Abstract
Although the human L-FABP T94A variant arises from the most commonly occurring SNP in the entire FABP family, there is a complete lack of understanding regarding the role of this polymorphism in human disease. It has been hypothesized that the T94A substitution results in complete loss of ligand binding ability and function analogous to L-FABP gene ablation. This possibility was addressed using recombinant human WT T94T and T94A variant L-FABP and cultured primary human hepatocytes. Non-conservative replacement of the medium sized, polar, uncharged T residue by a smaller, nonpolar, aliphatic A residue at position 94 of human L-FABP significantly increased L-FABP protein α-helical structure at the expense of β-sheet and concomitantly decreased thermal stability. T94A did not alter binding affinities for PPARα agonist ligands (phytanic acid, fenofibrate, fenofibric acid). While T94A did not alter the impact of phytanic acid and only slightly altered that of fenofibrate on human L-FABP secondary structure, the active metabolite fenofibric acid altered T94A secondary structure much more than that of WT T94T L-FABP. Finally, in cultured primary human hepatocytes the T94A variant exhibited significantly reduced fibrate-mediated induction of PPARα-regulated proteins such as L-FABP, FATP5, and PPARα itself. Thus, while T94A substitution did not alter the affinity of human L-FABP for PPARα agonist ligands, it significantly altered human L-FABP structure, stability, as well as conformational and functional response to fibrate.
Keywords: L-FABP, fibrate, liver, human
Liver fatty acid binding protein (L-FABP, FABP1) is the first discovered and arguably most unique member of the large FABP family (1-4). Unlike other family members, L-FABPs from a variety of species (e.g. rat, bovine, human) have a much larger ligand binding cavity that accommodates two instead of one ligand (4-14). L-FABP has broader specificity for binding not only straight-chain LCFA but also more potent peroxisome proliferator activated receptor-α (PPARα) agonists such as the naturally-occurring branched-chain LCFA phytanic acid (15-17) and less toxic xenobiotic fibrates (6,9,11,12,18,19) and phthalates (20). An intriguing result of structural (12,21,22), in vitro (23), cultured cell (24-27), and primary mouse hepatocyte studies (28-31) was the discovery that L-FABP provides a signaling pathway for both natural (i.e. very long chain polyunsaturated fatty acids) and xenobiotic (fibrates) ligands to the nucleus. In this pathway, L-FABP binds ligands (LCFA, fatty acid synthesis inhibitors, fibrates) (7-9,11,19,36,37) in the cytosol for cotransport into nuclei (32-35) and targets (23,25,26,28,37) bound ligands to nuclear PPARα to activate transcription of genes involved in LCFA uptake, transport, and metabolism (24,28-31).
X-ray and NMR demonstrated that human liver L-FABP has an even larger binding cavity than any other mammalian L-FABP, suggesting that results from other species’ L-FABP may not necessarily be straightforwardly extrapolated to the human L-FABP (12-14,38-42). The recent discovery of a SNP in the human L-FABP coding sequence that resulted in a single amino acid substitution, T94A, has added further complexity to this issue (43,44). This is especially important since the human L-FABP T94A variant is very common (26-38% minor allele freq.; 8.3±1.9% homozygous; MAF for 1000 genomes in NCBI dbSNP database; ALFRED database) and is the most prevalent polymorphism in the FABP family (43,45-50). Further, the L-FABP T94A variant has been associated with elevated plasma triglycerides (44,45), increased LDL cholesterol (45,49), atherothrombotic cerebral infarction (47), and non-alcoholic fatty liver disease (NAFLD) (49). The L-FABP ligand fenofibrate [most commonly prescribed fibrate activator of PPARα in the US and Canada (51)] is less effective in lowering elevated plasma triglyceride to basal levels in L-FABP T94A variant subjects (44). Resolving the molecular basis for this difference is important for future therapeutic studies whereby newer fibrates might be better targeted by L-FABP or T94A variant L-FABP to activate PPARα in the nucleus.
Based on the above findings we re-examined the literature to determine whether earlier structural studies were performed with recombinant human L-FABP derived WT T94T L-FABP or T94A variant coding cDNAs. Fortuitously, all previously cloned human L-FABPs were derived from human WT T94T L-FABP cDNA (52-54). Thus, all studies characterizing ligand specificity (9,12,40,55), structure (13,14,38,39), and mode of ligand binding (12-14) of human L-FABP protein were performed with the WT T94T L-FABP protein or the phenotype of the human L-FABP was not reported (41,42). While it has been suggested that the T94A substitution abolished ligand binding to human L-FABP (56), there have been no reports actually examining the structure or ligand binding specificity of the human L-FABP T94A variant protein. To begin to resolve these issues, studies with purified recombinant human WT and T94A variant L-FABP proteins were initiated. As shown by CD and UV spectroscopy as well as fluorescence displacement assays, human T94A variant L-FABP differed significantly in secondary structure, thermal stability, and structural response to fenofibric acid binding as compared to the human WT T94T L-FABP. Finally, T94A diminished fenofibrate-mediated induction of PPARα transcriptional activity in human hepatocytes. Such dissimilarities may contribute to the impaired therapeutic response of human T94A variant L-FABP expressing subjects to fenofibrate (44).
EXPERIMENTAL PROCEDURES
Materials
Luria-Bertani (LB) broth and LB agar were purchased from Becton, Dickinson Co. (Sparks, MD). Fenofibrate and fenofibric acid were obtained from Santa Cruz Biotechnology (Dallas, Texas). ANS (1-anilinonaphthalene-8-sulfonic acid) was purchased from Life Technologies (Grand Island, NY). Phytanic acid, ampicillin (sodium salt), isopropyl-β-D-galactoside (IPTG), ammonium sulfate, protamine sulfate, dithiothreitol (DTT), and all other common laboratory chemicals were obtained from Sigma-Aldrich (St. Louis, MO). Mini-PROTEAN TGX any kD precast polyacrylamide gels and Precision Plus Protein Dual Xtra Standards were purchased from Bio-Rad (Hercules, CA). SimplyBlue SafeStain was from Invitrogen (Carlsbad, CA). All reagents and solvents used were of the highest grade available.
Recombinant L-FABP Protein Expression in E. coli
Recombinant rat L-FABP was expressed in E. coli as described earlier (8,57). The cDNA for human L-FABP (NM_001443) was purchased from OriGene Technologies (Rockville, MD). Full-length human L-FABP was amplified using the following primers: 5’-CAGCCATATGAGTTTCTCCGGCAAGTAC-3’ and 5’-GGTGCTCGAGTTAAATTCTCTTGCTGATTCTC-3’ with restriction sites BamHI and HindIII, respectively, and was cloned into the pQE9-His vector (Qiagen, Valencia, CA). The cDNA purchased from OriGene was determined to be the human L-FABP T94A mutant (i.e. the 280th nucleotide was guanine instead of adenine resulting in an alanine substitution for threonine). The mutation was established by sequencing at the Research Technology Support Facility (RTSF, Michigan State University, East Lansing, MI). In order to obtain the T94T wild-type (WT) human L-FABP site-directed mutagenesis was performed with the following primers: 5’-CAATAAACTGGTGACAACTTTCAAAAACATCAAG-3’ and 5’-CTTGATGTTTTTGAAAGTTGTCACCAGTTTATTG-3’ utilizing PfuTurbo DNA polymerase (Agilent Technologies, Santa Clara, CA). The final T94T WT and T94A mutant constructs were transformed into E. coli C43 (Lucigen Corporation, Middleton, WI) for protein expression.
Recombinant L-FABP Protein Purification
Recombinant rat L-FABP was purified as described earlier (8,57). Briefly, transformed E. coli (expressing WT T94T L-FABP or T94A) were streaked onto LB agar plates containing ampicillin (50 μg/mL). Plates were incubated overnight at 37 °C, individual colonies were picked and inoculated into 100 mL of LB/ampicillin (50 μg/mL), and incubated overnight at 37 °C in a shaking incubator (250 rpm). Ten mL of the overnight culture was added to 1 L of LB/ampicillin (50 μg/mL) and incubated at 37 °C in a shaking incubator (250 rpm) until optical density (OD) at 600 nm reached 0.6 absorbance units (4-5 hr for E. coli C43). Protein expression was induced by adding IPTG (final concentration = 1 mM) to each 1-L culture. The cultures were incubated for 12 hr at 37 °C in a shaking incubator (250 rpm). Bacterial cells were harvested by centrifugation in a Beckman Avanti J-25 centrifuge (JA-14 rotor, 10000 × g, 15 min, 4 °C). Each cell pellet was resuspended in ice-cold NPND buffer (20 mM sodium phosphate (pH 7.4), 100 mM NaCl, l mM dithiothreitol) containing protease inhibitor cocktail (-EDTA, Sigma-Aldrich product number S8830, St. Louis, MO) at a concentration of 0.35 g cell pellet/mL buffer.
Bacterial cell lysis was achieved by homogenization utilizing a French Pressure Cell Press (SLM Aminco FA-078, high ratio, gage pressure = 1260) and the French Pressure Cell (Thermo FA-032, 1 in piston diameter, cell pressure = 20000 psi). Each 25-30 mL batch of bacterial cell suspension was processed 2x. Additional cell lysis was accomplished by sonication on ice utilizing a Fisher Scientific Sonic Dismembrator 550 equipped with micro-tip (Fisher Sci., Pittsburgh, PA). The sonication conditions were: setting 4, total processing time 15 min, on-time 15.0 sec, off-time 15.0 sec. Insoluble cell debris was removed by centrifugation (JA-25.50 rotor, 40000 × g, 20 min, 4 °C). The supernatant was slowly brought to 65% saturation with stirring at 4 °C using solid ammonium sulfate. After addition of all ammonium sulfate, the protein suspension was stirred slowly at 4 °C for 12 hr. Insoluble material was removed by centrifugation (JA25.50 rotor, 40000 × g, 20 min, 4 °C). The supernatant was desalted at 4 °C by chromatography through a Sephadex G-25 (GE Healthcare, Piscataway, NJ) column using NPND buffer (pH 7.4) as the mobile phase. DNA was precipitated from the desalted protein solution using protamine sulfate (0.1% w/v). The protamine sulfate was added slowly with stirring at 4 °C and this mixture was allowed to stir slowly at 4 °C for 12 hr. Insoluble material was removed by centrifugation as described above. The supernatant was concentrated by ultrafiltration utilizing an Amicon stirred cell (Model 402) at 4 °C (Millipore Ultrafiltration Membrane, Regenerated Cellulose, 76 mm diameter, NMWL = 1000 Da).
