MR-1 remains to be revealed.
Very recently, while our work was in progress, Zhou et al. [25] conducted a relevant study mainly based on gene expression providing a better understanding of the oxygen respiratory metabolism by S. oneidensis MR-1. However, because of translational repressor proteins, antisense RNA and posttranslational modifications, mRNA abundance level can be unrelated to active protein abundance level. Thus, many studies suggest that mRNA expression patterns are necessary but insufficient for quantitative description of biological systems [26]–[29]. For example, comparison of the transcriptome and proteome data for 27 proteins regulated by the ferric uptake regulator in S. oneidensis MR-1 revealed that the expression patterns was correlated with the gene expression data for about half of the proteins whereas microarray data and proteome data were not correlated or even inversely correlated for the others proteins [27].
In this work, we used a biochemical approach and directly measured oxidase activities coupled to mass-spectrometry analysis to further characterize the terminal segment of the respiratory chain of S. oneidensis MR-1 depending on the growth conditions. Measurements performed on solubilized membranes of wild-type and terminal oxidases deficient strains clearly indicate that the quinol oxidase activity, arising from the bd-type oxidase, is extremely higher under microaerobic conditions than under aerobic conditions which is consistent with the fact that bd-type oxidases are generally induced under O2 limited conditions. Surprisingly, cbb 3-type oxidase is the only cytochrome c oxidase present in S. oneidensis MR-1 membranes under both aerobic and microaerobic conditions. The overall results suggest that the A-type cytochrome c oxidase, predicted to have a low affinity for O2, is not present in the bacterium, in our growth conditions, whatever the oxygen tension. A thorough comparison between our results and those obtained by Zhou and colleagues [25] is developed in the discussion section.
Experimental Procedures
Bacterial Growth Conditions
The bacterial strains used in this study are listed in Table 1. All the strains were maintained on Luria-Bertani (LB) agar plates containing the appropriate antibiotic. Overnight cultures of the Escherichia coli strains CC118 λpir and 1047/pRK2013 used for conjugations were carried out in LB medium at 37°C and 250 rpm. S. oneidensis MR-1 and terminal oxidase-deficient strains were grown in LB medium at 30°C either in aerobic conditions or microaerobic conditions. A 30-L BIOSTAT® Cplus-C30-3 fermentor (Sartorius BBI Systems, Germany), controlled by a micro-DCU system and equipped with pH and pO2 sensors, was used for 30-L batch cultures. Bioreactor was inoculated at OD600 nm∼0.2 with 1 L of an overnight flask culture of S. oneidensis MR-1 in LB. The pH of the LB medium was regulated at 7.2 with 5 M H3PO4. The oxygen partial pressure sensor (InPro® 6820, Mettler-Toledo, Switzerland) was calibrated at 100% in air-saturated LB at 30°C which corresponds to an estimated oxygen concentration of 8 mg.L−1 (250 µM). For aerobic cultures, the pO2 was regulated at a minimum of 40% (100 µM O2) whereas for microaerobic cultures the pO2 was regulated at a maximum of 6.5% (16 µM O2). During growth, samples were taken from the cultures at the exponential phase and at the stationary phase, cells were harvested 10 min at 11,400×g and pellets were frozen at −80°C. For growth parameters measurements, strains were inoculated at a OD600 nm∼0.05 in 500 mL-flask with baffled base containing 100 mL LB and were aerobically grown at 30°C and 250 rpm. Samples were periodically taken in sterile conditions.
Table 1. Strains used in this study.
| Strain | Description | Reference or source |
| E. coli | ||
| DH5α | Competent cells for cloning | New England Biolabs |
| 1047/pRK2013 | Helper stain for conjugation | [76] |
| CC118λpir | Host strain for pKNG101 replication and donor strain for conjugation | [77] |
| S. oneidensis a | ||
| MR-1 | Wild-type | Laboratory collection |
| SLL01 | SO2364 deletion mutant/ΔccoN | This study |
| SLL02 | SO3286 deletion mutant/ΔcydA | This study |
| SLL03 | SO4607 deletion mutant/ΔcoxA | This study |
| SLL05 | SO4607 and SO3286 deletion mutant/ΔcoxAΔcydA | This study |
| SLL06 | SO4607 and SO2364 deletion mutant/ΔcoxAΔccoN | This study |
All S. oneidensis strains derived from the parental strain S. oneidensis MR-1.
Construction of the Deletion Mutants
S. oneidensis deletion mutants were constructed by double crossover events using the suicide plasmid pKNG101 as previously described [30]. Briefly, 500 bp regions upstream and downstream the gene to be deleted were amplified by overlapping PCR with Pfu DNA polymerase (Promega) and S. oneidensis MR-1 genomic DNA as template. Primers used for generating PCR products are listed in Table S1. The purified PCR product was cloned into the pGEM®-T Easy vector (Promega) and subcloned into pKNG101 using restriction sites SpeI and SalI. The resulting plasmid was transferred into the plasmid donor strain E. coli CC118λpir and mobilized into the appropriate S. oneidensis strain by conjugation with the helper strain E. coli 1047/pRK2013. Integration of the mutagenesis construct into the chromosome was selected by streptomycin resistance. Clones in which the double recombination events occurred were selected on LB agar plates containing 6% sucrose. Colonies that were resistant to sucrose and susceptible to streptomycin were screened for the deletion event by PCR and confirmed by DNA sequencing. Following this strategy, the SO2364, SO4607 and SO3286 genes encoding respectively the catalytic subunit of the annotated cbb 3-type cytochrome c oxidase, cytochrome c oxidase and cytochrome d ubiquinol oxidase were deleted. All the single and double deletion strains are listed in Table 1.
Preparation and Solubilization of Membranes
About 2 g of cells were resuspended in 20 mM Tris-HCl pH 7.6 supplemented with DNase I and protease inhibitors (Roche) and disrupted twice with an ultra high pressure cell homogenizer (Stansted Fluid Power). Debris and unbroken cells were removed by centrifugation for 10 min at 5,000×g and the supernatant was ultracentrifuged for 1 h at 230,000×g at 4°C to pellet the membranes. Membranes were gently resuspended in 20 mM Tris-HCl pH 7.6 with 5% glycerol to a protein concentration of 10 mg.mL−1 and n-dodecyl β-D-maltoside (DDM) was added to a final concentration of 1% (w/v). Solubilization was performed on a rotary tube mixer for 2 h at 4°C. After ultracentrifugation for 1 h at 230,000x g at 4°C, the supernatant containing the DDM-solubilized membranes was retained and protein concentration was determined with a bicinchoninic acid protein assay kit (Sigma) using bovine serum albumin as protein standard.
Spectral Analysis on Solubilized Membranes
Reduced minus oxidized difference absorbance spectra were recorded on a Lambda 25 UV/VIS spectrophotometer (PerkinElmer) at room temperature. 1 mM potassium ferricyanide was used as the oxidizing agent and a few grains of sodium dithionite was used as the reducing agent.
Cytochrome c Oxidase Activity and O2 Uptake Measurements
Cytochrome c from equine heart was reduced with 75 mM sodium ascorbate in 50 mM sodium phosphate buffer pH 7 with 5% glycerol (v/v). Excess of reductant was removed using a PD10 column (GE Healthcare) and reduced cytochrome c was stored at −20°C. Cytochrome c oxidase activity was spectrophotometrically measured, on a Lambda 25 UV/VIS spectrophotometer (PerkinElmer) at 550 nm at 30°C in a reaction mixture containing 50 mM sodium phosphate buffer pH 7.4 and 75 µM reduced cytochrome c. The reaction was initiated by the addition of solubilized membranes. O2 reductase activity on solubilized membranes was polarographically measured with a Clark-type oxygen electrode (Hansatech Instruments Oxygraph) in a stirred volume of 1 mL of 50 mM sodium phosphate buffer pH 7.4 using either ubiquinol-1 or N,N,N′,N′-tetramethyl-p-phenylenediamine (TMPD) as electron donor. Ubiquinol-1 was prepared from ubiquinone-1 (Sigma) by reduction with sodium dithionite and sodium borohydride as previously described [31]. Quinol oxidase activity was determined in the presence of 0.9 mM ubiquinol-1 and 10 mM DTT whereas TMPD oxidase activity assays were carried out with 100 µM TMPD and 10 mM sodium ascorbate. Reactions were initiated by the addition of DDM-solubilized membranes and the consumption of dioxygen was recorded at 30°C. Data were analyzed using the O2 View software (version 1.02; Hansatech Instruments Ltd.). For inhibition studies, membranes were pre-incubated with 50 µM KCN for 10 min and added to the reaction mixture containing 50 µM KCN.
