Abstract
The central structural feature of natural proteins is a tightly packed and highly ordered hydrophobic core. If some measure of exquisite, native-like core packing is necessary for enzymatic function, this would constitute a significant obstacle to the development of novel enzymes, either by design or by natural or experimental evolution. To test the minimum requirements for a core to provide sufficient structural integrity for enzymatic activity, we have produced mutants of the ribonuclease barnase in which 12 of the 13 core residues have together been randomly replaced by hydrophobic alternatives. Using a sensitive biological screen, we find that a strikingly high proportion of these mutants (23%) retain enzymatic activity in vivo. Further substitution at the 13th core position shows that a similar proportion of completely random hydrophobic cores supports enzyme function. Of the active mutants produced, several have no wild-type core residues. These results imply that hydrophobicity is nearly a sufficient criterion for the construction of a functional core and, in conjunction with previous studies, that refinement of a crudely functional core entails more stringent sequence constraints than does the initial attainment of crude core function. Since attainment of crude function is the critical initial step in evolutionary innovation, the relatively scant requirements contributed by the hydrophobic core would greatly reduce the initial hurdle on the evolutionary pathway to novel enzymes. Similarly, experimental development of novel functional proteins might be simplified by limiting core design to mere specification of hydrophobicity and using iterative mutation-selection to optimize core structure.
Full text
PDF
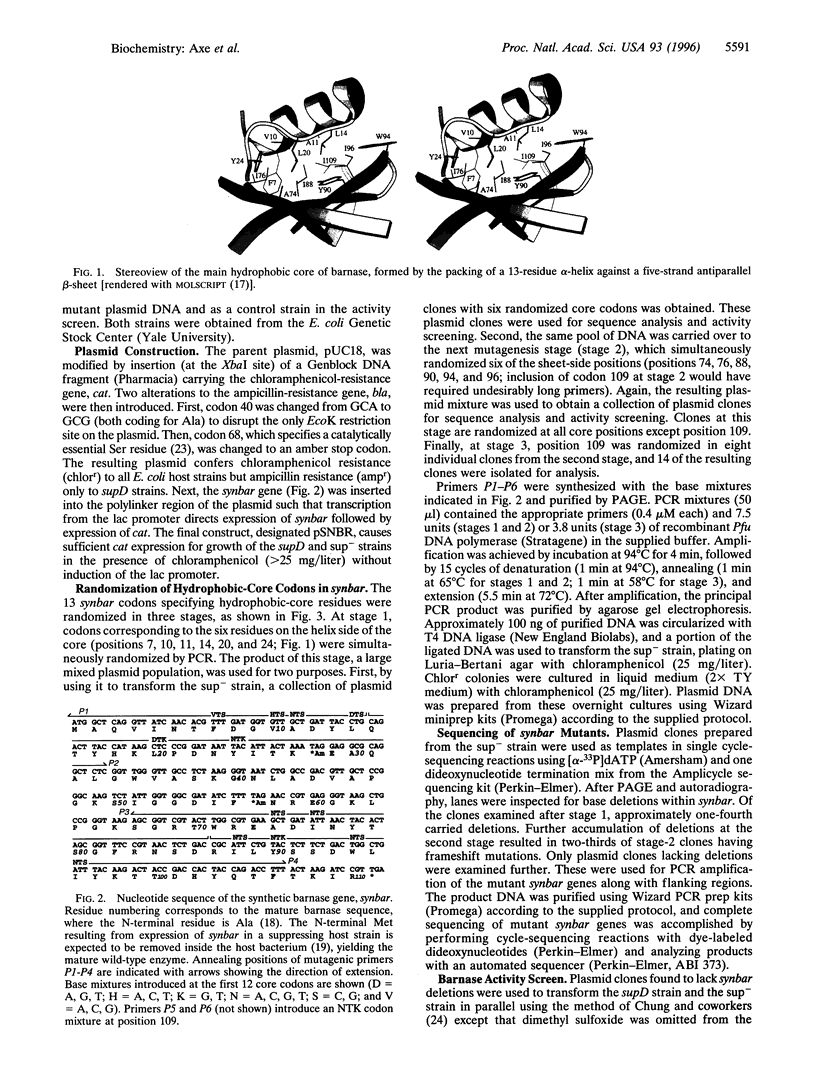



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bartel D. P., Szostak J. W. Isolation of new ribozymes from a large pool of random sequences [see comment]. Science. 1993 Sep 10;261(5127):1411–1418. doi: 10.1126/science.7690155. [DOI] [PubMed] [Google Scholar]
- Bashford D., Chothia C., Lesk A. M. Determinants of a protein fold. Unique features of the globin amino acid sequences. J Mol Biol. 1987 Jul 5;196(1):199–216. doi: 10.1016/0022-2836(87)90521-3. [DOI] [PubMed] [Google Scholar]
- Chen K., Arnold F. H. Tuning the activity of an enzyme for unusual environments: sequential random mutagenesis of subtilisin E for catalysis in dimethylformamide. Proc Natl Acad Sci U S A. 1993 Jun 15;90(12):5618–5622. doi: 10.1073/pnas.90.12.5618. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chothia C. Structural invariants in protein folding. Nature. 1975 Mar 27;254(5498):304–308. doi: 10.1038/254304a0. [DOI] [PubMed] [Google Scholar]