The concentrated protein solutions were buffer-exchanged into HisTrap binding buffer (20 mM sodium phosphate (pH 7.4)/0.5 M NaCl/30 mM imidazole) utilizing PD MidiTrap G-25 columns (GE Healthcare, Piscataway, NJ) as per the manufacturer’s directions. The buffer-exchanged protein mixture was loaded onto HisTrap FF (GE Healthcare, Piscataway, NJ) column at 25 °C pre-equilibrated with HisTrap binding buffer. The column was washed extensively with HisTrap binding buffer; column eluted with HisTrap elution buffer (20 mM sodium phosphate (pH 7.4)/0.5 M NaCl/0.5 M imidazole). The HisTrap eluted protein was buffer-exchanged into 50 mM sodium phosphate (pH 7.0)/1.0 M ammonium sulfate with PD MidiTrap G-25 columns as above and loaded onto a pre-equilibrated hydrophobic interaction column (HiTrap Phenyl HP, GE Healthcare, Piscataway, NJ) at 25 °C. Protein delipidation was achieved by eluting the protein from the HiTrap Phenyl HP column utilizing a linear buffer gradient from 50 mM sodium phosphate (pH 7.0)/1.0 M ammonium sulfate to 50 mM sodium phosphate (pH 7.0). Delipidated protein was buffer-exchanged (PD MidiTrap G-25) into 20 mM sodium phosphate (pH 7.4)/150 mM NaCl prior to His tag removal utilizing the TAGZyme Kit (Qiagen, Valencia, CA) according to the manufacturer’s directions. The protein was buffer-exchanged (PD MidiTrap G-25) into HisTrap binding buffer as described above prior to loading onto HisTrap FF column as above. The column wash (unbound) and the column eluate were examined by sodium dodecyl sulfate-polyacrylmide gel electrophoresis (SDS-PAGE). All L-FABP was found in the column wash material (data not shown) indicating complete His tag removal. Purified L-FABP (~1.5 mg/L culture) was buffer-exchanged (PD MidiTrap G-25) into 10 mM potassium phosphate (pH 7.4)/1 mM dithiothreitol, aliquoted, and stored at -80 °C.
Recombinant L-FABP Purity and Identity: Amino Acid Analysis and Mass Spectroscopy
Aliquots of purified rat L-FABP as well as human WT T94T and T94A mutant L-FABPs were analyzed by amino acid analysis and matrix-assisted laser desorption time-of-flight (MALDI-TOF) mass spectrometry (Dr. Larry Dangott, Protein Chemistry Laboratory, Texas A&M University, College Station, TX) for protein purity (>98%), concentration, and molecular weight. Mass spectrometry was performed utilizing a Shimadzu/Kratos Axima CFR MALDI-TOF mass spectrometer in reflectron mode. The matrix used was α-Cyano-4-hydroxycinnamic acid (Sigma-Aldrich, St. Louis, MO) and the instrument was calibrated using cytochrome C.
Fluorescence Spectra for Recombinant Proteins
L-FABP tyrosine fluorescence emission spectra were recorded at 24°C using a Varian Cary Eclipse Fluorescence Spectrophotometer (Varian, Inc., Palo Alto, CA), by scanning from 295 to 420 nm, with excitation wavelength 280 nm. Rat and human L-FABP have 3 and 1 tyrosine per protein molecule respectively. To obtain emission at equivalent quantities of tyrosine, protein concentrations used were: 200 nM for rat L-FABP and 600 nM for WT T94T and T94A.
Circular Dichroism Spectroscopy (CD) to Determine Recombinant Protein Secondary Structure
All CD spectroscopy experiments were performed utilizing a JASCO J-815 CD spectrometer (JASCO, Easton, MD) equipped with a Model PFD-425S Peltier Type FDCD attachment for temperature regulation. All experiments (temperature and ligand interaction) were done at a final protein concentration of 0.5 μM as determined by amino acid analysis (see above) in a buffer containing 10 mM potassium phosphate (pH 7.4) with or without 1% ethanol. The presence of ethanol was determined to have no effect on ligand binding, protein CD spectra, or resulting secondary structure determinations (data not shown). Each protein sample was incubated with stirring (250 rpm) at 25 °C for 10 min prior to scanning. The sample was scanned 10 times from 185 nm to 250 nm. The final CD spectrum obtained represented an average of ten scans, the spectrum was background-subtracted, and the spectrum was mathematically smoothed using the Means-Movement method with a convolution width = 5. CD spectra of each protein can be found at http://pcddb.cryst.bbk.ac.uk (PCDDB ID CD000413200-PCDDB ID CD000413400). Secondary structure analysis was performed utilizing the analysis software supplied with the CD spectrometer. SDP (soluble and denatured protein) 48 was used as the reference set. Spectra were analyzed by CONTIN, CDSSTR, and SELCON 3. CONTIN consistently resulted in lowest root-mean-square deviation (RMSD) of these algorithms (data not shown). The percent change in secondary structure between two protein samples was calculated using the following formula: [(% Secondary StructureSample 2 - % Secondary StructureSample 1) ÷ % Secondary StructureSample 1] * 100%.
Recombinant Protein Stability to Thermal Denaturation: CD Spectroscopy
Temperature CD studies of rat and human L-FABPs were performed as follows. Each sample containing 0.5 μM protein in 10 mM potassium phosphate (pH 7.4) was incubated with stirring at 25 °C for 10 min in the FDCD attachment and then CD spectra obtained as described above. Sample temperature was increased by 10 °C at a rate of 1 °C/min followed by a 10 min incubation prior to obtaining the CD spectrum. This procedure was repeated until the final sample analysis was performed as described at 95 °C. Upon completion of the temperature scans, all spectra were analyzed and secondary structure determinations were performed as described above.
Ligand Binding: ANS Fluorescence Displacement Assay
ANS is very weakly fluorescent in buffer, but its fluorescence increases upon binding to L-FABP. Therefore, ANS was excited at 380 nm and emission spectra recorded by scanning from 410-600 nm. Two types of titrations were performed: First, L-FABP (500 nM) was titrated with ANS (0-48 μM, forward titration). Second, ANS (100 nM) was titrated with increasing amounts of L-FABP (0-4 μM, reverse titration). The fluorescence intensity of ANS (per nM) when fully bound to L-FABP was calculated by curve fitting the reverse titration curve. This parameter was then used in forward titration to calculate the fractional saturation and free ANS concentration. Kd and Bmax were determined from the binding curve of fractional saturation (Y) vs free ANS concentration (X).
Ligand (phytanic acid, fenofibrate, and fenofibric acid) binding affinity to L-FABP was determined by ANS displacement assay. A solution of L-FABP (500 nM) and ANS (35 μM) was titrated with phytanic acid (0-6.4 μM) or fenofibrate (0-6 μM for rat L-FABP, and 0-4 μM for human L-FABP), or fenofibric acid (0-300 μM for rat L-FABP, 0-48 μM for human L-FABP). EC50 was obtained from the displacement curve. Ki was calculated from EC50/[ANS] = Ki/Kd.
Impact of Ligand Binding on Recombinant Protein Secondary Structure Determined by CD
Each sample contained 0.5 μM protein in 10 mM potassium phosphate (pH 7.4). Ligand (phytanic acid, fenofibrate or fenofibric acid) was added from a stock solution of 500 μM ligand in ethanol such that the final ligand concentration was 5 μM and the final ethanol concentration was 1%. The protein/ligand sample was incubated with stirring at 25 °C for 10 min in the FDCD attachment prior to obtaining the CD spectrum. Again, the final CD spectrum was an average of ten scans, background subtracted (buffer/ligand/ethanol), mathematically smoothed, and secondary structure determined as described. The % change in secondary structure between two protein samples was calculated using the following formula: [(% Secondary StructureSample 2 - % Secondary StructureSample 1) ÷ % Secondary StructureSample 1] * 100%.
Fenofibrate-mediated Induction of Transcription of PPARα-regulated Proteins in Cultured Primary Human Hepatocytes: Qrt-PCR
Cryopreserved primary human hepatocytes from female (50±3 yrs old) Caucasian donors were from Life Technologies (Grand Island, NY). Hepatocytes were genotyped to determine WT T94T (TT), heterozygous (TC), or T94A (CC) variant expression as in (45,46), thawed, plated and cultured overnight according to the supplier’s protocol (Life Technologies, Grand Island, NY). Human hepatocytes were then incubated with 40 μM BSA (fatty acid free) or 40 μM BSA/fenofibrate (1:1, mol/mol) complex for 24 hours in media prepared by adding 6 mM glucose, 100 nM insulin, and 10 nM dexamethasone to glucose free William’s E media (US Biological, Salem, MA). Total mRNA was obtained with RN-easy kit from Qiagen (Valencia, CA) and RN-ase free DNase set from Qiagen GmbH (Hilden, Germany). TaqMan, One-Step RT-PCR Master Mix reagents, TaqMan Gene Expression Assays for human mRNAs were from Applied Biosystems (by Life Technologies, Grand Island, NY): liver fatty acid binding protein (L-FABP), fatty acid transport protein-5 (FATP5), and peroxisome proliferator activated receptor-α (PPAR-α). Messenger RNA levels were determined according to the procedures provided by the manufacturer.
Statistics
Statistical analysis was performed by one-way analysis of variance (ANOVA) combined with the Newman-Keuls multiple-comparisons post-test (GraphPad Prism Version 3.03, San Diego, CA). Unless otherwise noted, data are expressed as means ± standard error of the mean (n = 4-6) and P is indicated as described in the figure legends. Graphical analysis was accomplished using SigmaPlot 2002 for Windows Version 8.02 (SPSS, Chicago, IL).
RESULTS
Amino Acid Sequence Homology between Rat, Human WT T94T, and Human T94A Variant L-FABPs
Human and murine L-FABPs mediate ligand signaling to their respective PPARαs (12,21,22,24,27,58). However, mRNA profiling of rodent (rat, mouse) and human hepatocytes revealed significant differences in response to fibrates (59-61). While this was attributed primarily to species differences in PPARα, species differences in L-FABP must also be considered. While rat (10,62) and human (13,14) L-FABPs share significant overall secondary structure, each having a ten-β-sheet β-barrel along with two α-helices and turns between them, differences in their 127 amino acid sequence suggest significant influences on secondary structures. The human WT L-FABP contains 13 (T94T in both rat and human WT L-FABP) non-conservative (Fig. 1, red) and 9 conservative (Fig. 1, green) substitutions such that it is only 82.7% identical and 89.8% similar to the rat L-FABP (63). Consequently, human L-FABPs have three fewer basic (positively-charged) amino acids than does rat L-FABP and the resultant pI, calculated as described (63), for the human T94T WT L-FABP is 6.60—considerably more acidic than that of the rat L-FABP with pI = 7.79. While human L-FABP T94A also has a pI = 6.60, this substitution results in replacement of a medium sized, polar, uncharged T residue by a smaller, nonpolar, aliphatic A residue (Fig. 1, *). The consequences of these differences to human L-FABP structure, stability, ligand binding, and function are detailed in the following sections.