Separation by Blue-native gel (BN-gel) Electrophoresis
BN-gels were performed using a mini VE cell electrophoresis apparatus (Amersham Pharmacia) according to the method of Schägger [32], [33], as described by Guiral et al. [34] except that the intensity was 10 mA/gel and the voltage 250 V. Running times were 3 h. The gels were run with the blue cathode buffer (50 mM Tricine, 7.5 mM imidazole pH 7.0, 0.02% Coomassie Blue G 250 (Serva Blue G)) until the front line had entered into half of the gels and then the cathode buffer was replaced by the one that contained only 0.002% Serva Blue G. The anode buffer was constituted of 25 mM imidazole pH 7.0. The apparent molecular mass of proteins was estimated using the native molecular mass markers NativeMark (Invitrogen).
In-gel Enzymatic Activities
Cytochrome c oxidase activity was revealed on the BN-gel at 30°C using 3,3′-diaminobenzidine and cytochrome c from equine heart as previously described [34]. TMPD oxidase activity was revealed on the BN-gel at 30°C using 1 mM TMPD as electron donor in 50 mM sodium phosphate buffer pH 7. Protein bands were cut out from the gel and stored at −20°C before mass spectrometry analysis.
Protein Identification by In-gel Digestion and Mass Spectrometry
Tryptic digestion experiments: a robotic workstation (Freedom EVO 100, TECAN) was used to perform automated sample preparation, including multiple steps: washes, reduction and alkylation, digestion by trypsin (PROMEGA, proteomics grade with 0.025% ProteasMax) and extraction of tryptic peptides.
Electrospray quadrupole time of flight (ESI-Q-ToF) analyses were performed on a Synapt G1 mass spectrometer (Waters, Manchester) equipped with a NanoLockSpray ion source and coupled to a nano flow UPLC nanoAcquity (Waters, Manchester). Tryptic peptides were dissolved in 3% acetonitrile/0.1% TFA in water and desalted on a C18 nano trap, before on line elution onto a C18 column (BEH 130 C18, 100 µm x 10 cm, 1.7 µm, Waters). Peptides were eluted with a linear gradient from 3% to 50% of mobile phase B (100% acetonitrile/0.1% formic acid) in A (0.1% formic acid in water) for 30 min. The peptides were detected into the mass spectrometer in a positive ion mode using the MSe mode. The doubly charged ion of GluFibrinopeptide (785,84 Da) was used as lock mass. Processing of the spectra and protein search were made by Protein Lynx Global Server 2.5.2 (Waters) using the following parameters: Shewanella oneidensis database (8 674 entries, with keratins and trypsin), trypsin enzyme, two miscleavages, carbamidomethylation of cysteine as a fixed modification, oxidation of methionine as an optional modification and a mass tolerance set as automatic. Proteins were considered as identified by passing these two filters: minimum 5 fragment consecutive ions from the b/y series matches per peptide and minimum 2 peptide matches per protein.
Sequence Analysis
Membrane-spanning helices were searched for using the program TMpred [35]. Multiples sequences were aligned using the program CLUSTAL W [36] and the HCO phylogenetic tree including various Shewanella A-type cytochrome c oxidases was built with MEGA5 (http://www.megasoftware.net) [37].
Results
Sequence Analysis of the S. oneidensis MR-1 Terminal Oxidases
The cbb 3-type oxidase (Fig. 1A)
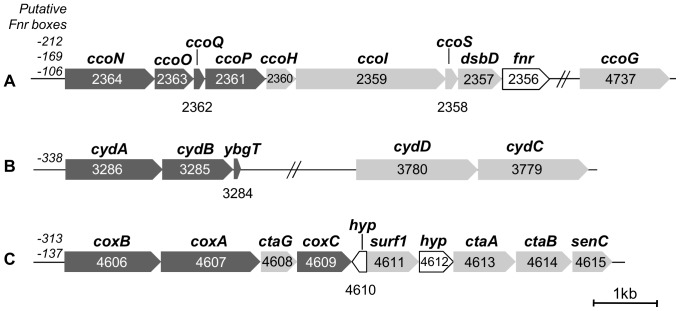
Figure 1. Gene clusters encoding the respiratory oxidases of S. oneidensis MR-1.
Genes encoding the structural subunits are shown in dark grey, genes involved in maturation and assembly of the oxidases are shown in light grey and genes encoding hypothetical proteins (hyp) or other proteins are shown in white. Upstream each gene cluster are indicated the positions of the putative Fnr boxes. The number indicated for each gene corresponds to the locus tag. Gene clusters encode the cbb 3 oxidase (A), the bd-type quinol oxidase (B) and the A-type cytochrome c oxidase Cox (C). The scale bar represents 1 kb.
The structural subunits of the cbb 3-oxidase are encoded by four genes (SO2364-SO2361) organized in the putative ccoNOQP operon. Sequence analysis of ccoN, encoding the catalytic subunit of 53.5 kDa, indicates 12 predicted transmembrane helices which harbor the conserved histidines required for ligating the heme b and the heme b 3/CuB binuclear center [38]. Inspection of the derived amino acid sequences revealed that ccoO and ccoP encode a monoheme c-type cytochrome containing a predicted N-terminal transmembrane helix and a diheme c-type cytochrome containing two predicted N-terminal helices respectively. ccoQ is predicted to encode a small protein of 6.5 kDa containing a transmembrane helix. The ccoGHIS (or fixGHIS) gene cluster whose expression is required for the assembly of a functional cbb 3 oxidase is found close to the ccoNOPQ operon in most bacteria expressing a cbb 3-type oxidase [39]–[41]. Interestingly, ccoG, encoding a putative polyferredoxin, is a monocistronic predicted gene (SO4737), not genomically adjacent to the cco genes. Moreover, an additional gene annotated disulfide bond oxidoreductase dsbD (SO2357), is found downstream of ccoS. Its predicted amino acid sequence exhibits significant similarity (77%) with the protein VC1435 from Vibrio cholerae located at the same position in the cco gene cluster and required for assembly of the cbb 3 oxidase [42].
The bd-type quinol oxidase (Fig. 1B)
The gene cluster SO3286–SO3284 comprises the genes coding for the two subunits CydA (58 kDa) and CydB (42 kDa). The deduced amino acid sequences reveal a high similarity to CydA and CydB from E. coli (71% and 66.5% of sequence identity, respectively). CydA is predicted to have nine transmembrane helices and sequence alignment clearly indicates that the residues ligating the low-spin heme b 558 and the high-spin heme b 595 are conserved in S. oneidensis MR-1. CydB is predicted to contain eight membrane-spanning helices like its counterpart in E. coli [5]. A third gene located in the 3′ region of cydAB and annotated ybgT in the S. oneidensis MR-1 genome encodes a very small protein of 38 aa predicted to contain a membrane spanning helix. This protein shares 58% sequence identity with the E. coli YbgT homologue designated as CydX which has recently been described as a subunit of the cytochrome bd oxidase complex required for complex activity [43]. A second gene cluster (SO3780–SO3779) is composed of cydC and cydD, two genes essential for the assembly of cytochrome bd-I in E. coli [44]–[47].
The A-type cytochrome c oxidase (Fig. 1C)
The cox gene cluster (SO4606–SO4609) is predicted to encode the three subunits of a third terminal oxidase (Cox) annotated aa 3-type cytochrome c oxidase in the S. oneidensis MR-1 genome. The predicted subunit CoxA (59 kDa) comprises 12 transmembrane helices and presents 76% of sequence similarity with the catalytic subunit of the aa 3 oxidase from Paracoccus denitrificans. The six histidines binding the heme-copper binuclear center and the low-spin heme are conserved. The conserved residues forming the D and K-channels and the GHPE260V motif allow to classify Cox in A-type oxidases in the A1 subfamily. The predicted subunit II CoxB (55 kDa) contains the five conserved ligands of the binuclear CuA center typical of cytochrome c oxidase. Sequence alignment with the subunit II of the aa 3 cytochrome c oxidase from P. denitrificans revealed an extension of about 230 amino acid residues in the C-terminal part of CoxB containing two c-type heme binding consensus sequences. This extra domain likely binds two c-type hemes, an uncommon feature among HCO. The predicted subunit III CoxC (37 kDa) comprises seven predicted transmembrane helices and does not contain any cofactor.