- Chung C. T., Niemela S. L., Miller R. H. One-step preparation of competent Escherichia coli: transformation and storage of bacterial cells in the same solution. Proc Natl Acad Sci U S A. 1989 Apr;86(7):2172–2175. doi: 10.1073/pnas.86.7.2172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Day A. G., Parsonage D., Ebel S., Brown T., Fersht A. R. Barnase has subsites that give rise to large rate enhancements. Biochemistry. 1992 Jul 21;31(28):6390–6395. doi: 10.1021/bi00143a005. [DOI] [PubMed] [Google Scholar]
- Dill K. A., Bromberg S., Yue K., Fiebig K. M., Yee D. P., Thomas P. D., Chan H. S. Principles of protein folding--a perspective from simple exact models. Protein Sci. 1995 Apr;4(4):561–602. doi: 10.1002/pro.5560040401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eriksson A. E., Baase W. A., Zhang X. J., Heinz D. W., Blaber M., Baldwin E. P., Matthews B. W. Response of a protein structure to cavity-creating mutations and its relation to the hydrophobic effect. Science. 1992 Jan 10;255(5041):178–183. doi: 10.1126/science.1553543. [DOI] [PubMed] [Google Scholar]
- Hartley R. W. Barnase and barstar. Expression of its cloned inhibitor permits expression of a cloned ribonuclease. J Mol Biol. 1988 Aug 20;202(4):913–915. doi: 10.1016/0022-2836(88)90568-2. [DOI] [PubMed] [Google Scholar]
- Hecht M. H. De novo design of beta-sheet proteins. Proc Natl Acad Sci U S A. 1994 Sep 13;91(19):8729–8730. doi: 10.1073/pnas.91.19.8729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hirel P. H., Schmitter M. J., Dessen P., Fayat G., Blanquet S. Extent of N-terminal methionine excision from Escherichia coli proteins is governed by the side-chain length of the penultimate amino acid. Proc Natl Acad Sci U S A. 1989 Nov;86(21):8247–8251. doi: 10.1073/pnas.86.21.8247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Joris B., Ghuysen J. M., Dive G., Renard A., Dideberg O., Charlier P., Frère J. M., Kelly J. A., Boyington J. C., Moews P. C. The active-site-serine penicillin-recognizing enzymes as members of the Streptomyces R61 DD-peptidase family. Biochem J. 1988 Mar 1;250(2):313–324. doi: 10.1042/bj2500313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kamtekar S., Hecht M. H. Protein Motifs. 7. The four-helix bundle: what determines a fold? FASEB J. 1995 Aug;9(11):1013–1022. doi: 10.1096/fasebj.9.11.7649401. [DOI] [PubMed] [Google Scholar]
- Kamtekar S., Schiffer J. M., Xiong H., Babik J. M., Hecht M. H. Protein design by binary patterning of polar and nonpolar amino acids. Science. 1993 Dec 10;262(5140):1680–1685. doi: 10.1126/science.8259512. [DOI] [PubMed] [Google Scholar]
