Abstract
Cholesterol is an important lipid of mammalian cells and plays a fundamental role in many biological processes. Its concentration in the various cellular membranes differs and is tightly regulated. Here, we present a novel alkyne cholesterol analog suitable for tracing both cholesterol metabolism and localization. This probe can be detected by click chemistry employing various reporter azides. Alkyne cholesterol is accepted by cellular enzymes from different biological species (Brevibacterium, yeast, rat, human) and these enzymes include cholesterol oxidases, hydroxylases, and acyl transferases that generate the expected metabolites in in vitro and in vivo assays. Using fluorescence microscopy, we studied the distribution of cholesterol at subcellular resolution, detecting the lipid in the Golgi and at the plasma membrane, but also in the endoplasmic reticulum and mitochondria. In summary, alkyne cholesterol represents a versatile, sensitive, and easy-to-use tool for tracking cellular cholesterol metabolism and localization as it allows for manifold detection methods including mass spectrometry, thin-layer chromatography/fluorography, and fluorescence microscopy.
Keywords: click reaction, analog, probe, oxysterols, alkyne lipid
Cholesterol is the major sterol of mammalian cell membranes (1). Beyond its role for discrete membrane structures such as caveolae (2) or microdomains (3), cholesterol also specifically interacts with numerous proteins (4, 5). Cholesterol metabolites are involved in signaling (6), transport (7), and lipid storage (8). Dysregulation of cholesterol levels causes a wide range of human diseases (9–11). Hence, mammalian cells tightly regulate sterol biosynthesis, turnover, and transport (1, 7, 12). To follow the cellular fate of cholesterol by fluorescence microscopy, various fluorescent analogs (13–16), sterol binding toxins (17), and anti-cholesterol antibodies (18) have been used. Isotope-labeled cholesterol (1) and isotope-labeled toxins (19) have been employed to follow metabolism or organization of cholesterol in membranes, respectively. A probe suitable for conveniently tracing both cholesterol metabolism and localization would be of great value.
With the advent of bioorthogonal chemistry (20), including click chemistry (21), the sensitive and specific detection of compounds containing azido groups or terminal alkynes has become possible. Lipids containing an alkyne moiety have been used to trace their metabolism and distribution (22–26) and to monitor protein lipidation (27–30) or protein-lipid interaction (31). Thus far, no alkyne-bearing analog of cholesterol has been used to trace cholesterol in mammalian cells.
Here we report on the synthesis and use of alkyne cholesterol to study cholesterol metabolism by in vitro assays, and in vivo by using various cells. These data are supplemented by cellular localization studies of the probe.
MATERIALS AND METHODS
Filipin and azide-PEG3-biotin conjugate were obtained from Sigma. Borondifluorodipyrromethene (Bodipy)-cholesterol [23-(dipyrrometheneboron-difluoride)-24-norcholesterol] was from Avanti. Deuterated d6-cholesterol, cholesterol-26,26,26,27,27,27-2H6, and 4-cholesten-3-one were from C/D/N Isotopes and Steraloids, respectively. LD540 was described before (32). Antibodies against calnexin (Stressgen, SPA-860), EGFR (Epitomics, 2235-1), GM130 (Epitomics, 1837-1), and HSP60 (Stressgen, SPA-806) were used.
Plasmids
pmRFP1-mito codes for the native presequence (MLRAALSTARRGPRLSRLL) of rat alcohol dehydrogenase (ALDH) followed by the first 17 amino acids of ALDH (SAAATSAVPAPNQQPEV), a linker (6 amino acids; LVPVAT), and mRFP1. pmRFP1-Golgi and pmRFP1-endoplasmic reticulum (ER) are described elsewhere (33). All constructs were sequence-verified and added to the lab database.
Chemical synthesis
The syntheses of alkyne cholesterol (Fig. 1) and the azido-reporter molecules are described in the supplementary text. Data from analyses of alkyne cholesterol by TLC, MS, and NMR spectroscopy are provided (supplementary Fig. I).
Fig. 1.
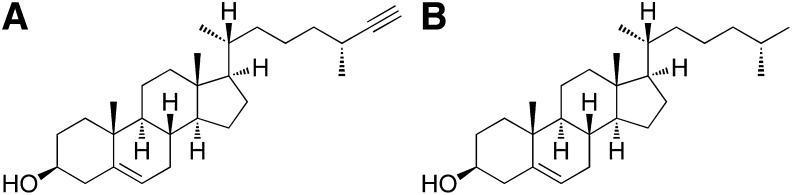
Schematic representation of alkyne cholesterol, (25R)-25-ethinyl-26-nor-3β-hydroxycholest-5-en (A) and cholesterol (B).
Generation of a Δhem1 yeast strain.
A Saccharomyces cerevisiae strain lacking HEM1 (delta-aminolevulinic acid synthase) was created (34). For that, the stain W303 (a gift from Simon Alberti: ade2-1; trp1-1; ura3-1; leu2-3,112; his3-11,15; can1-100) was transformed with the deletion plasmid pFA6a-KanMX6 (35) containing the targeted sequence of hem1 (hem1::KanMX6), introduced after performing a PCR using the following primers (GTT GTC CCT CAA TAA TCA TAA CAG TAC TTA GGT TTT TTT TTC AGT CGG ATC CCC GGG TTA ATT AA and AAA TTC CTT GTA CCT CTA TCT CAG CCC ATG CAT ATA TTG GTT GTT GAA TTC GAG CTC GTT TAA AC). The genotype of the resulting Δhem1 strain was verified by PCR.
Mammalian cell culture.
Human A172 glioblastoma cells (ATCC®; CRL-1620) were maintained in DMEM (Gibco) containing 10% (v/v) FCS. For primary cultures P0 to P2 Wistar rats (Charles River) were euthanized. The cortex was trypsinized at 37°C for 15 min and cells plated after mechanical dissociation. Cells were grown in DMEM containing 10% (v/v) FCS, Mito® plus serum extender (BD), and Pen/Strep for 10 days to obtain mixed glial cultures that were subsequently separated into astrocytes and oligodendrocyte precursor cells (OPCs). For this, adherent cells were shaken on an orbital shaker at 240 rpm for 20 h (36). Adherent astrocytes were propagated further, while detached OPCs were replated onto uncoated dishes and incubated in OPC medium [Neurobasal-A (Gibco), B27 supplement (Gibco), glutamine, and Pen/Strep] for 30 min. OPCs were retrieved from the supernatant, plated onto ornithine-coated dishes, and maintained in OPC medium containing 20 ng/ml FGF-basic-PDGF-AA (PeproTech) for 2 days. To differentiate OPCs to oligodendrocytes, fresh OPC medium containing 20 ng/ml triiodothyronine (Sigma) was applied daily for 5 days. For primary neurons, hippocampi from the above rats were papain-digested at 37°C for 1 h before adding trypsin inhibitor (Sigma) for 15 min and mechanical dissociation. Cells were plated onto poly-d-lysine-coated dishes and incubated in OPC medium for 2 days. All cells were maintained at 37°C and 5% (v/v) CO2.
Mammalian cell transfection.
