Glycosylated α-dystroglycan (α-DG) serves as cellular entry receptor for multiple pathogens, and defects in its glycosylation cause hereditary Walker-Warburg syndrome (WWS). At least eight Jae, page 2 proteins are critical to glycosylate α-DG, but many genes mutated in WWS remain unknown. To identify modifiers of α-DG, we performed a haploid screen for Lassa virus entry, a hemorrhagic fever virus causing thousands of deaths annually that hijacks glycosylated α-DG to enter cells. In complementary screens, we profiled cells for absence of α-DG carbohydrate chains or biochemically related glycans. This revealed virus host factors and a suite of glycosylation units, including all known Walker-Warburg genes and five additional factors critical for the modification of α-DG. Our findings accentuate the complexity of this posttranslational feature and point out genes defective in dystroglycanopathies.
In humans, α-dystroglycan (α-DG) links the extracellular matrix with the cytoskeleton and is extensively modified by sugar chains, including an unusual O-linked glycan (1). Mutations in genes required for α-DG glycosylation lead to congenital disorders, termed dystroglycanopathies. Notable is Walker-Warburg syndrome (WWS) (2), a severe muscular dystrophy with malformations of the eyes and brain, associated with defective binding of α-DG to its ligands, such as laminin (3). The O-linked carbohydrate unit is also used by pathogens to enter their host, including Mycobacterium leprae (leprosy) (4), Lassa virus (LASV), and other Old World arenaviruses (5, 6). At least eight potential glycosyltransferases are required to install the laminin-binding epitope on α-DG (7–9), but only ~50% of the WWS cases are explained by mutations in these genes (8).
We undertook a haploid genetic approach (10) to identify host factors essential for LASV entry. For this purpose, we replaced the glycoprotein of replication-competent vesicular stomatitis virus (VSV) with the Lassa virus glycoprotein (rVSV-GP-LASV) (fig. S1A). This virus infects normal human fibroblasts, whereas patient fibroblasts carrying mutations in the WWS gene ISPD (isoprenoid synthase domain containing) (8, 9) resist infection (fig. S1B). Likewise, haploid human HAP1 cells (11) are also infected and killed in an α-DG–dependent manner (fig. S2A and B).
Mutagenized HAP1 cells were exposed to rVSV-GP-LASV, and gene trap–insertion sites were analyzed in virus-resistant cells (12). Genes significantly enriched for mutagenic gene trap–insertion events include DAG1encoding α-DG, with 316 independent disruptive gene-trap insertions and 25 other genes that are predicted or known to be involved in glycosylation (Fig. 1 and figs. S3 and S4) (13). Among these are LARGE, ISPD, FKTN, FKRP, POMT1, POMT2, DPM3and C3orf39all of which cause dystroglycanopathies (2, 7, 14), and B3GNT1which was uncovered as a new WWS gene during preparation of this manuscript (15). Other hits include genes involved in sialic acid biosynthesis, the generation of uridine diphosphate (UDP)-glucuronic acid and UDP-xylose, N-glycosylation, mannose supply, and localization of glycosylating enzymes in the Golgi apparatus. Last, we found a number of potential enzymes that have not been linked to α-DG modification before (SGK196, TMEM5, PTAR1, ST3GAL4and B3GALNT2) and hits that did not readily connect to glycosylation. None of the enriched genes were identified when mutagenized HAP1 cells were selected with a recombinant VSV carrying the Ebola virus glycoprotein (11), which suggested that they are not required for biology related to the VSV vector. Thus, the haploid screen identifies host factors required for virus entry mediated by the Lassa glycoprotein, including the known entry receptor, known receptor modifiers, and a substantial number of additional genes.
