Abstract
Investigation of the possible role of Campylobacter concisus (C. concisus) in inflammatory bowel disease (IBD) is an emerging research area. Despite the association found between C. concisus and IBD, it has been difficult to explain how C. concisus, a bacterium that is commonly present in the human oral cavity, may contribute to the development of enteric diseases. The evidence presented in this review shows that some C. concisus strains in the oral cavity acquired zonula occludens toxin (zot) gene from a virus (prophage) and that C. concisus Zot shares conserved motifs with both Vibrio cholerae Zot receptor binding domain and human zonulin receptor binding domain. Both Vibrio cholerae Zot and human zonulin are known to increase intestinal permeability by affecting the tight junctions. Increased intestinal permeability is a feature of IBD. Based on these data, we propose that a primary barrier function defect caused by C. concisus Zot is a mechanism by which zot-positive C. concisus strains may trigger the onset and relapse of IBD.
Keywords: Campylobacter concisus, Inflammatory bowel disease, Zonula occludens toxin, Tight junctions, Intestinal permeability
Core tip: Campylobacter concisus (C. concisus) is an oral bacterium that was previously shown to be associated with inflammatory bowel disease (IBD). Evidence presented in this review shows that some strains of C. concisus acquired zonula occludens toxin (zot) gene from a virus (prophage), suggesting that a primary barrier function defect caused by C. concisus Zot is a mechanism by which zot-positive C. concisus strains may trigger the onset and relapse of IBD.
INTRODUCTION
Inflammatory bowel disease (IBD) is a chronic inflammatory condition of the gastrointestinal tract[1]. The two major clinical forms of IBD are Crohn’s disease (CD) and ulcerative colitis (UC). The etiology of IBD is not fully understood. Multiple contributors including genetic factors, environmental factors and intestinal microbiota have been suggested to play a role in the development of IBD[1]. The pathogenesis of IBD is thought to result from a dysregulated response of the intestinal mucosal immune system to luminal commensal microbes[1].
The highest incidence of IBD, both CD and UC, is in young adults[2]. This implies that in most of the patients with IBD, the intestinal mucosal immune system has maintained a non-hostile relationship with the intestinal commensal microbes for decades prior to the onset of the disease. Given this, the uncontrolled attack of the intestinal mucosal immune system to luminal commensal bacteria would have been initiated by a trigger. Such a trigger may not necessarily be a dominant intestinal bacterial species or a long-term intestinal resident microbe. Evidence presented in this review suggests that zonula occludens toxin gene (zot)-positive Campylobacter concisus (C. concisus) strains, a bacterium colonizing the human oral cavity, may be a trigger of IBD.
C. concisus
C. concisus is a Gram negative bacterium that requires microaerobic or anaerobic conditions enriched with H2 for growth[3]. Cells of C. concisus are curved, with a size of (0.5 - 1) × (2 - 6) μm. C. concisus is motile, driven by a single polarized flagellum (Figure 1).
Figure 1.
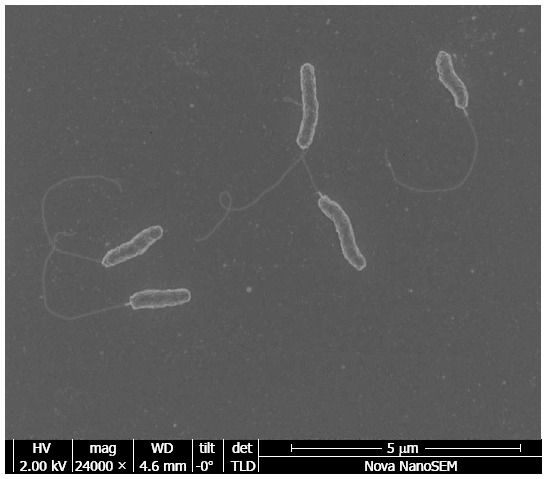
Electron microscopic image of Campylobacter concisus.
HUMANS ARE THE NATURAL HOST OF C. CONCISUS AND THE ORAL CAVITY IS THE PRIMARY COLONIZATION SITE
Tanner et al[4] first reported the isolation of C. concisus from patients with gingivitis in 1981. Zhang et al[5] examined the presence of C. concisus in saliva samples obtained from healthy individuals of different age groups and found that C. concisus is commonly present in the human oral cavity. In that study, C. concisus was detected in 97% (57/59) of saliva samples collected from healthy individuals aged 3-60 years old by polymerase chain reaction (PCR) targeting 16S rRNA gene and cultured from 75% (44/59) of these saliva samples using a filtration method. A study from Petersen et al[6] also detected a high prevalence of C. concisus in the human oral cavity: in this study C. concisus was detected in 100% of saliva samples (11/11) collected from healthy individuals by a PCR targeting 16S rRNA gene. Despite its high prevalence in the human oral cavity, C. concisus is not a dominant oral bacterial species[7].
In comparison to the high isolation of C. concisus from saliva samples, the isolation rates of C. concisus from fecal samples collected from healthy individuals were much lower. Using the filtration method, Engberg et al[8] isolated C. concisus from 2.8% (3/107) of fecal samples and Nielsen et al[9] did not isolate C. concisus from any of 108 fecal samples collected from healthy individuals. The low isolation rates of C. concisus from fecal samples suggest that the human intestinal tract of healthy individuals is a less optimal site for C. concisus colonization compared to the oral cavity.
To date, C. concisus has not been detected in any healthy animals. In a study examining the presence of Campylobacter species in fecal samples collected from 70 healthy pet dogs using a quantitative PCR targeting the 60 kDa chaperonin gene, Chaban et al[10] detected seven different campylobacter species but not C. concisus. Various campylobacter species have been detected in fecal samples collected from animals or birds, but C. concisus was not detected[3,10,11]. Lynch et al[12] isolated C. concisus from 10% (18/185) of chicken meat and 3% of beef meat (6/186) samples. However, whether chicken and cattle are natural hosts of C. concisus cannot be determined from these data.
C. concisus has been detected in some animals with gastrointestinal disorders. Petersen et al[6] detected C. concisus in 12.5% (1/8) of saliva samples from pet cats with dental diseases by PCR targeting the 16S rRNA gene[6]. In addition, Chaban et al[10] detected C. concisus in fecal samples of 9% of dogs with diarrhea (6/65).
