Abstract
Purpose
Molecular risk stratification of acute myeloid leukemia (AML) is largely based on genetic markers. However, epigenetic changes, including DNA methylation, deregulate gene expression and may also have prognostic impact. We evaluated the clinical relevance of integrating DNA methylation and genetic information in AML.
Methods
Next-generation sequencing analysis of methylated DNA identified differentially methylated regions (DMRs) associated with prognostic mutations in older (≥ 60 years) cytogenetically normal (CN) patients with AML (n = 134). Genes with promoter DMRs and expression levels significantly associated with outcome were used to compute a prognostic gene expression weighted summary score that was tested and validated in four independent patient sets (n = 355).
Results
In the training set, we identified seven genes (CD34, RHOC, SCRN1, F2RL1, FAM92A1, MIR155HG, and VWA8) with promoter DMRs and expression associated with overall survival (OS; P ≤ .001). Each gene had high DMR methylation and lower expression, which were associated with better outcome. A weighted summary expression score of the seven gene expression levels was computed. A low score was associated with a higher complete remission (CR) rate and longer disease-free survival and OS (P < .001 for all end points). This was validated in multivariable models and in two younger (< 60 years) and two older independent sets of patients with CN-AML. Considering the seven genes individually, the fewer the genes with high expression, the better the outcome. Younger and older patients with no genes or one gene with high expression had the best outcomes (CR rate, 94% and 87%, respectively; 3-year OS, 80% and 42%, respectively).
Conclusion
A seven-gene score encompassing epigenetic and genetic prognostic information identifies novel AML subsets that are meaningful for treatment guidance.
INTRODUCTION
Most adults with acute myeloid leukemia (AML) are not cured with current therapies, with only approximately 40% of younger (age < 60 years) and approximately 10% of older (age ≥ 60 years) patients achieving long-term survival.1,2 Therefore, novel strategies are needed to improve outcome and identify new therapeutic targets. To achieve this goal, recent studies have focused on identifying molecular features associated with clinical outcome and have applied them to patients' risk stratification and treatment guidance.3 To date, disease classification and prognostication for patients with AML have been largely based on cytogenetic and genetic testing, whereas other factors, including epigenetic changes, have not been considered.3,4
Epigenetics describes modifications of gene expression without alteration of DNA sequences.5,6 Aberrant DNA methylation is among the best characterized and therapeutically targetable of the epigenetic changes occurring in leukemia.5,6 However, most studies of aberrant DNA methylation in AML have focused on changes occurring only in subsets of genes, and few reports have analyzed whole-genome DNA methylation changes.7,8 Prognostic signatures based solely on DNA methylation have been reported,7,9 but to our knowledge, an integrated and clinically applicable biomarker in AML that encompasses both genetic and epigenetic information for prognostication and treatment response prediction is not available. Therefore, we developed a novel approach that used the prognostic methylated DNA profiles associated with known prognostic gene mutations to compute a clinically meaningful gene expression summary score. This score was studied in cytogenetically normal (CN) patients with AML, the largest cytogenetic AML subgroup.
METHODS
For detailed methods see the Data Supplement. For study strategy and glossary, see Figure 1 and Table 1, respectively.
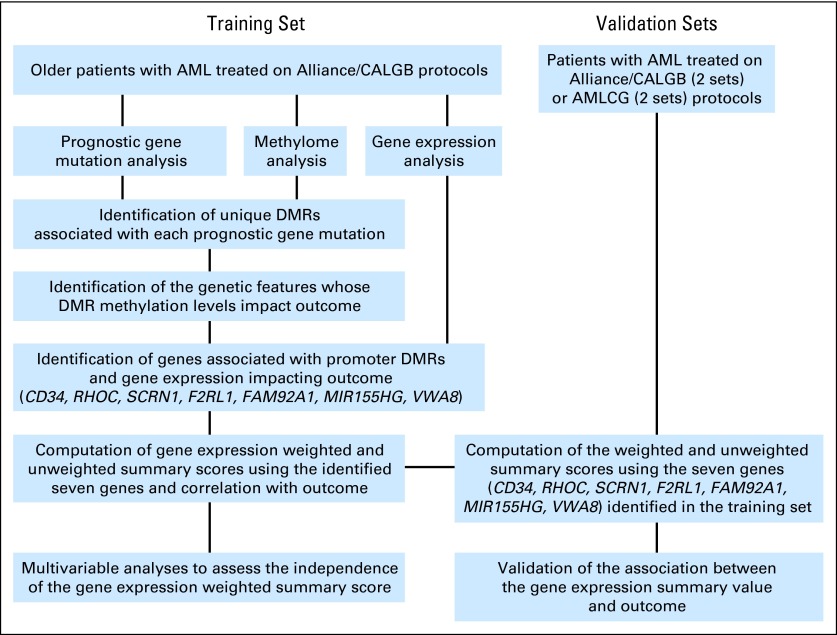
Fig 1.
Overview of the experimental strategy applied to a training set and validation sets of patients with cytogenetically normal acute myeloid leukemia (AML). Established AML prognostic mutations were studied for differentially methylated regions (DMRs) associated with each of the mutations. The association of the DMR methylation levels of distinct genomic features with outcome (ie, overall survival) was analyzed. Promoter DMRs were associated with outcome. Genes with DMRs and expression significantly associated with outcome at P ≤ .001 were identified. Using the expression levels of these genes, a weighted summary score was computed and its clinical impact was tested in the training set and validated in four independent sets of patients. AMLCG, German Acute Myeloid Leukemia Cooperative Group; CALGB, Cancer and Leukemia Group B.
Table 1.
Glossary
| Term | Description |
|---|---|
| Prognostic mutations |
NPM1, FLT3-ITD, DNMT3A (DNMT3A R882, DNMT3A non-R882), IDH1, IDH2 (IDH2 R140, IDH2 R172), TET2, RUNX1, CEBPA (CEBPA-2mut, CEBPA-1mut), FLT3-TKD, AXSL1, WT1, MLL-PTD. |
| DMRs | Regions with significant changes in the levels of DNA methylation when patients with mutations v wild-type patients were compared. The analysis to identify the DMRs was repeated for each of the prognostic mutations. |
| Unique DMRs | DMRs associated with only one of the prognostic gene mutations. |
| High-methylation DMRs | DMRs with methylation level higher in the mutated v wild-type patients. |
| Low-methylation DMRs | DMRs with methylation level lower in the mutated v wild-type patients. |
| Genomic features | Large genomic regions with distinct putative functions. They host DMRs. |
| Weighted summary score | A linear combination of the median dichotomized values of the expressions of seven genes whose promoter DMR methylation and expression levels were simultaneously predictive of clinical outcome. The weights for the linear combination were the overall survival Cox regression coefficients of the genes. The median value of the score was used to dichotomize patients into those with high and low scores for prognostic analyses. |
| Unweighted summary score | A linear combination of the median dichotomized values of the expressions of seven genes whose promoter DMR methylation and expression levels were both predictive of clinical outcome. This score indicated the number of genes that were highly expressed in each patient. |
Abbreviation: DMR, differentially methylated region.