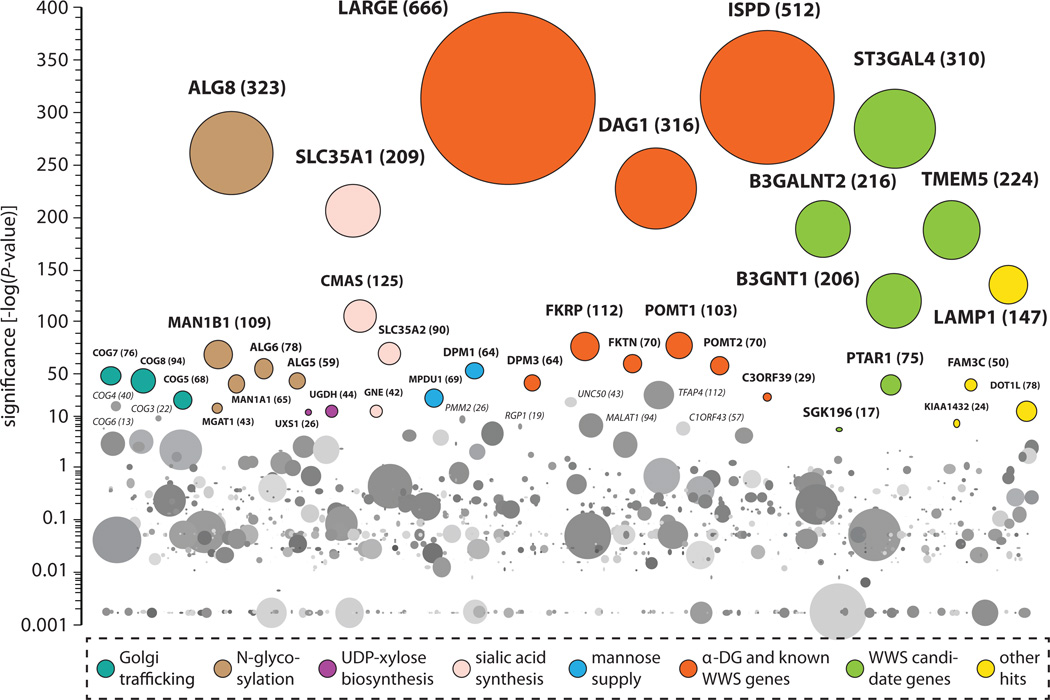
Fig. 1. Haploid genetic screen for cellular host factors required for rVSV-GP-LASV infection.
Significance of enrichment of gene-trap insertions in the virus-selected population Jae, page 7 compared with nonselected control cells is indicated on the y axis. Bubbles represent genes, and bubble size corresponds to the number of independent gene-trap events observed in the virus-selected population. Significant hits are grouped by function horizontally (other genes in random order). Genes carrying the majority of gene-trap insertions in introns were colored if they passed two statistical tests: enrichment of disruptive mutations compared with control cells (one-sided Fisher’s exact test, P ≤ 10−5) and bias for gene trap–insertion events in the transcriptional orientation of the affected gene (binomial test, P ≤ 0.05). Intron-poor genes were colored if they passed the former criterion using a stricter cut-off (one-sided Fisher’s exact test P ≤ 10−30) (13). Data are displayed until −log(P value) = 0.001.
As virus entry is a complex succession of events, we teased apart the roles of the identified genes through a series of comparative genetic screens. Principally, hits could have a Jae, page 3 specific role in α-DG glycosylation, they could affect glycosylation in general, or they could act in virus entry steps unrelated to receptor binding. We enriched mutagenized HAP1 cells for defective presentation of glycosylated α-DG at the cell surface (fig. S5A to C). This population showed a significant increase for haploid cells carrying gene-trap insertions in all known WWS genes, indicating that this mutagenesis screen was carried out at high coverage (Fig. 2A). Genes required for N-glycosylation and sialic acid biosynthesis were not enriched, in line with the notion that the laminin-binding epitope on α-DG is created through O-rather than Nglycosylation (1) and does not require the presence of sialic acid (16). An unexpected exception to this is SLC35A1which encodes a transporter for cytidine monophosphate (CMP)-sialic acid (17, 18). This may indicate that this gene is involved in the transport of other sugars needed for α-DG O-glycosylation or that it indirectly affects generation of the laminin-binding epitope. Together, this screen identifies genes required for α-DG modification and distinguishes them from host factors mediating LASV entry unrelated to α-DG binding (e.g., LAMP1 and genes required for sialic acid biosynthesis).
