Abstract
Background
Broccoli (Brassica oleracea var. italica), a member of Cruciferae, is an important vegetable containing high concentration of various nutritive and functional molecules especially the anticarcinogenic glucosinolates. The sprouts of broccoli contain 10–100 times higher level of glucoraphanin, the main contributor of the anticarcinogenesis, than the edible florets. Despite the broccoli sprouts’ functional importance, currently available genetic and genomic tools for their studies are very limited, which greatly restricts the development of this functionally important vegetable.
Results
A total of ∼85 million 251 bp reads were obtained. After de novo assembly and searching the assembled transcripts against the Arabidopsis thaliana and NCBI nr databases, 19,441 top-hit transcripts were clustered as unigenes with an average length of 2,133 bp. These unigenes were classified according to their putative functional categories. Cluster analysis of total unigenes with similar expression patterns and differentially expressed unigenes among different tissues, as well as transcription factor analysis were performed. We identified 25 putative glucosinolate metabolism genes sharing 62.04–89.72% nucleotide sequence identity with the Arabidopsis orthologs. This established a broccoli glucosinolate metabolic pathway with high colinearity to Arabidopsis. Many of the biosynthetic and degradation genes showed higher expression after germination than in seeds; especially the expression of the myrosinase TGG2 was 20–130 times higher. These results along with the previous reports about these genes’ studies in Arabidopsis and the glucosinolate concentration in broccoli sprouts indicate the breakdown products of glucosinolates may play important roles in the stage of broccoli seed germination and sprout development.
Conclusion
Our study provides the largest genetic resource of broccoli to date. These data will pave the way for further studies and genetic engineering of broccoli sprouts and will also provide new insight into the genomic research of this species and its relatives.
Introduction
Consumption of fruits and vegetables has long been associated with better health and lower incidence of a variety of diseases such as coronary heart disease, cancers, etc [1], [2]. Notably, a diet rich in cruciferous vegetables especially broccoli (Brassica oleracea var. italica) has been recognized as an efficient way to reduce the risk of getting many types of cancers. Epidemiological studies prior to 1996 showed an inverse relationship between cancer risk and cruciferous vegetable intake [3], [4]. Some newer studies demonstrate that this inverse relationship is mainly contributed by the breakdown products of glucosinolates [5], [6], [7]. Glucosinolates are a major group of sulfur-rich secondary metabolites specifically in Cruciferae, which are well-known by their breakdown products to display several bioactivities, including plant defense against pathogens and insects as well as anticarcinogenesis in mammals [8]. Based on their precursor amino acids, glucosinolates are divided into three major categories: aliphatic, indolic and aromatic glucosinolates [9], [10]. Among them, the degradation products of aliphatic glucosinolates are considered to have the higher phase 2 detoxication enzyme inducer ability than the other two groups which is effective in blocking chemical carcinogenesis; therefore, they are thought to be the main contributor to protection against carcinogenesis [11].
Although broccoli heads are generally used as the edible part, the sprouts have been suggested to be a better source for health benefits. A study in 1997 reported the sprouts of eight broccoli cultivars have phase 2 enzyme inducer potency (nearly all arose from glucosinolates) 10–100 times greater than that of mature plants [11]. During the first few days of germination, the inducer activity per unit plant weight declined from the maximum point in seeds in an exponential manner. The declining trend flattened after nine days, and finally approached the value in mature broccoli heads after about 15 days [11]. The most valuable information is that in sprouts the aliphatic glucosinolates are dominant, while in adult plant the indolic ones account for the most [12]. The high content of the aliphatic glucosinolates in broccoli sprouts is mainly attributed to glucoraphanin. Its hydrolytic product, sulforaphane, has been well studied with high anticancer activity. It can not only inhibit phase 1 enzymes but also induce phase 2 enzymes [13]. Besides, sulforaphane has an important ability to target the highly aggressive cancer stem cell population, which is responsible for tumor therapeutics and cannot be eliminated by conventional chemo- or radio-therapy [14], [15]. Another interesting fact is that no significant side effects were found in therapy with sulforaphane in the rapeutic concentrations in non-malignant cells or mice [14], [16]. In addition, glucoraphanin has an obvious effect on decreasing oxidative stress, hypertension and inflammation in the cardiovascular system of rats [17]. Based on these promising results, the first prospective clinical studies with cancer patients and sulforaphane-enriched broccoli sprouts have now been initiated in the United States. Therefore, broccoli sprouts should have played a more important role in human health than the mature inflorescences.
The currently available genetic and genomic tools for broccoli research are very limited. While most studies of broccoli focused on physiology, few have been done at the genetic level and functional genomic studies are still in the infancy. Conspicuous effects of ESTs have been reported to develop genetic engineering, including gene family expansion [18], [19], improving breeding programs by SNP and SSR markers [20], [21], facilitating genome annotation [22], and large-scale expression analysis [23], [24]. Currently, only 2,324 broccoli ESTs in the national center for biotechnology information (NCBI) database (http://www.ncbi.nlm.nih.gov/) are generated with the aim to identify gene expression profiles in microspore and floret bud. Most of them have no annotations. Despite of the growing demand and high-yield potential, the average yield of both fresh and processed broccoli has remained virtually unchanged in the United States in recent ten years [25]. Thus, very limited genomic resources of broccoli constitute a key limitation to the development of improved crops. The advent of next generation sequencing technologies has triggered a revolution in biological research, for it is cheaper and more rapid in providing genomic and transcriptomic data [26].
Here we performed a high-throughput Illumina Miseq sequencing to characterize the transcriptomes of five samples, including seeds, cotyledons of 3, 7, 11 day sprouts and euphyllas of 11 day sprouts. Since there is no available reference genome for broccoli, abundant short reads are required in order to perform de novo assembly. From the total of five libraries, we generated 557,094,098 raw reads with an average length of 251 bp containing 139,830,744,098 nucleotide bases. Formal research has suggested that to achieve 99% coverage of an mRNA, at least an 8X sequencing depth is required [27]. For this study, the sequencing depth is 50X, enough to get the maximum coverage. Using a de novo assembly method, 19,441 unigenes are obtained with an average length of 2133 bp. These unigenes are used for subsequent annotation analyses to provide a platform of transcriptome information for genes in broccoli sprouts. In this study, we focused our work on identification of glucosinolate metabolism genes in broccoli seed germination and sprout development. This will pave the way for further genetic engineering to improve this species’ agronomic traits.