Patients and Treatment
A training set of older (≥ 60 years) patients (n = 134) with untreated primary CN-AML who received first-line cytarabine-daunorubicin–based therapy on Cancer and Leukemia Group B (CALGB)/Alliance first-line protocols were sequenced for methylated DNA using a high-throughput approach.10 Of 134 patients, 126 also had gene expression data measured.11,12
Four independent sets comprising 355 patients were used for validation. Two sets comprised older and younger (< 60 years) adults with untreated, primary CN-AML receiving intensive cytarabine-daunorubicin–based therapy on CALGB/Alliance first-line protocols (Data Supplement). Two other sets comprised older and younger adult patients with CN-AML treated on the German AML Cooperative Group (AMLCG) 1999 trial.11
All patients provided written informed consent. Study protocols were in accordance with the Declaration of Helsinki and approved by institutional review boards at each center.
Genetic Molecular Analyses
For the CALGB/Alliance training and validation sets, prognostic mutations (NPM1, FLT3-ITD, DNMT3A [DNMT3A R882, DNMT3A non-R882], IDH1, IDH2 [IDH2 R140, IDH2 R172], TET2, RUNX1, CEBPA [CEBPA-2mut, CEBPA-1mut], FLT3-TKD, AXSL1, WT1, MLL-PTD; Fig 1; Data Supplement) and gene expression microarray data were analyzed centrally.11,12 For patients in the AMLCG validation sets, molecular status, gene expression, and outcome were previously reported.11
DNA Methylation Analysis
For the training set, high-throughput next-generation sequencing of methylated DNA fragments (MethylCap-seq; Diagenode, Denville, NJ; 150 to 200 base pairs) enriched by MBD2 protein (MethylMiner Methylated DNA Enrichment Kit; Invitrogen, Grand Island, NY) was performed on pretreatment bone marrow, as previously reported.10 Reduced representation bisulfite sequencing13 and MassArray14 were used to validate the MethylCap-seq findings (Data Supplement).
Differentially Methylated Regions
In the training set, DNA methylation patterns associated with each of the prognostic mutations were identified by comparing patients mutated for each mutation with patients wild-type for that mutation. Differentially methylated regions (DMRs) were defined as DNA sequences with significantly different DNA methylation levels between mutated and wild-type patients (Wilcoxon rank sum test). P values were adjusted for multiple comparisons using false discovery rate correction. Only regions with a false discovery rate cutoff of 0.05 were deemed DMRs. DMRs were considered unique if they were associated with only one prognostic mutation. If the same DMR was identified for two or more mutations, it was not included in further analyses. High- and low-methylation DMRs were DMRs with median methylation levels higher or lower, respectively, in the mutated than in the wild-type patients.
Definition of Genomic Features
Distribution of all unique DMRs among genomic features, large genomic regions with putative distinct functions, was evaluated. For this analysis, seven genomic features (Data Supplement) were selected for their potential impact on gene expression.10,15–22 The association of the summed DMR methylation levels with outcome was tested for each genomic feature. We found that the summed DMR methylation levels of the RefSeq gene promoter (hereafter called promoter) genomic features impacted on outcome (P ≤ .05).
Association of DMRs With Gene Expression
The promoter genomic features are part of genes. This enabled us to identify in the training set genes whose promoter DMR methylation levels and expression levels were associated with outcome (see Statistical Analyses). The genes with both methylation and expression levels associated with overall survival (OS) at P ≤ .001 were used to compute a gene expression weighted summary score.
Statistical Analyses
Definition of outcome end points and detailed statistical methods, including the derivation of the seven-gene expression weighted summary score, are reported in the Data Supplement (see also the footnote marked with a dagger symbol (†) in Table 4). The associations of the weighted summary score (low/high) with baseline clinical, demographic, and molecular features were compared using the Wilcoxon rank sum and Fisher's exact tests for continuous and categorical variables, respectively. For time-to-event analyses, survival estimates were calculated using the Kaplan-Meier method, and groups were compared using the log-rank test.
Table 4.
Multivariable Models for Clinical Outcome in the Training Set of Patients With Cytogenetically Normal Acute Myeloid Leukemia
| Variables in Final Model* | OR/HR | 95% CI | P |
|---|---|---|---|
| CR | |||
| Weighted summary score,† high v low | 0.17 | 0.07 to 0.41 | .01 |
| NPM1, mutated v wild-type | 2.37 | 1.00 to 5.59 | .05 |
| DFS | |||
| Weighted summary score,† high v low | 3.59 | 1.83 to 7.03 | < .001 |
| ERG expression, high v low | 1.93 | 1.04 to 3.55 | .04 |
| OS | |||
| Weighted summary score,† high v low | 3.41 | 2.13 to 5.47 | < .001 |
| DNMT3A, R882 v wild-type or non-R882 | 2.35 | 1.38 to 4.00 | .002 |
| BAALC expression, high v low | 1.73 | 1.10 to 2.71 | .02 |
| Platelets, continuous, 50-unit increase | 1.16 | 1.02 to 1.33 | .02 |
Abbreviations: CR, complete remission; DFS, disease-free survival; HR, hazard ratio; OR, odds ratio; OS, overall survival.
Variables that were significant at P < .20 in univariable models were considered for multivariable model inclusion and were as follows: for CR, weighted summary score (high v low), NPM1 (mutated v wild-type), IDH2 (R140 or R172 mutated v wild-type), ASXL1 (mutated v wild-type), RUNX1 (mutated v wild-type), BAALC expression (high v low), platelets (continuous, 50-unit increase), and WBC count (continuous, 50-unit increase); for DFS, weighted summary score (high v low), FLT3-ITD (present v absent), FLT3-TKD (present v absent), ASXL1 (mutated v wild-type), DNMT3A (R882 mutated v non-R882 mutated or wild-type), RUNX1 (mutated v wild-type), ERG expression (high v low), BAALC expression (high v low), age (continuous, 10-year increase), and sex (male v female); for OS, weighted summary score (high v low), NPM1 (mutated v wild-type), FLT3-ITD (present v absent), CEBPA (mutated v wild-type), ASXL1 (mutated v wild-type), DNMT3A (R882 mutated v non-R882 mutated or wild-type), RUNX1 (mutated v wild-type), ERG expression (high v low), BAALC expression (high v low), and platelets (continuous, 50-unit increase).
