SUMMARY
Many organisms, including plants, use the circadian clock to measure the duration of day and night. Daily rhythms in the plant circadian system are generated by multiple interlocked transcriptional/translational loops and also by spatial regulations such as nuclear translocation. GIGANTEA (GI), one of the key clock components in Arabidopsis, makes distinctive nuclear bodies like other nuclear-localized circadian regulators. However, little is known about the dynamics or roles of GI subnuclear localization. Here, we characterize GI subnuclear compartmentalization and identify unexpected dynamic changes under diurnal conditions. We further identify EARLY FLOWERING 4 (ELF4) as a regulator of GI nuclear distribution through a physical interaction. ELF4 sequesters GI from the nucleoplasm, where GI binds the promoter of CONSTANS (CO), to discrete nuclear bodies. We suggest that the subnuclear compartmentalization of GI by ELF4 contributes to the regulation of photoperiodic flowering.
RESULTS AND DISCUSSION
GI Makes Dynamic Subnuclear Organelles
Many of the Arabidopsis circadian clock components are localized both in the nucleus and the cytosol. As one of the important regulators in the Arabidopsis circadian system, GIGANTEA (GI) is present in both compartments and associates with a range of different partners (Huq et al., 2000; Kim et al., 2007). GI protein interacts with the cytosolic F box protein ZEITLUPE (ZTL), which acts to regulate endogenous rhythms through phase-specific proteolysis (Kim et al., 2007). FKF1, CDF1, and GI form a nuclear protein complex at the promoter of CONSTANS (CO), a flowering time integrator, to regulate CO expression (Sawa et al., 2007). Additionally, TEM1/TEM2 and SVP associate in the nucleus with GI to modulate FT expression (Sawa and Kay, 2011). GI also interacts with a nuclear ELF3/COP1 complex that destabilizes GI protein (Yu et al., 2008).
Interestingly, the GI/ELF3/COP1 complex exhibits a specific subnuclear localization in onion epidermal cells (Yu et al., 2008). To test whether GI localizes to nuclear bodies in Arabidopsis cells, we constitutively expressed GI-GFP transiently in Arabidopsis protoplasts and in transgenic plants. In both systems, GI distribution in the nucleus showed two typical patterns: a dispersed type, and a punctate type (Figure 1A). The characteristics of nuclear bodies can also be defined by their numbers, size, and dynamic movements (Lamond and Sleeman, 2003). In our experiments, the number of GI nuclear bodies ranged widely, from a few to tens of bodies both in transient expression and transgenic plants. Additionally, the size of GI nuclear bodies varied from 0.5 ± 0.025 μm in transient assays to 0.29 ± 0.0016 μm in transgenic plants (Table S1).
Figure 1. GI Forms Dynamic Subnuclear Structures.
(A) Nuclear distributions of GI in a transient expression system using Arabidopsis mesophyll cell protoplastsand in transgenic plants. Two prominent distribution types (dispersed and punctate) of GI-GFP were imaged. GFP was used as a control vector in the transient expression system, and DAPI was used as a nuclear marker.
(B–E) Subnuclear colocalization of GI with other subnuclear marker proteins: DAPI for chromatin (B), U2B for Cajal bodies (C), H2B for chromatin and nucleoli (D), and SCL28 for spliceosomes (E). Arabidopsis protoplasts were transfected with GI-GFP and stained with DAPI for 1 min before imaging. Arabidopsis protoplasts were transfected with a GI-GFP and H2B-RFP, U2B-YFP, or SCL28-RFP and kept in dim white light for 8 hr before examination. Intensity plots obtained across the nucleus (white arrows) are shown in each image set.
(F) Association and dissociation of GI nuclear bodies. Fluorescence intensities from GI-GFP in the nucleus were tracked over 2 hr on a microscope stage. Yellow arrowheads in the dispersed type of GI localization indicate newly emerged nuclear bodies.
(G) Variation in the proportion of dispersed and punctate types of GI subnuclear localization under LDs (16L/8D). Protoplasts isolated from plants grown under LDs at ZT 0 were transfected with GI-GFP and kept under LDs. White and black boxes represent day and night, respectively. The proportions of cells that express GI-GFP with each of the two nuclear distribution patterns were counted every 3 hr. Data were normalized to total cells counted. Open circle and black inverted triangle represent proportions of the dispersed and the punctate type, respectively. Error bars represent the SEM from four independent replicates.