The collective data suggest that humans are the natural host of C. concisus, with the human oral cavity being the primary colonization site (Table 1).
Table 1.
Detection of Campylobacter concisus in samples obtained from healthy individuals
| Samples | Cultivation of C. concisus | Detection of C. concisus DNA by PCR |
| Human saliva | 75% | 97%-100% |
| Human feces | 0%-2.8% | 33% |
| Human intestinal biopsies | 0% | 2%-38% |
Data shown in this table were obtained from references 5-10 and 17-21. To date, no studies have detected Campylobacter concisus (C. concisus) in samples obtained from healthy animals. The collective data suggest that humans are the natural host of C. concisus and the oral cavity is the primary colonization site. PCR: Polymerase chain reaction.
DIVERSITY OF C. CONCISUS STRAINS COLONIZING THE HUMAN ORAL CAVITY
C. concisus strains colonizing the human oral cavity are greatly diverse. On examination of oral C. concisus strains isolated from individual patients with IBD and healthy controls, it was found that C. concisus strains isolated from each individual had unique protein patterns on sodium dodecyl sulphate polyacrylamide gel electrophoresis[5,13,14]. Furthermore, some individuals were colonized with multiple C. concisus strains in the oral cavity, with as many as three different C. concisus strains having been isolated from individual patients with IBD or healthy controls[5,13,14]. A significantly higher number of patients with active IBD were colonized with multiple C. concisus strains in the oral cavity compared to healthy controls14].
COLONIZATION OF ORAL C. CONCISUS STRAINS IN THE INTESTINAL TRACT
It is estimated that 1-1.5 L of saliva is produced daily in humans, most of which is swallowed[15]. Thus, the human oral cavity is a constantly available source, transporting C. concisus from the oral cavity along with saliva to lower parts of the gastrointestinal tract. However, as mentioned previously, both the detection of C. concisus by PCR and the cultivation of C. concisus from fecal samples were much lower than that from saliva samples, indicating that C. concisus routinely transported from the oral cavity to the intestines does not commonly establish colonization there.
Nevertheless, intestinal colonization of oral C. concisus strains does occur in some individuals. For example, in a study comparing the housekeeping genes of C. concisus strains isolated from intestinal biopsies of patients with IBD and controls, Ismail et al[13] found that a C. concisus strain isolated from intestinal biopsies of a patient with UC had housekeeping genes identical to that of an oral C. concisus strain isolated from the same patient, providing evidence that the oral C. concisus strains are in fact able to colonize the human intestinal tract (Table 2).
Table 2.
Genetic relatedness of enteric and oral Campylobacter concisus strains
| Enteric C. concisus strains from patients with IBD | Genetically related C. concisus strains |
| A strain isolated from intestinal biopsies of patient No. 1 | An oral strain from patient No. 6 |
| A strain isolated from intestinal biopsies of patient No. 3 | An oral strain from patient No. 3 (identical) |
| A strain isolated from luminal fluid of patient No. 3 | An oral strain from healthy No. 1 |
Data shown in this table were obtained from reference 13. The genetic relatedness was assessed based on the sequences of six housekeeping genes. These data provide evidence showing that Campylobacter concisus (C. concisus) strains colonizing the intestinal tract of patients with IBD originating from either the patient’s own oral C. concisus strains or oral C. concisus strains from other individuals. IBD: Inflammatory bowel disease.
In addition to an individual’s own oral C. concisus, C. concisus detected in the intestinal tract may also come from a different source, most likely through materials contaminated with saliva from others. For example, in the above patient with UC, while the C. concisus strain isolated from intestinal biopsies had housekeeping genes identical to that of the patient’s own oral C. concisus strain, the C. concisus strain isolated from the luminal fluid of this patient had a genetic relationship closely related to an oral C. concisus strain from a healthy control, rather than to the patient’s own oral C. concisus (Table 2)[13].
The human intestinal environment is not optimal for C. concisus colonization in general, as suggested by the low isolation rate of C. concisus from fecal samples of healthy individuals (Table 1). Given this, C. concisus intestinal colonization is most likely a short-term event in most individuals. However, with the human oral cavity as a constantly available source of C. concisus, repeated intestinal colonization of C. concisus may occur.
Whether the characteristics of a given oral C. concisus strain or the intestinal environment of an individual, or both, plays the major role in determining whether or not intestinal colonization of oral C. concisus strains will occur is currently unknown. A previous study by Haag et al[16] showed that intestinal microbiota shifts towards elevated commensal Escherichia coli loads abrogated colonization resistance against Campylobacter jejuni in mice. Currently, it is not clear whether the dysbiosis associated with IBD plays a role in C. concisus intestinal colonization.
PREVALENCE OF C. CONCISUS IN THE INTESTINAL TRACT OF PATIENTS WITH IBD AND HEALTHY CONTROLS
A number of studies have examined the prevalence of C. concisus in the intestinal tract of patients with IBD using PCR methods. Most of these studies detected a significantly higher prevalence of C. concisus DNA in patients with IBD and controls[17-21]. The reported detection rates of C. concisus by PCR in enteric samples (biopsies and fecal samples) were 33%-69% in patients with IBD and 2%-38% in controls[17-21]. Analysis of the data in these studies revealed a number of interesting findings.
Firstly, different PCR strategies affect the detection of C. concisus in enteric samples. This was best seen in the study conducted by Man et al[18], who compared the prevalence of C. concisus in fecal samples collected from 54 children with CD, 33 healthy controls and 27 non-IBD controls using two different PCR methods. The first PCR method employed campylobacter genus specific PCR (primers C412F/C1288R) and sequencing the PCR products to determine campylobacter species. The second PCR method was a nested PCR, using campylobacter genus specific PCR (primers C412F/C1288R) followed by C. concisus specific PCR (primers Concisus F/Concisus R). These two PCR methods yielded very different results in detection of C. concisus in the same samples. The prevalence of C. concisus in children with CD, healthy controls and non-IBD controls detected by C. concisus genus specific PCR was 19% (10/54), 12% (4/33) and zero (0/27) respectively. The nested PCR greatly increased the detection of C. concisus in the same cohort of samples, with the prevalence of C. concisus being 65% (35/54) in children with CD, 33% (11/33) in healthy controls and 37% (10/27) in non-IBD controls. The nested PCR, but not the genus specific PCR, detected a significantly higher prevalence of C. concisus in children with CD as compared to healthy controls[18] (Table 3). Indeed, in studies revealing a significant difference in intestinal prevalence of C. concisus between patients with IBD and controls, nested PCR was used to examine all of the samples or part of the samples[17-20].