The weighted summary score is the Cox regression coefficient weighted sum of the median dichotomized values for the seven genes [weighted summary score = 0.71900(CD34) + 0.61927(MIR155HG) + 0.67258(RHOC) + 0.66929(SCRN1) + 0.65925(F2RL1) + 0.65777(FAM92A1) + 0.61491(VWA8)]. For each gene, high and low expression was based on the cohort-specific median cutoff. The weighted summary score was then dichotomized using a cohort-specific median cut to determine the low and high groups. An OR > 1 (< 1) corresponds to a higher (lower) odds of achieving a CR for the first level listed of a dichotomous variable. An HR > 1 corresponds to worse outcome for higher values of continuous variables and the first level listed of a dichotomous variable.
We compared the weighted summary score with other molecular prognosticators and previously reported profiles by computing the Akaike information criterion (AIC).23 Because the weights in the expression weighted summary score were nearly identical for all genes, we also tested an unweighted summary score that simply uses the number of individual genes with high expression. Additional detailed methodology on multivariable analyses and statistical methods are provided in the Data Supplement.
RESULTS
DMRs Associated With Established Prognostic Gene Mutations in CN-AML
To develop a molecular risk score that incorporated both epigenetic and genetic prognostic information (Fig 1), we postulated that these mutations in CN-AML are characterized by distinct DNA methylation patterns that complement their prognostic impact. Furthermore, because both mutations and DNA methylation changes may impact on gene expression, we also postulated that it is possible to derive a gene expression profile that depends on and encompasses the prognostic significance of both mutations and their corresponding DMRs.
To test these hypotheses, we analyzed a training set of older patients with CN-AML (n = 134) who were molecularly characterized for the most common prognostic mutations in AML (Data Supplement). The comparisons of mutated versus wild-type patients for each of the 15 mutations considered revealed 32,681 unique DMRs (Fig 2; Data Supplement). Comparative methylation analyses using alternative techniques (ie, reduced representation bisulfite sequencing and MassArray assays) validated our DMR calls (Data Supplement).
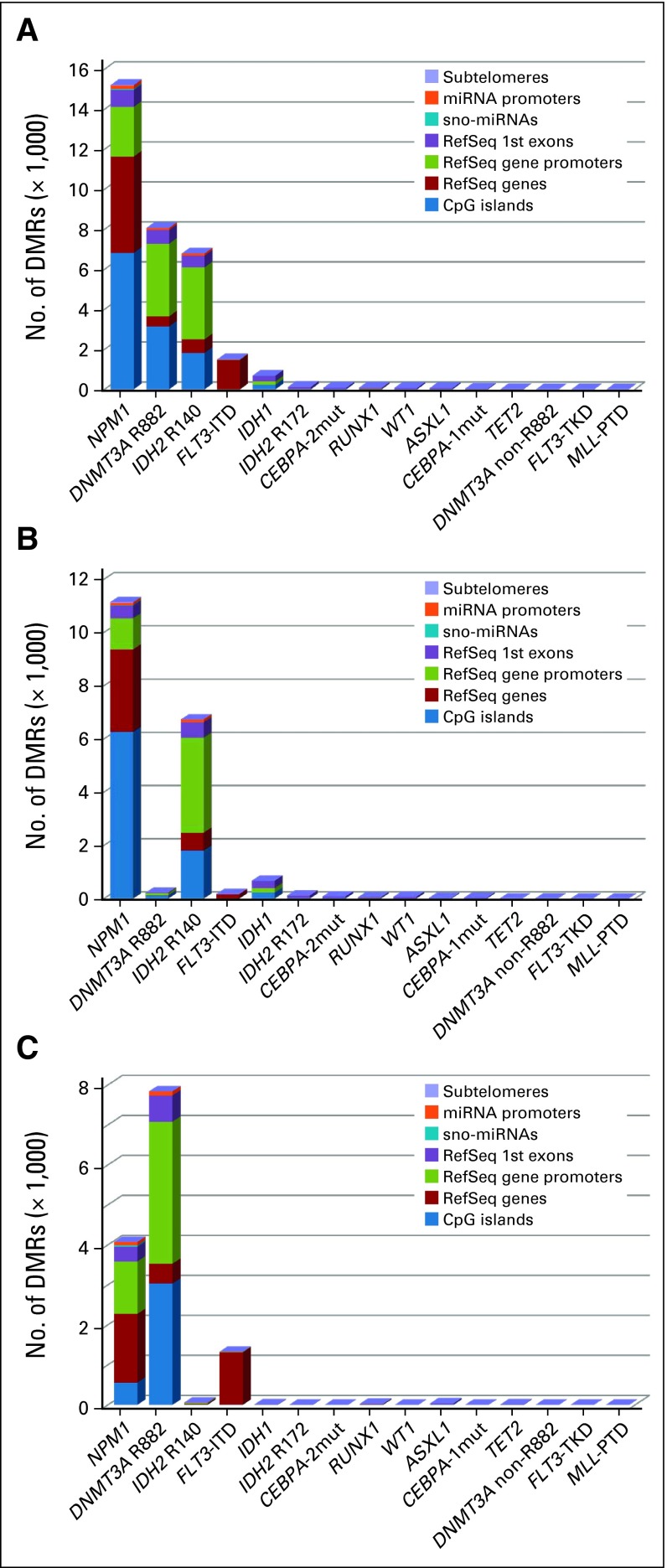
Fig 2.
Unique differentially methylated regions (DMRs) in distinct genomic features identified by comparing patients with cytogenetically normal acute myeloid leukemia with each of the prognostic gene mutations with wild-type patients for that gene mutation. The height of each color block depicts the number of DMRs identified in the respective genomic feature. Data are shown for (A) all DMRs, (B) high-methylation DMRs, and (C) low-methylation DMRs. Genomic regions were selected based on their reported function (Data Supplement), as follows: subtelomeres; microRNA (miRNA) promoters; sno-miRNA; RefSeq first exon; RefSeq gene promoters; RefSeq genes; and CpG islands.