The cox gene cluster includes genes involved in the maturation and assembly of the oxidase. SenC is known to bind copper and to be involved in the cytochrome c oxidase assembly in yeast or bacteria [48]–[50]. ctaA (SO4613) and ctaB (SO4614) encode putative heme a and heme o synthase respectively. Surf1 is involved in heme a incorporation during bacterial cytochrome c oxidase biogenesis [51] and CtaG is generally considered as the assembly factor that inserts copper into the CuB center [52], [53].
As Cox from S. oneidensis MR-1 has never been biochemically or spectroscopically investigated, the nature of its heme cofactors is unknown. In subunit I of cytochrome c oxidase from P. denitrificans, Arginine 54 is in interaction with the formyl group of heme a [54], [55]. This residue which is not present in oxidases known to incorporate a different type of heme at the “low-spin site” [2] is conserved in S. oneidensis MR-1 CoxA. The presence of this residue as well as the ctaA and ctaB genes required for the heme a synthesis in the cox gene cluster lead us to believe that an a-type heme is incorporated at the “low-spin site” in the subunit I of Cox.
Finally, the presence of the fnr (fumarate nitrate regulator) gene directly downstream of the cco genes and the presence of predicted Fnr-binding sites in the upstream promoter regions of the different gene clusters suggest a direct regulation by the Fnr transcriptional regulator.
Spectral Properties of S. oneidensis MR-1 Solubilized Membranes
To detect the different terminal oxidases, solubilized membranes from S. oneidensis MR-1 aerobically or microaerobically grown were studied by light absorption spectroscopy (Figure 2). Spectra show the presence of cytochrome c with peaks at 522 nm and 552 nm and cytochrome b with characteristic shoulders at 530 nm and 562 nm. These peaks are due to the contribution from diverse membrane cytochromes. The heme c/heme b ratio is higher (about twofold) in membranes isolated from cells grown under microaerobic conditions (Figure 2 A, B) than under aerobic conditions (Figure 2 C, D). The peak at 632 nm indicates the presence of a heme d, cofactor of the bd-type quinol oxidase and the trough around 650 nm most likely originates from a stable oxygenated cytochrome d species [56]. Furthermore, no a-type heme was detected on the spectra of membranes isolated from bacterial cells grown in microaerobic or aerobic conditions. Indeed, membranes isolated from this strain show no signal in the 440 nm-region (data not shown) and in the 600 nm-region. These results indicate the presence of the bd-type quinol oxidase in the bacterial membranes and the possible presence of the cbb 3 oxidase in all tested conditions. However, the fact that we did not spectroscopically detect heme a leads us to believe that Cox is not present in S. oneidensis MR-1 membranes even in aerobic conditions. Nevertheless, we cannot rule out the possibility that Cox does not contain any heme a as a cofactor. The possible lack of the A-type cytochrome c oxidase in the membranes must be confirmed with other techniques.
Figure 2. Reduced minus oxidized difference absorbance spectra of S. oneidensis MR-1 solubilized membranes.
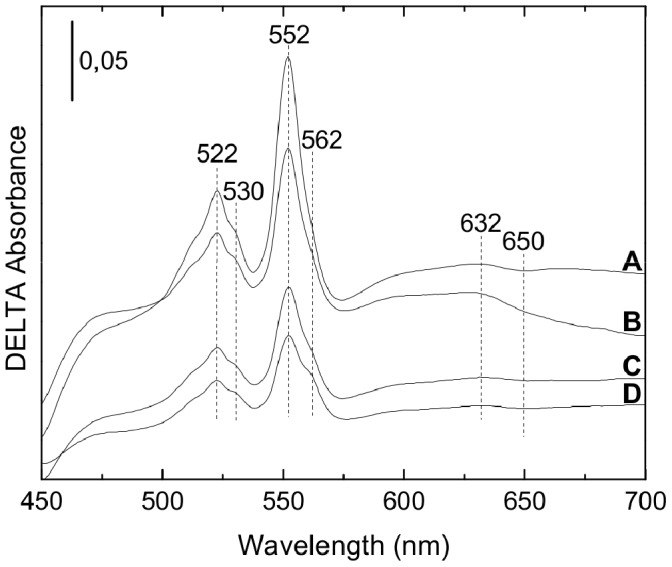
Solubilized membranes in 20-HCl pH 7.6, 5% glycerol, 1% DDM were oxidized using 1 mM potassium ferricyanide and reduced by adding few grains of sodium dithionite. Membranes were isolated from strain MR-1 grown under microaerobic conditions (spectra A and B) or aerobic conditions (spectra C and D). Bacterial cells were harvested in the exponential (spectra A and C) or stationary phase (spectra B and D) of growth. The concentration of proteins was 4.8 mg.mL−1. The spectra were recorded at room temperature. The vertical bar indicates the absorption scale.
TMPD, Cytochrome c and Quinol Oxidase Activities in Solubilized Membranes of S. oneidensis MR-1
To further investigate the role of the different terminal oxidases in aerobic respiration in S. oneidensis MR-1, catalytic activity of cytochrome c oxidases and bd-type quinol oxidase were spectrophotometrically and polarographically measured with solubilized membranes prepared from cells grown in LB medium under aerobic and microaerobic conditions (Table 2).
Table 2. TMPD, cytochrome c and quinol oxidase activities in solubilized membranes of S. oneidensis MR-1.
| Aerobic conditions | Microaerobic conditions | |||
| EPa | SPb | EP | SP | |
| TMPD oxidase activity (nmol O2.min−1.mg protein−1) | 90+/−6 | 95+/−6 | 94+/−4 | 84+/−3 |
| Cyt c oxidase activity (nmol cyt c.min−1.mg protein−1)c | 6.2+/−0.2 | 8.6+/−0.5 | 7.3+/−0.5 | 5.2+/−0.6 |
| Ubiquinol-1 oxidase activity (nmol O2.min−1.mg protein−1) | 9+/−1 | 20+/−1 | 364+/−18 | 456+/−17 |
EP: exponential phase of growth.
SP: stationary phase of growth.
cyt: cytochrome.
Listed values are averages of at least three separate experiments.
Cytochrome c oxidase activity was assessed by following oxidation of horse heart reduced cytochrome c by spectrophotometry and by monitoring O2 consumption in the presence of TMPD, an artificial electron donor capable of reducing cytochrome c. The values obtained by both methods indicate that cytochrome c oxidase activity is relatively constant, regardless of the tested culture conditions and growth phase.
Quinol oxidase activity of the cytochrome bd-type oxidase was determined by measuring O2 uptake with ubiquinol-1 as the electron donor. While the values obtained with membranes from cells grown in aerobic conditions were low and near to the detection limit, quinol-dependent O2 consumption was significantly higher with membranes from microaerobic cultures with a 20–40 fold increase depending on the growth phase. In addition, the oxygen consumption in the presence of ubiquinol-1 was lower with membranes prepared from cells harvested during the exponential phase of growth compared to the stationary phase under both aerobic and microaerobic conditions. Our data are consistent with the fact that bd-type oxygen reductases are generally expressed under O2-limited conditions i.e. in microaerobic conditions or when cultures enter the stationary phase, the decrease in oxygen tension resulting from the increase in cell density [14], [57].
Furthermore, the addition of 50 µM KCN strongly inhibited TMPD-dependent O2 consumption and cytochrome c oxidase activity (90–100%) whereas the quinol-dependent O2 uptake was slightly affected with a 10–15% inhibition (data not shown). This observation is in accord with the fact that a specific feature of cytochrome bd-type oxidase is its much lower sensitivity to cyanide than heme-copper oxygen reductases [5].