Table 3.
Detection rates of Campylobacter concisus in fecal samples by two polymerase chain reaction methods
| Campylobacter genus PCR | Nested PCR | |
| CD (n = 54) | 19% | 65% |
| Healthy controls (n = 33) | 12% | 33% |
| Non-IBD controls (n = 27) | 0 | 37% |
An example showing that different polymerase chain reaction (PCR) methods affect the detection of Campylobacter concisus (C. concisus) in intestinal samples. Data included in this table were from reference 18. Primers C412F and C1288R were used in campylobacter genus PCR. In nested PCR, PCR products amplified by primers C412F and C1288R were amplified again using C. concisus specific primers Concisus F and Concisus R. The nest PCR detected a significantly higher intestinal prevalence of C. concisus in patients with Crohn’s disease (CD) compared to both healthy controls and non-inflammatory bowel disease (IBD) controls. The genus PCR detected a significantly higher intestinal prevalence of C. concisus in patients with CD compared to non-IBD controls, but not healthy controls.
Secondly, collection of multiple intestinal biopsies increases the detection of intestinal prevalence of C. concisus. A study by Mahendran et al[20] showed that in comparison to the collection of one biopsy, the collection of four biopsies from each individual greatly increased the detection of C. concisus (Figure 2).
Figure 2.
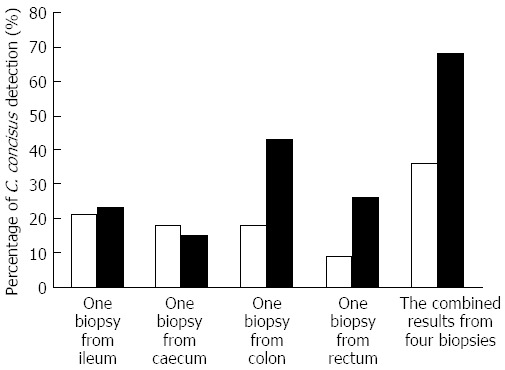
An example showing that collection of multiple intestinal biopsies increases the detection of intestinal prevalence of Campylobacter concisus. Data included in Figure 2 were from reference 20. In both patients with inflammatory bowel disease (IBD) (right column) and healthy controls (left column), the detection of intestinal prevalence of Campylobacter concisus (C. concisus) was greatly increased using four biopsies as compared to using one biopsy. In healthy controls, C. concisus detection rates in a single biopsy collected from ileum, caecum, colon and rectum were 21%, 18%, 18% and 9% respectively. If results from the four biopsies were used in determining the intestinal prevalence of C. concisus, the intestinal prevalence of C. concisus in healthy controls was 36%. Similarly, in patients with IBD, C. concisus detection rates in a single biopsy collected from ileum, caecum, colon and rectum were 23%, 15%, 43% and 26% respectively, and the intestinal prevalence of C. concisus in patients with IBD was 68% when four biopsies were taken into consideration.
Thirdly, despite the increased prevalence of C. concisus detected by PCR in the intestinal tract of patients with IBD, the isolation rates of C. concisus from intestinal biopsies of patients with IBD were low (3%-7.7%)[17,20,21]. This suggests that C. concisus detected in most of the enteric samples were at a low number or in a nonculturable state.
ADVERSE EFFECTS OF C. CONCISUS ON INTESTINAL EPITHELIAL CELLS
A number of adverse effects of C. concisus on intestinal epithelial cells have been described. Using in vitro cell culture models (Caco2 cells or HT-29 cells), epithelial adhesion and invasion, damage of the barrier function and up-regulation of Toll-like receptors-4 expression by C. concisus have been reported. Different strains showed varying degrees of ability to induce such adverse effects[13,22-25]. The underlying molecular mechanisms responsible for these effects have not yet been investigated.
C. CONCISUS ZOT GENE AND IBD
Mahendran et al[14] examined the prevalence of zot gene in 56 oral C. concisus strains isolated from saliva of 19 patients with IBD and 20 healthy controls. This study showed that 30% (17/56) of the oral C. concisus strains carried the zot gene. The zot-positive C. concisus strain was present in 55% (6/11) of patients with active IBD and 40% (8/20) of healthy controls. Some IBD patients with active disease 18% (2/11) were colonized with multiple zot-positive C. concisus strains in the oral cavity. Interestingly, polymorphic forms of the C. concisus zot gene resulting in the substitution of valine at amino acid position 270 were found to be associated with active IBD.
The zot gene was first discovered in Vibrio cholerae where it is carried by a filamentous prophage[26,27]. The zot gene in V. cholerae is required for phage production; a V. cholerae strain with zot gene mutation did not produce phage particles into the culture supernatant[28]. The V. cholerae Zot toxin was shown to increase intestinal permeability and to be associated with mild to moderate diarrhea[26,29,30].
Previous studies have found a human intestinal Zot analogue, namely zonulin, which is a physiological regulator that increases the intestinal permeability[31,32].
C. CONCISUS ZOT GENE IS A COMPONENT OF A CHROMOSOMALLY INTEGRATED PUTATIVE PROPHAGE
Stothard et al[33] established a visual database of bacterial chromosome maps in which C. concisus strain 13826 (Accession No. CP000792. 1) was included. Stothard et al[33] identified a region from nucleotide position 1576683 to 1615449 (38767 bp) in the genome of C. concisus strain 13826 as an “incomplete prophage”. In this region, 39 genes with open reading frames were identified, with four of these genes encoding integrases and 10 genes encoding phage-like proteins[33].