Nearly all DMRs (99.0%) were associated with NPM1 (46.5%), DNMT3A-R882 (24.7%), IDH mutations (23.2%), or FLT3-ITD (4.6%; Data Supplement). Of the 32,681 DMRs, 59% were high-methylation DMRs. IDH and NPM1 mutations were significantly associated with high-methylation DMRs (P < .001 for both), and DNMT3A-R882 mutation and FLT3-ITD were significantly associated with low-methylation DMRs (P < .001 for both). The findings were validated using permutation and equal patient number sampling tests (Data Supplement).
Integrating Epigenetic and Genetic Information in the Training Set
Distribution of the DMRs among the genomic features is shown in the Data Supplement. The summed methylation levels of DMRs in the promoter features impacted on outcome (data not shown). Summed high-methylation levels of promoter DMRs were associated with longer OS than low-methylation levels (P = .02; 3-year OS, 28% v 9%, respectively).
The promoter genomic feature is associated with gene coding sequences. We identified 82 individual genes whose promoter DMRs were associated with OS (P ≤ .001; Data Supplement). For 80 of them, higher DMR methylation was associated with longer OS (P ≤ .001).
Seven of the 82 genes (CD34, RHOC, SCRN1, F2RL1, FAM92A1, MIR155HG, and VWA8; Table 2; Data Supplement) had not only DMRs but also expression levels that were associated with outcome (P ≤ .001 for OS). For each of these seven genes, lower expression and higher DNA methylation were associated with longer OS. With the exception of VWA8, all of these genes had previously been associated with leukemia (CD34 and MIR155HG) or other cancer types (MIR155HG, RHOC, SCRN1, F2RL1, and FAM92A1).24–30
Table 2.
Seven Genes With Promoter DMRs and Gene Expression Associated With Overall Survival
| Gene* | Chromosome Location† | DMR Methylation |
Gene Expression |
Role in Cancer | ||||
|---|---|---|---|---|---|---|---|---|
| HR | 95% CI | P | HR | 95% CI | P | |||
| CD34 | 1q32 | 0.98 | 0.97 to 0.99 | < .001 | 2.05 | 1.41 to 2.99 | < .001 | High expression associated with poor prognosis in AML24 |
| RHOC | 1p13.1 | 0.96 | 0.93 to 0.98 | < .001 | 1.96 | 1.34 to 2.86 | < .001 | Involved in solid tumor growth and invasion25 |
| SCRN1 | 7p14.3-p14.1 | 0.93 | 0.88 to 0.97 | .001 | 1.95 | 1.34 to 2.85 | < .001 | High expression associated with poor prognosis in colon cancer26 |
| F2RL1 | 5q13 | 0.95 | 0.93 to 0.98 | < .001 | 1.93 | 1.33 to 2.82 | < .001 | High expression associated with tumor cell proliferation27 |
| FAM92A1 | 8q22.1 | 0.94 | 0.91 to 0.97 | < .001 | 1.93 | 1.32 to 2.81 | < .001 | Promotes cell growth in renal carcinogenesis28 |
| MIR155HG | 21q21.3 | 0.97 | 0.95 to 0.99 | .001 | 1.86 | 1.27 to 2.71 | .001 | Upregulated in aggressive types of CN-AML29 |
| VWA8 | 13q14.11 | 0.94 | 0.90 to 0.98 | .001 | 1.85 | 1.27 to 2.70 | .001 | Role in cancer unknown30 |
NOTE. For DMR methylation, HR < 1 indicates better survival with increasing methylation values. For gene expression, HR > 1 indicates worse survival with increasing gene expression values.
Abbreviations: AML, acute myeloid leukemia; CN, cytogenetically normal; DMR, differentially methylated region; HR, hazard ratio.
Genes are ordered according to decreasing gene expression HRs.
Methylation regions were as follows: chr1:206150306-206152306 for CD34, chr1:113050201-113052201 for RHOC, chr7:29995430-29997430 for SCRN1, chr5:76149588-76151588 for F2RL1, chr8:94780948-94782948 for FAM92A1, chr21:25855327-25857327 for MIR155HG, and chr13:41432221-41434221 for VWA8.
Computing a Prognostic Weighted Summary Score
A weighted summary score of the expression levels of the seven genes was computed and tested for its prognostic significance in the training set. Patients were dichotomized into low- and high-score groups using as a cutoff the median of the patients' score values. Low-score patients presented more frequently with prognostically favorable molecular markers (ie, NPM1 mutations and low BAALC, ERG, and MN1 expression) and less frequently with unfavorable markers (ie, FLT3-ITD, ASXL1, and RUNX1 mutations) compared with high-score patients (Data Supplement). Low-score patients had higher complete remission (CR) rates (P < .001) and longer disease-free survival (DFS; P < .001) and OS (P < .001) than high-score patients (Table 3, Fig 3A). When we compared the weighted summary score with previously reported prognostic molecular markers, the AIC value of the weighted summary score was lower than that of all other prognostic markers, suggesting that the weighted summary score provided the best prognostic information for all three clinical end points (Data Supplement).
Table 3.