Identification of Cytochrome c Oxidases by Mass Spectrometry
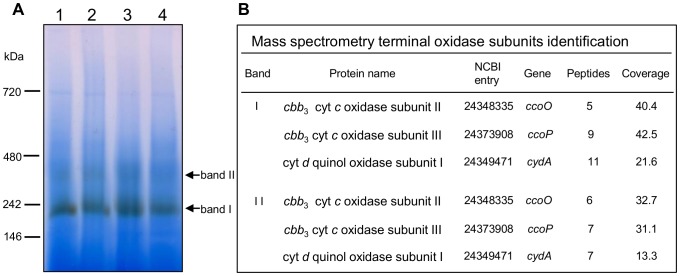
In order to identify enzymes responsible for the measured cytochrome c oxidase activity, solubilized S. oneidensis MR-1 membrane proteins were separated on a BN-gel electrophoresis followed by an in-gel detection of the cytochrome c oxidase activity (Fig. 3A). For this purpose, S. oneidensis MR-1 cells were grown aerobically or microaerobically and harvested during the exponential or the stationary phase of growth to determine the influence of the oxygen level and the growth phase on the respiratory cytochrome c oxidase content in the membranes. BN-gel of the solubilized membranes shows one major brown band of cytochrome c oxidase activity, of molecular mass around 240 kDa (Fig. 3A, band I). Another faint band of activity could be detected, of molecular mass around 450 kDa (Fig. 3A, band II). The pattern of cytochrome c oxidase activity bands was identical whatever the oxygen tension in the medium and the growth phase. When TMPD was used instead of horse heart cytochrome c to reveal cytochrome c oxidase activity, the same pattern was also observed (data not shown). With both electron donors, no band was revealed when the in-gel detection was performed in the presence of 50 µM potassium cyanide (data not shown).
Figure 3. Identification of cytochrome c oxidase(s) in S. oneidensis MR-1 membranes under aerobic and microaerobic conditions.
A. In-gel detection of cytochrome c oxidase activity in solubilized membranes on BN-gel. Membrane proteins were prepared from S. oneidensis MR-1 cells grown aerobically (lanes 1 and 2) or microaerobically (lanes 3 and 4) and harvested during the exponential (lanes 1 and 3) or the stationary (lanes 2 and 4) phase of growth. 130 µg of total proteins were loaded on a 5–15% polyacrylamide gel. The major brown band (band I) and the faint brown band (band II) of activity are indicated by arrows. B. Terminal oxidases subunits identified by ESI-Q-ToF mass spectrometry. Data correspond to identified proteins in bands I and II from exponentially grown cells under aerobic conditions (panel A, lane 1). Table heading: Band, roman figures refer to the protein bands from the BN gel shown in panel A; Protein name, name in NCBI database; NCBI entry, accession number; Peptides, number of unique peptides detected; Coverage, protein sequence coverage by the matching peptides (in %). Cyt: cytochrome.
To identify the cytochrome c oxidases, the protein band I and band II were directly cut out from the gel, for the four growth conditions, digested with trypsin and analyzed by ESI-Q-ToF mass spectrometry (Fig. 3B). Band I and band II were found to contain, among others proteins, CcoO and CcoP, the monohemic and dihemic cytochrome c subunits of the cbb 3 oxidase respectively. In addition, the CydA subunit of the bd-type quinol oxidase was also identified in band I and band II indicating that the cbb 3 oxidase and the bd-type quinol oxidase co-migrate in the BN gel. However, none of the three subunits of the Cox enzyme was identified in solubilized membranes of S. oneidensis MR-1 grown microaerobically or aerobically. These results strongly suggest that, against all expectation, the high O2-affinity cbb 3 oxidase is responsible for the cytochrome c oxidase activity measured in the membranes of microaerobically but also aerobically grown S. oneidensis MR-1 whereas the A-type cytochrome c oxidase Cox remains undetectable even under aerobic conditions.
Growth Parameters and TMPD Oxidase Activity of S. oneidensis MR-1 Wild Type and Oxidase Deletion Mutants
The first part of the study suggests that the bd-type and the cbb 3-type high-affinity oxidases are important under aerobic conditions while the A-type cytochrome c oxidase would be surprisingly dispensable despite its low affinity for O2. To confirm these data and determine the physiological significance of the different oxygen reductases under aerobic conditions, deletion mutants were constructed and characterized during the exponential phase of growth under aerobic conditions by measuring growth parameters and TMPD oxidase activities (Table 3). While the strain lacking the cbb 3-type terminal oxidase (SLL01) grew significantly more slowly and reached the stationary phase at lower cell density compared to the wild type, the single oxidase mutants lacking the bd-type terminal oxidase (SLL02) or the A-type cytochrome c oxidase Cox (SLL03) grew as well as the wild type. Firstly, the results indicate that none of the terminal oxidases are essential for aerobic growth in our conditions. Secondly, the reduced growth rate of SLL01 suggests that the cbb 3-type terminal oxidase is the predominant oxidase in exponentially growing S. oneidensis MR-1 cells under aerobic conditions.
Table 3. Growth parameters and TMPD-dependent oxidase activity of oxidase deletion mutants cultivated in aerobic conditions.
| Strain | Relevant genotype | Doubling timea | Relative yieldb | TMPD oxidase activityc |
| MR-1 | wild-type | 54.2+/−0.2 | 1.00 | 90+/−6 |
| SLL01 | ΔccoN | 78.0+/−0.7 | 0.77 | 0 |
| SLL02 | ΔcydA | 53.4+/−2.1 | 1.00 | 91+/−9 |
| SLL03 | ΔcoxA | 59.0+/−1.1 | 0.96 | 97+/−6 |
| SLL05 | ΔcydAΔcoxA | 56.5+/−2.7 | 0.97 | 101+/−9 |
| SLL06 | ΔccoNΔcoxA | 79.9+/−1.2 | 0.78 | 0 |
Expressed in min.
Relative yield is defined as the highest optical density in early stationary phase in mutant strains cultures relative to that in the wild-type strain cultures.
Expressed in nmol O2.min−1.mg protein−1.
Data were obtained from triplicate separated experiments.
The strains lacking two terminal oxidases were also characterized. Growth parameters of the strain containing only the cbb 3-type oxidase (SLL05) did not really differ from those of the wild-type indicating that the cbb 3-type oxidase is sufficient to support maximum growth rate in aerobic conditions. Conversely, doubling time of the strain containing only the bd-type oxidase (SLL06) was significantly longer and the growth yield lower than those of the wild-type strain, with values similar to those of the single oxidase mutants lacking cbb 3-type oxidase (SLL01). Interestingly, it was not possible, despite numerous efforts, to generate a double oxidase mutant lacking both bd-type and cbb 3-type oxidases suggesting that this mutant was not viable under aerobic conditions.
Taken together, these results demonstrate that the cbb 3-type or the bd-type oxidase is required for aerobic growth of S. oneidensis MR-1 and that the cbb 3-type terminal oxidase is the major oxidase in exponentially growing cells. These findings are consistent with the results obtained in this work with the wild type and also suggest that the low O2-affinity cytochrome c oxidase Cox is unexpectedly not involved in aerobic respiration in our experimental conditions.
Furthermore, TMPD-dependent oxidase activity was determined with S. oneidensis MR-1 wild type strain and oxidase deletion mutants cultivated under aerobic conditions. First, the strain SLL06 exhibited no activity indicating that TMPD was not oxidized by the bd-type oxidase. This result contrasts with the cytochrome bd oxidase from E. coli which can oxidize TMPD with a quite high turnover number [58]. However, several previously reported cytochrome bd-oxidases have been found to exhibit low TMPD oxidase activity as in Azotobacter vinelandii and Geobacillus thermodenitrificans K1041 (formerly Bacillus stearothermophilus K1041), or no TMPD oxidase activity as in Bacillus firmus OF4 [59]–[62]. Besides, TMPD oxidase activity was not affected in the single oxidase mutants lacking cydA (SLL02) or coxA (SLL03) compared to the wild-type strain whereas no activity was measured with the strain lacking ccoN (SLL01). Finally, TMPD oxidase activity did not differ from that of the wild type in the strain containing only the cbb 3-type oxidase (SLL05). These data indicate that TMPD oxidase activity arose from the cbb 3-type oxidase only suggesting that the cbb 3-type oxidase is the only HCO present in the aerobic membranes of S. oneidensis MR-1. Moreover, this could also explain the low values obtained from measurements of cytochrome c oxidase activity in Table 2 since a previous study revealed that horse heart cytochrome c was a poor substrate for the cbb 3-type oxidase in Vibrio cholerae, with a rate of oxygen reduction 10–20 fold lower than with the diheme cytochrome c 4 identified as a natural electron donor [63].