To develop an improved protocol of transfection of human A172 glioblastoma cells by cationic N1,N4-dioleyl-N1,N4-di-[2-hydroxy-3-(N-aminopropyl)]diaminobutane (37) without manipulating the cellular sterol balance, an equimolar formulation with dioleoylphosphatidylethanolamine was prepared, mixed with 4.5 parts of DNA, and used. Cells were transferred to Opti-MEM (Gibco, 31985) and incubated with the transfection mix for 2–3 h.
Yeast culture.
Cells (Δhem1) were maintained in complete synthetic medium (CSM) (MP Biomedicals) containing 0.01% (w/v) 5-aminolevulinic acid (ALA) and 200 μg/ml G418. For feeding experiments, cells were transferred to CSM containing G418 to deplete cellular ALA pools and precultured for 12 h (34). This starter culture was split into equal aliquots and the growth media were supplemented with 0.5% (v/v) Tween80, 5.6 μg/ml methionine, and 30 μM sterol or ALA. Cells were incubated aerobically with shaking at 30°C and growth was monitored using spectroscopy (optical density at 600 nm) for 22 h.
Sterol labeling.
A172 cells were grown in 3 cm dishes in DMEM containing FCS or delipidated FCS (38). Lipids were delivered by supplementing the respective fresh growth media with sterol from an ethanol stock solution to a final concentration of 10 μg/ml.
Cell collection and lipid extraction for TLC analysis.
Cells incubated with sterols were washed twice with PBS containing 10 mg/ml fatty acid-free BSA (Applichem) and once with PBS. Cells were scraped in 155 mM ammonium acetate and lipid extraction was performed as described (39), but the pH of the aqueous phase was not adjusted.
Click reaction, lipid analysis by TLC, and imaging of TLC plates.
The dry lipid pellet was redissolved by addition of 7 μl chloroform, followed by addition of 30 μl click reaction mixture (5 μl of 44.5 mM 3-azido-7-hydroxycoumarin, 500 μl of 10 mM [acetonitrile]4CuBF4 in acetonitrile, 2 ml ethanol). Click reaction, TLC separation, and imaging were performed as described (24). Lipid quantification by fluorometry and colorimetry was performed using a GelPro analyzer (Media Cybernetics) or ImageGaugeV3.3 software (Fuji), respectively.
Lipid analysis by mass spectrometry.
Details are provided in the supplementary text.
Cholesterol oxidase assay (in vitro).
Purified cholesterol oxidase from Brevibacterium sp. (Sigma, C8868) was dissolved in phosphate buffer (50 mM KH2PO4 (pH 7.5), 1 mg/ml lipid-free BSA) and stored in aliquots at −80°C. Incubation time, buffer composition, and the amount of enzyme in the assay were optimized to ensure linearity of the reaction rate in the kinetic studies. Alkyne cholesterol was incubated at 25°C with 2 ng enzyme in glass vials containing a total volume of 100 μl phosphate buffer supplemented with 0.1% (v/v) Triton X-100 while shaking (1,100 rpm) for 5 min. The reaction was stopped with 500 μl choloroform/methanol 3:1 (v/v), the vials centrifuged (500 g, 5 min), and the lower phase transferred to a reaction tube. The aqueous phase was washed with 200 μl chloroform and the combined chloroform phases were evaporated. Lipids were redissolved in 7 μl chloroform and reacted as above, but using a triple-concentrated click reaction mix (15 μl of 44.5 mM 3-azido-7-hydroxycoumarin, 500 μl of 10 mM [acetonitrile]4CuBF4 in acetonitrile, 2 ml ethanol). Detection and quantification of the signal from TLC plates, as well as the calculation of the Michaelis-Menten constant, Km, were performed as described previously (25).
Cholesterol oxidase assay (in vivo).
Cells incubated for 16 h in medium containing delipidated serum and equal amounts of alkyne cholesterol and deuterated d6-cholesterol (total 10 μg/ml) were chased in medium lacking lipids for 1 h to achieve a steady-state distribution of the provided sterols. Samples were either fixed for 1 h using glutaraldehyde to arrest cellular lipid transport or left untreated. All samples, living or fixed, were incubated with cholesterol oxidase in 310 mM sucrose and 0.5 mM Na-phosphate (pH 7.5) at 37°C for 30 min (40) before lipid extraction. Extracts were divided, deuterated sterols were quantified by MS, and alkyne sterols were quantified by MS and also by TLC.
Click reaction, lipid detection by fluorescence microscopy.
Cells incubated with medium containing delipidated FCS and 10 μg/ml alkyne cholesterol for 16 h were washed in PBS and fixed using 3.7% formalin in buffer A [0.1 M HEPES/KOH (pH 7.4)] for at least 16 h. Cells were washed once with 155 mM ammonium acetate and twice with buffer A. For the detection, 50 μM of the azido-reporter were dissolved in prewarmed buffer A and added to the samples. The click reaction was initiated by addition of 2 mM [acetonitrile]4CuBF4 in acetonitrile (final 2% acetonitrile) and performed at 43°C for 30 min while gently agitating. Samples were extensively washed using buffer A. If applying, samples were further incubated with fluorescent streptavidin, LD540, or 100 μg/ml freshly dissolved filipin, and mounted.
Filipin binding assay.
Liposomes from pure POPC were prepared by the reverse-phase evaporation method using ether and PBS solvents. The UV-absorbance of filipin in PBS containing the above liposomes was recorded before and after addition of 125 μM sterol or ethanol carrier to the mix using a Shimadzu UV-2450PC spectrophotometer.
Subcellular fractionation.
Cells were washed, scraped, homogenized (EMBL cell cracker; 7 strokes, 16 μm clearance), and centrifuged to obtain a postnuclear supernatant. The postnuclear supernatant was loaded onto a continuous sucrose gradient (1.4–0.3 M) and centrifuged at 110,000 g using a SW41 rotor for exactly 15 min (41). From the top, 12 fractions of 1 ml each were collected and lipids and proteins separated. Before performing lipid quantification by TLC as above (24), alkyne-phosphatidylcholine was added to the extracts as internal standard. The protein content was analyzed by Western blotting for the distribution of marker proteins of various cell organelles.
Microscopy.
Epifluorescence microscopy was performed using a Zeiss Observer.Z1 microscope (Carl Zeiss) equipped with a Plan-Apochromat 63× (1.40 NA) DIC and Fluar 40× (1.30 NA) and a Photometrics Coolsnap K4 camera. If applying, optical sectioning was performed using the apotome mode. The light source was a Polychrome V 150 W xenon lamp (Till Photonics). All images were processed employing Fiji (42) and Adobe Photoshop 6.0 (Adobe) software. Projections of z-stacks were calculated by summarizing corresponding pixel values.