The genetic structures of this “incomplete prophage” region are shown in Figure 3A and the proteins encoded by genes in this region are shown in Table 4. We compared the genes within this region using publicly available softwares[34,35]. A number of genes that have identical nucleotide sequences, including CCC13826_1099, CCC13826_0020, CCC13826_1102, CCC13826_0164, CCC13826_0185, CCC13826_2082 and CCC13826_2077 were annotated with identical protein names.
Figure 3.
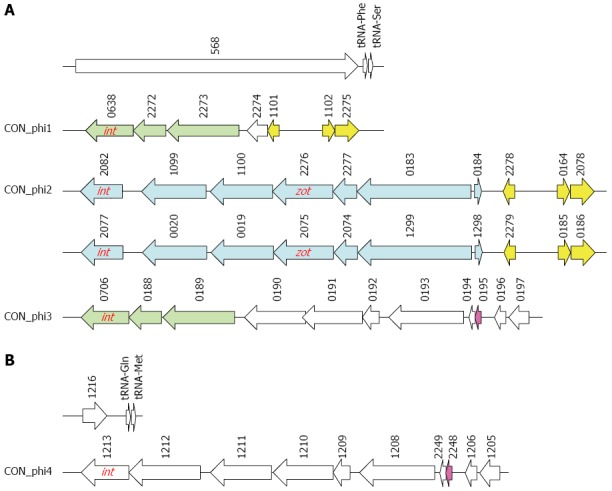
Genetic structures of putative prophage identified in Campylobacter concisus strain 13826. A: Multiple prophages at nucleotide position between 1576686 and 1614075; B: One putative prophage at nucleotide position between 939158 and 946901. Identical genes were indicated with the same color. int: A gene encodes phage integrase; zot: Zonula occludens toxin gene. The numbers are the locus tags.
Table 4.
Proteins encoded by putative prophages in Campylobacter concisus strain 13826
| Nucleotide position | Locus tag | Encoded proteins | Size (aa) |
| 1576686..1581986 | CCC13826_0568 | Hypothetical protein | 1766 |
| 1582376..1583269 | CCC13826_0638 | Phage integrase | 297 |
| 1583269..1583880 | CCC13826_2272 | Glutathionylspermidine synthase family | 203 |
| 1583895..1585235 | CCC13826_2273 | Bacteriophage replication gene A protein | 446 |
| 1585467..1585853 | CCC13826_2274 | Protein yitk | 128 |
| 1585894..1586115 | CCC13826_1101 | Phosphonate uptake transporter | 73 |
| 1586807..1587043 | CCC13826_1102 | Sensory box protein | 78 |
| 1587044..1587496 | CCC13826_2275 | Hypothetical protein | 150 |
| 1587598..1588506 | CCC13826_2082 | Phage integrase | 302 |
| 1588755..1589966 | CCC13826_1099 | Putative phage replication protein A | 403 |
| 1590015..1591184 | CCC13826_1100 | ATP binding protein (ABC) transporter | 389 |
| 1591228..1592352 | CCC13826_2276 | Zonula occludens toxin (Zot) family protein | 374 |
| 1592354..1592800 | CCC13826_2277 | Hypothetical protein | 148 |
| 1592797..1594923 | CCC13826_0183 | Hypothetical protein | 708 |
| 1594955..1595095 | CCC13826_0184 | Hypothetical protein | 46 |
| 1595477..1595698 | CCC13826_2278 | Phosphonate uptake transporter | 73 |
| 1596412..1596648 | CCC13826_0164 | Sensory box protein | 78 |
| 1596649..1597101 | CCC13826_2078 | Hypothetical protein | 150 |
| 1597203..1598111 | CCC13826_2077 | Phage integrase | 302 |
| 1598360..1599571 | CCC13826_0020 | Putative phage replication protein A | 403 |
| 1599620..1600789 | CCC13826_0019 | ABC transporter | 389 |
| 1600833..1601957 | CCC13826_2075 | Zot family protein | 374 |
| 1601959..1602405 | CCC13826_2074 | Hypothetical protein | 148 |
| 1602402..1604528 | CCC13826_1299 | Hypothetical protein | 708 |
| 1604560..1604700 | CCC13826_1298 | Hypothetical protein | 46 |
| 1605082..1605303 | CCC13826_2279 | Phosphonate uptake transporter | 73 |
| 1606017..1606253 | CCC13826_0185 | Sensory box protein | 78 |
| 1606254..1606706 | CCC13826_0186 | Hypothetical protein | 150 |
| 1606808..1607701 | CCC13826_0706 | Phage integrase | 297 |
| 1607701..1608312 | CCC13826_0188 | Glutathionylspermidine synthase family | 203 |
| 1608327..1609667 | CCC13826_0189 | Bacteriophage replication gene A protein | 446 |
| 1609899..1611041 | CCC13826_0190 | Type II and III secretion system protein | 380 |
| 1610992..1612116 | CCC13826_0191 | Hypothetical protein | 374 |
| 1612118..1612438 | CCC13826_0192 | Hypothetical protein | 106 |
| 1612537..1613946 | CCC13826_0193 | Hypothetical protein | 469 |
| 1613956..1614075 | CCC13826_0194 | Alkyl hydroperoxide reductase | 39 |
| 1614090..1614209 | CCC13826_0195 | Hypothetical protein | 39 |
| 1614489..1614707 | CCC13826_0196 | Hypothetical protein | 72 |
| 1614801..1615178 | CCC13826_0197 | Hypothetical protein | 125 |
aa: Amino acids.
Interestingly, the region that was considered as an “incomplete prophage” by Stothard et al[33] turned out to be four putative prophages, each beginning with a phage integrase (Figure 3 and Table 4). The first prophage had a genome size of 5.2 kb, which contained seven protein-encoding genes. The second prophage and third prophage were identical, each having a genome size of 9.6 kb consisting of 10 protein-encoding genes. The fourth prophage contained 11 protein-encoding genes with a genome size of 8.6 kb. We named the first prophage CON_phi1, the second prophage and third prophage CON_phi2 and the fourth prophage CON_phi3. The zot gene is a component of CON_phi2. Comparison of the proteins encoded by genes in CON_phi2 with that encoded by genes in CTX phage, the phage that carries the zot gene in V. cholera, did not show high similarities except for the Zot protein (data not shown), suggesting the CON_phi2 is a previously uncharacterised prophage.