Spectral Properties of S. oneidensis MR-1 Oxidase Deletion Mutants Solubilized Membranes
Solubilized membranes from aerobically grown S. oneidensis MR-1 and strains lacking terminal oxidases (single and double mutants) were studied by light absorption spectroscopy (Fig. 4). Dithionite-reduced minus ferricyanide-oxidized spectra demonstrate the presence of c-type and b-type cytochromes (peaks at 522, 552, and shoulders at 530 and 562 nm respectively) in membranes from all strains. The study of the mutants confirmed that the peak at 632 nm and the trough at 650 nm arise from the heme d of the bd-quinol oxidase since they are absent on spectra from SLL02 and SLL05, both strains lacking this quinol oxidase. Additionally, spectral signals for a-type heme (around 440 and 600 nm) were not detected in any of the strains confirming the absence of Cox in the membrane of S. oneidensis MR-1 in aerobic conditions.
Figure 4. Reduced minus oxidized difference absorbance spectra 0 of membranes from aerobically grown S. oneidensis MR-1 oxidase mutants.
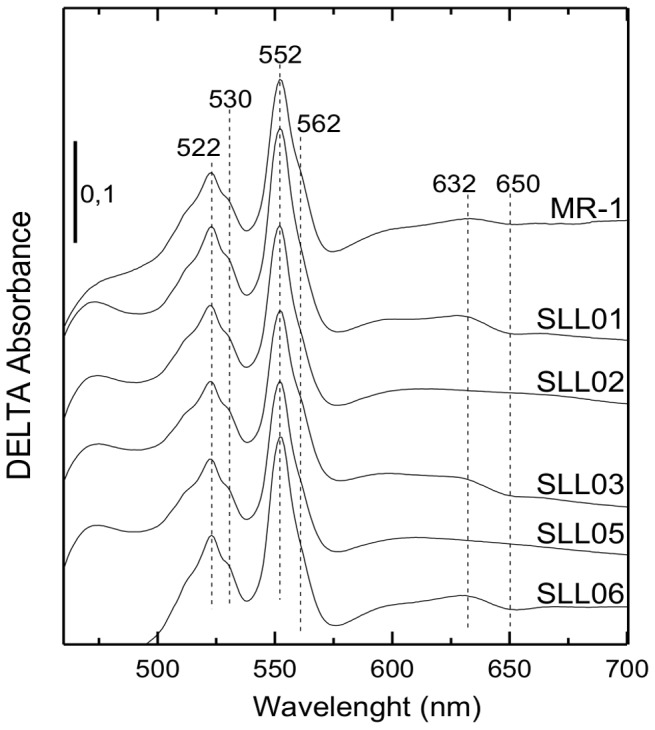
Solubilized membranes in 20-HCl pH 7.6, 5% glycerol, 1% DDM were oxidized using 1 mM potassium ferricyanide and reduced by adding few grains of sodium dithionite. Membranes were isolated from aerobically grown cells harvested in the exponential phase of growth. The concentration of proteins was 11 mg.mL−1. The spectra were recorded at room temperature. The vertical bar indicates the absorption scale.
Identification of Cytochrome c Oxidases in S. oneidensis MR-1 Oxidase Mutants Membranes by Mass Spectrometry
Membrane proteins from S. oneidensis MR-1 and oxidase mutant strains grown under aerobic conditions were separated on BN-gel electrophoresis and the cytochrome c oxidase activity was revealed (Fig. 5). One band of activity, of molecular mass around 240 kDa, was observed in all strains except for the strain lacking the cbb 3 oxidase (SLL01) and the strain lacking both the cbb 3 and the Cox oxidases (SLL06). If the BN-gel is incubated a longer time for the detection of the cytochrome c oxidase activity, a faint band of activity is detected, of molecular mass around 450 kDa (not shown) in all strains except for SLL01 and SLL06. These results strongly suggest that the cbb 3-oxidase is responsible for the bands of activity in the solubilized membranes of the bacterium. Furthermore, the analysis of these bands of activity by ESI-Q-ToF mass spectrometry (data not shown) confirmed that the cytochrome c oxidase responsible for in-gel activity is the cbb 3 oxidase since both CcoO and CcoP, the monohemic and dihemic cytochrome c subunits of the cbb 3 oxidase respectively are identified. In contrast, none of the A-type Cox subunits was identified. Collectively, these results undoubtedly demonstrate that under aerobic conditions, the cytochrome c oxidase activity detected in the membranes from S. oneidensis MR-1 arises from the high-affinity cbb 3 oxidase and that, on the contrary, the low-affinity A-type cytochrome c oxidase is surprisingly not required for aerobic respiration.
Figure 5. Detection of cytochrome c oxidase activity on BN-gel in aerobically grown S. oneidensis MR-1 oxidase mutants.
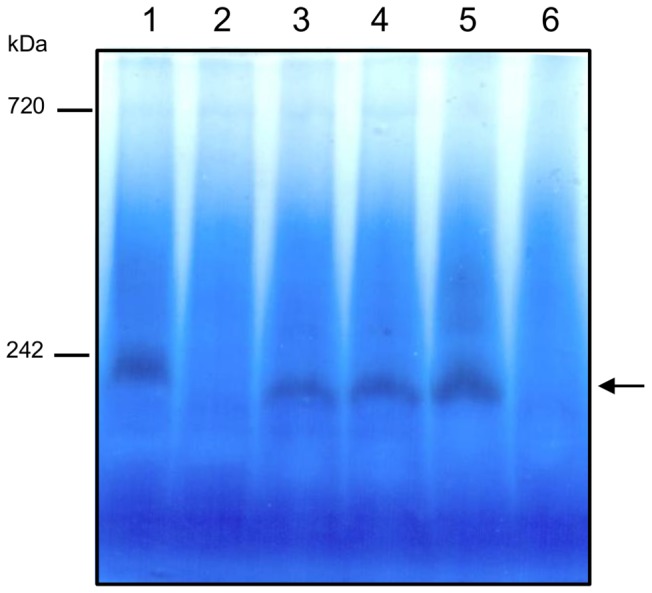
Membrane proteins were prepared from wild-type or oxidase mutants S. oneidensis MR-1 cells grown aerobically and harvested during the exponential phase of growth. 130 µg of total proteins were loaded on a 5–15% polyacrylamide gel. The band of activity is indicated by an arrow. Lane 1: S. oneidensis MR-1. Lane 2: strain SLL01. Lane 3: strain SLL02. Lane 4: strain SLL03. Lane 5: strain SLL05. Lane 6: strain SLL06.
Discussion
In this report, we investigated the aerobic respiratory pathway in S. oneidensis MR-1 to determine the physiological role of the three terminal oxidases in low and in high-O2 environments. Our results suggest that the high-affinity bd-type oxidase is weakly expressed in aerobic conditions and significantly induced under microaerobic conditions which is consistent with previous studies indicating that bd oxidases are typically used by bacteria for aerobic respiration under O2-limited conditions [5], [64], [65]. Taken together, our data also reveal that the cbb 3-type oxidase is the most important terminal oxidase under aerobic conditions and has a significant role under microaerobic conditions whereas the low affinity A-type cytochrome c oxidase Cox was not detected in the tested conditions even under aerobic conditions. In addition, characterization of the terminal oxidase deletion mutants confirms that the cbb 3-type or the bd-type oxidase is required for aerobic growth and that Cox is unexpectedly not involved in aerobic respiration in these experimental conditions.