RESULTS
Alkyne cholesterol rescues sterol auxotroph yeast
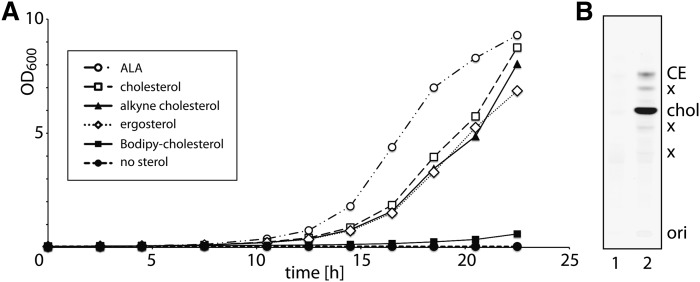
To trace cellular metabolism, we have synthesized an analog of cholesterol, alkyne cholesterol (Fig. 1, supplementary Fig. I). The alkyne moiety comprises the terminal part of the side chain and thus results in an analog of pronounced structural similarity to natural cholesterol. To test the bio-compliance of alkyne cholesterol, we generated a sterol auxotroph yeast strain lacking the hem1 gene (34). Under heme-depleted conditions, these cells are unable to synthesize sterols and depend on exogenous sources to meet the various demands and enable growth. We provided ergosterol, cholesterol, alkyne cholesterol, or ALA, the metabolic product of Hem1p, delta-aminolevulinic acid synthase to these cells and measured their growth (Fig. 2A). Alkyne cholesterol, as well as the natural sterols and ALA rescued the auxotroph, while an analog carrying a Bodipy-fluorophore in the side chain did not allow for rapid cell expansion (Fig. 2A). In our experiments yeast cells propagated by cell division, not fusion (43), and enzymatically converted alkyne cholesterol to the corresponding esters during growth (Fig. 2B). These data illustrate that alkyne cholesterol can substitute natural sterol in these cells.
Fig. 2.
A yeast sterol auxotroph (hem1) is rescued by and converts alkyne cholesterol. Before the experiments, the cellular pools of ALA were depleted by preincubating the cells in CSM lacking sterols or ALA. Subsequently cells were cultured in CSM supplemented with Tween80 as a source for oleic acid and 0.03 mM sterol or ALA. A: Cells were shaken aerobically at 30°C and growth was monitored for 22 h. The optical density at 600 nm (OD600) is plotted over time. B: Cellular alkyne sterols were analyzed by TLC after incubation with cholesterol (lane 1) or alkyne cholesterol (lane 2) for 22 h. The sterol auxotroph yeast metabolically converts alkyne cholesterol (chol) to sterol esters (CE). X denotes unidentified lipids, potentially oxysterols, for which we have not yet synthesized an according lipid standard.
Alkyne cholesterol mimics cholesterol in in vitro and in vivo enzymatic assays
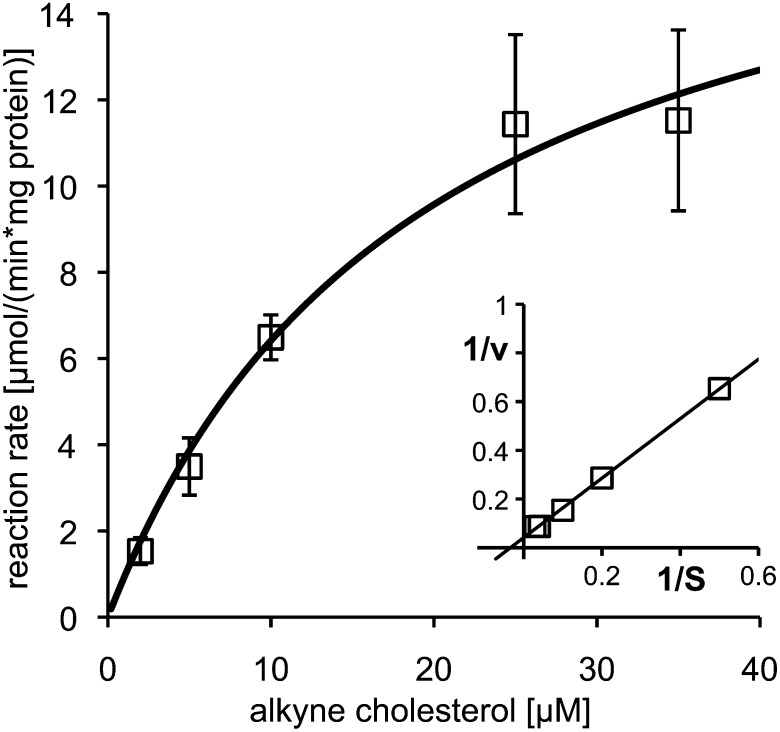
Next, we assayed the conversion of alkyne cholesterol to the corresponding cholest-4-en-3-one by purified cholesterol oxidase using our detection routine for alkyne lipids (24), recently optimized for in vitro assays (25). Cholesterol oxidase from Brevibacterium sp. accepted alkyne cholesterol as a substrate and the enzymatic reaction followed Michaelis-Menten kinetics (Fig. 3). The kinetic constant, Km, was determined as 19.4 μM by nonlinear regression.
Fig. 3.
Purified cholesterol oxidase converts alkyne cholesterol into the corresponding alkyne cholest-4-en-3-one. Michaelis-Menten kinetics were measured and the reaction rate directly (main) or double-reciprocally (insert) plotted against the substrate (S) concentration. Line graphs show the reaction rates calculated from the Michaelis-Menten equation using nonlinear (main: Km = 19.4 μM) or linear regression (insert). The experiment was performed in triplicate; error bars are shown in the direct plot only.