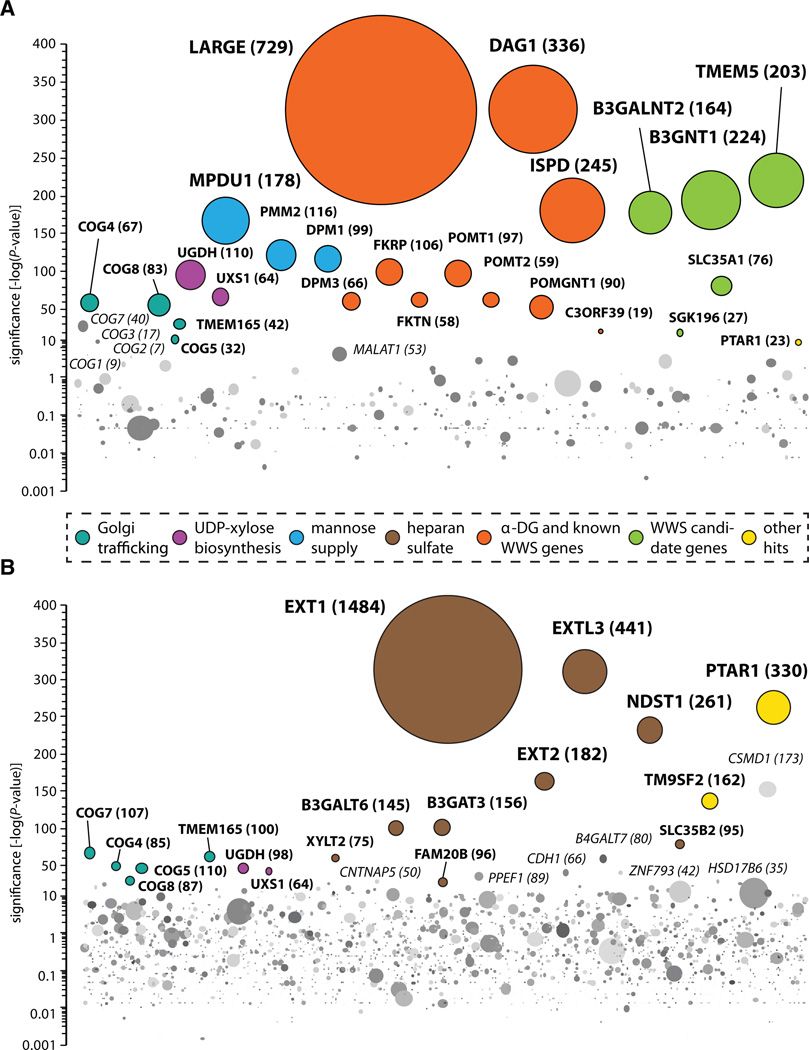
Fig. 2. Cell surface profiling of mutagenized haploid HAP1 cells.
(A) Genes enriched for mutations in a cell population depleted for glycosylated α-DG at the cell surface. The cell population enriched for mutants lacking glycosylated α-DG at the cell surface was analyzed and depicted as described in Fig. 1. (B) A mutant cell population selected for diminished cell surface heparan sulfate was obtained as described above. Data were analyzed as previously, except that, for intron-rich genes, the cut-off for disruptive mutations compared with control cells was adjusted (one-sided Fisher’s exact test, P ≤ 10−21) (13).