Scale bars represent 5 μm. See also Figure S1.
Compartmentalization of proteins in the nucleus is a well-known means to regulate processes such as transcription, DNA replication, splicing, and degradation (Lamond and Sleeman, 2003; Shaw and Brown, 2004). There are well-established markers for subnuclear organelles in Arabidopsis, including DAPI for chromatin, H2B for chromatin and nucleoli, SCL28 for spliceosomes, and U2B for Cajal bodies (Lorković et al., 2004). To test whether GI nuclear bodies colocalize with any of these markers, we transiently expressed GI-GFP in the presence of DAPI and also coexpressed GI-GFP with H2B-RFP, U2B-YFP, or SCL28-RFP. DAPI, H2B, and SCL28 display distinct and several to tens of nuclear bodies, whereas U2B is present as a single nuclear body (Figures 1B–1E). When we quantified the fluorescence intensities from GI-GFP and each marker, none of these markers colocalized with GI-GFP (right plots in Figures 1B–1E). These results suggest that GI nuclear body localization is distinct from these subnuclear compartments, and they may have little functional relationship with the processes indicated by these markers.
Nuclear bodies are also known to change dynamically (Lamond and Sleeman, 2003; Misteli, 2001). Thus, we tested the nuclear dynamics of GI over a diurnal time course by transiently expressing GI and following the changing distribution of the dispersed and punctate bodies in single-living cells. After transfection, we observed dispersed GI-GFP in the nucleus for 1 hr, with a few small nuclear bodies starting to form after 2 hr of observation (Figure 1F, upper panel). For punctate-type bodies, numerous GI nuclear bodies were present at the outset and then started to dissociate after 45 min of observation, with most of them absent after 2 hr (Figure 1F, lower panel). Thus, the dispersed and punctate types of GI nuclear distribution appeared to be interchangeable, indicating the dynamic nature of their assembly and disassembly.
We next measured the dynamics of the changing proportion of the dispersed and punctate types of GI nuclear expression under diurnal conditions to test the relationship of GI subnuclear distribution to its function. We counted the number of cells that expressed the two types every 3 hr under long days (LDs) and normalized these with total number of cells expressing GI-GFP at each observation point (Figure 1G). The relative proportion of the dispersed and punctate types of GI-GFP varied throughout the day. The dispersed type was high during the photoperiod and comprised the lowest portion at the end of night. In contrast, punctate GI was low during the day and was at the highest level by the end of night. We further measured the change of the nuclear distribution of GI under short days (Figure S1). The proportions of the dispersed and punctate types were also anticorrelated, but the peaks and troughs were not clear as seen in LDs. Interestingly, the point at which each nuclear distribution type changed relative to the other was phase advanced in short days compared to LDs and correlated well in both conditions with same number of hours after the light-to-dark transition (Figure S1). These data imply that the subnuclear distribution of GI might be coupled to the light-dark cycles and may provide a further regulatory mechanism to understand the photoperiodism in Arabidopsis.
GI Directly Interacts with ELF4 at the Nuclear Bodies
Nuclear bodies are large protein complexes that contain proteins with similar functions (Lamond and Sleeman, 2003). We next asked which additional proteins that have functions related to GI might complex with it. Among known circadian clock components, we screened for potential GI interaction proteins using yeast two-hybrid assays and isolated EARLY FLOWERING 4 (ELF4) as a GI interaction partner (Figure S2A). Both GI and ELF4 have a similar circadian phase (David et al., 2006; Fowler et al., 1999; Khanna et al., 2003; Kikis et al., 2005; Kim et al., 2007; Kolmos et al., 2009; McWatters et al., 2007; Mizoguchi et al., 2005; Nusinow et al., 2011; Park et al., 1999; Sawa et al., 2007) and genetically interact to affect similar circadian outputs such as seedling growth, flowering time, and expression of clock genes in a phase-specific manner over a diurnal cycle (Kim et al., 2012). GI localizes both in the nucleus and cytoplasm (Kim et al., 2007), and we performed immunoblot analyses on cell fractions using ELF4p∷ELF4-HA seedlings to establish its presence in both the nucleus and cytosol (Figure S2B). We next tested for potential interaction by transiently expressing GI and ELF4 in tobacco. Both proteins localized in the nucleus and the cytosol, which is consistent with previous reports (Kim et al., 2007) (Figure S2C). Additionally, in vivo coimmunoprecipitation (coIP) assays demonstrated both nuclear and cytosolic interactions (Figures 2A and S2C). We next used bifluorescence complementation (BiFC) assays to determine where subnuclear interactions occur and observed fluorescence at distinct nuclear bodies (Figure 2B). Additionally, GI-GFP and ELF4-RFP are able to form nuclear bodies independently and localize at the same bodies in which their interaction was observed (Figure 2C). Taken together, these data indicate that GI and ELF4 interact both in the nucleus and cytosol and within the same nuclear bodies.