A study by Kaakoush et al[36] found that two hypothetical proteins encoded by CCC13826_0191 and CCC13826_1210 have 47% and 46% similarity respectively to C. concisus Zot. Here we found that CCC13826_0191 is a gene of CON_phi3 and CCC13826_1210 is a gene of an additional putative prophage, which was named CON_phi4. A number of genes in CON_phi3 and CON_phi4 had high similarities, however, CON_phi4 did not contain the gene that corresponding to CCC13826_0188 in CON_phi3 (Table 5).
Table 5.
The similarity of proteins in CON_phi3 and CON_phi4
| CON_phi3 | CON_phi4 | Similarity1 |
| CCC13826_0706 | CCC13826_1213 | 23.23% |
| CCC13826_0188 | ||
| CCC13826_0189 | CCC13826_1212 | 93.72% |
| CCC13826_0190 | CCC13826_1211 | 98.95% |
| CCC13826_0191 | CCC13826_1210 | 92.25% |
| CCC13826_0192 | CCC13826_1209 | 92.03% |
| CCC13826_0193 | CCC13826_1208 | 58.90% |
| CCC13826_0194 | CCC13826_2249 | 100% |
| CCC13826_0195 | CCC13826_2248 | 97.44% |
| CCC13826_0196 | CCC13826_1206 | 61.11% |
| CCC13826_0197 | CCC13826_1205 | 95.24% |
Similarity is the percentage of identical amino acids.
IDENTIFICATION OF CONSERVED MOTIFS SHARED BY C. CONCISUS ZOT AND ZONULIN/ZOT RECEPTOR BINDING DOMAINS
Kaakoush et al[36] compared Zot sequences in C. concisus strain 13826 and V. cholerae strain 86015 and reported that the biological active domain (FCIGRL) previously found in V. cholerae Zot was not found in C. concisus Zot. In this review, we compared the sequence of C. concisus Zot with human zonulin receptor binding domain and V. cholerae Zot receptor binding domain previously reported[32,37]. Interestingly, we found that C. concisus Zot shares conserved motifs with both the human zonulin receptor binding domain and the V. cholerae Zot receptor binding domain (Table 6). These data suggest that C. concisus Zot may increase intestinal permeability using a mechanism that is similar to the human zonulin and V. cholerae Zot, affecting the tight junctions through proteinase activated receptor 2 activation[37,38]. The motif “GRFLSYHG” is located at amino acid position 123-130 in C. concisus Zot, which was found in all oral zot-positive C. concisus strains that we previously isolated as well as in the C. concisus strain 13826[14]. The polymorphisms of C. concisus zot gene that Mahendran et al[14] previously detected were not in the receptor binding domain, suggesting that these polymorphisms may impact on the function of C. concisus Zot, if there is any, using a different mechanism rather than affecting the binding of C. concicsus Zot to the receptor.
Table 6.
Motifs shared by Campylobacter concisus zonula occludens toxin and zonulin/zonula occludens toxin receptor binding domains
| C. concisus Zot1 | GRF LSYHG |
| Human adult intestine zonulin | GGXL |
| Human fetal intestine zonulin | GGVLVQPG |
| C. concisus Zot2 | GRF LSYHG |
| V. cholerae Zot | FCIGRLCVQDG |
Campylobacter concisus zonula occludens toxin (Zot) and zonulin binding domain shares a motif (bold letters): Non-polar G, variable, non-polar, non-polar L, variable, polar, variable and non-polar G;
C. concisus Zot and Vibrio cholerae Zot shares a motif (bold letters): non-polar G, basic polar R, non-polar, non-polar, variable, polar, variable and non-polar G. Comparison of C. concisus Zot and zonulin/Zot receptor was performed in this review. The amino acid sequences of human intestinal zonulin and V. cholerae Zot were obtained from reference 32.
INCREASED INTESTINAL PERMEABILITY IN PATIENTS WITH IBD
Increased intestinal permeability is a feature of both CD and UC[39-43]. While epithelial cell death and proinflammatory cytokines may damage the intestinal epithelial barrier during active disease, evidence shows that increased intestinal permeability may precede the initial onset or relapse of IBD. An early study from Hollander et al[39] reported that increased intestinal permeability was detected not only in patients with CD but also in their healthy relatives. A family history of IBD is a known risk factor for IBD[44]. Irvine et al[41] reported that an individual with a family history of CD had elevated intestinal permeability eight years prior to the onset of clinical symptoms and diagnosis of CD. Wyatt et al[43] measured the intestinal permeability in patients with quiescent CD and found that those with increased intestinal permeability were at a significantly higher risk of clinical relapse. These data suggest that increased intestinal permeability occurred prior to the onset and relapse of the disease may be a possible aetiological factor of IBD.
C. CONCISUS ZOT: A POTENTIAL TRIGGER OF IBD THROUGH CAUSING PRIMARY BARRIER DEFECT
The human zonulin and V. cholerae Zot toxin are known to increase intestinal permeability through affecting the tight junctions[31,32,37]. In this review, we found that C. concisus Zot has conserved motifs shared by the zonulin/Zot binding receptor domains. Given this, it is very likely that C. concisus Zot also affects the tight junctions.
Based on the information obtained from previous publications and the analysis that we have performed in this review, we propose a mechanism by which C. concisus, an oral bacterium, may trigger the onset or relapse of IBD: that some oral C. concisus strains acquire zot gene from a virus (prophage). With the human oral cavity as the reservoir of C. concisus, repeated intestinal colonization of C. concisus and release of C. concisus Zot due to prophage induction may occur, which is likely to result in a prolonged primary epithelial barrier defect and translocation of macromolecule such as luminal microbes and their products. In genetically susceptible individuals, this may trigger the development of IBD.
Damage to the intestinal epithelial tight junctions may also lead to the development of diarrhea. Indeed, in addition to its association with IBD, C. concisus has been frequently isolated from non-IBD-related diarrheal stool samples[9,45,46].