To distinguish general glycosylation genes from those required specifically for the generation of the laminin-binding epitope on α-DG, we probed mutagenized HAP1 cells for defects in the generation of heparan sulfate in a separate genetic screen (Fig. 2B and fig. S6). The carbohydrate chains present on α-DG or in heparan sulfate are both thought to contain xylose and glucuronic acid moieties, and indeed, genes required for their biogenesis (UGDH and UXS1) also stood out in this screen (19, 20). Other overlapping hits affect glycosylation globally, such as the COG complex and TMEM165 (21). PTAR1 constitutes a potential prenyltransferase that has not been implicated in glycosylation before but also appears to affect glycosylation globally (fig. S4). Finally, cells depleted for heparan sulfate on their surface were enriched for mutations in heparan sulfate biosynthesis genes (Fig. 2B and fig. S4) (19). This finding suggests that although there are biochemical similarities between heparan sulfate and the O-carbohydrate chains on α-DG, these are by and large installed by separate enzymes.
Using transcription activator–like effector nucleases (TALENs), we generated null alleles for a panel of selected genes in HAP1 cells (fig. S7) (22), and independent clones were isolated carrying frameshift mutations and/or premature stop codons (Fig. 3A and fig. S8). TALEN-induced mutations in all genes except for ST3GAL4 and LAMP1 affected α-DG glycosylation or its ability to interact with laminin (fig. S9A to C, and fig. S10, A and B). This is in agreement with the absence of ST3GAL4 and LAMP1 as hits in the α-DG antibody screen (see Fig. 2A and fig. S4). Mutant cell lines also showed increased resistance to viral infection, although this phenotype was less pronounced in the SGK196 mutants (Fig. 3B and fig. S10C). TALEN-induced phenotypes were reverted by complementation with the respective cDNAs (fig. S11, A and B). In summary, we conclude that TMEM5, B3GALNT2, B3GNT1, SLC35A1and SGK196 constitute genes required for the presentation of the laminin-binding carbohydrate feature present on α-DG, whereas ST3GAL4 and LAMP1 are likely involved in virus infection by means other than modification of α-DG.
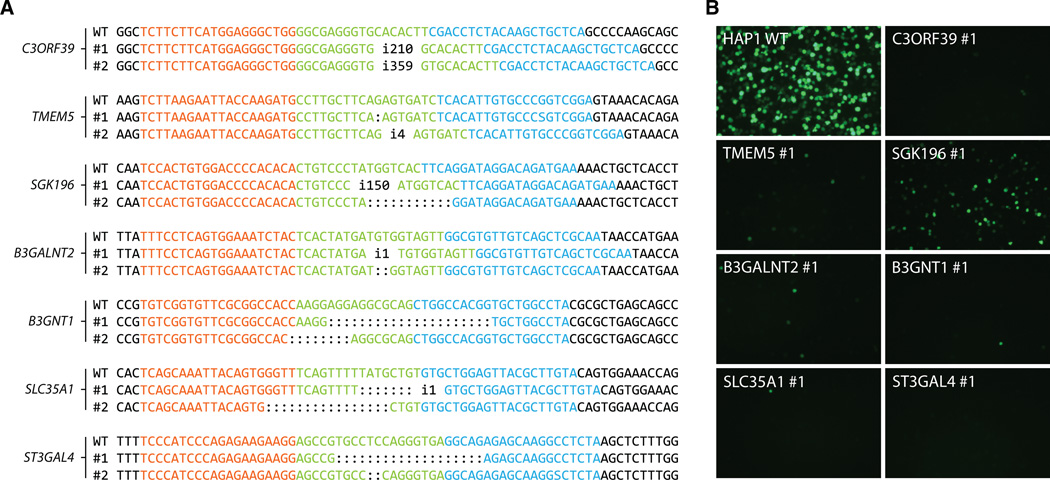
Fig. 3. TALEN-induced mutations in identified genes affect susceptibility to rVSV-GP-LASV.
(A) HAP1 cells transfected with TALENs display frameshift mutations and/or introduce premature stop codons in targeted genes. Sequences recognized by the TALENs are displayed in red and blue. (B) The HAP1 cell lines with TALEN-induced mutations in the corresponding genes and wild-type control cells were infected with rVSV-GP-LASV [infected cells express enhanced green fluorescent protein (eGFP)].