Figure 1. Amino acid sequence alignment of rat, human T94T WT, and human T94A variant L-FABPs.
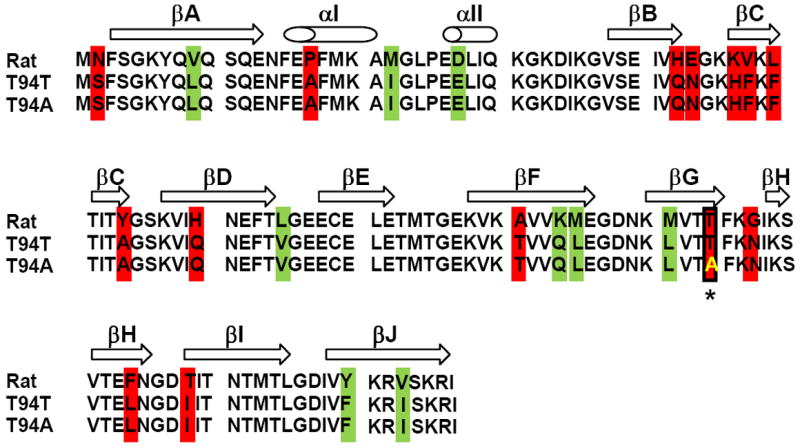
Multiple sequence alignment was performed utilizing Clustal W (embnet.vital-it.ch/software/ClustalW.html). Additional evaluation of amino acid substitutions was performed utilizing information in (63). Protein secondary structural elements are pictured above the aligned sequences; α-helices αI and αII are designated by the cylindrical cones, β-sheets βA-βJ are designated by the open arrows. Conservative amino acid substitution is indicated by green box; non-conservative amino acid substitution is indicated by red box; the T94A amino acid substitution in human L-FABP is indicated with a red box and *.
Purification of the Recombinant Human WT T94T and T94A Variant L-FABPs
Since all previously described human L-FABP cDNAs were of the human WT T94T L-FABP (52-54), it was necessary to prepare the T94A variant L-FABP as well as the WT T94T L-FABP proteins. Sequencing of a commercially available cDNA encoding the human L-FABP revealed that it encoded the T94A variant L-FABP. Therefore, the WT T94T L-FABP was obtained by site directed mutagenesis of T94A L-FABP variant cDNA as in Methods. Respective cDNAs were inserted in a bacterial expression vector for protein expression and purification (Fig 2A) also as detailed in Methods. Purified WT T94T and T94A variant L-FABP proteins were detected as single bands on SDS-PAGE gels (Figs 2B, D; lane 11). MALDI-TOF analysis of the final, Histag free human WT T94T L-FABP and T94A L-FABP variant detected main mass peaks at 14,208.5 Da (Fig. 2C) and 14,178.39 Da (Fig. 2E), respectively—consistent with molecular weights based on amino acid sequence (Fig. 1). Increasing the amount of L-FABP loaded on SDS-PAGE gel did not reveal significant additional bands (Fig 2F). Thus, these proteins were sufficiently pure (>98%) for structural and functional characterization.
Figure 2. Purification flow chart, SDS-PAGE analysis and mass spectra of human T94T WT and T94A variant L-FABPs.
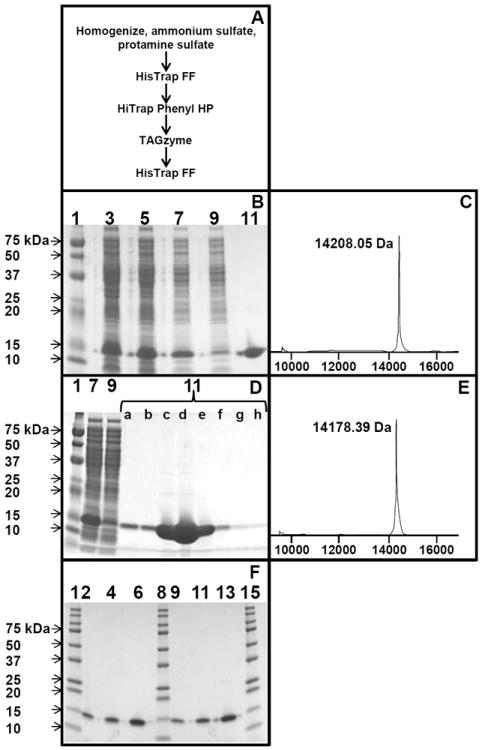
Panel A: Outline of human L-FABP purification procedure. Aliquots of human T94T WT (panels B, C) and T94A variant L-FABP (panels D, E) were examined by SDS-PAGE and mass spectrometry as described in Methods. Panel B: SDS-PAGE analysis of T94T WT human L-FABP; lane 1, molecular size markers; lane 3, cell homogenate; lane 5, cell homogenate supernatant; lane 7, ammonium sulfate/protamine sulfate supernatant; lane 9, HisTrap FF column wash (unbound); lane 11, HisTrap FF column eluate. Panel C: MALDI-TOF analysis of human T94T WT L-FABP after delipidation and His tag removal. Panel D: SDS-PAGE analysis of human T94A variant L-FABP; lane 1, molecular size markers; lane 7, ammonium sulfate/protamine sulfate supernatant; lane 9, HisTrap FF column wash (unbound); lanes 11 a-h, HisTrap FF column elution fractions. Panel E: MALDI-TOF analysis of human T94A variant L-FABP after delipidation and His tag removal. Panel F: SDS-PAGE analysis of purified T94T WT [lanes 2 (1 μg protein), 4 (2 μg), 6 (5 μg)] and T94A variant L-FABP [lanes 9 (1 μg protein), 11 (2 μg), 13 (5 μg)]. Lanes 1, 8, 15: molecular size markers.
Tyrosine Fluorescence Spectroscopy of Rat and Human L-FABPs
Human WT T94T L-FABP has a single Tyr residue at Y7, located in βA (part of the β-barrel ligand binding structure, N-terminal to the α-helical cap) (10). But, it is not known if the T94A substitution altered the polarity of the microenvironment wherein this Tyr residue resides. Although both WT T94T and T94A L-FABP had maximal fluorescence emission at 305 nm, the fluorescence efficiency of Tyr in both human L-FABPs was about 4-fold less than that of Tyr in rat L-FABP (not shown). While this suggested significant differences in rat vs human L-FABP Tyr microenvironments, T94A substitution did not alter this microenvironment.
Secondary Structure of Rat and Human L-FABP proteins
CD spectra of rat and human L-FABPs (Fig 3A) all displayed maxima and minima near 195 and 220 nm. As shown by T94T – rat L-FABP difference spectra, however, human WT T94T had less positive molar ellipticity at the 195-nm maximum and less negative molar ellipticity at the 220-nm minimum (Fig. 3B). By quantitative analysis human WT T94T had less α- helix of all types but more β-sheet of all types and more unordered structure as compared to rat L-FABP (Fig. 3C). In contrast, human T94A – T94T difference spectra had more positive molar ellipticity at the 195-nm maximum and more negative molar ellipticity at the 220-nm minimum (Fig 3D). Quantitative analysis showed that the human T94A variant L-FABP protein had significantly more α-helix of all types but less regular β-sheet and unordered structures as compared to human WT T94T L-FABP (Fig. 3E). The extent to which these differences impacted L-FABP stability, ligand binding, and conformational response to ligands was addressed below.
Figure 3. CD spectra and secondary structures of rat and human L-FABPs.
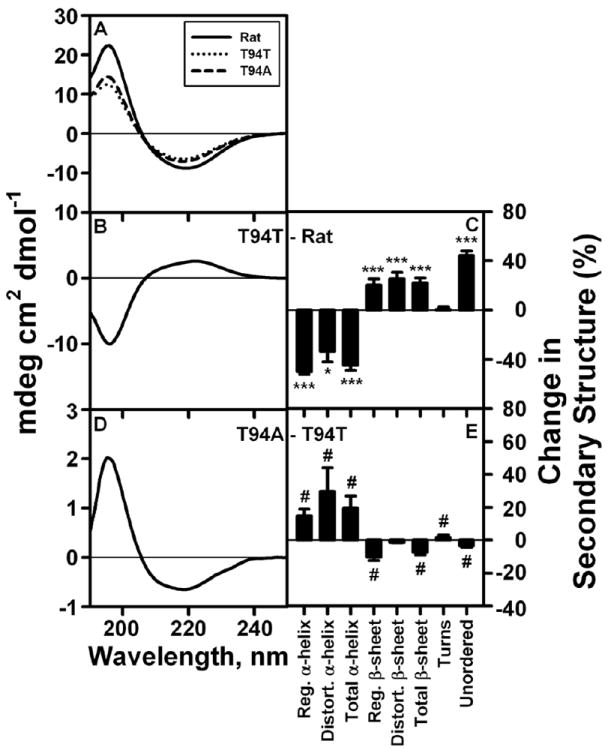
Rat and human L-FABPs were examined by CD spectroscopy and secondary structures were determined as described in Methods. Panel A: representative CD spectra of 0.5 μM rat, human T94T WT, and human T94A variant L-FABPs at 25 °C. Panel B: difference CD spectrum (human T94T WT - rat). Panel C: % change in secondary structure (human T94T WT - rat). Panel D: difference CD spectrum (T94A variant – T94T WT). Panel E: % change in secondary structure (T94A variant – T94T WT). *, P < 0.05 for human T94T vs rat secondary structure; ***, P < 0.001 for human T94T vs rat secondary structure; #, P < 0.05 for T94A vs T94T secondary structure.