Clinical Outcome Based on the Gene Expression Summary Scores in Five Independent Sets of Patients With CN-AML
| Score | Training Cohort CALGB/Alliance, ≥ 60 Years (n = 126) |
VS1 CALGB/Alliance, ≥ 60 Years (n = 72) |
VS2 CALGB/Alliance, < 60 Years (n = 134) |
VS3 AMLCG,* ≥ 60 Years (n = 65) |
VS4 AMLCG,* < 60 Years (n = 84) |
||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CR (%) | 3-Year DFS (%) |
3-Year OS (%) |
CR (%) | 3-Year DFS (%) |
3-Year OS (%) |
CR (%) | 3-Year DFS (%) |
3-Year OS (%) |
CR (%) | 3-Year DFS (%) |
3-Year OS (%) |
CR (%) | 3-Year DFS (%) |
3-Year OS (%) |
|||||||||||
| Rate | 95% CI | Rate | 95% CI | Rate | 95% CI | Rate | 95% CI | Rate | 95% CI | Rate | 95% CI | Rate | 95% CI | Rate | 95% CI | Rate | 95% CI | Rate | 95% CI | ||||||
| Weighted summary score† | |||||||||||||||||||||||||
| P‡ | < .001 | < .001 | < .001 | .12 | .04 | .03 | .004 | < .001 | < .001 | .20 | < .001 | .01 | .007 | .003 | < .001 | ||||||||||
| Low score | 86 | 26 | 15 to 38 | 30 | 19 to 54 | 81 | 21 | 8 to 37 | 33 | 19 to 49 | 94 | 54 | 41 to 65 | 66 | 53 to 76 | 70 | 31 | 14 to 51 | 42 | 24 to 58 | 85 | 58 | 39 to 73 | 70 | 51 to 82 |
| High score | 43 | 4 | 0 to 16 | 5 | 1 to 12 | 61 | 5 | 0 to 19 | 11 | 4 to 24 | 75 | 24 | 13 to 36 | 28 | 18 to 39 | 52 | 0 | 10 | 2 to 27 | 57 | 26 | 10 to 45 | 16 | 6 to 30 | |
| Unweighted summary score§ | |||||||||||||||||||||||||
| P‖ | < .001 | < .001 | < .001 | .01 | .05 | .007 | .02 | < .001 | < .001 | .20 | < .001 | .03 | .01 | .002 | < 001 | ||||||||||
| No. of genes and %¶ | |||||||||||||||||||||||||
| 0-1, 16% | 96 | 32 | 14 to 51 | 39 | 20 to 58 | 89 | NE | 44 | 14 to 72 | 91 | 65 | 40 to 82 | 82 | 59 to 93 | 69 | 42 | 11 to 71 | 46 | 19 to 70 | 100 | 60 | 25 to 83 | 76 | 30 to 94 | |
| 2-3, 40% | 80 | 22 | 10 to 37 | 24 | 14 to 39 | 77 | NE | 26 | 12 to 42 | 89 | 44 | 31 to 56 | 50 | 37 to 61 | 64 | 23 | 6 to 47 | 33 | 15 to 54 | 80 | 51 | 31 to 68 | 63 | 43 to 77 | |
| 4-5, 27% | 54 | 5 | 0 to 20 | 5 | 1 to 15 | 64 | NE | 14 | 3 to 31 | 77 | 25 | 10 to 43 | 29 | 15 to 45 | 60 | 0 | 10 | 1 to 36 | 55 | 42 | 15 to 67 | 18 | 5 to 38 | ||
| 6-7, 17% | 25 | 0 | 4 | 0 to 17 | 50 | NE | 10 | 1 to 36 | 71 | 17 | 3 to 41 | 24 | 7 to 45 | 50 | 0 | 12 | 1 to 38 | 53 | 13 | 1 to 42 | 7 | 0 to 28 | |||
NOTE. Median ages in the patient sets were as follows: training cohort, 69 years (range, 60 to 83 years); VS1, 69 years (range, 60 to 81 years); VS2, 46 years (range, 18 to 69 years); VS3, 67 years (range, 60 to 83 years); and VS4, 47 years (range, 17 to 59 years).
Abbreviations: AMLCG, German Acute Myeloid Leukemia Cooperative Group; CALGB, Cancer and Leukemia Group B; CN-AML, cytogenetically normal acute myeloid leukemia; CR, complete remission; DFS, disease-free survival; NE, data not evaluable as a result of small sample sizes (ie, if any of the categories have five or fewer patients for a given end point); OS, overall survival; VS, validation set.
Data from Metzeler et al.11 Data set accession number GSE12417. Patients with induction death were excluded.
The weighted summary score is the Cox regression coefficient weighted sum of the median dichotomized values for the seven genes [weighted summary score = 0.71900(CD34) + 0.61927(MIR155HG) + 0.67258(RHOC) + 0.66929(SCRN1) + 0.65925(F2RL1) + 0.65777(FAM92A1) + 0.61491(VWA8)]. For each gene, high and low expression was based on the cohort-specific median cutoff. The weighted summary score was then dichotomized using a cohort-specific median cut to determine the low and high groups.
For CR, the P value is from the Fisher's exact test; for DFS and OS, the P value is from the log-rank test.
The summary score is the unweighted sum of the median dichotomized values for the seven genes (CD34, MIR155HG, RHOC, SCRN1, F2RL1, FAM92A1, and VWA8). For each gene, high and low expression was based on the cohort-specific median cutoff. The value for the score can also be interpreted as the number of genes in the summary score with high expression.
P values are calculated using the continuous unweighted summary score. For CR, the P value is from the logistic regression model; for DFS and OS, the P value is from the Cox proportional hazards model.
Percentage of patients in each category out of the total set of patients analyzed (N = 481). See the Data Supplement for additional detail.
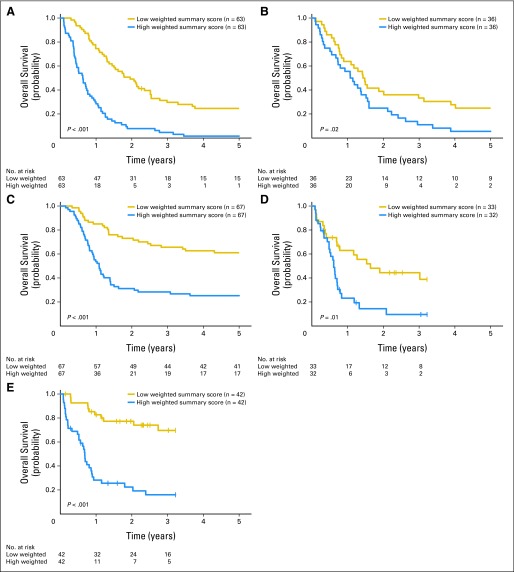
Fig 3.
Overall survival of patients with cytogenetically normal acute myeloid leukemia (CN-AML). (A) Training set. Validation sets (VSs) were as follows: (B) older (VS1) and (C) younger (VS2) Cancer and Leukemia Group B/Alliance patients with CN-AML; and (D) older (VS3) and (E) younger (VS4) German Acute Myeloid Leukemia Cooperative Group patients with CN-AML.9
In multivariable analyses, the weighted summary score was independently associated with higher CR rates (P = .01) and longer DFS (P < .001) and OS (P < .001), after adjusting for other clinical and molecular prognosticators (Table 4). Patients with low scores had a more than 80% increase in the odds of achieving CR and approximately 3.5-fold decrease in the risk of disease relapse or death compared with patients with high scores.
Validation of the Weighted Summary Score
The clinical impact of the weighted summary score was validated in four independent CN-AML patient sets (n = 355). Validation set 1 comprised older CALGB/Alliance patients with primary AML; patients with low scores had a trend for higher CR rates (P = .12) and longer DFS (P = .04) and OS (P = .02) than those with high scores (Table 3; Fig 3B). Validation set 2 comprised younger CALGB/Alliance patients with primary AML; patients with low scores had a higher CR rate (P = .004) and longer DFS (P < .001) and OS (P < .001) than those with high scores (Table 3; Fig 3C). Validation sets 3 and 4 comprised older and younger AMLCG patients with CN-AML, respectively.11 In validation set 3, a low score was associated with longer DFS (P < .001) and OS (P = .01) but not a significantly better CR rate (P = .20; Table 3; Fig 3D). In validation set 4, a low score was associated with a better CR rate (P = .007) and longer DFS (P = .003) and OS (P < .001; Table 3; Fig 3E).