Results and Discussion
Sequencing and Data Analysis
RNA sequencing of the five samples (seeds, 3 day cotyledons, 7 day cotyledons, 11 day cotyledons and 11 day euphyllas) produced a total of ∼85million 251 bp paired-end reads with an average of 17million reads for each sample. Cleaning and quality checks were performed to the raw data (cf. Materials and Methods). A total of ∼75million trimmed reads were obtained with useful data percentage in five time points ranging from 70.29% to 76.01% and the average length of each read was 207 bp (Table S1). Compared to the reads generated by the formal platforms, the longer length of Illumina Miseq sequencing reads greatly facilitated the accuracy of the subsequent de novo assembly. Using single k-mer assembler Velvet (http://www.ebi.ac.uk/~zerbino/velvet), assembly of reads generated 659,752 contigs with mean sizes of 254 bp and N50 of 222 bp (Table 1). The contigs with length more than 500 bp accounted for about 6.19%. Multiple K-mer assembler OASES (http://www.ebi.ac.uk/~zerbino/oases) was applied to produce 122,345 transcripts for 40,081 loci with average length of 1670 bp. Then, all the transcripts were blasted against the Arabidopsis database. For those “non-BLASTable” transcripts, we searched them against the NCBI non-redundant (nr) database, using BLASTx program with an E-value threshold of 1E-5. A total of 94,255 (77.04%) transcripts were significantly matched to known genes in Arabidopsis and 3,971 (3.25%) transcripts were matched to the nr database. The high percentage of transcripts matched to Arabidopsis database is due to the close relation of these two species. For the transcripts representing the same loci, the top hit ones were clustered as unigenes. Finally, 19,441 unigenes were generated, with the average length of 2,133 bp and ranging in size from 200 bp to 20,580 bp (Table 1). The size distribution of contigs, transcripts and unigenes were compiled (Figure S1). The sequencing data has been deposited into NCBI gene expression omnibus (GEO) and the accession number is GSE53298.
Table 1. Statistical summary of cDNA sequences of broccoli generated by the Illumina Miseq platform.
| Total length(bp) | Sequence No. | Max Length(bp) | Average Length(bp) | N50 | >N50 Reads No. | |
| Contigs | 167802146 | 659752 | 7524 | 254 | 222 | 205867 |
| Transcripts | 204377219 | 122345 | 20680 | 1670 | 2527 | 26568 |
| Unigenes | 41474146 | 19441 | 20680 | 2133 | 2631 | 5234 |
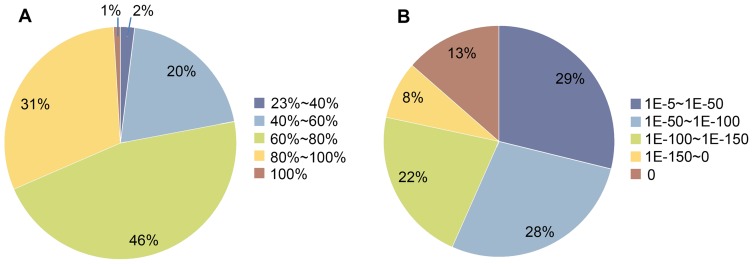
Variable efficiency of matching to sequences in the databases was found in assembled sequences of different lengths, with longer sequences showing higher match proportions (Figure S2). For sequences longer than 1500 bp, the match efficiency was 98.24%. But for sequences between 200–500 bp and 500–1000 bp in length, it was just 53.40% and 79.17%, respectively. E-value distribution of the top hits in the databases had shown 71.17% of matched sequenced with strong homology (<1.0e-50) (Figure 1). 30.54% of the transcripts had a similarity higher than 80%, while 46.47% showed similarity between 60%–80% in identity distribution pattern. The total 77.01% of the transcripts showing identity higher than 60% along with the high quality e-value distribution supported the reliability of the de novo assembly performed in the study.
Figure 1. Characterization of broccoli unigenes by searching against public database.
A. Identity distribution of unigenes blasted against public databases with E-value cutoff of 1E-5; B. E-value distribution of unigenes blasted against public databases with E-value cutoff of 1E-5.
Annotation and Classification
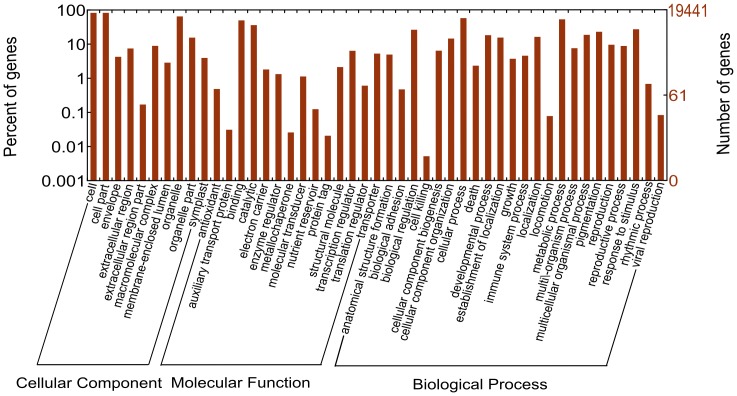
Since biologists have recognized that there is likely to be a single limited universe of genes and proteins are conserved in most, if not all living cells, the GO (gene ontology) Consortium was created as a joint project of many organism databases to produce a structured, precisely defined, common, controlled vocabulary for describing the functions of genes and gene products in any organisms [28]. To annotate the broccoli transcriptome, GO terms were assigned to broccoli unigenes based on their identity to known protein sequences in the Arabidopsis database and nr database. 19,441 unigenes were assigned to 47 functional groups with 134,938 functional terms using GO assignments (Figure 2). For the three main categories of GO classification scheme, the assignments to the “biological process” (61,583, 45.64%) made up the majority, followed by the “cellular component” (53,030, 39.30%) and the “molecular function” (20324, 15.06%). Among these GO groups, the high number of unigenes putatively involved in “cellular process” (11,129) and “metabolic process” (10,230) in the biological process category indicated that the broccoli tissues used in this study were undergoing exquisite metabolic activities, which coincided with the samples’ status. Interestingly, 5,220 unigenes were assigned to “response to stimulus”, showed that during the germination of broccoli seeds and the development of sprouts, there were some protective mechanisms for preparing for potential external and/or internal stresses. Under the category of cellular component, the “cell”, “cell part” and “organelle” were prominent groups. It is noteworthy that the unigenes were not gathered into few groups but were generally expressed. This might be due to the widespread requirements during seedling development.