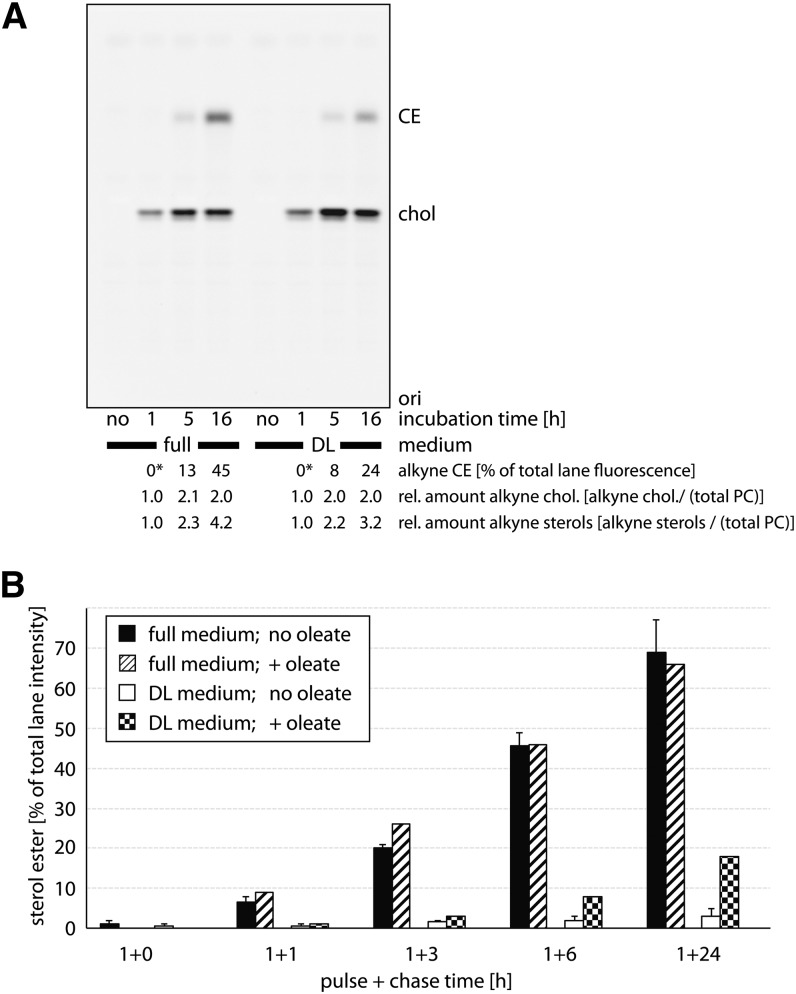
These enzymatic measurements were complemented by studies in living mammalian cells analyzing the conversion of alkyne cholesterol to alkyne cholesterol esters. Cultured human A172 glioblastoma cells were incubated with alkyne cholesterol and analyzed for cellular alkyne metabolites by click reaction and TLC (Fig. 4). Cells growing in regular culture medium took up the alkyne cholesterol and its cellular levels saturated after 5 h of incubation (Fig. 4A). Also, cells incubated in medium with delipidated FCS reached a plateau in alkyne cholesterol content between 5 and 16 h, but contained only half the amount of alkyne cholesterol esters after 16 h compared with cells in regular medium. At this time, the latter had esterified nearly half of the alkyne cholesterol. The total content of alkyne sterols, free and esterified, increased throughout the experiment for both feeding conditions. The rate of esterification in the different growth media was studied in more detail in pulse-chase experiments (Fig. 4B). After pulse-incubation of 1 h with alkyne cholesterol, hardly any esters could be detected. Cells chased in regular medium for up to 24 h progressively converted the absorbed alkyne cholesterol into esters reaching final levels of nearly 70%, a level that could not be further increased by supplementing the chase media with oleic acid to drive sterol ester formation. On the contrary, cells in delipidated medium accumulated levels of 4% or, in the presence of increased oleic acid levels, of 18% of alkyne cholesterol esters. The biosynthesis of sterol esters was also studied in primary cells of the rat brain (supplementary Fig. II). When incubated with alkyne cholesterol in the respective growth medium, astrocytes (panel A) and hippocampal neurons (panel B) were found to generate alkyne cholesterol esters to different extents, while oligodendrocytes (panel C) did not accumulate detectable amounts during the course of the experiments. After 20 h, astrocytes showed about 10-fold higher levels of alkyne cholesterol esters than neurons. After extended incubation times, oligodendrocytes, neurons, and astrocytes contained unidentified alkyne cholesterol metabolites, some of which potentially correspond to alkyne oxysterols synthesized by these cells (6).
Fig. 4.
Cellular acyltransferases synthesize alkyne cholesterol esters from alkyne cholesterol in A172 glioblastoma cells. Cells were grown in medium containing regular FCS (full) or delipidated FCS (DL) for 16 h prior to incubation with the respective medium containing 12 μM of alkyne cholesterol. A: After the indicated times, lipids of washed cells were extracted and analyzed by TLC for fluorescent metabolites, which were identified by comigrating lipid standards. Relative amounts of cellular alkyne cholesterol esters (CE) and alkyne cholesterol (chol) were determined by fluorography; total phosphatidylcholine was determined colorimetrically from the same TLC plates after charring. Asterisks denote lipid amounts below the detection limit. ori, origin. B: After 1 h of incubation, cells were washed twice with the respective medium containing delipidated BSA and incubated with fresh chase medium supplemented or not with 100 μM oleic acid for the indicated chase times. Lipids were analyzed as above. Data are mean ± range, n = 2 for solid colors, or single determinations.
To investigate the influence of alkyne cholesterol on cellular sterol balance, cells were incubated with alkyne cholesterol, cholesterol, or carrier in medium with delipidated FCS for 48 h and cellular sterols were analyzed by MS (Table 1). Compared with controls without any sterol supplement, cells showed reduced levels of latho-, lano-, and 24-dihydrolanosterol, all important precursors in cellular sterol biosynthesis, upon incubation with alkyne cholesterol. This finding is expected, as cholesterol feeding results in a downregulation of its endogenous synthesis. The cellular levels of cholestanol, an enzymatic product of cholesterol, followed the changes of the cholesterol concentration under the different feeding conditions. When feeding alkyne cholesterol, our experimental conditions yielded a ratio of alkyne cholesterol to cholesterol of 1:4.2. The total amount of cholesterol, originating from exogenous and endogenous sources, was increased if the medium was supplemented with either sterol. We also determined the cellular levels of oxysterols under the different feeding conditions and compared wild-type cells with cells transiently overexpressing cholesterol-24-hydroxylase CYP46A1 (Table 2). Increased amounts of the CYP46A1 enzyme resulted in a profound accumulation of the 24-hydroxylation products of both cholesterol and alkyne cholesterol that were identified by GLC-MS (supplementary Fig. III). Basal levels of 24-OH-alkyne cholesterol were also detectable in wild-type cells, indicating a basal expression of CYP46A1 in these cells.
TABLE 1.
Analysis of sterol metabolites in glioblastoma cells
| Alkyne Cholesterol (μg/mg) | Cholesterol (μg/mg) | Lanosterol (ng/mg) | 24-Dihydro-lanosterol (ng/mg) | Lathosterol (μg/mg) | Desmosterol (μg/mg) | Cholestanol (μg/mg) | |
| No supplement | ND | 13.80 ± 2.32 | 25.23 ± 3.82 | 9.52 ± 4.63 | 0.148 ± 0.002 | 0.038 ± 0.006 | 0.055 ± 0.016 |
| Plus alkyne chol | 3.04 ± 0.23 | 12.74 ± 0.92 | 12.07 ± 3.03 | 7.36 ± 4.15 | 0.074 ± 0.015 | 0.021 ± 0.002 | 0.044 ± 0.009 |
| Plus cholesterol | ND | 19.98 ± 2.43 | 8.64 ± 0.62 | 4.27 ± 0.85 | 0.074 ± 0.021 | 0.028 ± 0.007 | 0.170 ± 0.125 |
Glioblastoma cells were cultivated in medium containing delipidated FCS and supplemented with alkyne cholesterol, cholesterol, or carrier. Cellular sterol lipids were analyzed by GLC-MS. chol, cholesterol; ND, not determined. Data are mean ± SEM, n = 3.
TABLE 2.
Analysis of oxysterols in glioblastoma cells
| Wild Type |
+ CYP46A1 |
|||||
| No Supplement (ng/mg) | Plus Alkyne Cholesterol (ng/mg) | Plus Cholesterol (ng/mg) | No Supplement (ng/mg) | Plus Alkyne Cholesterol (ng/mg) | Plus Cholesterol (ng/mg) | |
| 24-OH-cholesterol | 4.17 ± 0.28 | 1.28 ± 0.15 | 2.70 ± 0.82 | 9.68 ± 3.14 | 3.06 ± 1.29 | 10.11 ± 5.26 |
| Ratio: 24-OH-alkyne chol ×103/d4-24-OH-cholesterol | ND | 14.23 ± 6.67 | ND | ND | 24.17 ± 3.90 | ND |
Glioblastoma cells were cultivated in medium containing delipidated FCS and supplemented with alkyne cholesterol, cholesterol, or carrier. Cells transiently expressing CYP46A1 were also examined. Cellular sterol lipids were analyzed by GLC-MS. chol, cholesterol; ND, not determined. Data are mean ± SEM, n = 3.