TMEM5 encodes a transmembrane protein that has not been assigned any function but that contains an exostosin family domain (E value 0.0002) (fig. S12) that is also present in the heparan sulfate biosynthesis enzymes EXT1, EXT2and EXTL3. SGK196 contains a kinaselike domain, and knockout mice develop hydrocephalus (23), reminiscent of the brain abnormalities observed in WWS patients. We sequenced the coding exons of TMEM5 and SGK196 in a panel of 28 patients with severe dystroglycanopathy, diagnosed with WWS or muscle-eye-brain Jae, page 4 disease (MEB), not carrying mutations in any known Walker-Warburg gene. Two families with patients that carried homozygous mutations in TMEM5 were identified. One mutant allele results in a stop codon at position Arg340 (c.1018C>T), the other family transmits an early frameshift mutation p.Ala47Argfs*42 (c.139delG) (Fig. 4A). The male patient with the p.Arg340* mutation died at the age of 22 months and had clinical manifestations suggestive of WWS (13). The female siblings carrying the frameshift mutation had a slightly milder phenotype suggestive of MEB. A cranial magnetic resonance image (MRI) of one of the affected girls recorded at the age of 1 year showed brainstem atrophy, dilated ventricles, widespread pachygyria, and substantial white matter involvement (Fig. 4B). During revision of this manuscript, mutations in TMEM5 have also been found in fetuses displaying cobblestone lissencephaly (24). A patient with compound heterozygous mutations (p.Leu137Arg, p.Gln258Arg) in SGK196 and typical WWS phenotype was identified in another family (Fig. 4C). To test causality of the identified mutations for the disease, we supplied HAP1 cells deficient for either SGK196 or TMEM5 with cDNAs encoding the patient-derived variants. Unlike their wild-type counterparts, these neither restored α-DG glycosylation (Fig. 4D) nor enhanced susceptibility to infection with rVSV-GP-LASV (fig. S13). Together, the detection and functional validation of TMEM5 and SGK196 loss-of-function mutations in families with WWS-MEB–type dystroglycanopathy underlines the relevance of the identified α-DG modifiers for human disease.
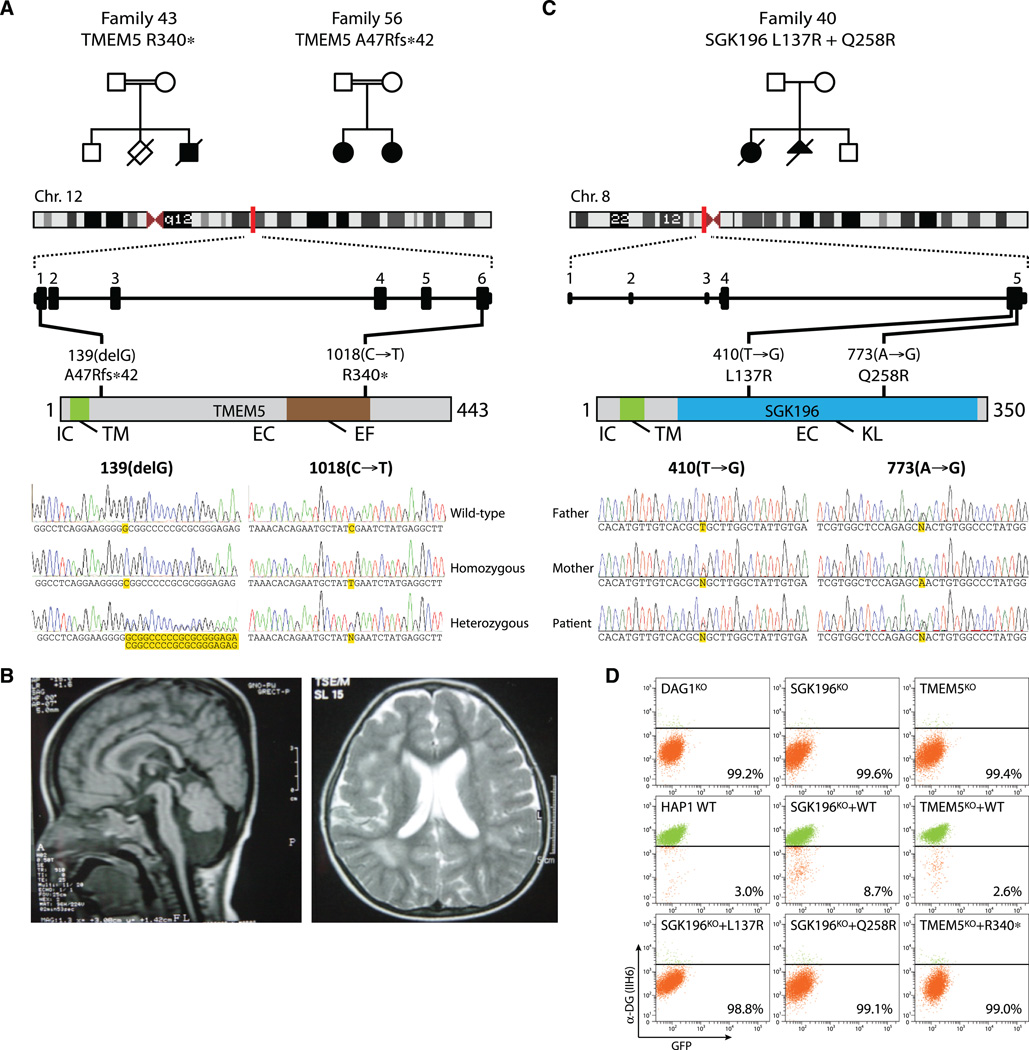