If some oral C. concisus strains are indeed involved in the development of human IBD, the question as to why the lesions of IBD occur more often in the intestinal tract rather than in the oral cavity arises. One explanation is that the virulence factors that are associated with IBD are more often expressed in the intestinal tract rather than in the oral cavity. For example, the expression of C. concisus Zot may require induction of prophage from the C. concisus genome. As prophage induction usually occurs when bacterial cells are under stressful conditions[47], the fact that C. concisus uses the human oral cavity as its primary colonization site suggests that the oral cavity is not a stressful site for C. concisus. However, as the C. concisus travels to the more hostile lower parts of the gastrointestinal tract, the stressful environment may trigger the induction of C. concisus prophage.
Another possible factor that may reduce the pathogenic effect of C. concisus Zot in the oral cavity is that the epithelium in the oral cavity is a stratified squamous epithelium, either keratinized or non-keratinized[48]. In contrast, the intestinal epithelium is a simple columnar epithelium[48]. The impact on permeability caused by Zot, even it is expressed in the oral cavity, in multiple layers of squamous epithelium may not be as evident as that in the single layered columnar epithelium.
C. CONCISUS ZOT: A POTENTIAL ENVIRONMENTAL FACTOR CONTRIBUTING TO THE INCREASED RISK OF IBD IN INDIVIDUALS WITH A FAMILY HISTORY OF IBD
A family history of IBD is a risk factor for developing IBD[44]. In addition to genetic factors, environmental factors have been shown to be involved in the increased incidence of IBD in members with a family history of this disease[49,50]. We suggest that C. concisus Zot is one such factor. This suggestion is based on the findings that the higher numbers of the relatives of patients with IBD have increased intestinal permeability and that some oral C. concisus strains carry the zot gene that encodes a toxin known to promote this[14,39,41]. This hypothesis remains to be further assessed by examining the correlation between colonization of zot-positive C. concisus strains and the increased intestinal permeability in family members of patients with IBD.
CONCLUSION
The evidence presented in this review shows that some C. concisus strains colonizing the human oral cavity acquired zot gene from a virus (prophage). We are currently examining the biologic activities of C. concisus Zot, the expression of Zot in zot-positive C. concisus strains isolated from patients with IBD and controls as well as the presence of C. concisus Zot in the oral cavity and intestinal tract of patients with IBD and controls, which will provide further information in understanding the role of C. concisus Zot in IBD and other human diseases.
ACKNOWLEDGMENTS
The authors would like to thank Vikneswari Mahendran and Jenny Norman for providing the scanning electron microscopic picture of C. concisus.
Footnotes
P- Reviewers: Actis GC, Azuma YT, Capasso R S- Editor: Gou SX L- Editor: A E- Editor: Wang CH
References
- 1.Khor B, Gardet A, Xavier RJ. Genetics and pathogenesis of inflammatory bowel disease. Nature. 2011;474:307–317. doi: 10.1038/nature10209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Bernstein CN, Wajda A, Svenson LW, MacKenzie A, Koehoorn M, Jackson M, Fedorak R, Israel D, Blanchard JF. The epidemiology of inflammatory bowel disease in Canada: a population-based study. Am J Gastroenterol. 2006;101:1559–1568. doi: 10.1111/j.1572-0241.2006.00603.x. [DOI] [PubMed] [Google Scholar]
- 3.Vandamme P, Dewhirst FE, Paster BJ, On SLW. Genus I. Campylobacter. In: Garrity GM, Brenner DJ, Krieg NR, Staley JT, editors. Bergey’s Manual of Syst Bacteriol. 2 ed. New York: Springer; 2005. pp. 1147–1160. [Google Scholar]
- 4.Tanner ACR, Badger S, Lai CH, Listgarten MA, Visconti RA, Socransky SS. Wolinella gen-nov, Wolinella-succinogenes (Vibrio-succinogenes-wolin et-al) comb-nov, and description of Bacteroides-gracilis sp-nov, Wolinella-recta sp-nov, Campylobacter-concisus sp-nov, and Eikenella-corrodens from humans with periodontal-disease. Int J Syst Bacteriol. 1981;31:432–445. [Google Scholar]
- 5.Zhang L, Budiman V, Day AS, Mitchell H, Lemberg DA, Riordan SM, Grimm M, Leach ST, Ismail Y. Isolation and detection of Campylobacter concisus from saliva of healthy individuals and patients with inflammatory bowel disease. J Clin Microbiol. 2010;48:2965–2967. doi: 10.1128/JCM.02391-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Petersen RF, Harrington CS, Kortegaard HE, On SL. A PCR-DGGE method for detection and identification of Campylobacter, Helicobacter, Arcobacter and related Epsilobacteria and its application to saliva samples from humans and domestic pets. J Appl Microbiol. 2007;103:2601–2615. doi: 10.1111/j.1365-2672.2007.03515.x. [DOI] [PubMed] [Google Scholar]
- 7.Aas JA, Paster BJ, Stokes LN, Olsen I, Dewhirst FE. Defining the normal bacterial flora of the oral cavity. J Clin Microbiol. 2005;43:5721–5732. doi: 10.1128/JCM.43.11.5721-5732.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Engberg J, On SL, Harrington CS, Gerner-Smidt P. Prevalence of Campylobacter, Arcobacter, Helicobacter, and Sutterella spp. in human fecal samples as estimated by a reevaluation of isolation methods for Campylobacters. J Clin Microbiol. 2000;38:286–291. doi: 10.1128/jcm.38.1.286-291.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Nielsen HL, Ejlertsen T, Engberg J, Nielsen H. High incidence of Campylobacter concisus in gastroenteritis in North Jutland, Denmark: a population-based study. Clin Microbiol Infect. 2013;19:445–450. doi: 10.1111/j.1469-0691.2012.03852.x. [DOI] [PubMed] [Google Scholar]