Figure 2. Histogram presentation of Gene Ontology classification of the assembled unigenes.
EggNOG (evolutionary genealogy of genes: Non-supervised Orthologous Groups) is a database providing orthologous groups for 943 Bacteria, 69 Archaea and 121 Eukaryotes [29]. According to the previous report, the proteins could be divided into 25 functional categories [30]. Out of 19,441 unigenes with significant identity with Arabidopsis database and nr database in this study, 11,242 could be classified into 24 eggNOG categories with only “Nuclear structure” having no annotated unigenes (Figure S3). The categories “function unknown” (2,088, 18.57%) and “general function prediction only” (2,050, 18.24%) were the two largest functional groups of the 25 eggNOG categories. The high percentage of unigenes classified into “general function prediction only” coincided with the transcriptome studies of other species [31], [32], [33]. But our newly noticed fact that so many unigenes were assigned to unknown functional group might indicate there are some interesting unknown mechanism during germination of broccoli seeds and the development of sprouts. Following the most abundant two groups were “transcription” (929, 8.26%), “replication”, “recombination and repair” (802, 7.13%), “signal transduction mechanisms”(797, 7.09%) and “posttranslational modification, protein turnover, chaperones”(680, 6.05%), whereas the two groups involving “cell motility” and “extracellular structures” consisted of a total of 10 unigenes (0.09%), representing the smallest eggNOG classifications. Noteworthily, 277 unigenes (2.64%) were classified into secondary metabolite biosynthesis group, including glucosinolate biosynthesis in broccoli sprouts.
The KEGG (Kyoto Encyclopedia of Genes and Genomes) is a database linking genomic information with higher order functional information by collecting manually drawn pathway maps representing current knowledge on cellular processes and standardized gene annotations [34]. A total of 9836 genes were classified into six main categories including 38 secondary pathways (Figure S4) in the five tested samples. “Metabolism” is the biggest category (3,624, 36.84%), followed by “human disease” (1,760, 17.89%), “Genetic Information Processing” (1,674, 17.02%), “Organismal Systems” (1,279, 13.00%) and “Cellular Processes” (909, 9.24%), whereas “environmental information processing” (590, 6.00%) containing only 3 sub-units (“membrane transport”, “signal transduction and signaling molecules and interaction”) was the smallest category. These results indicated that the broccoli sprouts were undertaken active metabolic and genetic processes and the functional classification of KEGG provided a valuable resource for investigating specific processes and pathways in broccoli sprouts.
Gene Expression Pattern
Gene expression patterns can provide important clues as to the roles of unknown genes in biological active processes [35]. While RPKM (reads aligned to gene per kilobase of exon per million mapped reads) was widely used to calculate gene expression values [36], we used a more accurate method called DESeq to estimate gene expression values in this analysis to infer differential expression signals with good statistical power [37]. K-means clustering analysis was performed using the software MeV edition 4.90 (http://www.tm4.prg/mev.html) to group unigenes with similar expression patterns under different time points and resulting in 10 different clusters (Figure 3). The most abundant cluster (IX) contained 3,659 genes with highest expression at the very beginning (i.e. seeds) and then their expression levels were down-regulated all through the development of sprouts, and reached the lowest point at the 11D euphyllas. These genes might greatly and specifically contribute to seed germination. Cluster I, II, III, VI and VIII comprised genes whose expression levels were very low in seeds but peaked at any one of the three points in cotyledons. Also, genes in cluster IV showed the highest expression level at 11D euphyllas with relatively low level at other time points.
Figure 3. Dynamic expression patterns during broccoli seed germination and sprout development.

K-means clustering was performed to identify 10 clusters, each containing various numbers of genes with similar expression pattern during broccoli seed germination and sprout development. The red lines show representative transcriptional regulators. The x-axis represents sequenced samples, and the y-axis represents normalized RNA-seq expression level. A. seeds; B. 3 day cotyledons; C. 7 day cotyledons; D. 11 day cotyledons; E. 11 day euphyllas.
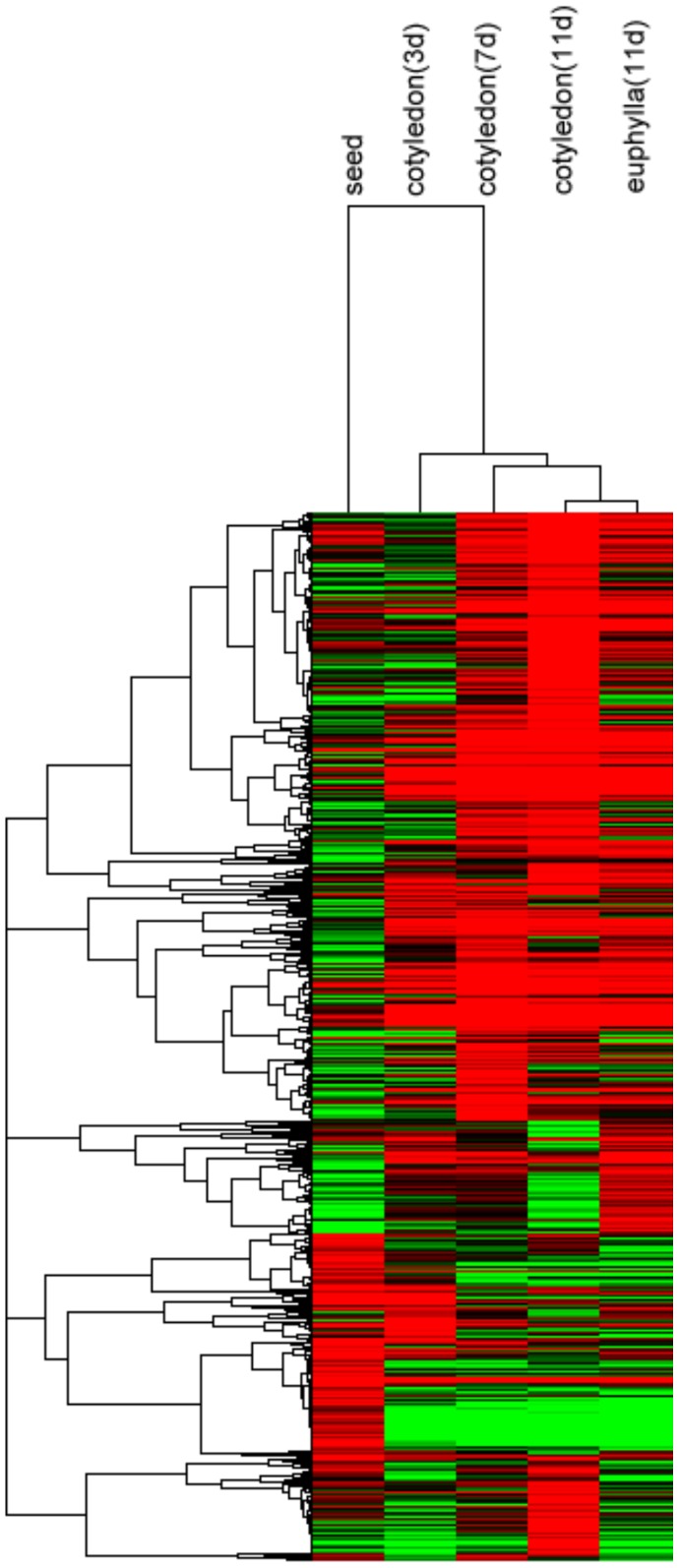
In order to identify differentially expressed genes between the five types of samples, we compared them with each other and picked out a total of 2675 genes, which were at least 2-fold up- or down-regulated between two samples with p-value smaller than 0.05. Then, hierarchical cluster was generated to gain a global view of the differential expressed genes (Figure 4). Obviously, the 11 day cotyledons showed closer relationship with the 11 day euphyllas than with the cotyledons of other time points. This indicated the similar function between the initial stage of euphyllas and the late stage of cotyledons. As expected, the three time points of cotyledons were more similar to each other than to seeds. Even though the 7th day is the mid-point of 3rd day and 11th day, the expression profile of 7 day cotyledons was more similar to that of 11 day cotyledons than that of 3 day cotyledons. This fact along with the big difference between seeds and 3 day cotyledons illustrated the 3 day might be the special point in broccoli sprout development.