Figure 2. ELF4 Interacts with GI within Nuclear Bodies.

(A) CoIP of GI and ELF4 in the nucleus. CsV∷GI-GFP and CsV∷ELF4-HA were transiently expressed in N. benthamiana, and GI-GFP and ELF4-HA were localized to both the cytosol and the nucleus. CoIP was performed with GFP antibody using the nuclear fraction. HSP90 and H3 were used as a marker for the cytosol and nucleus, respectively.
(B) BiFC analysis of the GI/ELF4 interaction. Green represents YFP from the GI/ELF4 interaction. Chloroplast autofluorescence is red.
(C) Subnuclear colocalization of GI and ELF4. GI-GFP and ELF4-RFP were transiently expressed in Arabidopsis protoplasts. Green indicates GI-GFP, red indicates ELF4-RFP, and blue indicates chloroplast autofluorescence.
Scale bars represent 10 μm. See also Figure S2.
ELF4 Regulates GI Nuclear Compartmentalization
Because GI and ELF4 interact within nuclear bodies, we tested whether they can regulate subnuclear localization of each other. GI-GFP and ELF4-RFP were transiently expressed in elf4 and gi-2 protoplasts, respectively, and their nuclear distribution patterns under LDs were observed. When we expressed GI-GFP in WT and elf4 protoplasts, the proportion of punctate bodies was lower in the elf4 background than in WT protoplasts, whereas the portion of the dispersed type was higher than in WT protoplasts at all time points tested (Figure 3A). This strongly increased proportion of the dispersed type and decreased proportion of the punctate type of GI nuclear localization in elf4, especially at night, led to a dampening in the cyclic changes in the proportion of GI subnuclear distribution type (Figure 3A). We also measured the nuclear distribution of ELF4-RFP in WT and gi-2 protoplasts. ELF4-RFP also displayed two prominent nuclear distribution types: dispersed, and punctate (Figure 3B). Approximately 80% of cells expressing ELF4-RFP formed nuclear bodies, and around 20% of these cells showed an even distribution of ELF4-RFP in WT nuclei with little change over a LD cycle. When we expressed ELF4-RFP in gi-2 protoplasts, there was no significant change in the nuclear distribution of ELF4-RFP compared to those in WT protoplasts (Figure 3B).
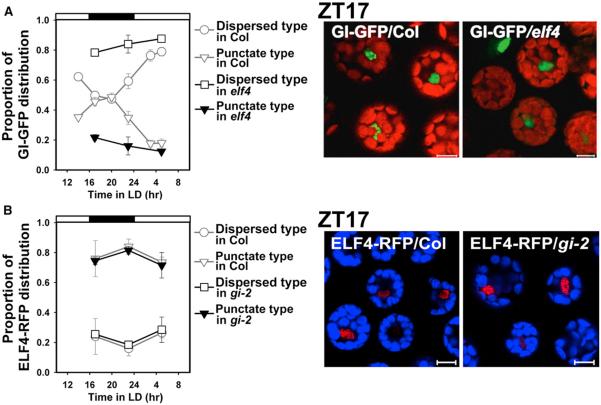
Figure 3. ELF4 Recruits GI to Nuclear Bodies.
(A) Altered distribution of GI-GFP in elf4 protoplasts. GI-GFP was expressed transiently in WT (Col) and elf4 protoplasts, and the proportion of each distribution type of GI was observed (left). Typical image of subnuclear distributions of GI-GFP in WT and elf4 protoplasts at ZT17 (right). Green indicates GI-GFP; red shows autofluorescence from chloroplasts.