Alkyne cholesterol and cellular metabolites can be visualized by fluorescence microscopy
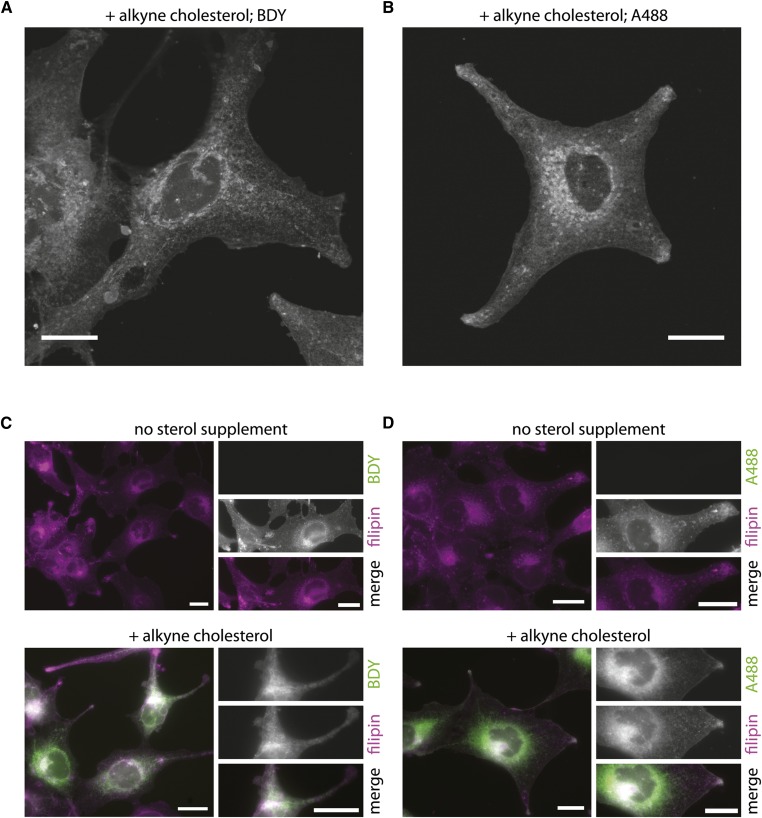
An important feature of alkyne cholesterol is its applicability to microscopy. Using the same analog, this allows for parallel investigations on cholesterol metabolism and distribution. To localize alkyne cholesterol by fluorescence microscopy in fixed cells, we developed a staining protocol employing different azido reporters. Importantly, the use of these detection agents allows for a great choice of bright photostable fluorophores with different spectral properties. In our microscopy studies, we detected either cellular alkyne cholesterol after click reaction with an azide coupled to a fluorescent Bodipy-dye (Fig. 5A) or an azide coupled to biotin and subsequent incubation with a fluorescent streptavidin-conjugate (Fig. 5B). With both approaches, we observed intense alkyne cholesterol staining at the plasma membrane and internal organelles. Parallel probing for free sterol using the UV-excitable filipin in a separate channel is possible and full channel separation could be achieved (Fig. 5C, D). Compared with cells in delipidated medium without any sterol supplement, the incubation with 10 μg/ml alkyne cholesterol increased the intensity of the cellular filipin signal and internal membranes showed a more pronounced labeling (Fig. 5C, D; supplementary Fig. IVA, D). This prompted us to investigate the binding of filipin to alkyne cholesterol using an in vitro approach, assaying the known spectral shift of the filipin absorbance upon sterol interaction (44). We found that filipin binds to both cholesterol and alkyne cholesterol, but could not detect an interaction with cholesteryl stearate (supplementary Fig. IVC). To characterize the subcellular distribution of alkyne cholesterol in more detail, we performed colocalization microscopy experiments using fluorescent marker proteins for different cellular organelles (Fig. 6). We detected strong costaining in the Golgi and ER, and some overlapping signals in mitochondria, but only minor alkyne fluorescence in lipid droplets. The plasma membrane pool of alkyne cholesterol was imaged in greater detail by structured illumination microscopy (supplementary Fig. VA) and separately analyzed by an in vivo assay employing cholesterol oxidase (supplementary Fig. VB). After cofeeding of alkyne cholesterol and deuterated d6-cholesterol, we quantified the respective 3-keto-sterols by MS and determined the ratio of enzymatic product to educt. A similar fraction of both supplemented sterols in the plasma membrane was accessible to oxidation by the exogenously administered enzyme. Living cells showed a higher ratio than fixed cells, supporting the notion that the sterol pool at the plasma membrane exchanges with internal pools by cellular transport. While impossible for d6-cholesterol, the enzymatic product derived from alkyne cholesterol was additionally quantified by TLC analysis and fluorometry. The obtained numbers support the measurements from the MS analysis. We also performed cellular fractionation experiments and have analyzed the distribution of alkyne and normal sterols in the different fractions of a velocity gradient (supplementary Fig. VI). We observed a similar distribution pattern for alkyne cholesterol and normal cholesterol on one hand, and their respective sterol esters on the other hand (supplementary Fig. VIA, B). As demonstrated by Western blotting for marker proteins of various cell organelles (supplementary Fig. VIC), the gradient was able to separate mitochondria and ER from each other, and from the plasma membrane/Golgi fraction. Regarding lipid localization, the experiments comparing both sterols suggest that the cells do not differentiate alkyne cholesterol from normal cholesterol.
Fig. 5.
Alkyne cholesterol distribution can be studied by fluorescence microscopy. Cells incubated with alkyne cholesterol or carrier in delipidated medium for 16 h were fixed and probed for sterols. Alkyne cholesterol was detected after click reaction using azides coupled to a Bodipy-dye (BDY) (A, C), or biotin in combination with Alexa-488-streptavidin (A488) (B, D). Cellular sterols were costained by filipin (C, D). The micrographs show individual slices from a z-stack (A, B), or epifluorescence images (C, D) depicting the signals of click reporter (green) and filipin (magenta) in merged images with colocalization resulting in a white appearance. The individual channels are shown as inserts (C, D); the full z-stacks of (A, B) are provided as supplementary Movies I and II, respectively. Bars, 20 μm.
Fig. 6.