Our results are overall consistent with those recently reported by Zhou et al. [25] who revealed, by measuring the mRNA abundance and the promoter activity in S. oneidensis MR-1, that cbb 3-type oxidase and bd-type oxidase are important under microaerobic conditions and the cbb 3-type is the major oxidase under aerobic conditions while Cox has no physiological significance. However, it is noteworthy that, in their study, the expression level of the cco genes encoding the cbb 3 oxidase was lower under microaerobic conditions than under aerobic conditions and directly proportional to the O2 level in aerobic cultures, suggesting that expression of cco is favored in O2-rich environments. On the contrary, we clearly present evidence that the oxidase activity arising from the cbb 3 oxidase was constant under the tested conditions and thus did not respond to growth conditions. Furthermore, if Zhou et al. [25] found that the cyd operon was slightly induced at low O2 concentration compared to high O2 conditions (<5-fold increase in mRNA abundance), the difference observed between aerobic and microaerobic conditions was much more significant at the protein level with a 40-fold increase observed by measuring quinol-dependent oxidase activity. Probably due to translational and/or post-translational regulation mechanisms, these discrepancies emphasize the importance of biochemical analysis at the protein level to understand the global metabolic and regulatory processes in addition to studies of expression at a transcriptional level.
These findings are intriguing since cbb 3 oxidases, given their high affinity for O2, are usually repressed at high oxygen tension in many bacteria [66]–[69]. Furthermore, in organisms carrying an aa 3-type and a cbb 3-type oxidase, the low affinity oxidase plays a dominant role under high oxygen conditions whereas the cbb 3 oxidase is induced only under low O2 concentrations [67]. In S. oneidensis MR-1, the uncommon expression pattern of the cytochrome c oxidases is reminiscent of the one found in Pseudomonas aeruginosa. In this bacterium, the cox genes encoding an aa 3-type oxidase are expressed at a very low level under O2-rich growth conditions in LB-medium while the expression level of the genes encoding the cbb 3-1 oxidase was high in the same conditions.
Interestingly, the cox genes were found to be significantly induced in P. aeruginosa under starvation of iron, carbon or nitrogen indicating that the aa 3-type oxidase might be involved in energy conservation under low nutrient conditions [70], [71]. The induction of the cox genes under nutrient starvation or other environmental stress conditions should thus be considered in S. oneidensis MR-1. Although a functional loss of the A-type Cox oxidase in S. oneidensis MR-1, as proposed by Zhou et al. [25], cannot be excluded, the presence of the cox genes as well as of genes involved in the maturation and assembly of the oxidase in all genome-sequenced Shewanella strains leads us to believe that the Cox oxidase has a significant role in some not yet defined conditions. Moreover, in the phylogenetic tree built with the sequences of the catalytic subunits of A, B and C-type oxidases from various Bacteria and Archaea, the A-type Cox from different Shewanella species, among them the one from S. oneidensis MR-1, clustered with bacterial A-type cytochrome c oxidases (data not shown). For Shewanella genes, the observed branch lengths did not indicate a high evolutionary rate characteristic of pseudogenes, which is again not in favour of the hypothesis of a functional loss of the A-type Cox.
In addition to the particularity of the membrane terminal oxidase content in S. oneidensis MR-1, examination of the derived amino acid sequences emphasized the singularity of the Cox oxidase. Indeed, if sequence analysis allowed to classify Cox in the A1 subfamily and to predict the incorporation of a-type heme at least at the low-spin site of CoxA, an uncommon extra C-terminal domain carrying two c-type heme binding consensus sequences was identified in the predicted subunit CoxB. Subunit II with an extra domain carrying only one c-type heme can be found in bacterial terminal oxidases such as in the caa 3 oxidase from Thermus thermophilus or Bacillus subtilis [72], [73]. Intriguingly, the C-terminal extension of CoxB from S. oneidensis MR-1 likely binds two c-type hemes suggesting that this enzyme is a ccaa 3-type HCO. Such an unusual feature was so far described only in Desulfovibrio species [74], [75] but we discover that the two c-type heme binding motifs were also conserved in some γ-proteobacteria such as Shewanella (except in S. violacea and S. benthica KT99) and some species of the genus Psychromonas, Colwellia and Methylosarcina (data not shown). As in Desulfovibrio species, the two heme c domains in S. oneidensis MR-1 are highly similar to each other (61% sequence identity) suggesting that the diheme domain evolved from a gene duplication event [74].
A study is currently underway to determine experimental conditions under which the ccaa 3-type oxidase Cox would be expressed. It should provide information about the physiological significance of this oxidase in S. oneidensis MR-1.
Supporting Information
Oligonucleotides used in this study.
(DOCX)
Acknowledgments
We thank Cécile JOURLIN-CASTELLI, Joshua ARMITANO and Vincent MEJEAN (CNRS, LCB, Marseilles, France) for expert technical assistance in bacterial conjugation experiments, for kindly providing the Shewanella oneidensis MR-1 strain and for helpful discussions. We acknowledge Wolfgang NITSCHKE (CNRS, BIP, Marseilles, France) for his help in phylogenetic analysis and Marianne GUIRAL for critical reading of the manuscript. The Proteomic platform is a member of Marseille Protéomique (http://map.univmed.fr/index.html) labeled IBiSA.
Funding Statement
This work was supported by research grants from the Agence Nationale de la Recherche (EngineeringH2cyano and Algo-H2 projects), the Centre National de la Recherche Scientifique (CNRS) and the Aix-Marseille Université. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1. Garcia-Horsman JA, Barquera B, Rumbley J, Ma J, Gennis RB (1994) The superfamily of heme-copper respiratory oxidases. J Bacteriol 176: 5587–5600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Pereira MM, Santana M, Teixeira M (2001) A novel scenario for the evolution of haem-copper oxygen reductases. Biochim Biophys Acta 1505: 185–208. [DOI] [PubMed] [Google Scholar]
- 3. Brochier-Armanet C, Talla E, Gribaldo S (2009) The multiple evolutionary histories of dioxygen reductases: Implications for the origin and evolution of aerobic respiration. Mol Biol Evol 26: 285–297. [DOI] [PubMed] [Google Scholar]
- 4. Junemann S (1997) Cytochrome bd terminal oxidase. Biochim Biophys Acta 1321: 107–127. [DOI] [PubMed] [Google Scholar]
- 5. Borisov VB, Gennis RB, Hemp J, Verkhovsky MI (2011) The cytochrome bd respiratory oxygen reductases. Biochim Biophys Acta 1807: 1398–1413. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Puustinen A, Finel M, Haltia T, Gennis RB, Wikstrom M (1991) Properties of the two terminal oxidases of Escherichia coli . Biochemistry 30: 3936–3942. [DOI] [PubMed] [Google Scholar]
- 7. Belevich I, Borisov VB, Zhang J, Yang K, Konstantinov AA, et al. (2005) Time-resolved electrometric and optical studies on cytochrome bd suggest a mechanism of electron-proton coupling in the di-heme active site. Proc Natl Acad Sci U S A 102: 3657–3662. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Borisov VB, Murali R, Verkhovskaya ML, Bloch DA, Han H, et al. (2011) Aerobic respiratory chain of Escherichia coli is not allowed to work in fully uncoupled mode. Proc Natl Acad Sci U S A 108: 17320–17324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Borisov VB, Forte E, Davletshin A, Mastronicola D, Sarti P, et al. (2013) Cytochrome bd oxidase from Escherichia coli displays high catalase activity: an additional defense against oxidative stress. FEBS Lett 587: 2214–2218. [DOI] [PubMed] [Google Scholar]
