Abstract
The bacterial second messenger cyclic diguanosine monophosphate (c-di-GMP) regulates cellular motility and the synthesis of organelles and molecules that promote adhesion to a variety of biological and nonbiological surfaces. These properties likely require tight spatial and temporal regulation of c-di-GMP concentration. We have developed genetically encoded fluorescence resonance energy transfer (FRET)–based biosensors to monitor c-di-GMP concentrations within single bacterial cells by microscopy. Fluctuations of c-di-GMP were visualized in diverse Gram-negative bacterial species and observed to be cell cycle dependent. Asymmetrical distribution of c-di-GMP in the progeny correlated with the time of cell division and polarization for Caulobacter crescentus and Pseudomonas aeruginosa. Thus, asymmetrical distribution of c-di-GMP was observed as part of cell division, which may indicate an important regulatory step in extracellular organelle biosynthesis or function.
Cyclic diguanosine monophosphate (c-di-GMP) has been implicated in regulating a variety of bacterial cellular characteristics, including antibiotic resistance, biofilm formation, extracellular carbohydrate and adhesin production, pilus- and flagellum-based motility, and cell cycle progression (1–3). Multiple diguanylate cyclases (DGCs) and phosphodiesterases (PDEs) synthesize and degrade c-di-GMP. Regulation of biological functions is accomplished by c-di-GMP's binding to a diverse array of receptors, including PilZ domain proteins (4, 5), cyclic diguanylate riboswitches (6), and transcription factors (7, 8). The apparent role of c-di-GMP in the cell cycle and the presence of many paralogous DGC, PDE, and PilZ proteins controlling diverse cellular functions indicate that there is likely tight spatial and temporal regulation of c-di-GMP.
To study the spatiotemporal dynamics of c-di-GMP fluctuations in individual living bacterial cells, we engineered a set of genetically encoded fluorescence resonance energy transfer (FRET)–based biosensors by fusing PilZ proteins between cyan and yellow fluorescent proteins (CFP and YFP, respectively) (fig. S1) (9). The FRET response of each biosensor to various c-di-GMP concentrations was characterized in vitro (table S1). Binding of c-di-GMP to the diguanylate receptor domain induced a conformational change (4, 10) that altered the relative orientation of the external fluorescent subunits, which decreased FRET efficiency. Thus, the fluorescence properties of these biosensors (i.e., the FRET/CFP emission ratio) could reflect cellular c-di-GMP levels. The biosensor derived from Salmonella enterica serovar Typhimurium protein YcgR exhibited the largest change in net FRET (nFRET) in vitro (–60.6%), with a binding constant of 195 nM for c-di-GMP and no detectable response to cyclic adenosine 3′,5′-monophosphate, cyclic GMP, or guanosine 5′-triphosphate (figs. S1 to S3).
For live-cell imaging of c-di-GMP signaling events in single bacterial cells, we expressed the YcgR-based c-di-GMP sensor in Caulobacter crescentus and measured FRET efficiencies by ratiometric dual-emission microscopy (Fig. 1, A and B). Caulobacter has asymmetric cell division, resulting in two morphologically distinct progeny: a surface-attached replication-competent stalk cell, and a motile swarmer cell with a polar flagellum. The two daughter cells exhibited different emission ratios, with an average nFRET/CFP ratio of 0.7424 ± 0.073 for swarmer cells and 0.5124 ± 0.054 for stalk cells. Correlating the emission ratio (FRET/CFP) with the binding curve derived from in vitro characterization (fig. S2) yielded a cytosolic c-di-GMP concentration below 100 nM for the flagellated swarmer cell and a concentration above 500 nM for the stalk cell (Fig. 1, A and B, and Fig. 2D). Thus, c-di-GMP is asymmetrically distributed in Caulobacter immediately after cell division, with low apparent levels in the flagellated swarmer cell and levels at least five times as high in the nonmotile stalk cell.
Fig. 1.
The asymmetrical distribution of c-di-GMP is dependent on cell division. (A) (Left) Phase-contrast and (right) dual-emission ratio microscopic images (FRET/CFP) of dividing C. crescentus cells reveal c-di-GMP distribution between stalk and swarmer progeny cells. Pseudocolors represent emission ratios (527/480 nm), reflective of differences in local c-di-GMP concentration, as indicated by the legend at left. Lines label the stalk poles (st), and arrows mark flagellated swarmer poles (sw) of mother and daughter cells, respectively. Caulobacter stalk cells had c-di-GMP levels above 500 nM, whereas levels were below 100 nM in the swarmer cell. (B) (Top) Profile of the ratio of net FRET (FRET – spectral bleed-through – direct YFP excitation) to CFP emission fluorescence and (bottom) emission intensity, along the stalk-swarmer polar cell axis of the dividing Caulobacter cell no. 1 from (A). (C) C-di-GMP polarization in a dividing P. aeruginosa cell. (Top) Dual-emission ratio microscopic images (FRET/CFP). (Bottom) Cells have been stained with Alexa Fluor 594; arrows indicate the position of the polar flagellum. (D) Mean nFRET/CFP ratios for flagellated and nonflagellated Caulobacter and Pseudomonas cells as indicated. Results shown are averages of single-cell measurements from at least three independent fields ± SD,*P < 0.005.