(B) Subnuclear distribution of ELF4-RFP in WT and gi-2 protoplasts. ELF4-RFP was transiently expressed in WT and gi-2 protoplasts, and the proportion of each distribution type of ELF4 was observed (left). Typical image of subnuclear distributions of ELF4-RFP in WT and gi-2 protoplasts at ZT17 (right). Red indicates ELF4-RFP; blue shows chloroplast autofluorescence. Data are mean ± SE from four biological replicates. Each dispersed or punctate form of GI-GFP and ELF4-RFP was counted and the relative proportion of each reported.
Scale bars represent 10 μm. See also Figure S3.
We next expressed GI-GFP with RFP or ELF4-RFP in elf4 protoplasts to test whether ELF4 could induce the localization of GI to nuclear bodies. The proportion of the dispersed type of GI-GFP when coexpressed with RFP was similar to the distribution pattern seen when GI-GFP was expressed alone in the elf4 mutant (Figure S3). In contrast, GI-GFP coexpression with ELF4-RFP decreased the proportion of the dispersed type by about 20%, whereas the proportion of the GI-GFP punctate type increased (Figure S3). From these results, we conclude that the subnuclear localization of GI is strongly controlled by ELF4. These results suggest that ELF4 can modulate GI function by regulating the subnuclear localization of GI.
ELF4 Modulates the CO Promoter Binding Affinity of GI
We then asked what the functional significance of ELF4 control of GI localization might be. ELF4 is known to act upstream of GI in photoperiodic flowering time regulation (Kim et al., 2012), and transient translocalization of GI to the nucleus induces flowering (Gunl et al., 2009). GI protein is destabilized by a complex of ELF3 and COP1 (Yu et al., 2008), and nuclear GI can bind the promoter of CO and FT directly and regulate their expression (Sawa and Kay, 2011; Sawa et al., 2007). These previous reports led us to consider two possibilities: (1) ELF4 might regulate GI protein stability at the nuclear bodies within an ELF3/COP1 complex, and (2) ELF4 might regulate GI action as a transcriptional modulator of photoperiodic flowering integrators. To test these possibilities, we generated transgenic plants expressing GI-GFP under a constitutive promoter (CsV∷GI-GFP) in an elf4-209 background to avoid possible transcriptional regulation of GI by ELF4 because ELF4 is known to suppress GI transcription (Kolmos et al., 2009). We first assayed the effects of the elf4 mutation on GI protein stability in the nucleus to avoid possible complication of a cytosolic effect of ELF4 on overall GI abundance. Nuclear GI showed weak cycling under light-dark cycles, and GI nuclear protein levels were not significantly affected by the elf4 mutation (Figures 4A and 4B). This result indicates that ELF4 has little effect on GI protein stability in the nucleus. Thus, the recruitment of GI to ELF4 nuclear bodies is unlikely to regulate GI protein abundance, and this complex seems to function differently from the GI/ELF3/COP1 complex, which affects GI protein levels (Yu et al., 2008). Cytosolic GI also showed very weak cycling under constitutive expression (Figure S4A), and cytosolic GI was decreased by the elf4 mutation about 30% throughout the day, indicating that ELF4 stabilizes GI in the cytosol through an unknown mechanism (Figure S4A). These results additionally imply that ELF4 acts on GI differently in the nucleus and the cytoplasm.
Figure 4. ELF4 Regulates Chromatin Accessibility of GI.
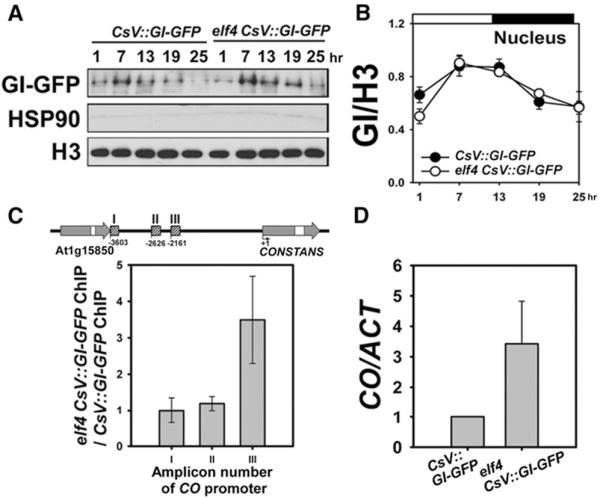
(A and B) Nuclear GI-GFPs in CsV∷GI-GFP and elf4CsV∷GI-GFP under 12/12 hr light-dark cycles. GI proteins were detected by anti-GFP. HSP90 and H3 were used as cytosolic and nuclear markers, respectively. GI protein abundance was normalized to H3. Data are mean ± SE from four biological replicates.