Figure 4. Cluster of differentially expressed genes during broccoli seed germination and sprout development.
Expression changes and cluster analysis of 2675 genes that were differentially expressed between any two of the five samples. Each row represents a differentially expressed gene, while each column represents a sample. Changes in expression levels are shown in color scales with saturation at >2.0-fold changes. Green and red color gradients indicate a decrease and increase in transcript abundance, respectively.
Putative Transcription Factors
Transcription factors (TFs) have been considered as one of the most important functional elements regulating gene expression that leads to developmental and other changes. It has been reported that in response to internal or external environment changes, TF genes exhibit more rapid expression changes than the bulk of the regulated genes [38]. Thus, the expression profile of TF genes may in some way reflect the subsequent transcription activities regulated by them. For their important roles, the key putative TFs involved in broccoli seed germination and sprout development were analyzed.
A study in 2003 has revealed that in Arabidopsis, most (84%) of TFs could be detected in six day old seedlings [38]. Currently, the Plant Transcription Factor Database (PlnTFDB) contains 2451 and 2162 distinct TF sequences from Arabidopsis and Arabidopsis lyrata, respectively, arranged in 81 families [39]. In the sequenced broccoli seeds and sprouts, 78 TF families including 1,633 putative TF genes had been identified with the five most expressed TF families being AP2-EREBP, bHLH, MYB, HB and C3H (Table S2). A total of 1,581 of the 1,633 genes accounting for 86.82% were annotated with sequences from the close related species Arabidopsis and Arabidopsis lyrata.
The biggest TF family in our study of broccoli seeds and sprouts was AP2-EREBP with 109 putative family members being detected. AP2-EREBP family is unique to plants and characterized by a conserved AP2 DNA-binding domain of about 60 amino acids [40]. AP2-EREBP genes have been found to play important roles throughout the life cycle including regulating several developmental processes especially leaf epidermal cell identity and forming part of the mechanisms used to respond to stress [41]. Some members of AP2-EREBP, like AP2, control seed mass and seed size in Arabidopsis which is very important to extended growth of cotyledons [42], [43]. The significantly large number of AP2-EREBP family members expressed in broccoli seeds and sprouts indicated the important function of these genes in this period as previously reported in other species [41], [42], [43].
The basic helix-loop-helix (bHLH) family members are involved in various process of seedling development such as light signaling, brassinosteroid and abscisic acid signaling, flavonoid biosynthesis, axillary meristem formation, stomatal patterning and trichome differentiation [44]. 99 unigenes identified to have bHLH-like sequences formed the second biggest TF family in the tested samples. Light is one of the most important elements in seed germination and seedling development, a subfamily containing 15 members are involved in light signaling. These bHLH proteins are known as PIF (phytochrome interacting factor) or PIL (phytochrome interacting factor-like) [45], [46], [47], [48]. These proteins play important roles in phytochrome signal transduction by interacting with phytochromes. Our study found that putative orthologs of bHLH56, PIF1, PIF3, PIF4, PIF5, PIL1, PIF7, SPT were expressed in broccoli seed germination and sprout development, suggesting their similar roles in light signaling during this period. SPATULA (SPT) was found as a leaf size regulator [49]. The SPT ortholog in broccoli sprouts showed highest expression level in the cotyledons of 3 day sprouts, indicating its possible regulation activity on cotyledon size in early broccoli sprouts.
The MYB TFs contain varying numbers of MYB domain repeats to bind DNA. The function of MYB proteins have been well studied in a variety of plant species to be involved in regulatory networks controlling development, metabolism and responses to biotic and abiotic stresses in Arabidopsis [50]. In Arabidopsis seedlings, MYB115 and MYB118 play important roles in embryogenesis [51]. MYB38 and MYB18 have been proposed to regulate hypocotyls elongation in response to blue [52] and far-red light, respectively [53]. Also, the MYB17 has shown activity in regulating seed germination [54]. Some other MYB proteins are involved in the control of cell wall biosynthesis like MYB58, MYB63, MYB85, MYB68 and MYB46 [55], [56], [57], [58]. In this study, a total of 84 putative MYB genes were detected including those orthologs involved in Arabidopsis seedling development. The many putative MYB TF genes expressed in the broccoli seed and sprouts indicated this important family also plays important roles in regulating the biological process during seed germination and sprout development.