Fig. 4. TMEM5 and SGK196 mutations found in patients with WWS and MEB lacking mutations in known WWS genes.
(A) Pedigree structure of consanguineous, respectively first and second cousins, families 43 and 56 segregating a TMEM5 mutation. Family 43 has an affected male with features of WWS and a stillbirth, without available clinical records. Family 56 has two affected females with clinical features reminiscent of MEB (13). A nonsense mutation in exon 6 was identified in family 43. Family 56 harbors a frameshift mutation in exon 1. Both mutations were homozygously present in the patient(s) and heterozygously in the parents. The unaffected boy in family 43 is heterozygous for the mutation. IC, intracellular domain; TM, transmembrane domain; EC, extracellular domain; EF, exostosin family domain. (B) Cranial MRI of the oldest affected female of family 56 at the age of 1 year; sagittal cut (T1-weighted image): atrophy of pons and cerebellum; axial cut (flair image): fronto-parietal pachygyria, enlarged ventricles, and abnormal white matter. (C) Compound heterozygosity of mutant SGK196 in an affected patient. Both nonconsanguineous parents are heterozygous carriers of either mutation. KL, kinase-like domain (D) HAP1 cells with TALEN-induced disruption of endogenous TMEM5 or SGK196 were complemented with cDNAs encoding the mutant variants observed in patients and analyzed for presence of the α-DG laminin-binding epitope using flow cytometry.
For decades, genes associated with Mendelian disorders have been discovered by studying pedigrees of affected individuals. Although expedited by robust sequencing strategies, the identification of causative mutations in genetically heterogeneous conditions remains problematic. Here, we apply a haploid genetic approach to capture the complexity of a severe hereditary disease in vitro. The resulting “glycosylome” of α-DG highlights the intricate nature of this posttranslational modification and identifies additional genes mutated in Walker-Warburg syndrome. Because polymorphisms associated with the human LARGE gene are under selective pressure in areas where LASV is endemic (25), it becomes of interest to examine the glycosylome genes in virus-exposed populations.