(C) CO promoter binding affinity of GI in the elf4 mutant relative to WT. Diagram for the CO amplicon in ChIP assay is indicated. Amplicons I, II, and III marked with hatched boxes correspond to amplicons 1, 3, and 4, respectively, as previously reported by Sawa et al. (2007). Ten-day-old seedlings were harvested 1 hr after lights on. The fold enrichment is calculated relative to ACT2.
(D) CO mRNA expression in CsV∷GI-GFP and elf4CsV∷GI-GFP. CO mRNA expression determined by quantitative RT-PCR with the same tissues used for ChIP. CO mRNA normalized relative to ACT2. Data are mean ± SE from six biological replicates.
See also Figure S4.
We next tested whether ELF4 might affect the activity of GI as a transcriptional modulator of CO. We performed chromatin immunoprecipitation (ChIP) assays using CsV∷GI-GFP and elf4 CsV∷GI-GFP seedlings. Tissues were sampled at ZT1 (ZT: hours after lights on), corresponding to the time of the highest proportion of dispersed nuclear GI, both in WT and the elf4 mutant (Figures 1G and 3A), and binding to amplicon III at the CO promoter was assayed (Figure 4C, diagram) (Sawa et al., 2007). When we compared CO promoter binding affinity of GI in these two backgrounds, GI-GFP was enriched 3-fold in the elf4 mutant, relative to WT at amplicon III, whereas there were no significant differences at the control amplicons (Figure 4C). Consistent with this finding, CO expression level in elf4 was significantly higher than WT (Figure 4D), and flowering time of elf4 CsV∷GI-GFP plants was earlier than that of CsV∷GI-GFP both in LDs and short days (Figure S4B). These results support the notion that ELF4 negatively regulates CO expression by sequestering GI from the CO promoter to specific ELF4-GI nuclear bodies.
Conclusions
Spatial regulation in cellular signaling is an important factor in understanding complex regulatory mechanisms (Herrero and Davis, 2012; Meier and Somers, 2011). Many circadian clock regulators can be localized at nuclear foci such as PRR5/TOC1 (Wang et al., 2010), ELF3/ELF4 (Herrero et al., 2012), ELF3/COP1/GI (Yu et al., 2008), and CKB4 (Portolés and Más, 2007). Diverse spatial regulations could, in principle, allow a single protein species to contribute to many signaling pathways. Specifically, subnuclear compartmentalization is characterized by a distinct set of resident proteins with dynamic changes in their size, shape, and number without the separation afforded by membranes (Lamond and Sleeman, 2003; Shaw and Brown, 2004). GI has been suggested to have differential roles in the nucleus and the cytosol (Gunl et al., 2009; Kim et al., 2007; Sawa and Kay, 2011; Sawa et al., 2007), and our results contribute to this notion by showing that GI forms a specific type of nuclear body into which it is sequestered by physical interaction with ELF4 from the CO promoter. GI nuclear body formation is induced by ELF4 under LDs (Figure 3), and this implies that the absolute levels of ELF4 protein can affect the proportion of GI nuclear body formation. Oscillation of ELF4 protein under diurnal conditions shows a broad peak at night that corresponds closely to that of GI (Figure S2B). This high expression of ELF4 at night may recruit GI to the nuclear bodies from the CO promoter, which is consistent with the cycling of GI subnuclear distributions (Figure 1F). However, CO transcript has a second peak at night (Suárez-López et al., 2001). This second CO peak does not correlate with GI/ELF4 nuclear body formation, and this discrepancy suggests that there are additional pathways to generate the CO waveform at night. Interestingly, ELF3/COP1/GI nuclear bodies have been suggested to form during darkness (Yu et al., 2008). Hence, there might be a competition between ELF4 and ELF3/COP1 to form a complex with GI. The ELF3/COP1 complex destabilizes GI proteins, which might act to shape the decreasing slope of the GI protein level during the night. In contrast, the GI/ELF4 nuclear body sequesters GI from the CO promoter, retaining GI within the nucleus for possible release in the morning, eliminating the need for de novo synthesis of GI. Taken together, the spatial regulation of GI with ELF4 in the nucleus can provide an additional mechanism that contributes to the currently known genetic and molecular interactions that control photoperiodic flowering.