Sixty-seven putative NAC TF family members were identified in seed and sprouts of broccoli. The large NAC transcription factor family has been implicated in a variety of plant developmental processes in many species including Arabidopsis and soy bean etc [59]. However, the molecular mechanisms of the family members are still unknown even in well studied species. It has been suggested that they have the ability to enable crosstalk between different pathways [60]. Cys2His2 (C2H2)-type zinc finger proteins are a group of widespread eukaryotic TFs. A majority of C2H2 zinc finger proteins are regarded as trans regulators of genes playing important roles in development, differentiation and suppression of malignant cell transformation [61]. In the sequenced tissues, 65 putative C2H2 zinc finger genes were identified.
Several other TF families were also found like 61 members in bZIP, 54 members in WRKY, 14 members in ARF, etc. Because of the importance of TFs in regulating the downstream genes in variety of pathways, further investigation of the putative TFs would provide interesting clues to the variety of activities in seed germination and sprout development of broccoli.
Glucosinolate Metabolic Pathways
The high contents of glucosinolates especially the much higher content of aliphatic glucosinolates in broccoli sprouts compared to mature tissues have attracted attention in past decade [11]. The biological basis of this trait especially whether the glucosinolate metabolic genes in Arabidopsis or Brassica rapa have the same functions in broccoli sprouts, remains an open question. In this study, a total of 36 unigenes were annotated as putative genes involved in aliphatic and indolic glucosinolate biosynthesis, degradation and regulation. By comparing these unigenes with the CDS of Arabidopsis ones and setting the identity cutoff of 60%, we abandoned 11 unigenes and finally got 25 putative broccoli glucosinolates metabolic genes sharing 62.04–89.72% nucleotide sequence identity with the Arabidopsis orthologs (Figure 5, Table 2).
Figure 5. Detected orthologs in the aliphatic and indolic glucosinolate biosynthetic and degradation pathways in broccoli seeds and sprouts.
Four stages of the pathways in Arabidopsis are showed separately for chain elongation, core structure biosynthesis, side chain modification and degradation. Othologs identified in Broccoli are marked in blue color. Predicted enzymes are marked by *.
Table 2. Putative genes involved in glucosinolate metabolic pathways in Broccoli.
| Name | Arabidopsis orthologs | Basemean of seeds | Basemean of 3 day cotyledons | Basemean of 7 day cotyledons | Basemean of 11 day cotyledons | Basemean of 11 day euphyllas | Identity |
| Aliphatic glucosinolates | |||||||
| Glucosinolate synthesis | |||||||
| BoIPMDH3** | AT1g31180 | 9.2 | 2.03 | 9.45 | 2.14 | 8.09 | 63.28% |
| BoBCAT3 | AT3g49680 | 274.54 | 635.68 | 722.67 | 482.47 | 678.31 | 86.31% |
| BoCYP83A1 | AT4g13770 | 1.42 | 54.92 | 922.24 | 345.23 | 427.55 | 88.20% |
| BoGSTF11** | AT3g03190 | 16.27 | 125.1 | 32.56 | 2.14 | 71.64 | 83.94% |
| BoFMO-GSOX2 | AT1g62540 | 9.91 | 8.14 | 534.65 | 11.79 | 9.24 | 76.50% |
| BoFMO-GSOX5 | AT1g12140 | 42.46 | 15.26 | 22.06 | 6.43 | 18.49 | 78.61% |
| BoSOT17 | AT1g18590 | 235.63 | 49.84 | 306.71 | 1627.53 | 116.71 | 83.00% |
| Transcription factor | |||||||
| BoMyb29 | AT5g07690 | 0 | 6.1 | 52.52 | 0 | 6.93 | 83.04% |
| Indolic glucosinolates | |||||||
| Tryptophan synthesis | |||||||
| BoTSB1 | AT5g54810 | 421.02 | 454.63 | 560.91 | 263.75 | 271.55 | 79.41% |
| Glucosinolate synthesis | |||||||
| BoCYP79B2 | AT4g39950 | 87.03 | 73.23 | 829.81 | 1029.27 | 145.6 | 85.41% |
| BoCYP79B3 | AT2g22330 | 0.71 | 13.22 | 281.51 | 306.64 | 18.49 | 89.72% |
| BoCYP83B1 | AT4g31500 | 961.62 | 515.66 | 1543.03 | 3386.94 | 479.55 | 86.67% |
| BoGSTF10** | AT2g30870 | 844.16 | 1554.1 | 3612.3 | 3070.66 | 1061.95 | 67.23% |
| BoCYP81F1 | AT4G37430 | 55.19 | 49.84 | 3.15 | 0 | 0 | 76.35% |
| BoCYP81F3 | AT4G37400 | 58.73 | 6.1 | 52.52 | 28.95 | 0 | 86.83% |
| BoCYP81F4 | AT4G37410 | 378.56 | 12.2 | 175.42 | 8.58 | 16.18 | 81.70% |
| BoIGMT1 | AT1G21100 | 89.16 | 8.14 | 52.52 | 700.12 | 27.73 | 82.44% |
| Transcription factor | |||||||
| BoMyb51 | AT1g18570 | 90.57 | 18.31 | 56.72 | 131.88 | 33.51 | 85.46% |
| Common to all glucosinolates | |||||||
| BoSOT18 | AT1g74090 | 79.25 | 39.67 | 53.57 | 164.04 | 55.47 | 64.22% |
| BoGGP1* | AT4g30530 | 780.47 | 474.98 | 1406.48 | 7330.34 | 746.48 | 81.14% |
| BoSUR1 | AT2g20610 | 179.02 | 58.99 | 257.35 | 536.08 | 104 | 75.56% |
| BoGSH1/PAD2 | AT4g23100 | 4680 | 1962.96 | 1898.06 | 5790.72 | 1453.68 | 68.61% |
| Glucosinolate degradation | |||||||
| BoPEN2 | AT2G44490 | 247.66 | 157.65 | 279.4 | 2437.01 | 294.66 | 70.49% |
| BoTGG1 | AT5G26000 | 55.19 | 42.72 | 74.58 | 111.5 | 132.89 | 68.58% |
| BoTGG2 | AT5G25980 | 38.21 | 1702.59 | 1590.29 | 818.06 | 4958.45 | 62.04% |
The glucosinolate biosynthesis proceeds through three independent stages: chain elongation (for aliphatic glucosinolates), core structure formation and side chain modification (Figure 5) [62]. For the 25 selected putative genes, 7 were uniquely involved in the aliphatic pathway including BoIPMDH3, BoBCAT3, BoCYP83A1, BoGSTF11, BoSOT17, BoFMOGS-OX2 and BoFMOGS-OX5. In the chain elongation stage, two genes (BoIPMDH3, BoBCAT3) were detected. BCAT3 encodes a chloroplast branched-chain amino acid aminotransferase [62] and IPMDH3 is one of the three isopropylmalate dehydrogenase genes in Arabidopsis whose isozyme IPMDH1 has been characterized as a functional gene involved in aliphatic glucosinolate chain elongation process [63]. Most of the Arabidopsis genes involved in biosynthesis of aliphatic glucosinolate core structure have orthologs expressed in the studied tissues except for CYP79F1, CYP79F2 and UGT74B1, UGT74C1 (Figure 5). It has been reported that in the double-knockout Arabidopsis mutant of CYP79F1 and CYP79F2, aliphatic glucosinolate biosynthesis is completely abolished, meaning that CYP79F1 and CYP79F2 are necessary for the pathway [64]. The contradiction between the high level of aliphatic glucosinolate content and the missing of both of CYP79F1 and CYP79F2 indicates that there may be unknown gene(s) performing the same function in broccoli sprouts. FMOGS-OX2 and FMOGS-OX5 are important genes performing S-oxygenation in side chain modification stage in Arabidopsis [65], [66], they may have the same function in broccoli. However, orthologs of other genes of this stage expressed in Arabidopsis have not been identified in our tissues. Notably, glucoraphanin is one of the products produced by FMOGS-OX2 [66]. The missing of the downstream genes of FMOGS-OX2 may explain the accumulation of glucoraphanin in broccoli sprouts.