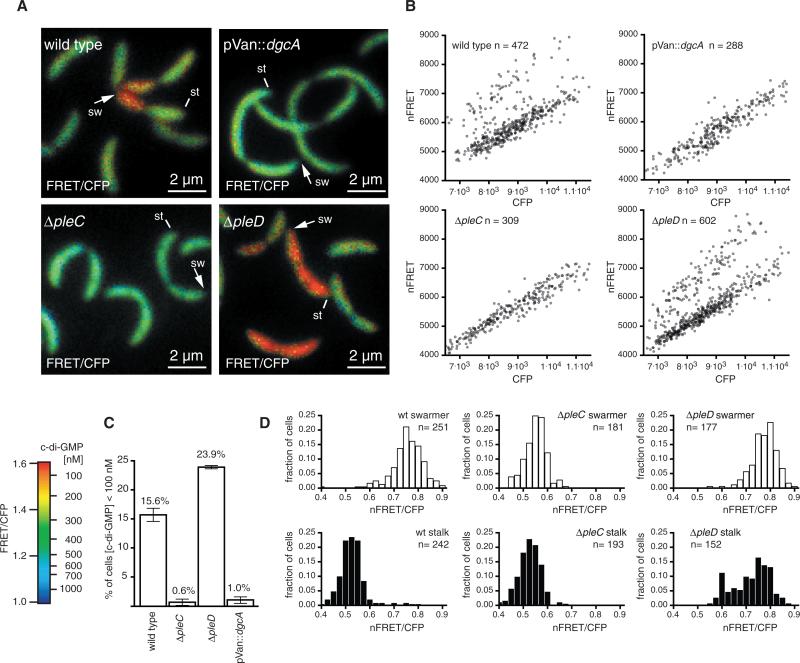
Fig. 2.
C-di-GMP signaling patterns in wild-type and mutant Caulobacter strains. (A) Dual-emission ratio FRET images of Caulobacter cells of a wild-type strain, a strain expressing a constitutively active diguanylate cyclase (pVan::dgcA), or pleC and pleD deletion mutants in logarithmically growing cultures. Pseudocolors represent emission ratios (FRET/CFP, 527/480 nm), which correlate to cellular c-di-GMP levels. Lines indicate stalk poles (st), and arrows indicate swarmer poles (sw). (B) Dot plots illustrating distributions of cellular c-di-GMP levels in logarithmically growing cultures of the indicated Caulobacter strains. Each data pointinthe graphsrepresents an individual bacterial cell for which the mean donor channel (CFP) signal is plotted against the mean net FRET (nFRET) intensity. Cells with relatively low c-di-GMP (<100 nM) are in the upper left half of each plot; cells with elevated c-di-GMP levels (>500 nM) are in the lower right. (C) Histogram showing the percentage of cells of the indicated strains from logarithmic growing cultures with a c-di-GMP concentration below 100 nM. For each condition, the average of three independent measurements ± SD is plotted. (D) Histogram depicting the nFRET/CFP ratio distribution in the swarmer cell and stalk cell subpopulation during the S to G2 transition of cells of the indicated Caulobacter strains.
Like Caulobacter, Pseudomonas produces a single polar flagellum that is partitioned to one daughter at cell division and can be visualized microscopically by labeling living cells with Alexa Fluor 594 conjugated to surface amine–specific succinimidyl ester (Fig. 1C, figs. S5 and S6, and movie S1). Before cell division, the two cell poles harbored similar c-di-GMP levels, probably because of rapid diffusion of the second-messenger molecule. When septum formation resulted in two distinct cells, c-di-GMP rose above 500 nM in the nonflagellated daughter cell and dropped in the flagellated cell below 100 nM (Fig. 1C). This asymmetrical second-messenger distribution was not a stochastic event: In both organisms, c-di-GMP levels were always significantly lower in the flagellated cell than in the nonflagellated one (Fig. 1D).