- 10. Borisov VB, Forte E, Konstantinov AA, Poole RK, Sarti P, et al. (2004) Interaction of the bacterial terminal oxidase cytochrome bd with nitric oxide. FEBS Lett 576: 201–204. [DOI] [PubMed] [Google Scholar]
- 11. Borisov VB, Forte E, Sarti P, Brunori M, Konstantinov AA, et al. (2007) Redox control of fast ligand dissociation from Escherichia coli cytochrome bd . Biochem Biophys Res Commun 355: 97–102. [DOI] [PubMed] [Google Scholar]
- 12. Giuffre A, Borisov VB, Mastronicola D, Sarti P, Forte E (2012) Cytochrome bd oxidase and nitric oxide: from reaction mechanisms to bacterial physiology. FEBS Lett 586: 622–629. [DOI] [PubMed] [Google Scholar]
- 13. Richardson DJ (2000) Bacterial respiration: a flexible process for a changing environment. Microbiology 146: 551–571. [DOI] [PubMed] [Google Scholar]
- 14. Poole RK, Cook GM (2000) Redundancy of aerobic respiratory chains in bacteria? Routes, reasons and regulation. Adv Microb Physiol 43: 165–224. [DOI] [PubMed] [Google Scholar]
- 15. Bueno E, Mesa S, Bedmar EJ, Richardson DJ, Delgado MJ (2012) Bacterial adaptation of respiration from oxic to microoxic and anoxic conditions: redox control. Antioxid Redox Signal 16: 819–852. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Morris RL, Schmidt TM (2013) Shallow breathing: bacterial life at low O2 . Nat Rev Microbiol 11: 205–212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Belevich I, Borisov VB, Bloch DA, Konstantinov AA, Verkhovsky MI (2007) Cytochrome bd from Azotobacter vinelandii: evidence for high-affinity oxygen binding. Biochemistry 46: 11177–11184. [DOI] [PubMed] [Google Scholar]
- 18. Belevich I, Borisov VB, Konstantinov AA, Verkhovsky MI (2005) Oxygenated complex of cytochrome bd from Escherichia coli : stability and photolability. FEBS Lett 579: 4567–4570. [DOI] [PubMed] [Google Scholar]
- 19. Heidelberg JF, Paulsen IT, Nelson KE, Gaidos EJ, Nelson WC, et al. (2002) Genome sequence of the dissimilatory metal ion-reducing bacterium Shewanella oneidensis . Nat Biotechnol 20: 1118–1123. [DOI] [PubMed] [Google Scholar]
- 20. Liu C, Gorby YA, Zachara JM, Fredrickson JK, Brown CF (2002) Reduction kinetics of Fe(III), Co(III), U(VI), Cr(VI), and Tc(VII) in cultures of dissimilatory metal-reducing bacteria. Biotechnol Bioeng 80: 637–649. [DOI] [PubMed] [Google Scholar]
- 21. Gralnick JA, Vali H, Lies DP, Newman DK (2006) Extracellular respiration of dimethyl sulfoxide by Shewanella oneidensis strain MR-1. Proc Natl Acad Sci U S A 103: 4669–4674. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Le Laz S, Kpebe A, Lorquin J, Brugna M, Rousset M (2013) H2-dependent azoreduction by Shewanella oneidensis MR-1: involvement of secreted flavins and both [Ni-Fe] and [Fe-Fe] hydrogenases. Appl Microbiol Biotechnol. In Press. 10.1007/s00253-013-5208-z. [DOI] [PubMed]
- 23. Fu H, Chen H, Wang J, Zhou G, Zhang H, et al. (2013) Crp-dependent cytochrome bd oxidase confers nitrite resistance to Shewanella oneidensis . Environ Microbiol 15: 2198–2212. [DOI] [PubMed] [Google Scholar]
- 24. Marritt SJ, McMillan DG, Shi L, Fredrickson JK, Zachara JM, et al. (2012) The roles of CymA in support of the respiratory flexibility of Shewanella oneidensis MR-1. Biochem Soc Trans 40: 1217–1221. [DOI] [PubMed] [Google Scholar]
- 25. Zhou G, Yin J, Chen H, Hua Y, Sun L, et al. (2013) Combined effect of loss of the caa 3 oxidase and Crp regulation drives Shewanella to thrive in redox-stratified environments. ISME J 7: 1752–1763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Gygi SP, Rochon Y, Franza BR, Aebersold R (1999) Correlation between protein and mRNA abundance in yeast. Mol Cell Biol 19: 1720–1730. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Wan XF, Verberkmoes NC, McCue LA, Stanek D, Connelly H, et al. (2004) Transcriptomic and proteomic characterization of the Fur modulon in the metal-reducing bacterium Shewanella oneidensis . J Bacteriol 186: 8385–8400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Nie L, Wu G, Zhang W (2006) Correlation of mRNA expression and protein abundance affected by multiple sequence features related to translational efficiency in Desulfovibrio vulgaris: a quantitative analysis. Genetics 174: 2229–2243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Nie L, Wu G, Zhang W (2006) Correlation between mRNA and protein abundance in Desulfovibrio vulgaris: a multiple regression to identify sources of variations. Biochem Biophys Res Commun 339: 603–610. [DOI] [PubMed] [Google Scholar]
- 30. Kaniga K, Delor I, Cornelis GR (1991) A wide-host-range suicide vector for improving reverse genetics in gram-negative bacteria: inactivation of the blaA gene of Yersinia enterocolitica . Gene 109: 137–141. [DOI] [PubMed] [Google Scholar]
- 31. Rieske JS (1967) Preparation and properties of reduced coenzyme Q-cytochrome c reductase (complex III of the respiratory chain). Methods Enzymol 10: 239–245. [Google Scholar]
- 32. Schagger H, von Jagow G (1991) Blue native electrophoresis for isolation of membrane protein complexes in enzymatically active form. Anal Biochem 199: 223–231. [DOI] [PubMed] [Google Scholar]
- 33. Schagger H, Cramer WA, von Jagow G (1994) Analysis of molecular masses and oligomeric states of protein complexes by blue native electrophoresis and isolation of membrane protein complexes by two-dimensional native electrophoresis. Anal Biochem 217: 220–230. [DOI] [PubMed] [Google Scholar]
- 34. Guiral M, Prunetti L, Lignon S, Lebrun R, Moinier D, et al. (2009) New insights into the respiratory chains of the chemolithoautotrophic and hyperthermophilic bacterium Aquifex aeolicus . J Proteome Res 8: 1717–1730. [DOI] [PubMed] [Google Scholar]
- 35. Claros MG, von Heijne G (1994) TopPred II: an improved software for membrane protein structure predictions. Comput Appl Biosci 10: 685–686. [DOI] [PubMed] [Google Scholar]
- 36. Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22: 4673–4680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Tamura K, Peterson D, Peterson N, Stecher G, Nei M, et al. (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28: 2731–2739. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Buschmann S, Warkentin E, Xie H, Langer JD, Ermler U, et al. (2010) The structure of cbb 3 cytochrome oxidase provides insights into proton pumping. Science 329: 327–330. [DOI] [PubMed] [Google Scholar]
- 39. Preisig O, Zufferey R, Hennecke H (1996) The Bradyrhizobium japonicum fixGHIS genes are required for the formation of the high-affinity cbb 3-type cytochrome oxidase. Arch Microbiol 165: 297–305. [DOI] [PubMed] [Google Scholar]
- 40. Koch HG, Winterstein C, Saribas AS, Alben JO, Daldal F (2000) Roles of the ccoGHIS gene products in the biogenesis of the cbb 3-type cytochrome c oxidase. J Mol Biol 297: 49–65. [DOI] [PubMed] [Google Scholar]
- 41. Kulajta C, Thumfart JO, Haid S, Daldal F, Koch HG (2006) Multi-step assembly pathway of the cbb 3-type cytochrome c oxidase complex. J Mol Biol 355: 989–1004. [DOI] [PubMed] [Google Scholar]
- 42. Braun M, Thony-Meyer L (2005) Cytochrome c maturation and the physiological role of c-type cytochromes in Vibrio cholerae . J Bacteriol 187: 5996–6004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. VanOrsdel CE, Bhatt S, Allen RJ, Brenner EP, Hobson JJ, et al. (2013) The Escherichia coli CydX protein is a member of the CydAB cytochrome bd oxidase complex and is required for cytochrome bd oxidase activity. J Bacteriol 195: 3640–3650. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Georgiou CD, Fang H, Gennis RB (1987) Identification of the cydC locus required for expression of the functional form of the cytochrome d terminal oxidase complex in Escherichia coli . J Bacteriol 169: 2107–2112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Poole RK, Hatch L, Cleeter MW, Gibson F, Cox GB, et al. (1993) Cytochrome bd biosynthesis in Escherichia coli: the sequences of the cydC and cydD genes suggest that they encode the components of an ABC membrane transporter. Mol Microbiol 10: 421–430. [PubMed] [Google Scholar]