In broccoli seeds and sprouts, the indolic glucosinolate biosynthetic pathway showed a high colinearity with Arabidopsis. Eight genes involved in the indolic pathway were detected with BoCYP79B2, BoCYP79B3, BoCYP83B1 and BoGSTF10 in core structure formation and BoCYP81F4, BoCYP81F1, BoCYP81F3 and BoIGMT1 in side chain modification. The enzyme UGT74B1 transforming the Indolylmethyl-thiohydroximate to the Indolylmethyl-desulfoglucosinolate in the indolic glucosinolate pathway was missing. In Arabidopsis, when the indolylmethyl-glucosinolates were formed, there were two ways for them to be modified. Part of them would be transformed to 1-methoxy-3indolylmethyl-glucosinolates by CYP81F4 and the others would be transformed to 4-methoxy-3indolylmethyl-glucosinolates by CYP81F1, CYP81F2 or CYP81F3 and then the IGMT1 or IGMT2 would modify them to 4-methoxy-3-indolylmethyl-glucosinolates [67]. The CYP81F2 and IGMT2 were not detected in this study.
Beside these genes uniquely expressed in indolic or aliphatic glucosinolate pathway, the three genes involved in both the two pathways including GGP1, SUR1 and SOT18 were all identified in the studied tissues too. In Arabidopsis, TSB1 is a tryptophan synthesis gene [68] and GSH1 is a crucial gene to form GSH, which is considered as the sulfur donor to be conjugated with the activated aldoxime [69], [70]. The orthologs in broccoli with these two genes not involved in glucosinolate biosynthesis directly but having important roles for the forming of glucosinolates were also identified.
Some transcription factors of MYB family are crucial in regulating glucosinolate biosynthesis pathways of Arabidopsis, in which MYB28, MYB29 and MYB76 [71], [72], [73] are involved in aliphatic glucosinolates biosynthesis whereas MYB51, MYB34 and MYB 122 [74], [75] could strongly enhance the expression of indolic glucosinolate biosynthesis genes. In broccoli seeds and sprouts, only BoMYB29 and BoMYB51 were detected.
In the glucosinolate degradation pathways, the PEN2, TGG1 and TGG2 were well studied in Arabidopsis. PEN2 was reported to cleave indolic glucosinolates as a myrosinase [76]. TGG1 and TGG2 were two important myrosinases identified long time ago. The double mutant of these two genes showed nearly no aliphatic glucosinolate degradation but only reduced indolic glucosinolate myrosinase activity [77]. In addition, the tgg1tgg2 double mutant still exhibited a wild type callose response to fungi simulation which required the degradation products of indolic glucosinolates [78]. These facts indicated the TGG1 and TGG2 mainly degrade the aliphatic glucosinolates and had slight effects on indolic ones. In the sequenced tissues, the three myrosinase genes’ orthologs were all identified.
It is interesting to note that the expression levels of many broccoli glucosinolate related genes were expressed higher in sprouts than in seeds. Some previous studies had indicated the glucosinolate concentration decreased exponentially after germination [11], [12]. This contradiction between the decreased concentration level of glucosinolates and the increased level of biosynthesis genes might due to the high consumption of glucosinolates and this dramatic degradation of glucosinolates possibly played an important role in the stage of broccoli seed germination and sprout development.
Besides, the putative genes involved in indolic glucosinolate synthesis have higher expression levels than those involved in aliphatic glucosinolate synthesis in general. This was more obvious in the expression of TFs. The only identified transcription factor BoMYB29 in aliphatic glucosinolate synthesis had no expression in seeds and 11 day cotyledons; in the other three time points, the expression level was also relatively low (Table 2). The expression values of BoMYB51 were much higher compared to the expression of BoMYB29. Furthermore, we noticed that the expression value of BoTGG2, was 45-fold higher in 3 day cotyledons than in seeds (Table 2). The expression value decreased slowly along with the development of sprouts and got to the lowest point in the 11 day cotyledons, which was still about 21-fold higher than that in seeds. Notably, at the time of 11th day, the expression values of BoTGG2 in the new forming euphyllas astonishingly increased to about 130-fold. While the expression values of the indolic glucosinolate degradation gene BoPEN2 were relatively low compared to those of BoTGG2 and not too much different in our sequenced tissues except for the 11 day cotyledons (Table 2). These results demonstrated that in glucosinolate sprouts, the exponentially decreased levels of glucosinolates were mainly due to the degradation of aliphatic glucosinolates especially in the young stage of tissues, while the indolic glucosinolates might be constantly synthesized and stored. The exception was the expression value of BoPEN2 in late stage cotyledons increased to 10-fold higher than that in seeds. This might indicate the degraded indolic glucosinolates had special roles at the old stage of cotyledons.