EXPERIMENTAL PROCEDURES
Plant Materials and Transgenic Plants
The gi-2 was reported previously (Park et al., 1999), and elf4-209 (Kolmos et al., 2009) was backcrossed three times. CsV∷GI-GFP was constructed using LR recombination (GATEWAY; Invitrogen, Carlsbad, CA, USA).
Preparation of Protoplasts
For transient expression in Arabidopsis, protoplasts were prepared from fourth, fifth, and sixth leaves of 1-month-grown Col-0 under LDs (16L/8D) or short days (8L/16D) at 22°C, as described previously (Kim et al., 2008). Transfected protoplasts were maintained and observed under the stated conditions of each experiment.
Vector Construction
The CsV∷GI-GFP, GI cDNA clone was generated by PCR using the primers 5′-GGA TCC GAT GGC TAG TTC ATC-3′ and 5′-GAT GGA TCC TTG GGA CAA GGA TAT AGT ACA GCC GAG-3′, which resulted in removal of the termination codon. GI cDNA digested with EcoRI and BamHI was fused with pCsV-eGFP-N-999 (Kim et al., 2008). For transient expression of CsV∷GI-GFP, the gene cassette was moved to the pBlueSKII+ vector using NotI.
For SCL28∷mRFP, SCL28 cDNA was obtained by PCR using primers 5′-AGA GAA TTC ATG AGG GGA AGG AGC TAC-3′ and 5′-GAT GGA TCC TGC TTC TTC TAG GGC TGG-3′. pCsV-mRFP-N-999 was generated by substitution of eGFP from pCsV-eGFP-N-999 to monomeric RFP from pDsRed-Monomer-N1 (Clonetech Laboratories, Mountain View, CA, USA).
U2B∷YFP was kindly donated by Drs. Yuda Fang and David L. Spector from Cold Spring Harbor Laboratory (Cold Spring Harbor, NY, USA). The CsVMV∷ELF4-HA vector was prepared by recombination of an entry clone that contained the ELF4 cDNA and pCsVMV-HA-1300. The ELF4-RFP vector was also generated by recombination with pCsVMV-HcRed1-999.
To confirm the cellular localization of ELF4 in planta, we generated ELF4p∷ELF4-HA plants. ELF4p∷ELF4 was amplified with primers 5′-CTT CTG CAG CTC ATG ATT TCC TGC GGTAAT-3′ and 5′-CTT AGG CCT AGC TCT AGT TCC GGC AGC A-3′ from Arabidopsis genomic DNA. This genomic gene cassette was moved to an entry vector and then transferred to an HA-tagged binary vector that has no promoter. The ELF4p∷ELF4-HA clone was transformed into the elf4-209 mutant.
Microscopic Analysis
Fluorescence images were obtained using an inverted confocal microscope (LSM5 LIVE; Carl Zeiss, Germany). Subcellular and subnuclear localizations of fluorescence fusion protein were observed with × 100 and × 63 oil-immersion apochromat objectives, respectively (Carl Zeiss). Images from each experiment were pseudocolored and plotted with LSM5 LIVE software.
Immunoblot Analysis and CoIP
Immunoblot analyses for GI, HSP90, and H3 protein detection were done as reported (Kim et al., 2007). Polyclonal HSP90 antibodies (rabbit) were made by W.Y.K. and were diluted 1:5,000 for detection. For cytosolic interaction between GI and ELF4, CsV∷GI-GFP and CsV∷ELF4-HA were transiently expressed in N. benthamiana, and immunoprecipitations were performed as reported (Wang et al., 2010).
Flowering Time Measurement/mRNA Expression
These experiments were performed as described (Kim et al., 2008).
ChIP
Chromatin binding affinity of GI on CO promoter was performed as reported by Sawa et al. (2007).