Secondary Structure Stability of Rat and Human L-FABP proteins
Murine and human L-FABPs bind their respective PPARαs to induce structural alterations that alter coregulator recruitment and induce PPARα activation (12,21,22,24,58). It is not known how differences between rat and human WT T94T L-FABP and T94A variant impact folding stability that may be required for ligand transfer to and/or activation of PPARα. Rat L-FABP secondary structures were relatively stable to increasing temperature up to about 65 °C (Fig 4A, C, E, G). Human WT T94T L-FABP was more stable to unfolding, which did not occur until 85 °C (Fig 4B, D, F, H; ●). As compared to the human WT T94T L-FABP, the T94A variant was more rapidly unfolded beginning at 65 °C (Fig 4B, D, F, H; ○). Based on thermal induced increase in unordered structure, stability of the three L-FABPs was in the order: human WT T94T (Fig 4H; ●) > rat (Fig 4G) > human T94A (Fig 4H; ○). These findings with human and rat L-FABPs may contribute, in part, to the species differences in PPARα transcriptional regulation (61).
Figure 4. Sensitivity of rat and human L-FABP secondary structure to temperature induced unfolding.
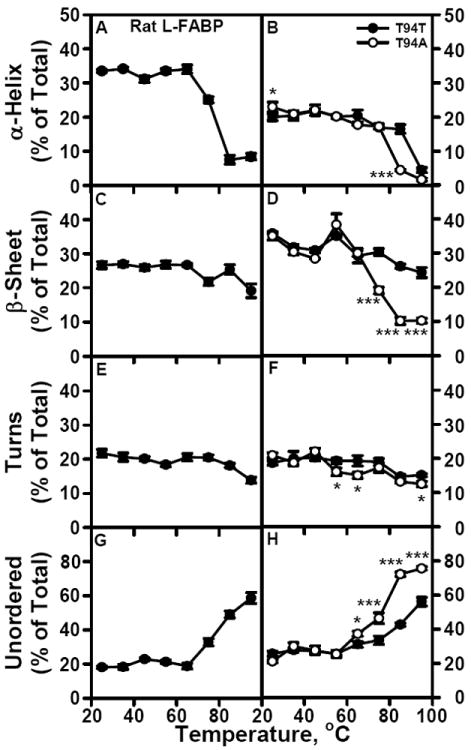
CD spectra and secondary structure analysis of rat and human L-FABPs were determined as a function of sample temperature as described in Methods. Panels A, C, E, and G show the amount of rat L-FABP total α-helix, total β-sheet, turn, and unordered secondary structure, respectively, as a percentage of the total secondary structure. Panels B, D, F. and H show the amount of human T94T WT L-FABP (●) or human T94A variant L-FABP (○) total α-helix, total β-sheet, turn, and unordered secondary structure, respectively, as a percentage of the total secondary structure. *, P < 0.05 for T94A vs T94T; ***, P < 0.001 for T94A vs T94T.
Ligand Binding Specificity of Rat, Human WT T94T, and Human T94A Variant L-FABP: ANS Fluorescence Displacement
While T94A substitution has been hypothesized to abolish ligand binding to human L-FABP, this has not been shown (56). Phytanic acid, a naturally-occurring branched-chain fatty acid from which less toxic fibrate analogues were subsequently developed (64,65), displaced ANS from all three L-FABPs (Fig 5A) with similar Ki’s (Table 1). Fenofibrate and fenofibric acid are the most commonly prescribed fibrates in the US and Canada (51). However, fibrate binding differed markedly between the three L-FABPs (Fig 5B, C; Table 1): i) rat L-FABP bound fenofibrate and fenofibric acid nearly 3- and 150-fold more weakly than phytanic acid; ii) human WT T94T and T94A variant bound fenofibrate 20-fold more strongly than fenofibric acid; iii) human WT T94T and T94A bound fenofibrate with 7-fold higher affinity than rat L-FABP; iv) human WT T94T and T94A variant bound fenofibric acid more strongly with > 20-fold higher affinity than rat L-FABP. Thus, T94A did not abolish or alter binding of phytanic acid, fenofibrate, or fenofibric acid—all potent PPARα agonists (16,66-69). Further, rat and human L-FABPs differed significantly in their ability to bind these ligands.
Figure 5. Ligand binding to rat and human L-FABPs: ANS displacement.
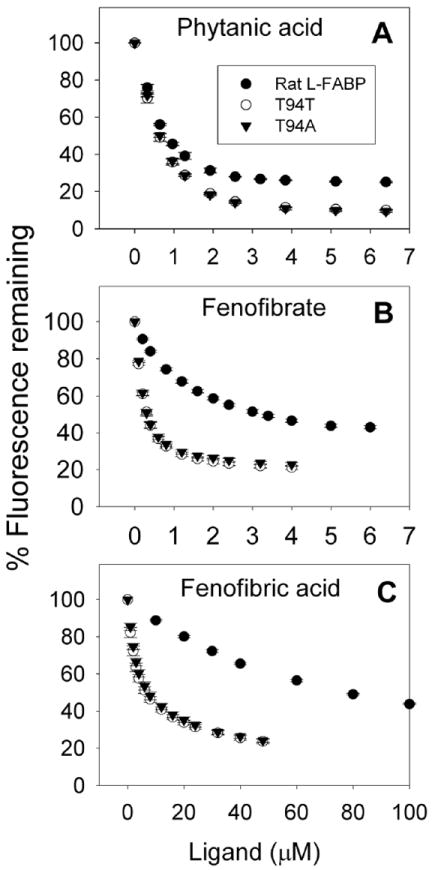
Displacement curves of phytanic acid (panel A), fenofibrate (panel B), and fenofibric acid (panel C) of L-FABP-bound ANS were obtained as described in Methods (L-FABP, 500 nM; ANS, 35 μM) (●), rat L-FABP; (○), human WT T94T L-FABP; (▼), human T94A variant L-FABP. All data presented as mean ± SE, (n = 4-5).
Table 1. Comparison of ANS displacement coefficients of phytanic acid and fibrates from murine and human L-FABPs.
Ki’s were obtained by analysis of multiple ANS displacement curves similar to those in Fig. 5 as described in Methods.
| Ligand | ANS displacement Ki (μM)a | ||
|---|---|---|---|
| Rat L-FABP | T94T | T94A | |
| Phytanic acid | 0.037 ± 0.002 | 0.035 ± 0.003 | 0.038 ± 0.002 |
| Fenofibrate | 0.10 ± 0.01† | 0.015 ± 0.001*,† | 0.015 ± 0.001*,† |
| Fenofibric acid | 5.8 ± 0.5†,# | 0.25 ± 0.03*,†,# | 0.31 ± 0.03*,†,# |
Value represent mean ± SE (n=3-5).
P<0.05 vs rat L-FABP;
P<0.05 vs phytanic acid;
P<0.05 vs fenofibrate.
Ligand Binding Differentially Altered Rat, Human WT T94T, and Human T94A Variant L-FABP Protein Secondary Structure
Ligand-induced conformational changes in L-FABP may significantly impact the α-helical portal region, interaction with PPARα, and possibly transfer of activating ligand to PPARα (12). However, the impact of ligand binding on rat L-FABP secondary structure was modest (5-10% change) and highly specific for each ligand. For example, phytanic acid stabilized the regular α-helical region of the rat L-FABP (Fig 6A) while fenofibrate and its active metabolite fenofibric acid did not (Fig 6D, G). Consistent with this finding, phytanic acid (Fig 6C), but not fenofibrate or fenofibric acid (Fig 6F, I) decreased the proportion of unordered structure in rat L-FABP. However, all three ligands similarly altered rat L-FABP β-sheet structures (Fig 6B, E, H).
Figure 6. Change in rat L-FABP secondary structure upon interaction with phytanic acid, fenofibrate, or fenofibric acid.
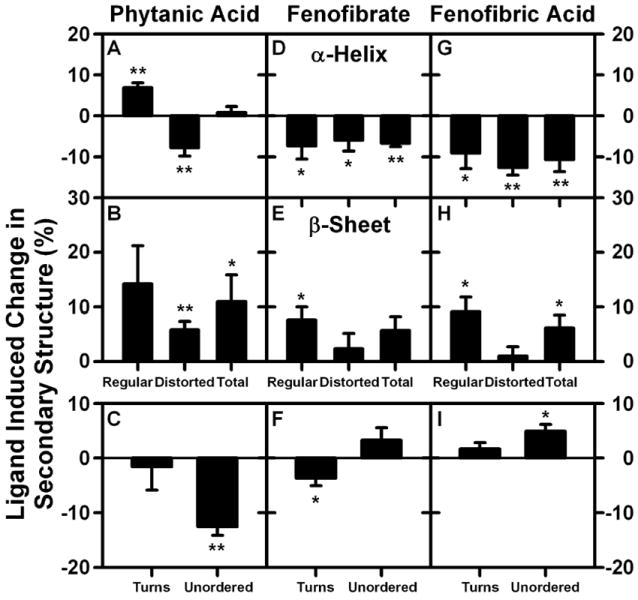
Rat L-FABP (0.5 μM), with or without 5 μM ligand was examined by CD spectroscopy and subsequent secondary structure analysis as described in Methods. Panels A (α-helix), B (β-sheet), and C (turn/unordered) show the % change in secondary structure (L-FABP/ligand – L-FABP only) induced by phytanic acid interaction with rat L-FABP. Panels D (α-helix), E (β-sheet), and F (turn/unordered) show the % change in secondary structure (L-FABP/ligand – L-FABP only) induced by fenofibrate interaction with rat L-FABP. Panels G (α-helix), H (β-sheet), and I (turn/unordered) show the % change in secondary structure (L-FABP/ligand – L-FABP only) induced by fenofibric acid interaction with rat L-FABP. *, P < 0.05 for L-FABP/ligand secondary structure vs L-FABP only secondary structure; **, P < 0.01 for L-FABP/ligand secondary structure vs L-FABP only secondary structure.
In contrast, these ligands more dramatically (10-35%) and differentially altered human WT T94T and T94A variant L-FABP secondary structures. Phytanic acid binding similarly altered the secondary structure of human WT T94T and T94A variant L-FABP (Fig 7A, B, C). In contrast, the fibrate ligands differentially altered human WT T94T L-FABP secondary structure—both quantitatively and with regards to fibrate specificity (Fig 7, black bars). For example, fenofibrate had little, if any, effect on human WT T94T L-FABP proportion of α-helix (Fig 7D), modestly increased β-sheet (Fig 7E) and turns (Fig 7F), and decreased significantly the amount of unordered (Fig 7F) structures. In contrast, the active metabolite fenofibric acid markedly increased the proportion of α-helix (Fig 7G) and turns (Fig 7I), modestly decreased β-sheet (Fig 7H), and significantly decreased the amount of unordered (Fig 7I) structures.