Actually, previous studies have reported that the expression of TGG1, redundantly functioning with TGG2 in Arabidopsis, is higher in young developing tissues than older tissues [79], [80], [81] and PEN2 is unlikely to function in glucosinolate turnover during seedling development [82], which are coordinated to our results. Degraded glucosinolates have been proposed to regulate the cellular signaling in response to abiotic stress which is based on the observation that TGG1 is highly enriched in stomatal guard cells and regulate the stomatal opening or closing by affecting the ABA and MeJA signaling [83], [84]. So the myrosinases’ high level in young seedling may be necessary for proper protection due to lower physical strength barriers. In our study, BoTGG2 was the predominantly expressed myrosinase rather than TGG1 in Arabidopsis. Considering that TGG1 and TGG2 redundantly function in glucosinolate degradation [77], the predominantly functional gene may be BoTGG2 in broccoli sprouts instead of BoTGG1. Also, glucosinolates has been suggested to represent up to 30% of total sulfur content of plant organs and glucosinolate content has been observed to decrease during sulfur deprivation [85]. So we can hypothesis that the enriched degradation products of glucosinolates may contribute greatly to the defense system as a compensation for the lower physical strength barrier in broccoli sprouts and they may also play an important role as potential sulfur donor during broccoli seed germination and sprout development.
Conclusion
In this study, we performed a transcriptome sequencing of seeds, 3 day, 7 day and 11 day cotyledons and 11day euphyllas to identify the transcripts and quantify their levels of expression in broccoli seed germination and sprout developments. In total, we obtained 19441 unigenes annotated by homologs with sequences in the public databases. 2675 genes differentially expressed between any two of the five samples and 1,633 putative TFs were detected. These genes may be closely related to the seed germination and sprout development in broccoli. Twenty-five unigenes with high identity to Arabidopsis glucosinolate metabolic genes have been investigated. This established a high colinearity between Arabidopsis and broccoli in glucosinolate metabolic pathways for further studies. In addition, expression analysis of these glucosinolate metabolic genes showed contradiction between the increased expression of the candidate synthesis genes and the previously reported decreased concentration of glucosinolate content after germination. The ortholog of TGG2, which mainly degrade aliphatic glucosinolates in Arabidopsis, expressed astonishingly higher after germination. These results indicate the breakdown products of glucosinolates may play important roles in the stage of broccoli seed germination and sprout development. The results here represent the largest genetic resource for broccoli and will provide new insight into the genomic research of this species and its relatives.
Materials and Methods
Sample Preparation
Seeds of broccoli cultivar “Qingxiu” with wide suitability of temperature and soil were germinated and grown in trays containing a soil mixture (peat: vermiculite, 2∶3, v/v). Plants were adequately watered with Hoagland’s solution and grown in a culture room with the following settings: 24°C, light regime of 16 h light and 8 h dark, 70% relative humidity and a constant illumination of 100 µmol·m−2·s−1. For the sample of seeds, they were incubated in 5% NaClO with shaking for 8 min and then were washed six times using sterile water with once 30 s. Subsequently, seeds were placed on moist filter paper in petri dishes for one night. Finally, samples of equal weight were harvested for seeds, cotyledons at the 3rd day, 7th day and 11th day and euphyllas at 11th day (Figure 6). To minimize biological variance, each sample was harvested in three independent biological replicates with equal weight and subsequently pooled for sequencing. Samples were immediately frozen in liquid nitrogen and stored at −80°C until RNA was extracted. Total RNA of each sample was isolated using E.Z.N.A. Plant RNA Kit (OMEGA bio-tek, GA) according to the instructions from the manufacturer. RNA quality was characterized on an agarose gels electrophoresis and spectrophotometry. High quality RNA with 28S:18S more than 1.5 and absorbance 260/280 ratios between 1.8 and 2.2 was used for library construction and sequencing.
Figure 6. Images of the sampled tissues.
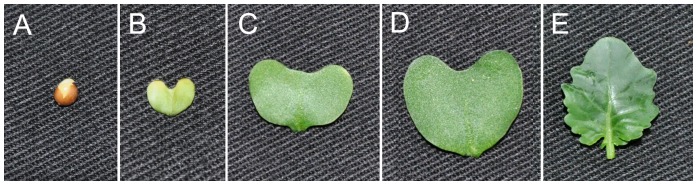

A. seed; B. 3 day cotyledon; C. 7 day cotyledon; D. 11 day cotyledon; E. 11 day euphylla.
cDNA Library Construction and Sequencing
Illumina Miseq library construction was performed according to the manufacturer’s instructions (Illumina, San Diego, CA). Magnetic beads with poly T oligos attached were used for purifying the mRNA from the total RNA. Fragmentation buffer was added to cleave mRNA into short fragments. The fragments were used to synthesize first-strand cDNA using random hexamerprimers, which was transformed into double stranded cDNA with RHase H and DNA polymerase I. A paired-end library was constructed from the cDNA synthesized with Genomic Sample Prep Kit (Illumina). Fragments in desirable lengths were purified with QIAquick PCR (Qiagen) Extraction Kit, end repaired and linked with sequencing adapters (Margulies et al., 2005). AMPureXP beads were used to remove the unsuitable fragments, then the sequencing library was constructed with PCR amplification. After being checked with Pico green staining and fluorospectrophotometry and quantified with Agilent 2100, the multiplexed DNA libraries were mixed by equal volume with normalized 10 nM concentration. The sequencing library was then sequenced with Illumina Miseq platform (Shanghai Personal Biotechnology Cp., Ltd. Shanghai, China).
Data Filtering and de novo Assembly
Raw sequencing reads of five samples were mixed together to perform the following filtration using a stringent process and subsequent de novo assembly. The adaptor contamination was removed, the reads were screened from the 3′ to 5′ to trim the bases with a quality score of Q<20 using 5 bp windows and the reads with final length less than 25 bp were removed. De novo transcriptome assembly was performed by Velvet [86] followed by Oases [87] with default settings except for K-mer value to get contigs and transcripts. Velvet was run using single k-mer length of 69 and OASES was then run with the preliminary Velvet assemblies as input. Because the results after merging with multiple k-mers used in OASES program prompted severe assembly redundancy, we use the same single k-mer in OASES program. High quality reads of every sample were remapped to transcripts to get the abundance of transcripts using Bowtie program [88]. Those transcripts with no reads mapped in all five samples were considered error and removed. All the transcripts were searched against Arabidopsis database, for those with no hits were then searched to NCBI non-redundant (nr) database (ftp://ftp.ncbi.nlm.nih.gov/blast/db/) with BLAST program (E-value <1E-5), and the top-hit transcripts were selected as unigenes. For the unigenes failed to be aligned to the databases, the software GetORF [89] was used to predict their open reading frames (ORFs) and ascertain their sequence directions, with default settings except for the parameter “–find” being set 1.
