Abstract
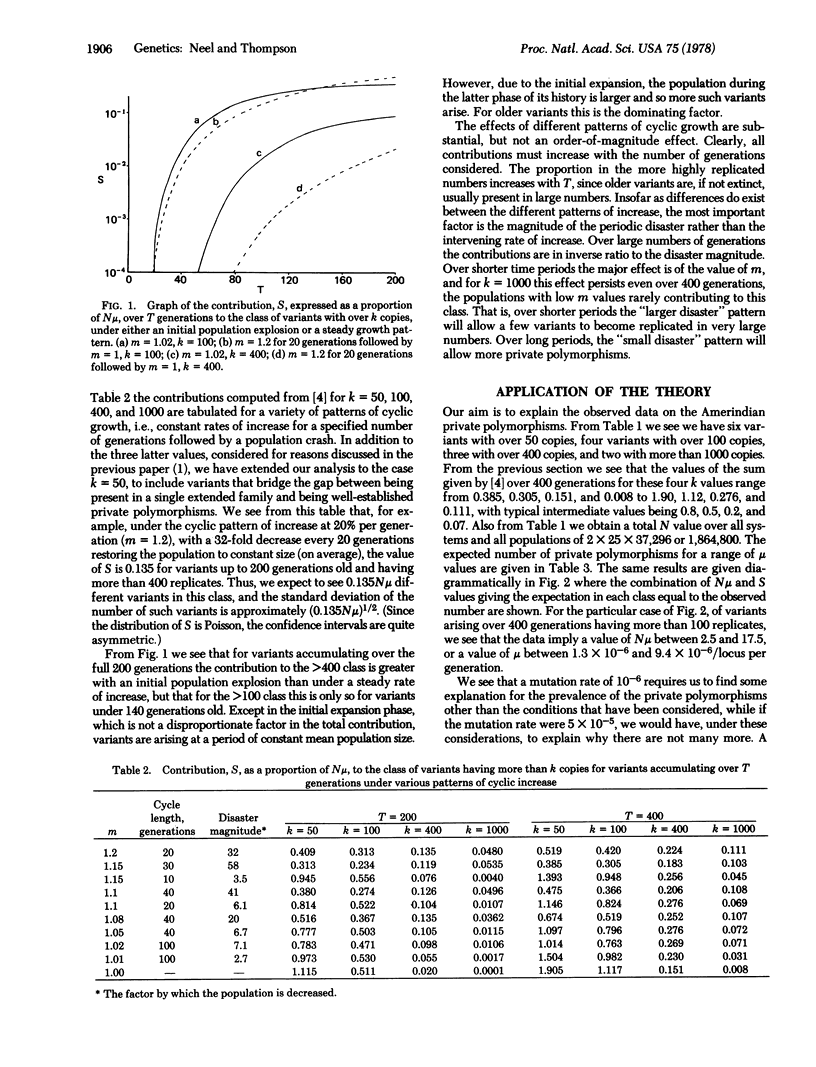
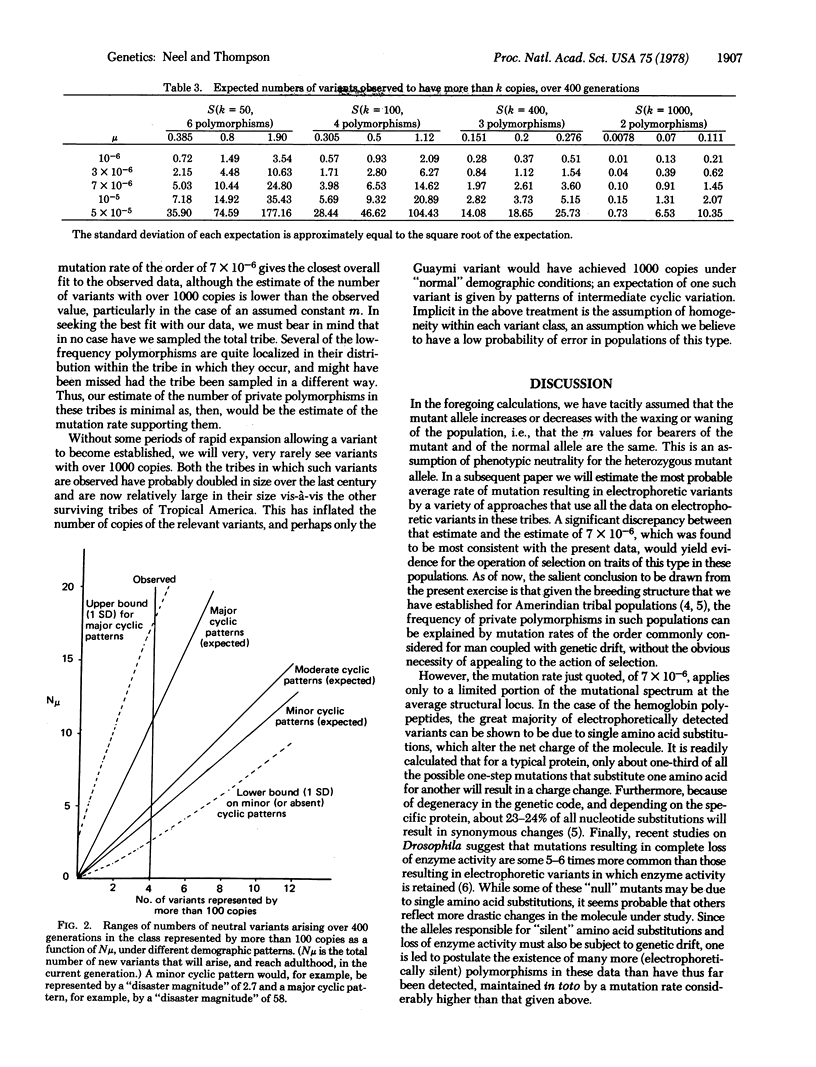
In studies extending over the past dozen years, we have observed eight examples of "private" genetic polymorphisms in 12 Amerindian tribes surveyed for electrophoretic variants of an average of 25 proteins. Each of these is presumed to trace to a single mutation. In a preceding communication [Thompson, E.A. & Neel, J.V. (1978) Proc. Natl. Acad. Sci. USA 75, 1442-1445] the statistical theory was developed for estimating the likelihood of such a founder effect in a tribal population of this type. In this paper that theory is applied to the distribution defined by these eight variants. It is demonstrated that on the assumption that the phenotypes in question are selectively neutral, such findings are most compatible with a mutation rate of 7 X 10(-6)/locus per generation. This figure applies only to variants that can be detected by the electrophoretic technique.
Full text
PDF


Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Mukai T., Cockerham C. C. Spontaneous mutation rates at enzyme loci in Drosophila melanogaster. Proc Natl Acad Sci U S A. 1977 Jun;74(6):2514–2517. doi: 10.1073/pnas.74.6.2514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Neel J. V., Weiss K. M. The genetic structure of a tribal population, the Yanomama Indians. XII. Biodemographic studies. Am J Phys Anthropol. 1975 Jan;42(1):25–51. doi: 10.1002/ajpa.1330420105. [DOI] [PubMed] [Google Scholar]
- Thompson E. A., Neel J. V. Probability of founder effect in a tribal population. Proc Natl Acad Sci U S A. 1978 Mar;75(3):1442–1445. doi: 10.1073/pnas.75.3.1442. [DOI] [PMC free article] [PubMed] [Google Scholar]