When we considered all patients together (training and validation sets; n = 481), those with low scores, compared with those with high scores, had a higher CR rate (85% v 58%, respectively; P < .001) and longer DFS (3-year DFS, 40% v 15%, respectively; P < .001) and OS (3-year OS, 48% v 15%, respectively; P < .001; Data Supplement). In multivariable models, the weighted summary score remained an independent predictor for CR (P < .001), DFS (P < .001), and OS (P < .001), after adjusting for other clinical and molecular variables (Data Supplement).
When we compared the weighted summary score with previously reported prognostic gene expression profiles, the AIC values of the weighted summary score were lower than the AIC values for the leukemia stem-cell profile of Eppert et al31 and the AML profile of Li et al32 for all clinical end points, suggesting that the score provides a better prognostic model for CN-AML than other previously reported gene expression profiles (Data Supplement).
Clinical Applicability of the Unweighted Summary Score
Although the weighted summary score allowed for the concurrent assessment of the prognostic impact of expression of all seven genes, for prospective evaluation and risk stratification of individual patients, it is simpler to compute an unweighted summary score reporting the number of the genes with high expression. Thus, we computed and tested the unweighted summary score for the seven genes. Patients in the training set with no highly expressed genes or one highly expressed gene had 96% CR, 32% 3-year DFS, and 39% 3-year OS rates, compared with rates of 25%, 0%, and 4%, respectively, in patients with six or seven highly expressed genes (Table 3). The unweighted summary score also proved to be a better model compared with all other prognostic markers (Data Supplement).
Furthermore, patients with no highly expressed genes or only one highly expressed gene had an excellent outcome compared with patients with six or seven highly expressed genes in all four validation sets (Table 3). Older patients in validation sets 1 and 3 with no highly expressed genes or one highly expressed gene had CR rates of 89% and 69%, respectively, compared with 50% for patients with six or seven highly expressed genes, and had 3-year OS rates of 44% and 46%, respectively, compared with 10% and 12%, respectively, for patients with six or seven highly expressed genes. Younger patients in validation sets 2 and 4 with no highly expressed genes or one highly expressed gene had CR rates of 91% and 100%, respectively, compared with 71% and 53%, respectively, for patients with six or seven highly expressed genes, and had 3-year OS rates of 82% and 76%, respectively, compared with 24% and 7%, respectively, for patients with six or seven highly expressed genes. The unweighted summary score's AIC values were also better than the AIC values of the other previously reported gene expression profiles (Data Supplement).31,32
DISCUSSION
We report here a gene expression score that incorporates prognostic genetic and epigenetic information. The score was derived from a training set of older patients with CN-AML and validated in four independent older and younger patient sets.
Our approach is novel and relatively straightforward and provides a strategy to integrate epigenetic and genetic information. Different from studies that have been mainly descriptive of the concurrent presence of aberrant genetic and epigenetic patterns in AML,8 our work provides insight into the clinical relevance of prognostic mutations and the mutation-associated gene DNA methylation promoter and expression patterns. In fact, we postulated that each prognostic gene mutation had a defined methylation profile that distinguished mutated from wild-type patients. The methylation profile comprised DMRs, regions with significantly different methylated levels. The DMRs associated with prognostic gene mutations were distributed among different genomic features. DMRs in the promoter feature had a prognostic impact; high methylation levels of these DMRs were associated favorably with OS. Considering individual DMRs, we found seven genes whose promoter DMRs and expression levels were both significantly associated with outcome. These genes could not have been identified without the initial steps of relating prognostic mutations with their corresponding DNA methylation patterns. In fact, none of the seven genes were previously reported in other prognostic studies based exclusively on gene methylation status.7,9 However, it should be noted that methodologic differences prevented a meaningful, direct comparison of our integrated genetic-epigenetic score with previously reported epigenetic signatures.7,9
For all seven genes, high methylation and low expression levels were associated with better outcome, thereby supporting an inverse functional relationship between hypermethylated promoters, usually resulting in epigenetic gene silencing, and low gene expression. Notably, of the seven genes in the score, six (CD34, RHOC, SCRN1, F2RL1, FAM92A1, and MIR155HG) have been associated with an aggressive tumor phenotype, supporting not only a prognostic but also a biologic relevance of the expression levels of these genes that could be regarded as putative oncogenes.24–30
The expression levels of the seven genes were used to compute a weighted summary score. A low score corresponding to low expression levels of the seven genes was associated with better outcome in the training set of older patients with CN-AML. The weighted summary score not only predicted CR rates and DFS and OS duration, but also provided better prognostic information than other molecular prognosticators, including NPM1 mutation, one of the strongest prognosticators in older patients with CN-AML.33 The prognostic value of the weighted summary score was validated in four independent sets of older and younger patients with CN-AML. Furthermore, the excellent prognostic value of the weighted summary score was also supported by the comparison of the score with other recently reported prognostic gene expression profiles, including a stem-cell profile.30,31 The lowest AIC value of the weighted summary score among these gene expression profiles indicated that it provided the best prognostic model.
Importantly, all seven genes in the weighted summary score had a similar prognostic impact. This enabled us to compute an unweighted summary score to report simply the number of the seven highly expressed genes for each patient. In all five sets of patients analyzed, we observed a similar trend, where patients with no highly expressed genes or only one highly expressed gene, representing approximately 15% of patients with CN-AML, had an excellent outcome (Data Supplement). These results support the usefulness for treatment guidance of the unweighted summary score for identifying a relatively small, but clinically significant subset of patients who respond better to chemotherapy. This could be particularly valuable for older patients with AML. These patients have an unfavorable prognosis and often are treated with reduced-intensity treatment or best supportive care to spare them the toxicity of intense, but frequently ineffective, chemotherapy. In our study, older patients with no or only one highly expressed gene had excellent CR rates (range, 69% to 96%) and 3-year OS rates (range, 39% to 46%), suggesting that the seven-gene score can select for chemotherapy-responsive older patients. Furthermore, low-score patients may also be excluded from hypomethylating treatments. In fact, a low score indicated low expression of genes that are highly methylated and partially repressed. Thus, DNA hypomethylating agents could harmfully reactivate one or more of the seven genes, thereby unleashing their leukemogenic potential.