- 10.Chaban B, Ngeleka M, Hill JE. Detection and quantification of 14 Campylobacter species in pet dogs reveals an increase in species richness in feces of diarrheic animals. BMC Microbiol. 2010;10:73. doi: 10.1186/1471-2180-10-73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Moore JE, Corcoran D, Dooley JS, Fanning S, Lucey B, Matsuda M, McDowell DA, Mégraud F, Millar BC, O’Mahony R, et al. Campylobacter. Vet Res. 2005;36:351–382. doi: 10.1051/vetres:2005012. [DOI] [PubMed] [Google Scholar]
- 12.Lynch OA, Cagney C, McDowell DA, Duffy G. Occurrence of fastidious Campylobacter spp. in fresh meat and poultry using an adapted cultural protocol. Int J Food Microbiol. 2011;150:171–177. doi: 10.1016/j.ijfoodmicro.2011.07.037. [DOI] [PubMed] [Google Scholar]
- 13.Ismail Y, Mahendran V, Octavia S, Day AS, Riordan SM, Grimm MC, Lan R, Lemberg D, Tran TA, Zhang L. Investigation of the enteric pathogenic potential of oral Campylobacter concisus strains isolated from patients with inflammatory bowel disease. PLoS One. 2012;7:e38217. doi: 10.1371/journal.pone.0038217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Mahendran V, Tan YS, Riordan SM, Grimm MC, Day AS, Lemberg DA, Octavia S, Lan R, Zhang L. The Prevalence and Polymorphisms of Zonula Occluden Toxin Gene in Multiple Campylobacter concisus Strains Isolated from Saliva of Patients with Inflammatory Bowel Disease and Controls. PLoS One. 2013;8:e75525. doi: 10.1371/journal.pone.0075525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Humphrey SP, Williamson RT. A review of saliva: normal composition, flow, and function. J Prosthet Dent. 2001;85:162–169. doi: 10.1067/mpr.2001.113778. [DOI] [PubMed] [Google Scholar]
- 16.Haag LM, Fischer A, Otto B, Plickert R, Kühl AA, Göbel UB, Bereswill S, Heimesaat MM. Intestinal microbiota shifts towards elevated commensal Escherichia coli loads abrogate colonization resistance against Campylobacter jejuni in mice. PLoS One. 2012;7:e35988. doi: 10.1371/journal.pone.0035988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Zhang L, Man SM, Day AS, Leach ST, Lemberg DA, Dutt S, Stormon M, Otley A, O’Loughlin EV, Magoffin A, et al. Detection and isolation of Campylobacter species other than C. jejuni from children with Crohn’s disease. J Clin Microbiol. 2009;47:453–455. doi: 10.1128/JCM.01949-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Man SM, Zhang L, Day AS, Leach ST, Lemberg DA, Mitchell H. Campylobacter concisus and other Campylobacter species in children with newly diagnosed Crohn’s disease. Inflamm Bowel Dis. 2010;16:1008–1016. doi: 10.1002/ibd.21157. [DOI] [PubMed] [Google Scholar]
- 19.Mukhopadhya I, Thomson JM, Hansen R, Berry SH, El-Omar EM, Hold GL. Detection of Campylobacter concisus and other Campylobacter species in colonic biopsies from adults with ulcerative colitis. PLoS One. 2011;6:e21490. doi: 10.1371/journal.pone.0021490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Mahendran V, Riordan SM, Grimm MC, Tran TA, Major J, Kaakoush NO, Mitchell H, Zhang L. Prevalence of Campylobacter species in adult Crohn’s disease and the preferential colonization sites of Campylobacter species in the human intestine. PLoS One. 2011;6:e25417. doi: 10.1371/journal.pone.0025417. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Hansen R, Berry SH, Mukhopadhya I, Thomson JM, Saunders KA, Nicholl CE, Bisset WM, Loganathan S, Mahdi G, Kastner-Cole D, et al. The microaerophilic microbiota of de-novo paediatric inflammatory bowel disease: the BISCUIT study. PLoS One. 2013;8:e58825. doi: 10.1371/journal.pone.0058825. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Kalischuk LD, Inglis GD. Comparative genotypic and pathogenic examination of Campylobacter concisus isolates from diarrheic and non-diarrheic humans. BMC Microbiol. 2011;11:53. doi: 10.1186/1471-2180-11-53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Man SM, Kaakoush NO, Leach ST, Nahidi L, Lu HK, Norman J, Day AS, Zhang L, Mitchell HM. Host attachment, invasion, and stimulation of proinflammatory cytokines by Campylobacter concisus and other non-Campylobacter jejuni Campylobacter species. J Infect Dis. 2010;202:1855–1865. doi: 10.1086/657316. [DOI] [PubMed] [Google Scholar]
- 24.Nielsen HL, Nielsen H, Ejlertsen T, Engberg J, Günzel D, Zeitz M, Hering NA, Fromm M, Schulzke JD, Bücker R. Oral and fecal Campylobacter concisus strains perturb barrier function by apoptosis induction in HT-29/B6 intestinal epithelial cells. PLoS One. 2011;6:e23858. doi: 10.1371/journal.pone.0023858. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Ismail Y, Lee H, Riordan SM, Grimm MC, Zhang L. The effects of oral and enteric Campylobacter concisus strains on expression of TLR4, MD-2, TLR2, TLR5 and COX-2 in HT-29 cells. PLoS One. 2013;8:e56888. doi: 10.1371/journal.pone.0056888. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Fasano A, Baudry B, Pumplin DW, Wasserman SS, Tall BD, Ketley JM, Kaper JB. Vibrio cholerae produces a second enterotoxin, which affects intestinal tight junctions. Proc Natl Acad Sci USA. 1991;88:5242–5246. doi: 10.1073/pnas.88.12.5242. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Waldor MK, Mekalanos JJ. Lysogenic conversion by a filamentous phage encoding cholera toxin. Science. 1996;272:1910–1914. doi: 10.1126/science.272.5270.1910. [DOI] [PubMed] [Google Scholar]