- Karpusas M., Baase W. A., Matsumura M., Matthews B. W. Hydrophobic packing in T4 lysozyme probed by cavity-filling mutants. Proc Natl Acad Sci U S A. 1989 Nov;86(21):8237–8241. doi: 10.1073/pnas.86.21.8237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kellis J. T., Jr, Nyberg K., Sali D., Fersht A. R. Contribution of hydrophobic interactions to protein stability. Nature. 1988 Jun 23;333(6175):784–786. doi: 10.1038/333784a0. [DOI] [PubMed] [Google Scholar]
- Lim W. A., Sauer R. T. Alternative packing arrangements in the hydrophobic core of lambda repressor. Nature. 1989 May 4;339(6219):31–36. doi: 10.1038/339031a0. [DOI] [PubMed] [Google Scholar]
- Lim W. A., Sauer R. T. The role of internal packing interactions in determining the structure and stability of a protein. J Mol Biol. 1991 May 20;219(2):359–376. doi: 10.1016/0022-2836(91)90570-v. [DOI] [PubMed] [Google Scholar]
- Milla M. E., Sauer R. T. Critical side-chain interactions at a subunit interface in the Arc repressor dimer. Biochemistry. 1995 Mar 14;34(10):3344–3351. doi: 10.1021/bi00010a025. [DOI] [PubMed] [Google Scholar]
- Mossakowska D. E., Nyberg K., Fersht A. R. Kinetic characterization of the recombinant ribonuclease from Bacillus amyloliquefaciens (barnase) and investigation of key residues in catalysis by site-directed mutagenesis. Biochemistry. 1989 May 2;28(9):3843–3850. doi: 10.1021/bi00435a033. [DOI] [PubMed] [Google Scholar]
- Parsell D. A., Sauer R. T. The structural stability of a protein is an important determinant of its proteolytic susceptibility in Escherichia coli. J Biol Chem. 1989 May 5;264(13):7590–7595. [PubMed] [Google Scholar]
- Richards F. M., Lim W. A. An analysis of packing in the protein folding problem. Q Rev Biophys. 1993 Nov;26(4):423–498. doi: 10.1017/s0033583500002845. [DOI] [PubMed] [Google Scholar]
- Sandberg W. S., Terwilliger T. C. Influence of interior packing and hydrophobicity on the stability of a protein. Science. 1989 Jul 7;245(4913):54–57. doi: 10.1126/science.2787053. [DOI] [PubMed] [Google Scholar]
- Shortle D., Stites W. E., Meeker A. K. Contributions of the large hydrophobic amino acids to the stability of staphylococcal nuclease. Biochemistry. 1990 Sep 4;29(35):8033–8041. doi: 10.1021/bi00487a007. [DOI] [PubMed] [Google Scholar]
- Skolnick J., Kolinski A., Yaris R. Dynamic Monte Carlo study of the folding of a six-stranded Greek key globular protein. Proc Natl Acad Sci U S A. 1989 Feb;86(4):1229–1233. doi: 10.1073/pnas.86.4.1229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vuilleumier S., Sancho J., Loewenthal R., Fersht A. R. Circular dichroism studies of barnase and its mutants: characterization of the contribution of aromatic side chains. Biochemistry. 1993 Oct 5;32(39):10303–10313. doi: 10.1021/bi00090a005. [DOI] [PubMed] [Google Scholar]
- Wilson C., Szostak J. W. In vitro evolution of a self-alkylating ribozyme. Nature. 1995 Apr 27;374(6525):777–782. doi: 10.1038/374777a0. [DOI] [PubMed] [Google Scholar]