Figure 7. Impact of T94A substitution on human L-FABP secondary structural response to phytanic acid, fenofibrate, or fenofibric acid.
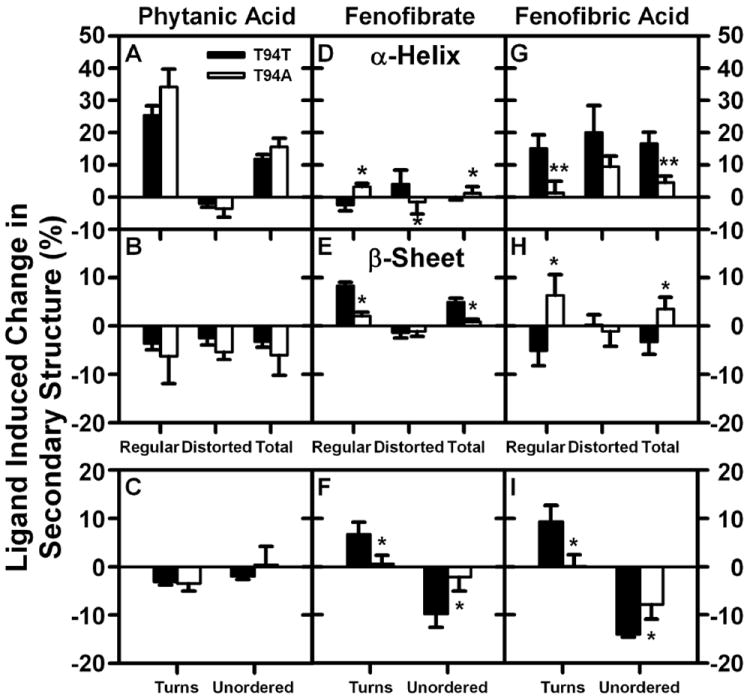
Human T94T WT and T94A variant L-FABPs (0.5 μM) were examined by CD spectroscopy and subsequent secondary structure analysis in the absence or presence of 5 μM phytanic acid, fenofibrate, or fenofibric acid as described in Methods. Panels A (α-helix), B (β-sheet), and C (turn/unordered) show the % change in secondary structure (L-FABP/ligand – L-FABP only) upon phytanic acid interaction with human L-FABP. Panels D (α-helix), E (β-sheet), and F (turn/unordered) show the % change in secondary structure (L-FABP/ligand – L-FABP only) upon fenofibrate interaction with human L-FABP. Panels G (α-helix), H (β-sheet), and I (turn/unordered) show the % change in secondary structure (L-FABP/ligand – L-FABP only) upon fenofibric acid interaction with human L-FABP. *, P < 0.05 for T94A L-FABP/ligand secondary structure vs T94T L-FABP/ligand secondary structure; **, P < 0.01 for T94A L-FABP/ligand secondary structure vs T94T L-FABP/ligand secondary structure.
The T94A substitution in the human T94A variant L-FABP significantly altered the ability of fenofibrate and fenofibric acid to impact secondary structure. As compared to its effect on the human WT T94T L-FABP, fenofibrate modestly increased proportions of α-helix (Fig 7D) and β-sheet (Fig 7E) while somewhat decreasing unordered structures (Fig 7F). The human T94A substitution decreased the ability of fenofibric acid to affect the proportion of α-helical structure (Fig 7G), increased the proportion of β-sheet (Fig 7H), and decreased unordered structures (Fig 7I).
T94A Expression Impaired Fenofibrate-mediated Transcription of PPARα-regulated Proteins in Primary Human Hepatocytes
Human T94A variant L-FABP protein exhibited altered conformational response to fenofibric acid, the active metabolite of fenofibrate (Fig 7). Therefore, the functional impact of fenofibrate was examined in cultured primary human hepatocytes that had been genotyped and segregated into homozygous WT T94T L-FABP (TT), heterozygous T94T/T94A (TC), and homozygous T94A variant (CC) L-FABP expressors as described in Methods. T94A expressing hepatocytes had impaired response to fenofibrate-mediated transcription of PPARα regulated proteins: PPARα itself, L-FABP, and FATP5 (Fig 8). Taken together, these findings indicated that the T94A substitution not only altered structure and structural response to fibrate binding, but also diminished its ability to function in mediating fibrate signaling to PPARα.
Figure 8. Effect of T94A on ligand-induced transcription of PPARα-regulated proteins.
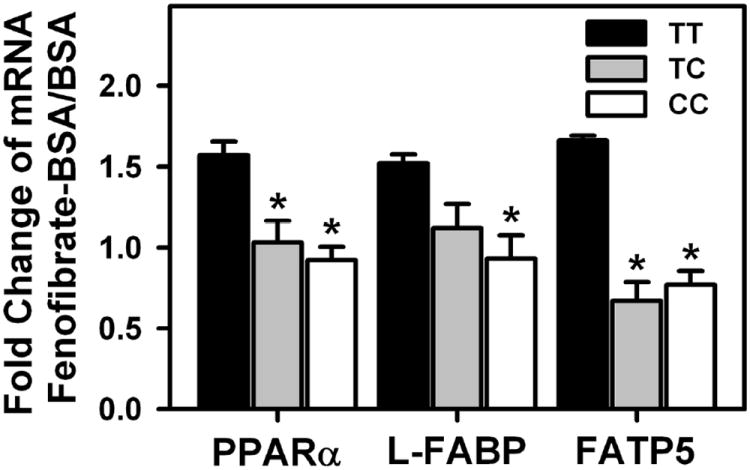
Primary human hepatocytes were treated as described in Methods and then incubated for 24 hr with fatty acid-free BSA (Alb) or BSA/fenofibrate (40 μM, FF) in 6 mM glucose-containing medium as described (30,31). Quantitative real-time (rt) PCR for human PPARα, L-FABP, and FATP5 determined mRNA levels that were normalized to an internal control (18S RNA). Values = the fold change induced by FF/alb complex relative to albumin only. All data presented as mean ± SE, (n = 8-10 in each group). *, P < 0.05, for homozygous (CC) or heterozygous (TC) T94A variants vs T94T WT (TT).
DISCUSSION
Although both murine and human liver fatty acid binding proteins (L-FABP) were cloned nearly 30 years ago, subsequent progress focused primarily on resolving the structure (10,11,19,62,70) and function [rev. in (2,3)] of the murine L-FABP. More recent studies have begun to examine the tertiary structure (14,38,39,41,42,71,72) and xenobiotic ligand (fibrate, phthalates, phenoxy herbicides) binding specificity (12,40) of the human WT T94T L-FABP. Although a highly common SNP in the human L-FABP coding sequence results in a single amino acid substitution, T94A, almost nothing is known about its impact on structure, ligand binding affinity, or function (43,44). Studies herein provide the following new insights:
First, rat and human WT T94T L-FABP differed significantly in secondary structure, structural stability, fibrate binding affinity/specificity, and conformational response. Furthermore, the human L-FABPs exhibited much greater affinity for fibrates than the rat L-FABP. In contrast, there was no difference in affinities for phytanic acid, the naturally-occurring fatty acid from which less toxic fibrate analogues were first developed (64,65). Transcriptional profiling studies also demonstrated significant species differences in mRNA induction by PPARα agonists in cultured rat, mouse, and human primary hepatocytes (59-61). These findings underscore the need for structure/activity studies with human as well as rat L-FABP.
Second, the T94A substitution in human L-FABP significantly altered the secondary structure and decreased its stability to unfolding. CD spectral analysis is less sensitive to β-sheet structure and therefore overestimates human L-FABP proportion of α-helix (Fig 4, about 20%) as compared to NMR or X-ray (about 12%) (14,62,72). Nevertheless, there are clear differences between human WT T94T L-FABP and T94A variant L-FABP proteins. The T94A substitution increased the proportion of α-helical structure and concomitantly decreased the thermal stability of the human L-FABP by nearly 10 °C from 85 °C to 65-75 °C. Interestingly, an earlier infrared spectroscopic study of native human L-FABP isolated from liver of unknown phenotype showed that at temperatures higher than 65-75°C the proportion of α-helix and β-sheet significantly decreased (73). This suggests that the latter native human L-FABP may have been isolated from an individual expressing the T94A variant L-FABP. Structural differences can impact L-FABP interaction with other proteins as evidenced by L-FABP conformers differentially enhancing bound LCFA-CoA utilization by microsomal glycerol-3-phosphate acyltransferase (GPAT) (74-76). Likewise, while single amino acid substitutions in the cytosolic domain of carnitine palmitoyl acyltransferase-1A (CPT1A, the rate limiting enzyme in fatty acid β-oxidation) do not alter ligand binding affinity, they significantly alter secondary structure and inhibit binding of L-FABP to thereby inhibit mitochondrial fatty acid β-oxidation (77).
Third, although the T94A substitution in human L-FABP did not alter the affinity for potent PPARα agonists (phytanic acid, fenofibrate, fenofibric acid), the T94A substitution diminished the ability of fibrate ligands (especially fenofibric acid) to alter the secondary structure of the human L-FABP.
Fourth, T94A expression in cultured primary human hepatocytes significantly impaired fenofibrate-mediated transcription of PPARα-regulated proteins such as FATP5. L-FABP directly interacts with FATP5 at the mouse hepatocyte plasma membrane—suggesting that this interaction may facilitate ligand uptake (78). Indeed, L-FABP overexpression enhances uptake of phytanic acid and other fatty acids while L-FABP gene ablation inhibits uptake in cultured cells, mouse hepatocytes, and in vivo (35,78-80). Whether T94A substitution impacts fenofibrate uptake is unknown, but fenofibrate has been reported to be less effective in lowering elevated plasma triglyceride to basal levels in L-FABP T94A variant human subjects (44).
In summary, T94A substitution in human L-FABP significantly altered the secondary structure, stability, conformational response to fibrate binding, and fenofibrate activation of PPARα transcriptional activity in human hepatocytes. Fibrate binding induces L-FABP redistribution into nuclei for interaction with and activation of PPARα in mouse primary hepatocytes (30). Within the nucleus, L-FABP binds PPARα (12,21,22), likely to facilitate ligand transfer (12), and induces PPARα transcription of multiple proteins in fatty acid metabolism in mouse primary hepatocytes (24,28-31,81). The net effect of these actions is to lower plasma triglyceride (69,82-89), primarily by inducing hepatic PPARα transcription of proteins in long chain fatty acid (LCFA) uptake (59,60,90,91) and β-oxidation (59,60,82) as well as extrahepatic PPARα regulated proteins in plasma VLDL triglyceride hydrolysis (59,60,92). While the impact of the human L-FABP T94A substitution on these pathways remains to be resolved, our findings of altered conformation and conformational response to fibrate binding suggest significant influence. Finally, the data presented herein indicate that the human L-FABP T94A variant represents an altered-function rather than abolition-of-function mutation. In contrast, L-FABP gene ablation completely abolishes the contribution of L-FABP to ligand binding as well as the ability of fibrates (fenofibrate, bezafibrate) to induce PPARα transcriptional activity only in murine hepatocytes (30).