Supplementary Material
ACKNOWLEDGMENTS
We appreciate Drs. Yuda Fang and David L. Spector for the kind donation of the U2B-YFP clone. We also thank K.H. Suh, Y.S. Park, and B.H. Kim for technical assistance and E.H. Cha for supporting the settlement in DGIST. This work was supported by grants from the National Research Foundation of Korea funded by the Korean government (MEST), the Research Center Program of IBS (CA1208), the National Researcher Support Program (no. 20100020417), the World Class University Program (no. R31-2008-000-10105-0 to D.E.S. and no. R32-10148), and the National Institutes of Health (R01GM093285 to D.E.S.).
Footnotes
SUPPLEMENTAL INFORMATION
Supplemental Information includes four figures and one table and can be found with this article online at http://dx.doi.org/10.1016/j.celrep.2013.02.021.
REFERENCES
- David KM, Armbruster U, Tama N, Putterill J. Arabidopsis GIGANTEA protein is post-transcriptionally regulated by light and dark. FEBS Lett. 2006;580:1193–1197. doi: 10.1016/j.febslet.2006.01.016. [DOI] [PubMed] [Google Scholar]
- Fowler S, Lee K, Onouchi H, Samach A, Richardson K, Morris B, Coup-land G, Putterill J. GIGANTEA: a circadian clock-controlled gene that regulates photoperiodic flowering in Arabidopsis and encodes a protein with several possible membrane-spanning domains. EMBO J. 1999;18:4679–4688. doi: 10.1093/emboj/18.17.4679. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gunl M, Liew EF, David K, Putterill J. Analysis of a post-translational steroid induction system for GIGANTEA in Arabidopsis. BMC Plant Biol. 2009;9:141. doi: 10.1186/1471-2229-9-141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Herrero E, Davis SJ. Time for a nuclear meeting: protein trafficking and chromatin dynamics intersect in the plant circadian system. Mol. Plant. 2012;5:554–565. doi: 10.1093/mp/sss010. [DOI] [PubMed] [Google Scholar]
- Herrero E, Kolmos E, Bujdoso N, Yuan Y, Wang M, Berns MC, Uhlworm H, Coupland G, Saini R, Jaskolski M, et al. EARLY FLOWERING4 recruitment of EARLY FLOWERING3 in the nucleus sustains the Arabidopsis circadian clock. Plant Cell. 2012;24:428–443. doi: 10.1105/tpc.111.093807. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huq E, Tepperman JM, Quail PH. GIGANTEA is a nuclear protein involved in phytochrome signaling in Arabidopsis. Proc. Natl. Acad. Sci. USA. 2000;97:9789–9794. doi: 10.1073/pnas.170283997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khanna R, Kikis EA, Quail PH. EARLY FLOWERING 4 functions in phytochrome B-regulated seedling de-etiolation. Plant Physiol. 2003;133:1530–1538. doi: 10.1104/pp.103.030007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kikis EA, Khanna R, Quail PH. ELF4 is a phytochrome-regulated component of a negative-feedback loop involving the central oscillator components CCA1 and LHY. Plant J. 2005;44:300–313. doi: 10.1111/j.1365-313X.2005.02531.x. [DOI] [PubMed] [Google Scholar]
- Kim J, Kim Y, Yeom M, Kim JH, Nam HG. FIONA1 is essential for regulating period length in the Arabidopsis circadian clock. Plant Cell. 2008;20:307–319. doi: 10.1105/tpc.107.055715. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim WY, Fujiwara S, Suh SS, Kim J, Kim Y, Han L, David K, Putterill J, Nam HG, Somers DE. ZEITLUPE is a circadian photoreceptor stabilized by GIGANTEA in blue light. Nature. 2007;449:356–360. doi: 10.1038/nature06132. [DOI] [PubMed] [Google Scholar]
- Kim Y, Yeom M, Kim H, Lim J, Koo HJ, Hwang D, Somers D, Nam HG. GIGANTEA and EARLY FLOWERING 4 in Arabidopsis exhibit differential phase-specific genetic influences over a diurnal cycle. Mol. Plant. 2012;5:678–687. doi: 10.1093/mp/sss005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kolmos E, Nowak M, Werner M, Fischer K, Schwarz G, Mathews S, Schoof H, Nagy F, Bujnicki JM, Davis SJ. Integrating ELF4 into the circadian system through combined structural and functional studies. HFSP J. 2009;3:350–366. doi: 10.2976/1.3218766. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lamond AI, Sleeman JE. Nuclear substructure and dynamics. Curr. Biol. 2003;13:R825–R828. doi: 10.1016/j.cub.2003.10.012. [DOI] [PubMed] [Google Scholar]