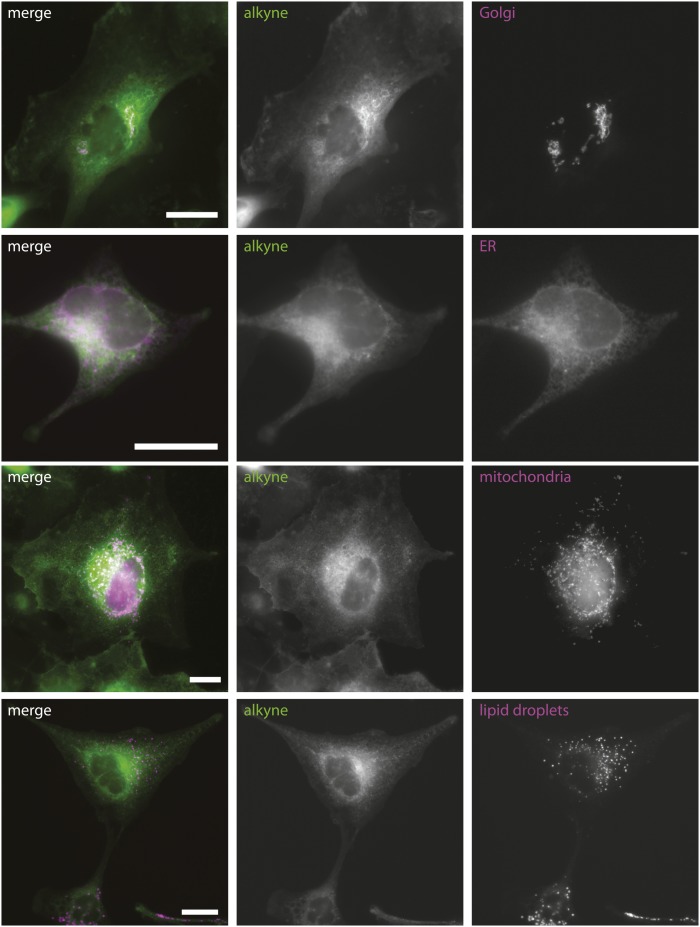
Co-localization analysis of alkyne cholesterol and fluorescent organelle marker proteins. Cells transiently expressing marker proteins were incubated with alkyne cholesterol in delipidated medium for 16 h, fixed, and probed for alkyne cholesterol by a click reaction using an azide coupled to biotin. Alexa-488-streptavidin was used for detection. Epifluorescence micrographs depict the signals of click reporter (green) and marker proteins (magenta) in merged images with colocalization resulting in a white appearance. The individual channels are shown as gray scale images. Bars, 20 μm.
DISCUSSION
To trace the cellular metabolism and localization of cholesterol, the alkyne analog presented here offers several advantages over cholesterol radioisotopes. Its handling is easier, as specialized laboratories and official permissions are not needed. Because experimental samples involving alkyne cholesterol are free of radioactivity, they are easily compatible with modern analysis methods. Alkyne cholesterol and other alkyne lipids can be detected from lipid extracts by MS and, as we showed before for alkyne fatty acids, by TLC and fluorography with high sensitivity after click reaction with a reporter azide (24). When designing the cholesterol analog, we chose to implement the small alkyne moiety at the terminus of the side chain. This allows for a minimal impact on the overall biophysical and biological properties, as all important features of the cholesterol molecule are preserved (45) and introduction of heteroatoms (46) is avoided. From the structural aspect, alkyne cholesterol shares its high similarity to cholesterol with the fluorescent dehydroergosterol (DHE) and cholestatrienol (CTL) (13–15, 47). Both these probes contain additional double bonds in the sterol rings and in CTL, but not DHE, the unmodified side chain of cholesterol is preserved. While for many applications alkyne cholesterol will prove superior, experiments requiring a probe with an unchanged side chain especially will benefit from CTL as a complementary tool.
To validate our alkyne cholesterol probe, we employed various biological systems. Alkyne cholesterol is accepted by enzymes from different biological species (Brevibacterium, yeast, rat, human), and these enzymes include cholesterol oxidases, hydroxylases, and acyl transferases. We have demonstrated the use of alkyne cholesterol in in vitro and in vivo experiments and followed its enzymatic conversion to various metabolites. Also, alkyne cholesterol allowed for the survival and proliferation of a yeast strain vitally dependent on exogenous sources of sterols. A related yeast strain that also is auxotroph for exogenous sterol has been used before for validation of DHE (13).
A major advantage of the alkyne cholesterol analog is its applicability to microscopy. After a click reaction employing a group of versatile reporter azides and bright photostable fluorophores, alkyne cholesterol was conveniently localized at subcellular resolution. As alkyne cholesterol binds to filipin, a popular cholesterol probe, an additional detection option is available. In a localization experiment using filipin, or both filipin and click detection, we detected profound sterol labeling of the plasma membrane in accordance with the notion that the majority of cellular cholesterol is located there (48). Additionally, a significant costaining in the perinuclear region was observed. As the intensity of the filipin signal, especially at internal organelles, increased upon alkyne cholesterol feeding, we argue that filipin also interacts with alkyne cholesterol in cells, in agreement with our in vitro binding data. These cellular pools of cholesterol have been imaged in many studies employing filipin, DHE, CTL (13–15, 47), and BCΘ, a derivative of the sterol binding toxin perfringolysin O (17, 18, 49). Using alkyne cholesterol and fluorescently labeled marker proteins, we confirmed positive costaining of the Golgi, ER, and mitochondria. Supportive subcellular fractionation analysis was performed to complement the imaging experiments and demonstrate equality of alkyne and normal cholesterol localization. Thus far, ER and mitochondria labeling have not been highlighted in microscopy studies using filipin or DHE. Contrarily, BCΘ did not label the ER in an electron microscopy analysis (49). It is currently unclear whether the addition of alkyne cholesterol to cells lifts the overall sterol levels, e.g., at the ER above a certain threshold (50, 51), that if met otherwise, would allow for labeling by BCΘ also. A dependence of the toxins’ interaction on the cholesterol levels has been reported (18, 50). Filipin is known to fail to label some sterol-containing membranes (52), and the sub-optimal spectral properties of DHE and CTL (47) might prevent detection of lower levels of sterol in specific cellular membranes. However, the ER is known to contain an important finely-tuned pool of cholesterol (51). Our findings on mitochondrial cholesterol in turn are supported by MS experiments that show considerable amounts of ergosterol or cholesterol in mitochondria from the yeast Pichia pastoris (53) and in mitochondria from mouse brains (54), respectively.
In summary, the novel alkyne cholesterol presented here proves useful for tracking cellular cholesterol metabolism and localization. It represents a versatile, sensitive, and easy-to-use tool allowing for manifold modes of detection such as MS, TLC/fluorography, and fluorescence microscopy. Despite several important advantages, microscopy studies using alkyne cholesterol are not free of limitations. As applying to all analogs, great attention must be paid to determine whether a certain probe represents a suitable tool for the specific scientific question addressed. One obvious concern when localizing alkyne cholesterol and metabolites by microscopy using our protocol is the need for an additional chemical detection reaction involving copper ions. While alkyne cholesterol is metabolically converted to the respective sterol esters that are typically stored in lipid droplets, the copper ions will unlikely be able to catalyze the click reaction with a reporter azide within the hydrophobic core of these organelles. Accordingly, staining of lipid droplets in our micrographs is likely underrepresented. Importantly, this only applies to microscopy, not to assays on cholesterol ester metabolism, and as cholesterol esters usually contribute very little to the total cellular sterol content, the majority of the cellular cholesterol remains accessible by our imaging method. As the current protocol includes a fixation step, it does not allow for lipid tracking in living cells. However, this limitation will likely be overcome in the near future.