Acknowledgments
Funding Information: This work was supported by the USPHS, National Institutes of Health Grants DK41402 (F.S. and A.B.K.) and DK70965 (B.P.A.).
Abbreviations
- CD
circular dichroism
- ANS
1-anilinonaphthalene-8-sulfonic acid
- FF
fenofibrate
- FA
fenofibric acid
- L-FABP
liver fatty acid binding protein or FABP1
- WT T94T
wild type human L-FABP
- L-FABP T94A
human L-FABP T94A variant
- LCFA
long chain fatty acid
- PPARα -β/δ, or -γ
peroxisome proliferator-activated receptor alpha, beta/delta, or gamma
- SNP
single nucleotide polymorphism
References
- 1.Ockner RK, Manning JA, Poppenhausen RB, Ho WK. A binding protein for fatty acids in cytosol of intestinal mucosa, liver, myocardium, and other tissues. Science. 1972;177:56–58. doi: 10.1126/science.177.4043.56. [DOI] [PubMed] [Google Scholar]
- 2.Atshaves BP, Martin GG, Hostetler HA, McIntosh AL, Kier AB, Schroeder F. Liver fatty acid binding protein (L-FABP) and Dietary Obesity. Journal of Nutritional Biochemisty. 2010;21:1015–1032. doi: 10.1016/j.jnutbio.2010.01.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Storch J, Corsico B. The emerging functions and mechanisms of mammalian fatty acid binding proteins. Annu Rev Nutr. 2008;28:18.1–18.23. doi: 10.1146/annurev.nutr.27.061406.093710. [DOI] [PubMed] [Google Scholar]
- 4.Thompson J, Reese-Wagoner A, Banaszak L. Liver fatty acid binding protein: species variation and the accomodation of different ligands. Biochim Biophys Acta. 1999;1441:117–130. doi: 10.1016/s1388-1981(99)00146-8. [DOI] [PubMed] [Google Scholar]
- 5.Miller KR, Cistola DP. Titration calorimetry as a binding assay for lipid-binding proteins. Mol Cell Biochem. 1993;123:29–37. doi: 10.1007/BF01076472. [DOI] [PubMed] [Google Scholar]
- 6.Rolf B, Oudenampsen-Kruger E, Borchers T, Faergeman NJ, Knudsen J, Lezius A, Spener F. Analysis of the ligand binding properties of recombinant bovine liver-type fatty acid binding protein. Biochim Biophys Acta. 1995;1259:245–253. doi: 10.1016/0005-2760(95)00170-0. [DOI] [PubMed] [Google Scholar]
- 7.Richieri GV, Ogata RT, Kleinfeld AM. Equilibrium constants for the binding of fatty acids with fatty acid binding proteins from adipocyte, intestine, heart, and liver measured with the flourscent probe ADIFAB. J Biol Chem. 1994;269:23918–23930. [PubMed] [Google Scholar]
- 8.Frolov A, Cho TH, Murphy EJ, Schroeder F. Isoforms of rat liver fatty acid binding protein differ in structure and affinity for fatty acids and fatty acyl CoAs. Biochemistry. 1997;36:6545–6555. doi: 10.1021/bi970205t. [DOI] [PubMed] [Google Scholar]
- 9.Wolfrum C, Borchers T, Sacchettini JC, Spener F. Binding of fatty acids and perooxisome proliferators to orthologous fatty acid binding proteins from human, murine, and bovine liver. Biochemistry. 2000;39:1469–1474. doi: 10.1021/bi991638u. [DOI] [PubMed] [Google Scholar]
- 10.He Y, Yang X, Wang H, Estephan R, Francis F, Kodukula S, Storch J, Stark RE. Solution-state molecular structure of apo and oleate-liganded liver fatty acid binding protein. Biochemistry. 2007;46:12543–12556. doi: 10.1021/bi701092r. [DOI] [PubMed] [Google Scholar]
- 11.Chuang S, Velkov T, Horne J, Wielens J, Chalmers DK, Porter CJH, Scanlon MJ. Probing fibrate binding specificity of rat liver fatty acid binding protein. J Med Chem. 2009;52:5344–5355. doi: 10.1021/jm801349e. [DOI] [PubMed] [Google Scholar]
- 12.Velkov T. Interactions between human liver fatty acid binding protein and peroxisome proliferator activated receptor drugs. PPAR Research. 2013:1–14. doi: 10.1155/2013/938401. doi.org/10.1155/2013/938401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Sharma A, Sharma A. Fatty acid induced remodeling within the human liver fatty acid binding protein. J Bio Chem. 2011;286 doi: 10.1074/jbc.M111.270165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Cai J, Lucke C, Chen Z, Qiao Y, Klimtchuk E, Hamilton JA. Solution structure and backbone dynamics of human liver fatty acid binding protein: fatty acid binding revisited. Biophys J. 2012;102:2585–2594. doi: 10.1016/j.bpj.2012.04.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Frolov A, Miller K, Billheimer JT, Cho T-C, Schroeder F. Lipid specificity and location of the sterol carrier protein-2 fatty acid binding site: A fluorescence displacement and energy transfer study. Lipids. 1997;32:1201–1209. doi: 10.1007/s11745-997-0154-5. [DOI] [PubMed] [Google Scholar]
- 16.Wolfrum C, Ellinghaus P, Fobker M, Seedorf U, Assmann G, Borchers T, Spener F. Phytanic acid is ligand and transcriptional activator of murine liver fatty acid binding protein. J Lipid Res. 1999;40:708–714. [PubMed] [Google Scholar]
- 17.Hanhoff T, Benjamin S, Borchers T, Spener F. Branched-chain fatty acids as activators of peroxisome proliferators. Eur J Lip Sci Technol. 2005;107:716–729. [Google Scholar]
- 18.Maatman RG, van Moerkerk HT, Nooren IM, van Zoelen EJ, Veerkamp JH. Expression of human liver fatty acid-binding protein in Escherichia coli and comparative analysis of its binding characteristics with muscle fatty acid-binding protein. Biochim Biophys Acta. 1994;1214:1–10. doi: 10.1016/0005-2760(94)90002-7. [DOI] [PubMed] [Google Scholar]
- 19.Chuang S, Velkov T, Horne J, Porter CJH, Scanlon MJ. Characterization of the drug binding specificity of rat liver fatty acid binding protein. J Med Chem. 2008;51:3755–3764. doi: 10.1021/jm701192w. [DOI] [PubMed] [Google Scholar]
- 20.Kanda T, Ono T, Matsubara Y, Muto T. Possible role of rat fatty acid binding proteins in the intestine as carriers of phenol and phthalate derivatives. Biochem Biophy Res Comm. 1990;168:1053–1058. doi: 10.1016/0006-291x(90)91136-g. [DOI] [PubMed] [Google Scholar]
- 21.Hostetler HA, McIntosh AL, Atshaves BP, Storey SM, Payne HR, Kier AB, Schroeder F. Liver type Fatty Acid Binding Protein (L-FABP) interacts with peroxisome proliferator activated receptor-a in cultured primary hepatocytes. J Lipid Res. 2009;50:1663–1675. doi: 10.1194/jlr.M900058-JLR200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Hostetler HA, Balanarasimha M, Huang H, Kelzer MS, Kaliappan A, Kier AB, Schroeder F. Glucose regulates fatty acid binding protein interaction with lipids and PPARa. J Lipid Res. 2010;51:3103–3116. doi: 10.1194/jlr.M005041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Lawrence JW, Kroll DJ, Eacho PI. Ligand dependent interaction of hepatic fatty acid binding protein with the nucleus. J Lipid Res. 2000;41:1390–1401. [PubMed] [Google Scholar]
- 24.Wolfrum C, Borrmann CM, Borchers T, Spener F. Fatty acids and hypolipidemic drugs regulate PPARalpha and PPARgamma gene expresion via L-FABP: a signaling path to the nucleus. Proc Natl Acad Sci. 2001;98:2323–2328. doi: 10.1073/pnas.051619898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Huang H, Starodub O, McIntosh A, Kier AB, Schroeder F. Liver fatty acid binding protein targets fatty acids to the nucleus: real-time confocal and multiphoton fluorescence imaging in living cells. J Biol Chem. 2002;277:29139–29151. doi: 10.1074/jbc.M202923200. [DOI] [PubMed] [Google Scholar]
- 26.Huang H, Starodub O, McIntosh A, Atshaves BP, Woldegiorgis G, Kier AB, Schroeder F. Liver fatty acid binding protein colocalizes with peroxisome proliferator receptor alpha and enhances ligand distribution to nuclei of living cells. Biochemistry. 2004;43:2484–2500. doi: 10.1021/bi0352318. [DOI] [PubMed] [Google Scholar]
- 27.Schroeder F, Petrescu AD, Huang H, Atshaves BP, McIntosh AL, Martin GG, Hostetler HA, Vespa A, Landrock K, Landrock D, Payne HR, Kier AB. Role of fatty acid binding proteins and long chain fatty acids in modulating nuclear receptors and gene transcription. Lipids. 2008;43:1–17. doi: 10.1007/s11745-007-3111-z. [DOI] [PubMed] [Google Scholar]
- 28.McIntosh AL, Atshaves BP, Hostetler HA, Huang H, Davis J, Lyuksyutova OI, Landrock D, Kier AB, Schroeder F. Liver type fatty acid binding protein (L-FABP) gene ablation reduces nuclear ligand distribution and peroxisome proliferator activated receptor-alpha activity in cultured primary hepatocytes. Arch Biochem Biophys. 2009;485:160–173. doi: 10.1016/j.abb.2009.03.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Huang H, McIntosh AL, Martin GG, Petrescu AD, Landrock K, Landrock D, Kier AB, Schroeder F. Inhibitors of fatty acid synthesis induce PPARa-regulated fatty acid b-oxidative enzymes: synergistic roles of L-FABP and glucose. PPAR Research. 2013:1–22. doi: 10.1155/2013/865604. Article ID 865604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Petrescu AD, McIntosh AL, Storey SM, Huang H, Martin GG, Landrock D, Kier AB, Schroeder F. High glucose potentiates liver fatty acid binding protein (L-FABP) mediated fibrate induction of PPARa in mouse hepatocytes. Biochim Biophys Acta Volume. 2013;2013:1–22. doi: 10.1016/j.bbalip.2013.05.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Petrescu AD, Huang H, Martin GG, McIntosh AL, Storey SM, Landrock D, Kier AB, Schroeder F. Impact of L-FABP and glucose on polyunsaturated fatty acid induction of PPARa regulated b-oxidative enzymes. Am J Physiol Gastrointest and Liver Phys. 2012;304:G241–G256. doi: 10.1152/ajpgi.00334.2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Weisiger RA. Cytosolic fatty acid binding proteins catalyze two distinct steps in intracellular transport of their ligands. Mol Cell Biochem. 2005;239:35–42. [PubMed] [Google Scholar]