- 46. Bebbington KJ, Williams HD (1993) Investigation of the role of the cydD gene product in production of a functional cytochrome d oxidase in Escherichia coli . FEMS Microbiol Lett 112: 19–24. [DOI] [PubMed] [Google Scholar]
- 47. Poole RK, Gibson F, Wu G (1994) The cydD gene product, component of a heterodimeric ABC transporter, is required for assembly of periplasmic cytochrome c and of cytochrome bd in Escherichia coli . FEMS Microbiol Lett 117: 217–223. [DOI] [PubMed] [Google Scholar]
- 48. Krummeck G, Rodel G (1990) Yeast SCO1 protein is required for a post-translational step in the accumulation of mitochondrial cytochrome c oxidase subunits I and II. Curr Genet 18: 13–15. [DOI] [PubMed] [Google Scholar]
- 49. Nittis T, George GN, Winge DR (2001) Yeast Sco1, a protein essential for cytochrome c oxidase function is a CuI-binding protein. J Biol Chem 276: 42520–42526. [DOI] [PubMed] [Google Scholar]
- 50. Swem DL, Swem LR, Setterdahl A, Bauer CE (2005) Involvement of SenC in assembly of cytochrome c oxidase in Rhodobacter capsulatus . J Bacteriol 187: 8081–8087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Hannappel A, Bundschuh FA, Ludwig B (2012) Role of Surf1 in heme recruitment for bacterial COX biogenesis. Biochim Biophys Acta 1817: 928–937. [DOI] [PubMed] [Google Scholar]
- 52. Carr HS, George GN, Winge DR (2002) Yeast Cox11, a protein essential for cytochrome c oxidase assembly, is a CuI-binding protein. J Biol Chem 277: 31237–31242. [DOI] [PubMed] [Google Scholar]
- 53. Hiser L, Di Valentin M, Hamer AG, Hosler JP (2000) Cox11p is required for stable formation of the CuB and magnesium centers of cytochrome c oxidase. J Biol Chem 275: 619–623. [DOI] [PubMed] [Google Scholar]
- 54. Iwata S, Ostermeier C, Ludwig B, Michel H (1995) Structure at 2.8 Å resolution of cytochrome c oxidase from Paracoccus denitrificans . Nature 376: 660–669. [DOI] [PubMed] [Google Scholar]
- 55. Ostermeier C, Harrenga A, Ermler U, Michel H (1997) Structure at 2.7 A resolution of the Paracoccus denitrificans two-subunit cytochrome c oxidase complexed with an antibody FV fragment. Proc Natl Acad Sci U S A 94: 10547–10553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Poole RK, Kumar C, Salmon I, Chance B (1983) The 650 and chromophore in Escherichia coli is an “oxy-” or oxygenated compound, not the oxidized form of cytochrome oxidase d: an hypothesis. Journal of General Microbiology 129: 1335–1344. [DOI] [PubMed] [Google Scholar]
- 57. Georgiou CD, Dueweke TJ, Gennis RB (1988) Regulation of expression of the cytochrome d terminal oxidase in Escherichia coli is transcriptional. J Bacteriol 170: 961–966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Yang K, Borisov VB, Konstantinov AA, Gennis RB (2008) The fully oxidized form of the cytochrome bd quinol oxidase from E. coli does not participate in the catalytic cycle: direct evidence from rapid kinetics studies. FEBS Lett 582: 3705–3709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59. Sone N, Tsukita S, Sakamoto J (1999) Direct correlationship between proton translocation and growth yield: an analysis of the respiratory chain of Bacillus stearothermophilus . J Biosci Bioeng 87: 495–499. [DOI] [PubMed] [Google Scholar]
- 60. Sakamoto J, Matsumoto A, Oobuchi K, Sone N (1996) Cytochrome bd-type quinol oxidase in a mutant of Bacillus stearothermophilus deficient in caa 3-type cytochrome c oxidase. FEMS Microbiol Lett 143: 151–158. [DOI] [PubMed] [Google Scholar]
- 61. Ng TC, Laheri AN, Maier RJ (1995) Cloning, sequencing, and mutagenesis of the cytochrome c 4 gene from Azotobacter vinelandii: characterization of the mutant strain and a proposed new branch in the respiratory chain. Biochim Biophys Acta 1230: 119–129. [DOI] [PubMed] [Google Scholar]
- 62. Gilmour R, Krulwich TA (1997) Construction and characterization of a mutant of alkaliphilic Bacillus firmus OF4 with a disrupted cta operon and purification of a novel cytochrome bd . J Bacteriol 179: 863–870. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63. Chang HY, Ahn Y, Pace LA, Lin MT, Lin YH, et al. (2010) The diheme cytochrome c 4 from Vibrio cholerae is a natural electron donor to the respiratory cbb 3 oxygen reductase. Biochemistry 49: 7494–7503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64. Tseng CP, Albrecht J, Gunsalus RP (1996) Effect of microaerophilic cell growth conditions on expression of the aerobic (cyoABCDE and cydAB) and anaerobic (narGHJI, frdABCD, and dmsABC) respiratory pathway genes in Escherichia coli . J Bacteriol 178: 1094–1098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65. Winstedt L, Yoshida K, Fujita Y, von Wachenfeldt C (1998) Cytochrome bd biosynthesis in Bacillus subtilis: characterization of the cydABCD operon. J Bacteriol 180: 6571–6580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66. Preisig O, Zufferey R, Thony-Meyer L, Appleby CA, Hennecke H (1996) A high-affinity cbb 3-type cytochrome oxidase terminates the symbiosis-specific respiratory chain of Bradyrhizobium japonicum . J Bacteriol 178: 1532–1538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67. Mouncey NJ, Kaplan S (1998) Oxygen regulation of the ccoN gene encoding a component of the cbb 3 oxidase in Rhodobacter sphaeroides 2.4.1T: involvement of the FnrL protein. J Bacteriol 180: 2228–2231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68. Baker SC, Ferguson SJ, Ludwig B, Page MD, Richter OM, et al. (1998) Molecular genetics of the genus Paracoccus: metabolically versatile bacteria with bioenergetic flexibility. Microbiol Mol Biol Rev 62: 1046–1078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69. Pitcher RS, Watmough NJ (2004) The bacterial cytochrome cbb 3 oxidases. Biochim Biophys Acta 1655: 388–399. [DOI] [PubMed] [Google Scholar]
- 70. Kawakami T, Kuroki M, Ishii M, Igarashi Y, Arai H (2010) Differential expression of multiple terminal oxidases for aerobic respiration in Pseudomonas aeruginosa . Environ Microbiol 12: 1399–1412. [DOI] [PubMed] [Google Scholar]
- 71. Arai H (2011) Regulation and Function of Versatile Aerobic and Anaerobic Respiratory Metabolism in Pseudomonas aeruginosa . Front Microbiol 2: 103 10.3389/fmicb.2011.00103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72. Lauraeus M, Haltia T, Saraste M, Wikstrom M (1991) Bacillus subtilis expresses two kinds of haem-A-containing terminal oxidases. Eur J Biochem 197: 699–705. [DOI] [PubMed] [Google Scholar]
- 73. Mather MW, Springer P, Fee JA (1991) Cytochrome oxidase genes from Thermus thermophilus. Nucleotide sequence and analysis of the deduced primary structure of subunit IIc of cytochrome caa 3 . J Biol Chem 266: 5025–5035. [PubMed] [Google Scholar]
- 74. Lobo SA, Almeida CC, Carita JN, Teixeira M, Saraiva LM (2008) The haem-copper oxygen reductase of Desulfovibrio vulgaris contains a dihaem cytochrome c in subunit II. Biochim Biophys Acta 1777: 1528–1534. [DOI] [PubMed] [Google Scholar]
- 75. Lamrabet O, Pieulle L, Aubert C, Mouhamar F, Stocker P, et al. (2011) Oxygen reduction in the strict anaerobe Desulfovibrio vulgaris Hildenborough: characterization of two membrane-bound oxygen reductases. Microbiology 157: 2720–2732. [DOI] [PubMed] [Google Scholar]
- 76. Figurski DH, Helinski DR (1979) Replication of an origin-containing derivative of plasmid RK2 dependent on a plasmid function provided in trans. Proc Natl Acad Sci U S A 76: 1648–1652. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77. Herrero M, de Lorenzo V, Timmis KN (1990) Transposon vectors containing non-antibiotic resistance selection markers for cloning and stable chromosomal insertion of foreign genes in gram-negative bacteria. J Bacteriol 172: 6557–6567. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Oligonucleotides used in this study.
(DOCX)