Supplementary Material
Acknowledgments
The authors are grateful to Drs. David Russell and Henry Weiner for providing the plasmids encoding for human CYP46A1 and rat ALDH, respectively. They thank Dr. Michael Gütschow for help with the elementary analysis. The members of their labs are acknowledged for critical reading of the manuscript.
Footnotes
Abbreviations:
- ALA
- 5-aminolevulinic acid
- ALDH
- alcohol dehydrogenase
- CSM
- complete synthetic medium
- CTL
- cholestatrienol
- DHE
- dehydroergosterol
- ER
- endoplasmic reticulum
- OPC
- oligodendrocyte precursor cell
This work was supported by grants from the Deutsche Forschungsgemeinschaft (SFB-TRR 83) to C.T. and L.K.
The online version of this article (available at http://www.jlr.org) contains supplementary data in the form of text, six figures, one table, and two movies.
REFERENCES
- 1.Brown M. S., Goldstein J. L. 2009. Cholesterol feedback: from Schoenheimer's bottle to Scap's MELADL. J. Lipid Res. 50(Suppl): S15–S27 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Parton R. G., Simons K. 2007. The multiple faces of caveolae. Nat. Rev. Mol. Cell Biol. 8: 185–194 [DOI] [PubMed] [Google Scholar]
- 3.Lingwood D., Simons K. 2010. Lipid rafts as a membrane-organizing principle. Science. 327: 46–50 [DOI] [PubMed] [Google Scholar]
- 4.Thiele C., Hannah M. J., Fahrenholz F., Huttner W. B. 2000. Cholesterol binds to synaptophysin and is required for biogenesis of synaptic vesicles. Nat. Cell Biol. 2: 42–49 [DOI] [PubMed] [Google Scholar]
- 5.Hulce J. J., Cognetta A. B., Niphakis M. J., Tully S. E., Cravatt B. F. 2013. Proteome-wide mapping of cholesterol-interacting proteins in mammalian cells. Nat. Methods. 10: 259–264 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Russell D. W. 2000. Oxysterol biosynthetic enzymes. Biochim. Biophys. Acta. 1529: 126–135 [DOI] [PubMed] [Google Scholar]
- 7.Russell D. W. 2009. Fifty years of advances in bile acid synthesis and metabolism. J. Lipid Res. 50(Suppl): S120–S125 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Ghosh S., Zhao B., Bie J., Song J. 2010. Macrophage cholesteryl ester mobilization and atherosclerosis. Vascul. Pharmacol. 52: 1–10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Brown M. S., Goldstein J. L. 1986. A receptor-mediated pathway for cholesterol homeostasis. Science. 232: 34–47 [DOI] [PubMed] [Google Scholar]
- 10.Vance J. E. 2012. Dysregulation of cholesterol balance in the brain: contribution to neurodegenerative diseases. Dis. Model. Mech. 5: 746–755 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Moore K. J., Sheedy F. J., Fisher E. A. 2013. Macrophages in atherosclerosis: a dynamic balance. Nat. Rev. Immunol. 13: 709–721 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Gill S., Chow R., Brown A. J. 2008. Sterol regulators of cholesterol homeostasis and beyond: the oxysterol hypothesis revisited and revised. Prog. Lipid Res. 47: 391–404 [DOI] [PubMed] [Google Scholar]
- 13.Mukherjee S., Zha X., Tabas I., Maxfield F. R. 1998. Cholesterol distribution in living cells: fluorescence imaging using dehydroergosterol as a fluorescent cholesterol analog. Biophys. J. 75: 1915–1925 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Wüstner D. 2007. Fluorescent sterols as tools in membrane biophysics and cell biology. Chem. Phys. Lipids. 146: 1–25 [DOI] [PubMed] [Google Scholar]
- 15.McIntosh A. L., Atshaves B. P., Huang H., Gallegos A. M., Kier A. B., Schroeder F. 2008. Fluorescence techniques using dehydroergosterol to study cholesterol trafficking. Lipids. 43: 1185–1208 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Marks D. L., Bittman R., Pagano R. E. 2008. Use of Bodipy-labeled sphingolipid and cholesterol analogs to examine membrane microdomains in cells. Histochem. Cell Biol. 130: 819–832 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Fujimoto T., Hayashi M., Iwamoto M., Ohno-Iwashita Y. 1997. Crosslinked plasmalemmal cholesterol is sequestered to caveolae: analysis with a new cytochemical probe. J. Histochem. Cytochem. 45: 1197–1205 [DOI] [PubMed] [Google Scholar]
- 18.Ohno-Iwashita Y., Shimada Y., Hayashi M., Iwamoto M., Iwashita S., Inomata M. 2010. Cholesterol-binding toxins and anti-cholesterol antibodies as structural probes for cholesterol localization. Subcell. Biochem. 51: 597–621 [DOI] [PubMed] [Google Scholar]
- 19.Das A., Goldstein J. L., Anderson D. D., Brown M. S., Radhakrishnan A. 2013. Use of mutant 125I-perfringolysin O to probe transport and organization of cholesterol in membranes of animal cells. Proc. Natl. Acad. Sci. USA. 110: 10580–10585 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Bertozzi C. R. 2011. A decade of bioorthogonal chemistry. Acc. Chem. Res. 44: 651–653 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Kolb H. C., Finn M. G., Sharpless K. B. 2001. Click chemistry: diverse chemical function from a few good reactions. Angew. Chem. Int. Ed. Engl. 40: 2004–2021 [DOI] [PubMed] [Google Scholar]
- 22.Jao C. Y., Roth M., Welti R., Salic A. 2009. Metabolic labeling and direct imaging of choline phospholipids in vivo. Proc. Natl. Acad. Sci. USA. 106: 15332–15337 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Neef A. B., Schultz C. 2009. Selective fluorescence labeling of lipids in living cells. Angew. Chem. Int. Ed. Engl. 48: 1498–1500 [DOI] [PubMed] [Google Scholar]
- 24.Thiele C., Papan C., Hoelper D., Kusserow K., Gaebler A., Schoene M., Piotrowitz K., Lohmann D., Spandl J., Stevanovic A., et al. 2012. Tracing fatty acid metabolism by click chemistry. ACS Chem. Biol. 7: 2004–2011 [DOI] [PubMed] [Google Scholar]
- 25.Gaebler A., Milan R., Straub L., Hoelper D., Kuerschner L., Thiele C. 2013. Alkyne lipids as substrates for click chemistry-based in vitro enzymatic assays. J. Lipid Res. 54: 2282–2290 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Haberkant P., Raijmakers R., Wildwater M., Sachsenheimer T., Brugger B., Maeda K., Houweling M., Gavin A. C., Schultz C., van Meer G., et al. 2013. In vivo profiling and visualization of cellular protein-lipid interactions using bifunctional fatty acids. Angew. Chem. Int. Ed. Engl. 52: 4033–4038 [DOI] [PubMed] [Google Scholar]