- 33.McArthur MJ, Atshaves BP, Frolov A, Foxworth WD, Kier AB, Schroeder F. Cellular uptake and intracellular trafficking of long chain fatty acids. J Lipid Res. 1999;40:1371–1383. [PubMed] [Google Scholar]
- 34.Murphy EJ. L-FABP and I-FABP expression increase NBD-stearate uptake and cytoplasmic diffusion in L-cells. Am J Physiol. 1998;275:G244–G249. doi: 10.1152/ajpgi.1998.275.2.G244. [DOI] [PubMed] [Google Scholar]
- 35.Atshaves BP, McIntosh AL, Lyuksyutova OI, Zipfel WR, Webb WW, Schroeder F. Liver fatty acid binding protein gene ablation inhibits branched-chain fatty acid metabolism in cultured primary hepatocytes. J Biol Chem. 2004;279:30954–30965. doi: 10.1074/jbc.M313571200. [DOI] [PubMed] [Google Scholar]
- 36.Paulussen RJA, Veerkamp JH. Intracellular fatty acid-binding proteins characteristics and function. In: Hilderson HJ, editor. Subcellular Biochemistry. Plenum Press; New York: 1990. pp. 175–226. [DOI] [PubMed] [Google Scholar]
- 37.McIntosh AL, Huang H, Atshaves BP, Wellburg E, Kuklev DV, Smith WL, Kier AB, Schroeder F. Fluorescent n-3 and n-6 very long chain polyunsaturated fatty acids: three photon imaging and metabolism in living cells overexpressing liver fatty acid binding protein. J Biol Chem. 2010;285:18693–18708. doi: 10.1074/jbc.M109.079897. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Long D, Yang D. Millisecond timescale dynamics of human liver fatty acid binding protein: testing of its relevance to the ligand entry process. Biophys J. 2011;98:3054–3061. doi: 10.1016/j.bpj.2010.03.047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Long D, Yang D. Buffer interference with protein dynamics: a case study on human liver fatty acid binding protein. Biophys J. 2009;96:1482–1488. doi: 10.1016/j.bpj.2008.10.049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Carbone V, Velkov T. Interaction of phthalates and phonxy acid herbicide environmental pollutants with intestinal intracellular lipid binding proteins. Chem Res in Tox. 2013;26:1240–1250. doi: 10.1021/tx400170t. [DOI] [PubMed] [Google Scholar]
- 41.Favretto F, Assfalg M, Gallo M, Cicero DO, D’Onofrio M, Molinari H. Ligand binding promiscuity and human liver fatty acid binding protein: structural and dynamic insights from an interaction study with glycocholate and oleate. ChemBioChem. 2013;14:1807–1819. doi: 10.1002/cbic.201300156. [DOI] [PubMed] [Google Scholar]
- 42.Santambrogio C, Favretto F, D’Onofrio M, Assfalg M, Grandori R, Molinari H. Mass spectrometry and NMR analysis of ligand binding by human liver fatty acid binding protein. J Mass Spectrometry. 2013;48:895–903. doi: 10.1002/jms.3237. [DOI] [PubMed] [Google Scholar]
- 43.Robitaille J, Brouillette C, Lemieux S, Perusse L, Gaudet D, Vohl M-C. Plasma concentrations of apolipoprotein B are modulated by a gene-diet interaction effect between the L-FABP T94A polymorphism and dietary fat intake in French-Canadian men. Mol Gen and Metab. 2004;82:296–303. doi: 10.1016/j.ymgme.2004.06.002. [DOI] [PubMed] [Google Scholar]
- 44.Brouillette C, Bose Y, Perusse L, Gaudet D, Vohl M-C. Effect of liver fatty acid binding protein (FABP) T94A missense mutation on plasma lipoprotein responsiveness to treatment with fenofibrate. J Hum Gen. 2004;49:424–432. doi: 10.1007/s10038-004-0171-2. [DOI] [PubMed] [Google Scholar]
- 45.Fisher E, Weikert C, Klapper M, Lindner I, Mohlig M, Spranger J, Boeing H, Schrezenmeir J, Doring F. L-FABP T94A is associated with fasting triglycerides and LDL-cholesterol in women. Mol Gen and Metab. 2007;91:278–284. doi: 10.1016/j.ymgme.2007.03.002. [DOI] [PubMed] [Google Scholar]
- 46.Weikert MO, Loeffelholz Cv, Roden M, Chandramouli V, Brehm A, Nowotny P, Osterhoff MA, Isken F, Spranger J, Landau BR, Pfeiffer A, Mohlig M. A Thr94Ala mutation in human liver fatty acid binding protein contributes to reduced hepatic glycogenolysis and blunted elevation of plasma glucsoe levels in lipid-exposed subjects. Am J Physiol Endocrinol Metab. 2007;293:E1078–E1084. doi: 10.1152/ajpendo.00337.2007. [DOI] [PubMed] [Google Scholar]
- 47.Yamada Y, Kato K, Oguri M, Yoshida T, Yokoi K, Watanabe S, Metoki N, Yoshida H, Satoh K, Ichihara S, Aoyagi Y, Yasunaga A, Park H, Tanaka M, Nozawa Y. Association of genetic variants with atherothrombotic cerebral infarction in Japanese individuals with metabolic syndrome. Int J Mol Med. 2008;21:801–808. [PubMed] [Google Scholar]
- 48.Bu L, Salto LM, De Leon KJ, De Leon M. Polymorphisms in fatty acid binding protein 5 show association with type 2 diabetes. Diabetes Res Clin Prac. 2011;92:82–91. doi: 10.1016/j.diabres.2011.01.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Peng X-E, Wu YL, Lu Q-Q, Ju Z-J, Lin X. Two genetic variants in FABP1 and susceptibility to non-alcoholic fatty liver disease in a Chinese population. Gene. 2012;500:54–58. doi: 10.1016/j.gene.2012.03.050. [DOI] [PubMed] [Google Scholar]
- 50.Mansego ML, Martinez F, Martinez-Larrad MT, Zabena C, Rojo G, Morcillo S, Soriguer F, Martin-Escudero JC, Serrano-Rios M, Redon J, Chaves FJ. Common variants of the liver fatty acid binding protein gene influence the risk of Type 2 Diabetes and insulin resistance in Spanish population. PLoS ONE. 2012;7:e31853. doi: 10.1371/journal.pone.0031853. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Jackevicius CA, Tu JV, Ross JS, Ko DT, Carreon D, Krumholz HM. Use of fibrates in the United States and Canada. JAMA. 2012;305:1217–1224. doi: 10.1001/jama.2011.353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Lowe JB, Boguski MS, Sweetser DA, Elshourbagy N, Taylor JM, Gordon JI. Human liver fatty acid binding protein: Isolation of a full length cDNA and comparative sequence analyses of orthologous and paralogous proteins. J Biol Chem. 1985;260:3417. [PubMed] [Google Scholar]
- 53.Chan L, Wei CF, Li WH, Yang CY, Ratner P, Pownall H, Gotto AM, Jr, Smith LC. Human liver fatty acid binding protein cDNA and amino acid sequence. Functional and evolutionary implications. J Biol Chem. 1985;260:2629–2632. [PubMed] [Google Scholar]
- 54.Maatman RG, van de Westerlo EM, Van Kuppevelt TH, Veerkamp JH. Molecular identification of the liver- and the heart-type fatty acid-binding proteins in human and rat kidney. Use of the reverse transcriptase polymerase chain reaction. Biochem J. 1992;288:285–290. doi: 10.1042/bj2880285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Maatman RG, Degano M, van Moerkerk HT, Van Marrewijk WJ, Van der Horst DJ, Sacchettini JC, Veerkamp JH. Primary structure and binding characteristics of locust and human muscle fatty-acid-binding proteins. Eur J Biochem. 1994;221:801–810. doi: 10.1111/j.1432-1033.1994.tb18794.x. [DOI] [PubMed] [Google Scholar]
- 56.Gao N, Qu X, Yan J, Huang Q, Yuan HY, Ouyang D-S. L-FABP T94A decreased fatty acid uptake and altered hepatic triglyceride and cholesterol accumulation in Chang liver cells stably transfected with L-FABP. Mol Cell Biochem. 2010;345:207–214. doi: 10.1007/s11010-010-0574-7. [DOI] [PubMed] [Google Scholar]
- 57.Chao H, Zhou M, McIntosh A, Schroeder F, Kier AB. Acyl CoA binding protein and cholesterol differentially alter fatty acyl CoA utilization by microsomal acyl CoA: cholesterol transferase. J Lipid Res. 2003;44:72–83. doi: 10.1194/jlr.m200191-jlr200. [DOI] [PubMed] [Google Scholar]
- 58.Smathers RL, Galligan JJ, Shearn CT, Fritz KS, Mercer K, Ronis M, Orlicky DJ, Davidson NO, Petersen DR. Susceptibility of L-FABP -/- mice to oxidative stress in early-stage alcoholic liver. J Lipid Res. 2013;54:1335–1345. doi: 10.1194/jlr.M034892. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Richert L, Lamboley C, Viollon-Abadie C, Grass P, Hartmann N, Laurent S, Heyd B, Mantion G, Chibout S-D, Staedtler F. Effects of clofibric acid on mRNA expression profiles in primary cultures of rat, mouse, and human hepatocytes. Toxicol Appl Pharmacol. 2003;191:130–146. doi: 10.1016/s0041-008x(03)00231-x. [DOI] [PubMed] [Google Scholar]