From a biologic standpoint, we also observed that FLT3-ITD and DNMT3A-R882 mutations were preferentially associated with low-methylation DMRs, and NPM1 and IDH mutations were associated with high-methylation DMRs. Our results are consistent with those reported by other groups.7,8,33–35 However, concerning DNMT3A mutations,8,36 we noted that only the DNMT3A-R882 mutations, and not the DNMT3A non-R882 mutations, were associated with low-methylation DMRs, supporting the notion that loss of function may be restricted to the former. Because we showed that low methylation of the promoter features was associated with worse outcome, these results are consistent with our previous report showing that the DNMT3A-R882 mutation, which may lead to a decrease of DNA methylating activity of the encoded protein, also has a worse prognostic impact in older patients with CN-AML.37
In summary, integration of epigenetic and genetic prognostic information is feasible and revealed a novel seven-gene summary score that identified clinically meaningful groups of patients with AML who responded differently to chemotherapy regardless of age. Considering the relatively small number of genes making up the score, it should be feasible to rapidly develop and implement a simple quantitative assay that allows identification of the number of the seven genes with high expression and, in turn, prospective risk stratification of individual patients. Gene expression is indeed already used in other types of cancer (ie, breast).38 Moreover, except for MIR155HG and CD34, the remaining five genes in the score have not been previously associated with AML, suggesting that they may be involved in novel leukemogenic pathways and thus represent therapeutic targets that warrant future investigation.
Supplementary Material
Footnotes
Supported in part by Grants No. CA101140, CA114725, CA31946, CA33601, CA16058, CA77658, CA129657, and CA140158 from the National Cancer Institute; the Coleman Leukemia Research Foundation; the Deutsche Krebshilfe-Dr Mildred Scheel Cancer Foundation (H.B.); the Pelotonia Fellowship Program (A.-K.E.); and the Conquer Cancer Foundation (J.H.M.).
Authors' disclosures of potential conflicts of interest and author contributions are found at the end of this article.
AUTHORS' DISCLOSURES OF POTENTIAL CONFLICTS OF INTEREST
The author(s) indicated no potential conflicts of interest.
AUTHOR CONTRIBUTIONS
Conception and design: Guido Marcucci, Pearlly Yan, Ralf Bundschuh, Clara D. Bloomfield
Financial support: Guido Marcucci, Clara D. Bloomfield
Administrative support: Guido Marcucci, Clara D. Bloomfield
Provision of study materials or patients: Guido Marcucci, Maria R. Baer, Meir Wetzler, Thomas H. Carter, Bayard L. Powell, Jonathan E. Kolitz, John C. Byrd, Michael A. Caligiuri, Richard M. Stone, Clara D. Bloomfield
Collection and assembly of data: Guido Marcucci, Yue-Zhong Wu, Donna Bucci, John P. Curfman, Susan P. Whitman, Ann-Kathrin Eisfeld, Jason H. Mendler, Sebastian Schwind, Heiko Becker, Constance Bär, Andrew J. Carroll, Maria R. Baer, Meir Wetzler, Thomas H. Carter, Bayard L. Powell, Jonathan E. Kolitz, Richard M. Stone
Data analysis and interpretation: Guido Marcucci, Kati Maharry, David Frankhouser, Deedra Nicolet, Klaus H. Metzeler, Jessica Kohlschmidt, Krzysztof Mrózek, John C. Byrd, Christoph Plass, Ramiro Garzon, Michael A. Caligiuri, Stefano Volinia, Ralf Bundschuh, Clara D. Bloomfield
Manuscript writing: All authors
Final approval of manuscript: All authors
REFERENCES
- 1.Estey E. AML in older patients: Are we making progress? Best Pract Res Clin Haematol. 2009;22:529–536. doi: 10.1016/j.beha.2009.08.007. [DOI] [PubMed] [Google Scholar]
- 2.Dombret H. Gene mutation and AML pathogenesis. Blood. 2011;118:5366–5367. doi: 10.1182/blood-2011-09-379081. [DOI] [PubMed] [Google Scholar]
- 3.Marcucci G, Haferlach T, Döhner H. Molecular genetics of adult acute myeloid leukemia: Prognostic and therapeutic implications. J Clin Oncol. 2011;29:475–486. doi: 10.1200/JCO.2010.30.2554. [DOI] [PubMed] [Google Scholar]
- 4.Mrózek K, Marcucci G, Nicolet D, et al. Prognostic significance of the European LeukemiaNet standardized system for reporting cytogenetic and molecular alterations in adults with acute myeloid leukemia. J Clin Oncol. 2012;30:4515–4523. doi: 10.1200/JCO.2012.43.4738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Plass C, Oakes C, Blum W, et al. Epigenetics in acute myeloid leukemia. Semin Oncol. 2008;35:378–387. doi: 10.1053/j.seminoncol.2008.04.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Blum W, Garzon R, Klisovic RB, et al. Clinical response and miR-29b predictive significance in older AML patients treated with a 10-day schedule of decitabine. Proc Natl Acad Sci U S A. 2010;107:7473–7478. doi: 10.1073/pnas.1002650107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Figueroa ME, Lugthart S, Li Y, et al. DNA methylation signatures identify biologically distinct subtypes in acute myeloid leukemia. Cancer Cell. 2010;17:13–27. doi: 10.1016/j.ccr.2009.11.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Cancer Genome Atlas Research Network. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N Engl J Med. 2013;368:2059–2074. doi: 10.1056/NEJMoa1301689. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Deneberg S, Guardiola P, Lennartsson A, et al. Prognostic DNA methylation patterns in cytogenetically normal acute myeloid leukemia are predefined by stem cell chromatin marks. Blood. 2011;118:5573–5582. doi: 10.1182/blood-2011-01-332353. [DOI] [PubMed] [Google Scholar]
- 10.Yan P, Frankhouser D, Murphy M, et al. Genome-wide methylation profiling in decitabine-treated patients with acute myeloid leukemia. Blood. 2012;120:2466–2474. doi: 10.1182/blood-2012-05-429175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Metzeler KH, Hummel M, Bloomfield CD, et al. An 86-probe-set gene-expression signature predicts survival in cytogenetically normal acute myeloid leukemia. Blood. 2008;112:4193–4201. doi: 10.1182/blood-2008-02-134411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Mendler JH, Maharry K, Radmacher MD, et al. RUNX1 mutations are associated with poor outcome in younger and older patients with cytogenetically normal acute myeloid leukemia and with distinct gene and microRNA expression signatures. J Clin Oncol. 2012;30:3109–3118. doi: 10.1200/JCO.2011.40.6652. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Gu H, Smith ZD, Bock C, et al. Preparation of reduced representation bisulfite sequencing libraries for genome-scale DNA methylation profiling. Nat Protoc. 2011;6:468–481. doi: 10.1038/nprot.2010.190. [DOI] [PubMed] [Google Scholar]