- 28.Uzzau S, Cappuccinelli P, Fasano A. Expression of Vibrio cholerae zonula occludens toxin and analysis of its subcellular localization. Microb Pathog. 1999;27:377–385. doi: 10.1006/mpat.1999.0312. [DOI] [PubMed] [Google Scholar]
- 29.Baudry B, Fasano A, Ketley J, Kaper JB. Cloning of a gene (zot) encoding a new toxin produced by Vibrio cholerae. Infect Immun. 1992;60:428–434. doi: 10.1128/iai.60.2.428-434.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Levine MM, Kaper JB, Herrington D, Losonsky G, Morris JG, Clements ML, Black RE, Tall B, Hall R. Volunteer studies of deletion mutants of Vibrio-cholerae O1 prepared by recombinant techniques. Infect Immun. 1988;56(1):161–167. doi: 10.1128/iai.56.1.161-167.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Fasano A, Not T, Wang W, Uzzau S, Berti I, Tommasini A, Goldblum SE. Zonulin, a newly discovered modulator of intestinal permeability, and its expression in coeliac disease. Lancet. 2000;355:1518–1519. doi: 10.1016/S0140-6736(00)02169-3. [DOI] [PubMed] [Google Scholar]
- 32.Wang W, Uzzau S, Goldblum SE, Fasano A. Human zonulin, a potential modulator of intestinal tight junctions. J Cell Sci. 2000;113 Pt 24:4435–4440. doi: 10.1242/jcs.113.24.4435. [DOI] [PubMed] [Google Scholar]
- 33.Stothard P, Van Domselaar G, Shrivastava S, Guo A, O’Neill B, Cruz J, Ellison M, Wishart DS. BacMap: an interactive picture atlas of annotated bacterial genomes. Nucleic Acids Res. 2005;33:D317–D320. doi: 10.1093/nar/gki075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol. 2011;28:2731–2739. doi: 10.1093/molbev/msr121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, et al. Clustal W and Clustal X version 2.0. Bioinformatics. 2007;23:2947–2948. doi: 10.1093/bioinformatics/btm404. [DOI] [PubMed] [Google Scholar]
- 36.Kaakoush NO, Man SM, Lamb S, Raftery MJ, Wilkins MR, Kovach Z, Mitchell H. The secretome of Campylobacter concisus. FEBS J. 2010;277:1606–1617. doi: 10.1111/j.1742-4658.2010.07587.x. [DOI] [PubMed] [Google Scholar]
- 37.Goldblum SE, Rai U, Tripathi A, Thakar M, De Leo L, Di Toro N, Not T, Ramachandran R, Puche AC, Hollenberg MD, et al. The active Zot domain (aa 288-293) increases ZO-1 and myosin 1C serine/threonine phosphorylation, alters interaction between ZO-1 and its binding partners, and induces tight junction disassembly through proteinase activated receptor 2 activation. FASEB J. 2011;25:144–158. doi: 10.1096/fj.10-158972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Tripathi A, Lammers KM, Goldblum S, Shea-Donohue T, Netzel-Arnett S, Buzza MS, Antalis TM, Vogel SN, Zhao A, Yang S, et al. Identification of human zonulin, a physiological modulator of tight junctions, as prehaptoglobin-2. Proc Natl Acad Sci USA. 2009;106:16799–16804. doi: 10.1073/pnas.0906773106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Hollander D, Vadheim CM, Brettholz E, Petersen GM, Delahunty T, Rotter JI. Increased intestinal permeability in patients with Crohn‘s disease and their relatives. A possible etiologic factor. Ann Intern Med. 1986;105:883–885. doi: 10.7326/0003-4819-105-6-883. [DOI] [PubMed] [Google Scholar]
- 40.May GR, Sutherland LR, Meddings JB. Is small intestinal permeability really increased in relatives of patients with Crohn’s disease? Gastroenterology. 1993;104:1627–1632. doi: 10.1016/0016-5085(93)90638-s. [DOI] [PubMed] [Google Scholar]
- 41.Irvine EJ, Marshall JK. Increased intestinal permeability precedes the onset of Crohn’s disease in a subject with familial risk. Gastroenterology. 2000;119:1740–1744. doi: 10.1053/gast.2000.20231. [DOI] [PubMed] [Google Scholar]
- 42.Welcker K, Martin A, Kölle P, Siebeck M, Gross M. Increased intestinal permeability in patients with inflammatory bowel disease. Eur J Med Res. 2004;9:456–460. [PubMed] [Google Scholar]
- 43.Wyatt J, Vogelsang H, Hübl W, Waldhöer T, Lochs H. Intestinal permeability and the prediction of relapse in Crohn’s disease. Lancet. 1993;341:1437–1439. doi: 10.1016/0140-6736(93)90882-h. [DOI] [PubMed] [Google Scholar]
- 44.Halme L, Paavola-Sakki P, Turunen U, Lappalainen M, Farkkila M, Kontula K. Family and twin studies in inflammatory bowel disease. World J Gastroenterol. 2006;12:3668–3672. doi: 10.3748/wjg.v12.i23.3668. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Lindblom GB, Sjögren E, Hansson-Westerberg J, Kaijser B. Campylobacter upsaliensis, C. sputorum sputorum and C. concisus as common causes of diarrhoea in Swedish children. Scand J Infect Dis. 1995;27:187–188. doi: 10.3109/00365549509019006. [DOI] [PubMed] [Google Scholar]
- 46.Lastovica AJ. Emerging Campylobacter spp.: the tip of the iceberg. Clin Microbiol Newsl. 2006;28:49–56. [Google Scholar]
- 47.Imamovic L, Muniesa M. Characterizing RecA-independent induction of Shiga toxin2-encoding phages by EDTA treatment. PLoS One. 2012;7:e32393. doi: 10.1371/journal.pone.0032393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Eroschenko VP. Di Fiore’s atlas of histology with functional correlations. 9 ed. Canada: Susan Katz; 2003. [Google Scholar]
- 49.Bennett RA, Rubin PH, Present DH. Frequency of inflammatory bowel disease in offspring of couples both presenting with inflammatory bowel disease. Gastroenterology. 1991;100:1638–1643. doi: 10.1016/0016-5085(91)90663-6. [DOI] [PubMed] [Google Scholar]
- 50.Comes MC, Gower-Rousseau C, Colombel JF, Belaïche J, Van Kruiningen HJ, Nuttens MC, Cortot A. Inflammatory bowel disease in married couples: 10 cases in Nord Pas de Calais region of France and Liège county of Belgium. Gut. 1994;35:1316–1318. doi: 10.1136/gut.35.9.1316. [DOI] [PMC free article] [PubMed] [Google Scholar]