- 60.Rakhshandehroo M, Hooiveld G, Muller M, Kersten S. Comparative analysis of gene regulation by the transcription factor PPARa between mouse and human. PLoS ONE. 2009;4:e6796. doi: 10.1371/journal.pone.0006796. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Ito Y, Nakamura T, Yanagiba Y, Ramdhan DH, Yamagishi N, Naito H, Kamijima M, Gonzalez FJ, Nakajima T. Plasticizers may activate human hepatic peroxisome proliferator activated receptor alpha less than that of a mouse but may activate constitutive androstane receptor in liver. PPAR Research. 2012;2012:11. doi: 10.1155/2012/201284. Article ID 201284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Thompson J, Winter N, Terwey D, Bratt J, Banaszak L. The crystal structure of the liver fatty acid-binding protein. J Biol Chem. 1997;272:7140–7150. doi: 10.1074/jbc.272.11.7140. [DOI] [PubMed] [Google Scholar]
- 63.Betts MJ, Russell RB. Amino Acid Properties and Conseaquences of Substitutions. In: Barnes MR, Gray IC, editors. Bioinformatics for Geneticists. 2003. pp. 289–316. [Google Scholar]
- 64.Zomer AWM, van der Burg B, Jansen GA, Wanders RJA, Poll-The BT, van der Saag PT. Pristanic acid and phytanic acid: naturally occurring ligands for the nuclear receptor peroxisome proliferator activated receptor alpha. J Lipid Res. 2000;41:1801–1807. [PubMed] [Google Scholar]
- 65.Wolfrum C, Spener F. Fatty acids as regulators in lipid metabolism. Eur J Lip Sci Technol. 2000;102:746–762. [Google Scholar]
- 66.Chapman MJ. Fibrates in 2003: therapeutic action in atherogenic dyslipidemia and future perspectives. Atherosclerosis. 2003;171:1–13. doi: 10.1016/s0021-9150(03)00156-4. [DOI] [PubMed] [Google Scholar]
- 67.Dayspring T, Pokrywka G. Fibrate therapy in patients with metabolic syndrome and diabetes mellitus. Current Atherosclerosis Reports. 2006;8:356–364. doi: 10.1007/s11883-006-0032-x. [DOI] [PubMed] [Google Scholar]
- 68.Desvergne B, Michalik L, Wahli W. Be fit or be sick: peroxisome proliferator-activated receptors are down the road. Mol Endocrinology. 2004;18:1321–1332. doi: 10.1210/me.2004-0088. [DOI] [PubMed] [Google Scholar]
- 69.Staels B, Maes M, Zambon A. Fibrates and future PPARa agonists in the treatment of cardiovascular disease. Nature Clin Pract Cardiovasc Med. 2008;5:542–553. doi: 10.1038/ncpcardio1278. [DOI] [PubMed] [Google Scholar]
- 70.Winter NS, Gordon JI, Banaszak LJ. Characterization of crystalline rat liver fatty acid binding protein produced in Escherichia coli. J Biol Chem. 1990;265:10955–10958. [PubMed] [Google Scholar]
- 71.Cai J, Lucker C, Chen Z, Klimtchuk E, Qiao Y, Hamilton JA. Human liver fatty acid binding protein: Solutoiin structure and ligand binding. Biophys J. 2009;96:600a. doi: 10.1016/j.bpj.2012.04.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Cai J, Lucke C, Qiao Y, Klimtchuk E, Hamilton JA. Solution structure and backbone dynamics of human liver fatty acid binding protein. Biophys J. 2010;98:238a. doi: 10.1016/j.bpj.2012.04.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Tanfani F, Kochan Z, Swierczynski J, Zydowo MM, Bertoli E. Structural properties and thermal stability of human liver and heart fatty acid binding proteins: a Fourier transform IR spectroscopy study. Biopolymers. 1995;36:569–577. doi: 10.1002/bip.360360503. [DOI] [PubMed] [Google Scholar]
- 74.Jolly CA, Hubbell T, Behnke WD, Schroeder F. Fatty acid binding protein: Stimulation of microsomal phosphatidic acid formation. Arch Biochem Biophys. 1997;341:112–121. doi: 10.1006/abbi.1997.9957. [DOI] [PubMed] [Google Scholar]
- 75.Jolly CA, Murphy EJ, Schroeder F. Differential influence of rat liver fatty acid binding protein isoforms on phospholipid fatty acid composition: phosphatidic acid biosynthesis and phospholipid fatty acid remodeling. Biochim Biophys Acta. 1998;1390:258–268. doi: 10.1016/s0005-2760(97)00186-0. [DOI] [PubMed] [Google Scholar]
- 76.Schroeder F, Jolly CA, Cho TH, Frolov AA. Fatty acid binding protein isoforms: structure and function. Chem Phys Lipids. 1998;92:1–25. doi: 10.1016/s0009-3084(98)00003-6. [DOI] [PubMed] [Google Scholar]
- 77.Hostetler HA, Lupas D, Tan Y, Dai J, Kelzer MS, Martin GG, Woldegiorgis G, Kier AB, Schroeder F. Acyl-CoA binding proteins interact with the acyl-CoA binding domain of mitochondrial carnitine palmitoyltransferase I. Mol Cell Biochem. 2011;355:135–148. doi: 10.1007/s11010-011-0847-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Storey SM, McIntosh AL, Huang H, Martin GG, Landrock KK, Landrock D, Payne HR, Kier AB, Schroeder F. Loss of intracellular lipid binding proteins differentially impacts saturated fatty acid uptake and nuclear targeting in mouse hepatocytes. Am J Physiol Gastrointest and Liver Phys. 2012;303:G837–G850. doi: 10.1152/ajpgi.00489.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Atshaves BP, Storey SM, Petrescu AD, Greenberg CC, Lyuksyutova OI, Smith R, Schroeder F. Expression of fatty acid binding proteins inhibits lipid accumulation and alters toxicity in L-cell fibroblasts. Am J Physiol. 2002;283:C688–C703. doi: 10.1152/ajpcell.00586.2001. [DOI] [PubMed] [Google Scholar]
- 80.Martin GG, Danneberg H, Kumar LS, Atshaves BP, Erol E, Bader M, Schroeder F, Binas B. Decreased liver fatty acid binding capacity and altered liver lipid distribution in mice lacking the liver fatty acid binding protein (L-FABP) gene. J Biol Chem. 2003;278:21429–21438. doi: 10.1074/jbc.M300287200. [DOI] [PubMed] [Google Scholar]
- 81.Wolfrum C, Buhlman C, Rolf B, Borchers T, Spener F. Variation of liver fatty acid binding protein content in the human hepatoma cell line HepG2 by peroxisome proliferators and antisense RNA affects the rate of fatty acid uptake. Biochim Biophys Acta. 1999;1437:194–201. doi: 10.1016/s1388-1981(99)00008-6. [DOI] [PubMed] [Google Scholar]
- 82.Oosterveer MH, Grefhourst A, van Dijk TH, Havinga R, Staels B, Kuipers F, Groen AK, Reijngoud D-J. Fenofibrate simultaneously induces hepatic fatty acid oxidation, synthesis, and elongation in mice. J Biol Chem. 2009;284:34036–34044. doi: 10.1074/jbc.M109.051052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Sekiya M, Yhagi N, Matsuzaka T, Najima Y, Nakakuki M, Nagai R, Ishibashi S, Osuga J-I, Yamada N, Shimano H. Polyunsaturated fatty acids ameliorate hepatic steatosis in obese mice by SREBP-1 suppression. Hepatology. 2003;38:1529–1539. doi: 10.1016/j.hep.2003.09.028. [DOI] [PubMed] [Google Scholar]
- 84.Froyland L, Madsen L, Vaagenes H, Totland GK, Auwerx J, Kryvi H, Staels B, Berge RK. Mitochondrion is the principal target for nutritional and pharmacological control of triglyceride metabolism. J Lipid Res. 1997;38:1851–1858. [PubMed] [Google Scholar]
- 85.Bijland S, Pieterman EJ, Maas ACE, van der Hoorn JWA, van Erk MJ, van Klinken JB, Havekes LM, van Dijk KW, Princen HM, Rensen PCN. Fenofibrate increases VLDL triglyceride production despite reducing plasma triglyceride levels in APOE3-Leiden.CETP mice. J Biol Chem. 2010;285:25168–25175. doi: 10.1074/jbc.M110.123992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Robins SJ. Fibrates and coronary heart disease reduction. Curr Op in Endocrin & Diabetes. 2002;9:312–322. [Google Scholar]
- 87.Saha SA, Arora RR. Fibrates in the prevention of cardiovascular disease in pateints with type 2 diabetes mellitus--A pooled meta-analysis of randomized placebo-controlled clinical trials. Int J Cardiol. 2010;141:157–166. doi: 10.1016/j.ijcard.2008.11.211. [DOI] [PubMed] [Google Scholar]
- 88.Frederiksen KS, Wulf EM, Wassermann K, Sauerberg P, Fleckner J. Identification of hepatic transcriptional changes in insulin-resistant rats treated with peroxisome proliferator activated receptor-alpha agonists. J Mol Endocrinol. 2003;30:317–329. doi: 10.1677/jme.0.0300317. [DOI] [PubMed] [Google Scholar]
- 89.Otvos JD, Collins D, Freedman DS, Shaluarova I, Schaefer EJ, McNamara JR, Bloomfield HE, Robins SJ. Low-density lipoprotein and high-density lipoproteion particle subclasses predict coronary events and are favorably changed by gemfibrozil therapy in the Veterans Affairs high density lipoprotein intervention trial. Circulation. 2006;113:1556–1563. doi: 10.1161/CIRCULATIONAHA.105.565135. [DOI] [PubMed] [Google Scholar]
- 90.Heinaniemi M, Uski JO, Degenhart T, Carlberg C. Meta-analysis of primary target genes of peroxisome proliferator-activated receptors. Genome Biology. 2007;8:R147. doi: 10.1186/gb-2007-8-7-r147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Rakhshandehroo M, Knoch B, Muller M, Kersten S. Peroxisome proliferator-activated receptor alpha target genes. PPAR Research 2010. 2010 doi: 10.1155/2010/612089. Article ID 612089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Atshaves BP, Payne HR, McIntosh AL, Tichy SE, Russell D, Kier AB, Schroeder F. Sexually dimorphic metabolism of branched chain lipids in C57BL/6J mice. J Lipid Res. 2004;45:812–830. doi: 10.1194/jlr.M300408-JLR200. [DOI] [PubMed] [Google Scholar]