- 14.Claus R, Hackanson B, Poetsch AR, et al. Quantitative analyses of DAPK1 methylation in AML and MDS. Int J Cancer. 2012;131:E138–E142. doi: 10.1002/ijc.26429. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Roh T-Y, Cuddapah S, Zhao K. Active chromatin domains are defined by acetylation islands revealed by genome-wide mapping. Genes Dev. 2005;19:542–552. doi: 10.1101/gad.1272505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Riethman H, Ambrosini A, Paul S. Human subtelomere structure and variation. Chromosome Res. 2005;13:505–515. doi: 10.1007/s10577-005-0998-1. [DOI] [PubMed] [Google Scholar]
- 17.Deaton AM, Bird A. CpG islands and the regulation of transcription. Genes Dev. 2011;25:1010–1022. doi: 10.1101/gad.2037511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Jones PA. Functions of DNA methylation: Islands, start sites, gene bodies and beyond. Nat Rev Genet. 2012;13:484–492. doi: 10.1038/nrg3230. [DOI] [PubMed] [Google Scholar]
- 19.Baylin SB, Chen WY. Aberrant gene silencing in tumor progression: Implications for control of cancer. Cold Spring Harb Symp Quant Biol. 2005;70:427–433. doi: 10.1101/sqb.2005.70.010. [DOI] [PubMed] [Google Scholar]
- 20.Baer C, Claus R, Frenzel LP, et al. Extensive promoter DNA hypermethylation and hypomethylation is associated with aberrant microRNA expression in chronic lymphocytic leukemia. Cancer Res. 2012;72:3775–3785. doi: 10.1158/0008-5472.CAN-12-0803. [DOI] [PubMed] [Google Scholar]
- 21.Brenet F, Moh M, Funk P, et al. DNA methylation of the first exon is tightly linked to transcriptional silencing. PLoS One. 2011;6:e14524. doi: 10.1371/journal.pone.0014524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Suzuki H, Maruyama R, Yamamoto E, et al. DNA methylation and microRNA dysregulation in cancer. Mol Oncol. 2012;6:567–578. doi: 10.1016/j.molonc.2012.07.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Burnham KP, Anderson DR. Multimodel inference: Understanding AIC and BIC in model selection. Sociol Methods Res. 2004;33:261–304. [Google Scholar]
- 24.Raspadori D, Lauria F, Ventura MA, et al. Incidence and prognostic relevance of CD34 expression in acute myeloblastic leukemia: Analysis of 141 cases. Leuk Res. 1997;21:603–607. doi: 10.1016/s0145-2126(97)00015-5. [DOI] [PubMed] [Google Scholar]
- 25.Karlsson R, Pedersen ED, Wang Z, et al. Rho GTPase function in tumorigenesis. Biochim Biophys Acta. 2009;1796:91–98. doi: 10.1016/j.bbcan.2009.03.003. [DOI] [PubMed] [Google Scholar]
- 26.Miyoshi N, Ishii H, Mimori K, et al. SCRN1 is a novel marker for prognosis in colorectal cancer. J Surg Oncol. 2010;101:156–159. doi: 10.1002/jso.21459. [DOI] [PubMed] [Google Scholar]
- 27.Han N, Jin K, He K, et al. Protease-activated receptors in cancer: A systematic review. Oncol Lett. 2011;2:599–608. doi: 10.3892/ol.2011.291. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Liang S, Gong F, Zhao X, et al. Prokaryotic expression, purification of a new tumor-relative protein FAM92A1-289 and its characterization in renal cell carcinoma. Cancer Lett. 2009;276:81–87. doi: 10.1016/j.canlet.2008.10.043. [DOI] [PubMed] [Google Scholar]
- 29.Marcucci G, Maharry KS, Metzeler KH, et al. Clinical role of microRNAs in cytogenetically normal acute myeloid leukemia: miR-155 upregulation independently identifies high-risk patients. J Clin Oncol. 2013;31:2086–2093. doi: 10.1200/JCO.2012.45.6228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Anney R, Klei L, Pinto D, et al. A genome-wide scan for common alleles affecting risk for autism. Hum Mol Genet. 2010;19:4072–4082. doi: 10.1093/hmg/ddq307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Eppert K, Takenaka K, Lechman ER, et al. Stem cell gene expression programs influence clinical outcome in human leukemia. Nat Med. 2011;17:1086–1093. doi: 10.1038/nm.2415. [DOI] [PubMed] [Google Scholar]
- 32.Li Z, Herold T, He C, et al. Identification of a 24-gene prognostic signature that improves the European LeukemiaNet risk classification of acute myeloid leukemia: An international collaborative study. J Clin Oncol. 2013;31:1172–1181. doi: 10.1200/JCO.2012.44.3184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Becker H, Marcucci G, Maharry K, et al. Favorable prognostic impact of NPM1 mutations in older patients with cytogenetically normal de novo acute myeloid leukemia and associated gene- and microRNA-expression signatures: A Cancer and Leukemia Group B study. J Clin Oncol. 2010;28:596–604. doi: 10.1200/JCO.2009.25.1496. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Figueroa ME, Abdel-Wahab O, Lu C, et al. Leukemic IDH1 and IDH2 mutations result in a hypermethylation phenotype, disrupt TET2 function, and impair hematopoietic differentiation. Cancer Cell. 2010;18:553–567. doi: 10.1016/j.ccr.2010.11.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Shih AH, Abdel-Wahab O, Patel JP, et al. The role of mutations in epigenetic regulators in myeloid malignancies. Nat Rev Cancer. 2012;12:599–612. doi: 10.1038/nrc3343. [DOI] [PubMed] [Google Scholar]
- 36.Ley TJ, Ding L, Walter MJ, et al. DNMT3A mutations in acute myeloid leukemia. N Engl J Med. 2010;363:2424–2433. doi: 10.1056/NEJMoa1005143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Marcucci G, Metzeler KH, Schwind S, et al. Age-related prognostic impact of different types of DNMT3A mutations in adults with primary cytogenetically normal acute myeloid leukemia. J Clin Oncol. 2012;30:742–750. doi: 10.1200/JCO.2011.39.2092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Paik S. Is gene array testing to be considered routine now? Breast. 2011;20(suppl 3):S87–S91. doi: 10.1016/S0960-9776(11)70301-0. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.