Bacterial genomes encode multiple DGCs and PDEs in proteins with signal-sensing domains (fig. S4), some of which exhibit distinct sub-cellular localization (11–13). Thus, asymmetrical distribution of c-di-GMP might be caused by spatially restricted expression or activation of individual DGC and PDE enzymes within each daughter cell. We reasoned that overexpression of a DGC would abolish this asymmetrical distribution by increasing c-di-GMP, and we therefore compared the cellular c-di-GMP distribution patterns in various Caulobacter strains. In a strain expressing the constitutively active DGC DgcA (14), c-di-GMP concentrations in the swarmer cells also rose (Fig. 2), which indicated that a localized decrease in DGC activity could cause the drop in c-di-GMP in wild-type swarmer cells. One likely candidate for a DGC that becomes inactivated in the swarmer cell is the diguanylate cyclase PleD. The DGC output activity of PleD is regulated via phosphorylation of its N-terminal receiver domain (11) by the histidine kinase DivJ and the phosphatase PleC, which localize to opposite poles of the predivisional cell (15). Upon cell division, PleC becomes restricted to the swarmer cell, where it dephosphorylates and subsequently inactivates PleD (3). Consistent with this model, a ΔpleC mutant yielded equivalent nFRET/CFP ratios for the swarmer cell (0.5318 ± 0.042) and for the stalked cell (0.5317 ± 0.046) (Fig. 2, A and D), which indicated that pleC is normally required to lower c-di-GMP concentrations in swarmer cells. In contrast, for a ΔpleD mutant, the average nFRET/CFP ratio in the stalk cell increased to 0.7057 ± 0.076, whereas the swarmer cell maintained its normally high ratio of 0.7631 ± 0.051 (Fig. 2, A and D), which resulted in c-di-GMP concentrations below 100 nM in both cells.
Time-lapse experiments using wild-type and mutant Caulobacter strains further defined c-di-GMP dynamic changes during cell division (for parallel measurements in Pseudomonas see figs. S5 and S6 and movie S1). Wild-type Caulobacter cells before cell division had equivalent c-di-GMP levels (Fig. 3A and movie S2), and upon septation, c-di-GMP decreased in the flagellated swarmer cell below 100 nM (Fig. 3, A and B). Note that the swarmer cell–specific drop in c-di-GMP was a transient phenomenon, and the levels of c-di-GMP reverted within less than 20 min to those found in the stalk cell. In contrast, the ΔpleD mutant demonstrated a drop in both cells during the transition from S to G2 phases and extended the period of decreased c-di-GMP levels in the swarmer cell (Fig. 3, A and B, and movie S3). Consistent with PleC function as a negative regulator of PleD activity, cellular c-di-GMP levels remained constitutively high in ΔpleC cells throughout the cell cycle (Fig. 3, A and B, and movie S4). Thus, pleC is required for asymmetrical distribution of c-di-GMP during cell division, most likely by spatially restricting PleD phosphorylation, and its DGC activity, to the stalk cell. Nevertheless, deletion of pleD itself did not result in a complete loss of cellular c-di-GMP, which suggested that additional DGC enzymes contribute to cell cycle–dependent c-di-GMP fluctuations.
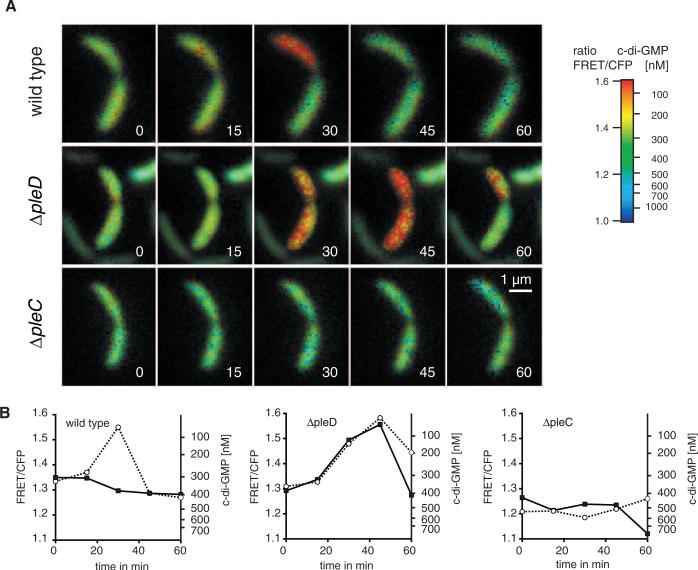
Fig. 3.