- Lorković ZJ, Hilscher J, Barta A. Use of fluorescent protein tags to study nuclear organization of the spliceosomal machinery in transiently transformed living plant cells. Mol. Biol. Cell. 2004;15:3233–3243. doi: 10.1091/mbc.E04-01-0055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McWatters HG, Kolmos E, Hall A, Doyle MR, Amasino RM, Gyula P, Nagy F, Millar AJ, Davis SJ. ELF4 is required for oscillatory properties of the circadian clock. Plant Physiol. 2007;144:391–401. doi: 10.1104/pp.107.096206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meier I, Somers DE. Regulation of nucleocytoplasmic trafficking in plants. Curr. Opin. Plant Biol. 2011;14:538–546. doi: 10.1016/j.pbi.2011.06.005. [DOI] [PubMed] [Google Scholar]
- Misteli T. Protein dynamics: implications for nuclear architecture and gene expression. Science. 2001;291:843–847. doi: 10.1126/science.291.5505.843. [DOI] [PubMed] [Google Scholar]
- Mizoguchi T, Wright L, Fujiwara S, Cremer F, Lee K, Onouchi H, Mouradov A, Fowler S, Kamada H, Putterill J, Coupland G. Distinct roles of GIGANTEA in promoting flowering and regulating circadian rhythms in Arabidopsis. Plant Cell. 2005;17:2255–2270. doi: 10.1105/tpc.105.033464. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nusinow DA, Helfer A, Hamilton EE, King JJ, Imaizumi T, Schultz TF, Farré EM, Kay SA. The ELF4-ELF3-LUX complex links the circadian clock to diurnal control of hypocotyl growth. Nature. 2011;475:398–402. doi: 10.1038/nature10182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Park DH, Somers DE, Kim YS, Choy YH, Lim HK, Soh MS, Kim HJ, Kay SA, Nam HG. Control of circadian rhythms and photoperiodic flowering by the Arabidopsis GIGANTEA gene. Science. 1999;285:1579–1582. doi: 10.1126/science.285.5433.1579. [DOI] [PubMed] [Google Scholar]
- Portolés S, Más P. Altered oscillator function affects clock resonance and is responsible for the reduced day-length sensitivity of CKB4 over-expressing plants. Plant J. 2007;51:966–977. doi: 10.1111/j.1365-313X.2007.03186.x. [DOI] [PubMed] [Google Scholar]
- Sawa M, Kay SA. GIGANTEA directly activates Flowering Locus T in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA. 2011;108:11698–11703. doi: 10.1073/pnas.1106771108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sawa M, Nusinow DA, Kay SA, Imaizumi T. FKF1 and GIGANTEA complex formation is required for day-length measurement in Arabidopsis. Science. 2007;318:261–265. doi: 10.1126/science.1146994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shaw PJ, Brown JW. Plant nuclear bodies. Curr. Opin. Plant Biol. 2004;7:614–620. doi: 10.1016/j.pbi.2004.09.011. [DOI] [PubMed] [Google Scholar]
- Suárez-López P, Wheatley K, Robson F, Onouchi H, Valverde F, Coupland G. CONSTANS mediates between the circadian clock and the control of flowering in Arabidopsis. Nature. 2001;410:1116–1120. doi: 10.1038/35074138. [DOI] [PubMed] [Google Scholar]
- Wang L, Fujiwara S, Somers DE. PRR5 regulates phosphorylation, nuclear import and subnuclear localization of TOC1 in the Arabidopsis circadian clock. EMBO J. 2010;29:1903–1915. doi: 10.1038/emboj.2010.76. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu JW, Rubio V, Lee NY, Bai S, Lee SY, Kim SS, Liu L, Zhang Y, Irigoyen ML, Sullivan JA, et al. COP1 and ELF3 control circadian function and photoperiodic flowering by regulating GI stability. Mol. Cell. 2008;32:617–630. doi: 10.1016/j.molcel.2008.09.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.