Supplementary Material
Acknowledgements
We thank T. Sixma, J. Roix, S. Mukherjee, S. Hill, and S. Nijman for discussions and S. Radoshitzky and M. Farzan for providing the plasmid encoding the LASV glycoprotein. Supported by The Netherlands Genomics Initiative (NGI), Netherlands Organization for Scientific Research (Vidi-91711316), and the European Research Council (ERC) starting grant (ERC- 2012-StG 309634) to T.R.B., Prinses Beatrix Fonds (W.OR09-15) to D.J.L. and H.V.B., European Union Framework Programme 7 Health Programme (241995 GENCODYS) to H.V.B., and NIH grants AI081842 and AI057159 to S.P.W. T.R.B. is a cofounder of Haplogen GmbH, S.P.W. is inventor on a patent describing the reverse-genetics system for VSV (International Patent no: 5,789,229), J.E.C. and T.R.B. are inventors on a patent on mutagenesis in haploid or near-haploid cells (U.S. Patent Application no: 2012/0190,011), and materials will be made available to the academic community under a Materials Transfer Agreement. The study was approved by the ethical board of the Radboud University Nijmegen Medical Centre, Commissie Mensgebonden Onderzoek Regio Arnhem-Nijmegen Approval 2011/155 (9612-1812). Deep sequencing data have been deposited in the NCBI Sequence Read Archive (http://www.ncbi.nlm.nih.gov/sra) under accession number SRP018361.
References and Notes
- 1.Yoshida-Moriguchi T, et al. O-Mannosyl phosphorylation of alpha-dystroglycan is required for laminin binding. Science. 2010;327:88. doi: 10.1126/science.1180512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Godfrey C, Foley AR, Clement E, Muntoni F. Dystroglycanopathies: Coming into focus. Curr. Opin. Genet. Dev. 2011;21:278. doi: 10.1016/j.gde.2011.02.001. [DOI] [PubMed] [Google Scholar]
- 3.Michele DE, et al. Post-translational disruption of dystroglycan-ligand interactions in congenital muscular dystrophies. Nature. 2002;418:417. doi: 10.1038/nature00837. [DOI] [PubMed] [Google Scholar]
- 4.Rambukkana A, et al. Role of alpha-dystroglycan as a Schwann cell receptor for Mycobacterium leprae . Science. 1998;282:2076. doi: 10.1126/science.282.5396.2076. [DOI] [PubMed] [Google Scholar]
- 5.Cao W, et al. Identification of alpha-dystroglycan as a receptor for lymphocytic choriomeningitis virus and Lassa fever virus. Science. 1998;282:2079. doi: 10.1126/science.282.5396.2079. [DOI] [PubMed] [Google Scholar]
- 6.Kunz S, et al. Posttranslational modification of alpha-dystroglycan, the cellular receptor for arenaviruses, by the glycosyltransferase LARGE is critical for virus binding. J. Virol. 2005;79:14282. doi: 10.1128/JVI.79.22.14282-14296.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Manzini MC, et al. Exome sequencing and functional validation in zebrafish identify GTDC2 mutations as a cause of Walker-Warburg syndrome. Am. J. Hum. Genet. 2012;91:541. doi: 10.1016/j.ajhg.2012.07.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Roscioli T, et al. Mutations in ISPD cause Walker-Warburg syndrome and defective glycosylation of α-dystroglycan. Nat. Genet. 2012;44:581. doi: 10.1038/ng.2253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Willer T, et al. ISPD loss-of-function mutations disrupt dystroglycan O-mannosylation and cause Walker-Warburg syndrome. Nat. Genet. 2012;44:575. doi: 10.1038/ng.2252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Carette JE, et al. Haploid genetic screens in human cells identify host factors used by pathogens. Science. 2009;326:1231. doi: 10.1126/science.1178955. [DOI] [PubMed] [Google Scholar]
- 11.Carette JE, et al. Ebola virus entry requires the cholesterol transporter Niemann-Pick C1. Nature. 2011;477:340. doi: 10.1038/nature10348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Carette JE, et al. Global gene disruption in human cells to assign genes to phenotypes by deep sequencing. Nat. Biotechnol. 2011;29:542. doi: 10.1038/nbt.1857. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Materials and methods are available as supplementary material on Science Online.