- 27.Martin B. R., Cravatt B. F. 2009. Large-scale profiling of protein palmitoylation in mammalian cells. Nat. Methods. 6: 135–138 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Charron G., Wilson J., Hang H. C. 2009. Chemical tools for understanding protein lipidation in eukaryotes. Curr. Opin. Chem. Biol. 13: 382–391 [DOI] [PubMed] [Google Scholar]
- 29.Hannoush R. N., Sun J. 2010. The chemical toolbox for monitoring protein fatty acylation and prenylation. Nat. Chem. Biol. 6: 498–506 [DOI] [PubMed] [Google Scholar]
- 30.Yap M. C., Kostiuk M. A., Martin D. D., Perinpanayagam M. A., Hak P. G., Siddam A., Majjigapu J. R., Rajaiah G., Keller B. O., Prescher J. A., et al. 2010. Rapid and selective detection of fatty acylated proteins using omega-alkynyl-fatty acids and click chemistry. J. Lipid Res. 51: 1566–1580 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Best M. D., Rowland M. M., Bostic H. E. 2011. Exploiting bioorthogonal chemistry to elucidate protein-lipid binding interactions and other biological roles of phospholipids. Acc. Chem. Res. 44: 686–698 [DOI] [PubMed] [Google Scholar]
- 32.Spandl J., White D. J., Peychl J., Thiele C. 2009. Live cell multicolor imaging of lipid droplets with a new dye, LD540. Traffic. 10: 1579–1584 [DOI] [PubMed] [Google Scholar]
- 33.Kuerschner L., Ejsing C. S., Ekroos K., Shevchenko A., Anderson K. I., Thiele C. 2005. Polyene-lipids: a new tool to image lipids. Nat. Methods. 2: 39–45 [DOI] [PubMed] [Google Scholar]
- 34.Crisp R. J., Pollington A., Galea C., Jaron S., Yamaguchi-Iwai Y., Kaplan J. 2003. Inhibition of heme biosynthesis prevents transcription of iron uptake genes in yeast. J. Biol. Chem. 278: 45499–45506 [DOI] [PubMed] [Google Scholar]
- 35.Wach A., Brachat A., Alberti-Segui C., Rebischung C., Philippsen P. 1997. Heterologous HIS3 marker and GFP reporter modules for PCR-targeting in Saccharomyces cerevisiae. Yeast. 13: 1065–1075 [DOI] [PubMed] [Google Scholar]
- 36.McCarthy K. D., de Vellis J. 1980. Preparation of separate astroglial and oligodendroglial cell cultures from rat cerebral tissue. J. Cell Biol. 85: 890–902 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Chu Y., Masoud M., Gebeyehu G., inventors 2008. Transfection reagents. United States patent US 7470817 B2 [Google Scholar]
- 38.Cham B. E., Knowles B. R. 1976. A solvent system for delipidation of plasma or serum without protein precipitation. J. Lipid Res. 17: 176–181 [PubMed] [Google Scholar]
- 39.Bligh E. G., Dyer W. J. 1959. A rapid method of total lipid extraction and purification. Can. J. Biochem. Physiol. 37: 911–917 [DOI] [PubMed] [Google Scholar]
- 40.Lange Y., Matthies H., Steck T. L. 1984. Cholesterol oxidase susceptibility of the red cell membrane. Biochim. Biophys. Acta. 769: 551–562 [DOI] [PubMed] [Google Scholar]
- 41.Wang Y., Thiele C., Huttner W. B. 2000. Cholesterol is required for the formation of regulated and constitutive secretory vesicles from the trans-Golgi network. Traffic. 1: 952–962 [DOI] [PubMed] [Google Scholar]
- 42.Schindelin J., Arganda-Carreras I., Frise E., Kaynig V., Longair M., Pietzsch T., Preibisch S., Rueden C., Saalfeld S., Schmid B., et al. 2012. Fiji: an open-source platform for biological-image analysis. Nat. Methods. 9: 676–682 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Aguilar P. S., Heiman M. G., Walther T. C., Engel A., Schwudke D., Gushwa N., Kurzchalia T., Walter P. 2010. Structure of sterol aliphatic chains affects yeast cell shape and cell fusion during mating. Proc. Natl. Acad. Sci. USA. 107: 4170–4175 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Milhaud J., Bolard J., Benveniste P., Hartmann M. A. 1988. Interaction of the polyene antibiotic filipin with model and natural membranes containing plant sterols. Biochim. Biophys. Acta. 943: 315–325 [DOI] [PubMed] [Google Scholar]
- 45.Schroeder F. 1984. Fluorescent sterols: probe molecules of membrane structure and function. Prog. Lipid Res. 23: 97–113 [DOI] [PubMed] [Google Scholar]
- 46.Heal W. P., Jovanovic B., Bessin S., Wright M. H., Magee A. I., Tate E. W. 2011. Bioorthogonal chemical tagging of protein cholesterylation in living cells. Chem. Commun. (Camb.). 47: 4081–4083 [DOI] [PubMed] [Google Scholar]
- 47.Maxfield F. R., Wustner D. 2012. Analysis of cholesterol trafficking with fluorescent probes. Methods Cell Biol. 108: 367–393 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Lange Y., Swaisgood M. H., Ramos B. V., Steck T. L. 1989. Plasma membranes contain half the phospholipid and 90% of the cholesterol and sphingomyelin in cultured human fibroblasts. J. Biol. Chem. 264: 3786–3793 [PubMed] [Google Scholar]
- 49.Möbius W., Ohno-Iwashita Y., van Donselaar E. G., Oorschot V. M., Shimada Y., Fujimoto T., Heijnen H. F., Geuze H. J., Slot J. W. 2002. Immunoelectron microscopic localization of cholesterol using biotinylated and non-cytolytic perfringolysin O. J. Histochem. Cytochem. 50: 43–55 [DOI] [PubMed] [Google Scholar]
- 50.Sokolov A., Radhakrishnan A. 2010. Accessibility of cholesterol in endoplasmic reticulum membranes and activation of SREBP-2 switch abruptly at a common cholesterol threshold. J. Biol. Chem. 285: 29480–29490 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Radhakrishnan A., Goldstein J. L., McDonald J. G., Brown M. S. 2008. Switch-like control of SREBP-2 transport triggered by small changes in ER cholesterol: a delicate balance. Cell Metab. 8: 512–521 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Gimpl G., Gehrig-Burger K. 2011. Probes for studying cholesterol binding and cell biology. Steroids. 76: 216–231 [DOI] [PubMed] [Google Scholar]
- 53.Wriessnegger T., Leitner E., Belegratis M. R., Ingolic E., Daum G. 2009. Lipid analysis of mitochondrial membranes from the yeast Pichia pastoris. Biochim. Biophys. Acta. 1791: 166–172 [DOI] [PubMed] [Google Scholar]
- 54.Kiebish M. A., Han X., Seyfried T. N. 2009. Examination of the brain mitochondrial lipidome using shotgun lipidomics. Methods Mol. Biol. 579: 3–18 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.