Kinetics of fluorescence ratio changes (527/480 nm), reflecting c-di-GMP levels recorded in Caulobacter cells during the S to G2 transition. (A) Time-lapse dual-emission ratiometric FRET microscopy of representative cells of the indicated strains recorded at intervals of 15 min (see also movies S2 to S4). (B) Corresponding plots of the emission ratio changes for the indicated strains over time in the swarmer cell (open circles) and the stalk cell (filled squares).
Impairing the cellular distribution of c-di-GMP has major consequences for Caulobacter polar organelle development and function. Deletion of pleC and overexpression of dgcA both lead to a nonmotile phenotype, most likely as a result of a generalized increase in c-di-GMP and activation of DgrA, which can in turn inhibit motility by binding to flagellar cytoplasmic components (4, 5, 16). Conversely, a pleD deletion mutant is hypermotile and expresses functional flagella at both cell poles (2), which indicates that the observed drop in c-di-GMP at cell division (Fig. 3) promotes rapid motility of flagellated cells immediately after septum formation. In addition to their use in Pseudomonas and Caulobacter, we have successfully used FRET-based c-di-GMP biosensor constructs in peritrichously flagellated Salmonella enterica serovar Typhimurium and nonflagellated Klebsiella pneumoniae. We observed that these organisms also exhibit asymmetric second-messenger distribution upon cell division (fig. S7), which suggests that this phenomenon is not unique to Pseudomonas and Caulobacter, and cell properties other than motility are regulated by asymmetrical second-messenger distribution upon cell division.
Supplementary Material
Acknowledgments
We thank R. Jones from Rutgers University, Piscataway, NJ, for c-di-GMP and T. Ohashi from Duke University, Durham, NC, for the pET15b::mCYPet_12AA_mYPet plasmids. This research was supported by the National Institute of Allergy and Infectious Diseases, NIH (1R21NS067579-01 and U54AI057141 to S.I.M; KO8AI066251 to L.R.H and 1R21NS067579-01 to H.D.K), NSF (Graduate Research Fellowship to B.R.K), Swiss National Foundation (PA00P3-124140 and PBBSA-120489 to M.C. and PA00P3-126243 to B.C), Novartis Foundation to M.C., and Cystic Fibrosis Foundation (HOFFMA04L0 to L.R.H).
Footnotes
Supporting Online Material
References and Notes
- 1.Hoffman LR, et al. Nature. 2005;436:1171. doi: 10.1038/nature03912. [DOI] [PubMed] [Google Scholar]
- 2.Hecht GB, Newton A. J. Bacteriol. 1995;177:6223. doi: 10.1128/jb.177.21.6223-6229.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Aldridge P, Paul R, Goymer P, Rainey PB, Jenal U. Mol. Microbiol. 2003;47:1695. doi: 10.1046/j.1365-2958.2003.03401.x. [DOI] [PubMed] [Google Scholar]
- 4.Christen M, et al. Proc. Natl. Acad. Sci. U.S.A. 2007;104:4112. [Google Scholar]
- 5.Paul K, Nieto V, Carlquist WC, Blair DF, Harshey RM. Mol. Cell. 2010;38:128. doi: 10.1016/j.molcel.2010.03.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Sudarsan N, et al. Science. 2008;321:411. doi: 10.1126/science.1159519. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Leduc JL, Roberts GP. J. Bacteriol. 2009;191:7121. doi: 10.1128/JB.00845-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Krasteva PV, et al. Science. 2010;327:866. doi: 10.1126/science.1181185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Ohashi T, Galiacy SD, Briscoe G, Erickson HP. Protein Sci. 2007;16:1429. doi: 10.1110/ps.072845607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Benach J, et al. EMBO J. 2007;26:5153. doi: 10.1038/sj.emboj.7601918. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Chan C, et al. Proc. Natl. Acad. Sci. U.S.A. 2004;101:17084. doi: 10.1073/pnas.0406134101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Güvener ZT, Harwood CS. Mol. Microbiol. 2007;66:1459. doi: 10.1111/j.1365-2958.2007.06008.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Huitema E, Pritchard S, Matteson D, Radhakrishnan SK, Viollier PH. Cell. 2006;124:1025. doi: 10.1016/j.cell.2006.01.019. [DOI] [PubMed] [Google Scholar]
- 14.Christen B, et al. J. Biol. Chem. 2006;281:32015. doi: 10.1074/jbc.M603589200. [DOI] [PubMed] [Google Scholar]
- 15.Wheeler RT, Shapiro L. Mol. Cell. 1999;4:683. doi: 10.1016/s1097-2765(00)80379-2. [DOI] [PubMed] [Google Scholar]
- 16.Boehm A, et al. Cell. 2010;141:107. doi: 10.1016/j.cell.2010.01.018. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.