- 14.Lefeber DJ, et al. Deficiency of Dol-P-Man synthase subunit DPM3 bridges the congenital disorders of glycosylation with the dystroglycanopathies. Am. J. Hum. Genet. 2009;85:76. doi: 10.1016/j.ajhg.2009.06.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Buysse K, et al. Missense mutations in β-1,3-N-acetylglucosaminyltransferase 1 (B3GNT1) cause Walker-Warburg syndrome. Hum. Mol. Genet. 2013 doi: 10.1093/hmg/ddt021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Combs AC, Ervasti JM. Enhanced laminin binding by alpha-dystroglycan after enzymatic deglycosylation. Biochem. J. 2005;390:303. doi: 10.1042/BJ20050375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Patnaik SK, Stanley P. Lectin-resistant CHO glycosylation mutants. Methods Enzymol. 2006;416:159. doi: 10.1016/S0076-6879(06)16011-5. [DOI] [PubMed] [Google Scholar]
- 18.Eckhardt M, Mühlenhoff M, Bethe A, Gerardy-Schahn R. Expression cloning of the Golgi CMP-sialic acid transporter. Proc. Natl. Acad. Sci. U.S.A. 1996;93:7572. doi: 10.1073/pnas.93.15.7572. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Hari RJL, Garg G, Hales Charles A. Chemistry and Biology of Heparin and Heparan Sulfate. Kidlington, Oxford, UK: Elsevier; 2005. [Google Scholar]
- 20.Inamori K, et al. Dystroglycan function requires xylosyl- and glucuronyltransferase activities of LARGE. Science. 2012;335:93. doi: 10.1126/science.1214115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Foulquier F, et al. TMEM165 deficiency causes a congenital disorder of glycosylation. Am. J. Hum. Genet. 2012;91:15. doi: 10.1016/j.ajhg.2012.05.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Sanjana NE, et al. A transcription activator-like effector toolbox for genome engineering. Nat. Protoc. 2012;7:171. doi: 10.1038/nprot.2011.431. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Vogel P, et al. Congenital hydrocephalus in genetically engineered mice. Vet. Pathol. 2012;49:166. doi: 10.1177/0300985811415708. [DOI] [PubMed] [Google Scholar]
- 24.Vuillaumier-Barrot S, et al. Identification of mutations in TMEM5 and ISPD as a cause of severe cobblestone lissencephaly. Am. J. Hum. Genet. 2012;91:1135. doi: 10.1016/j.ajhg.2012.10.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Sabeti PC, et al. International HapMap Consortium, Genome-wide detection and characterization of positive selection in human populations. Nature. 2007;449:913. doi: 10.1038/nature06250. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Whelan SP, Ball LA, Barr JN, Wertz GT. Efficient recovery of infectious vesicular stomatitis virus entirely from cDNA clones. Proc. Natl. Acad. Sci. U.S.A. 1995;92:8388. doi: 10.1073/pnas.92.18.8388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Wong AC, Sandesara RG, Mulherkar N, Whelan SP, Chandran K. A forward genetic strategy reveals destabilizing mutations in the Ebolavirus glycoprotein that alter its protease dependence during cell entry. J. Virol. 2010;84:163. doi: 10.1128/JVI.01832-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Lefeber DJ, et al. Autosomal recessive dilated cardiomyopathy due to DOLK mutations results from abnormal dystroglycan O-mannosylation. PLoS Genet. 2011;7:e1002427. doi: 10.1371/journal.pgen.1002427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Banerjee A, Sammarco MC, Ditch S, Wang J, Grabczyk E. A novel tandem reporter quantifies RNA polymerase II termination in mammalian cells. PLoS ONE. 2009;4:e6193. doi: 10.1371/journal.pone.0006193. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Collier LS, Carlson CM, Ravimohan S, Dupuy AJ, Largaespada DA. Cancer gene discovery in solid tumours using transposon-based somatic mutagenesis in the mouse. Nature. 2005;436:272. doi: 10.1038/nature03681. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.