Gene Annotation and Analysis
To further annotate the unigenes in this study, we used the Blast2GO program [90], [91], [92], [93] to get GO annotation based on GO terms related to the Arabidopsis and nr database annotation. EggNOG (evolutionary genealogy of genes: Non-supervised Orthologous Groups) is a database of orthologous groups of genes. To annotate genes with common denominators or functional categories, the unigenes were also aligned to the eggNOG database (http://eggnog.embl.de/version_3.0/). To summarize the pathways information involved in broccoli seeds and sprouts, the KEGG database were used to perform the pathway annotation (http://www.genome.jp/kegg/). To identify putative TFs presented in this research, we searched the unigenes against the complete TF gene sequences of the Plant Transcription Factor Database (http://plntfdb.bio.uni-potsdam.de/) using BLAST program with an E-value cutoff of 1E-5. To identify the putative sequences related to glucosinolate pathways, the unigenes annotated by putative glucosinolate biosynthetic and regulator genes according to previous studies were chosen [9], [66]. Then, CDS of Arabidopsis glucosinolate biosynthesis and regulator genes were aligned to broccoli homologs using DNAMAN6.0 (http://www.lynnon.com/) and unigenes with identity larger than 60% were selected.
Comparative Expression Analysis
The R package DESeq was performed to identify differential gene expression [94]. This method represents the widely accepted and accurate analysis approaches of RNA-seq data. We first mapped high-quality reads to unigenes to calculate the number of reads mapped to each unigene in five samples. These raw read counts were then used as the input of DESeq to get the normalized signal for each unigene, and the fold change of unigene expression values with p-values compared to each other of the five samples was used to report differential expression. Those with p-value<0.05 were considered as significant differential expression. We performed cluster analysis of gene expression patterns with the Cluster [95], MeV [96] and Java treeview software packages [97].
Supporting Information
Length distribution of contigs, transcripts and unigenes.
(TIF)
Matching percentage of broccoli unigenes with different lengths to entries in public databases.
(TIF)
EggNOG classification of the broccoli seed and sprout transcriptome.
(TIF)
Classification of unigenes based on KEGG categorization.
(TIF)
Characterization of raw data and trimmed data.
(PDF)
Transcription factor members of every family detected in the broccoli seeds and sprouts.
(PDF)
Funding Statement
The national Natural Science Foundation of China 31370334 http://www.nsfc.gov.cn/Portal0/default152.htm; Opening project of Key Laboratory of breed improvement and physiecology of cold region crops, College of Heilongjiang Province http://61.167.33.11/kjgz/Index.asp?page=2; Heilongjiang Provincial University Science and Technology Innovation Team Building Program 2011TD005; http://61.167.33.11/kjgz/Index.asp?page=2. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.US Food and Drug Administration. 2005. Code of Federal Regulations: 21 CFR101.78. Health claims: fruits and vegetables and cancer.
- 2.US Food and Drug Administration. 2012. Code of Federal Regulations: 21 CFR101.77. Health claims: fruits, vegetables, and grain products that contain fiber, particularly soluble fiber, and risk of coronary heart disease.
- 3. Van Poppel G, Verhoeven DT, Verhagen H, Goldbohm RA (1999) Brassica vegetables and cancer prevention. Epidemiology and mechanisms. Adv Exp Med Biol. 472: 159–168. [DOI] [PubMed] [Google Scholar]
- 4. Verhoeven DT, Goldbohm RA, van Poppel G, Verhagen H, van den Brandt PA (1996) Epidemiological studies on brassica vegetablesand cancer risk, Cancer Epidemiol Biomarkers Prev. 5: 733–748. [PubMed] [Google Scholar]
- 5. Michaud DS, Spiegelman D, Clinton SK, Rimm EB, Willett WC, et al. (1999) Fruit and vegetable intake and incidence of bladder cancer in a male prospective cohort. J Natl Cancer Inst. 91: 605–613. [DOI] [PubMed] [Google Scholar]
- 6. Rose P, Faulkner K, Williamson G, Mithen R (2000) 7-Methylsulfinylheptyl and 8-methylsulfinyloctyl isothiocyanates from watercress are potent inducers of phase II enzymes. Carcinogenesis. 21: 1983–1988. [DOI] [PubMed] [Google Scholar]
- 7. Verhoeven DT, Verhagen H, Goldbohm RA, van den Brandt PA, van Poppel G (1997) A review of mechanisms underlying anticarcinogenicity by brassica vegetables. Chem Biol Interact. 103: 79–129. [DOI] [PubMed] [Google Scholar]
- 8. Chen Y, Yan X, Chen S (2011) Bioinformatic analysis of molecular network of glucosinolate biosynthesis. Comput Biol Chem. 35: 10–18. [DOI] [PubMed] [Google Scholar]
- 9. Yan X, Chen S (2007) Regulation of plant glucosinolate metabolism. Planta. 226: 1343–1352. [DOI] [PubMed] [Google Scholar]
- 10. Sønderby IE, Geu-Flores F, Halkier BA (2010) Biosynthesis of glucosinolates–gene discovery and beyond. Trends Plant Sci. 15: 283–290. [DOI] [PubMed] [Google Scholar]
- 11. Fahey JW, Zhang Y, Talalay P (1997) Broccoli sprouts: An exceptionally rich source of inducers of enzymes that protect against chemical carcinogens. Proc Natl Acad Sci U S A. 94: 10367–10372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. S Pérez-Balibrea, DA Moreno, C García-Viguera (2008) Influence of light on health-promoting phytochemicals of broccoli sprouts. J Sci Food Agric. 88: 904–910. [Google Scholar]
- 13. Fahey JW, Talalay P (1999) Antioxidant functions of sulforaphane: a potent inducer of Phase II detoxication enzymes. Food Chem Toxicol. 37: 973–979. [DOI] [PubMed] [Google Scholar]
- 14. Kallifatidis G, Rausch V, Baumann B, Apel A, Beckermann BM, et al. (2009) Sulforaphane targets pancreatic tumour-initiating cells by NF-kappaBinduced antiapoptotic signalling. Gut. 58: 949–963. [DOI] [PubMed] [Google Scholar]
- 15. Abbott A (2006) Cancer: the root of the problem. Nature. 442: 742–743. [DOI] [PubMed] [Google Scholar]
- 16. Kallifatidis G, Labsch S, Rausch V, Mattern J, Gladkich J, et al. (2011) Sulforaphane increases drug-mediated cytotoxicity towards cancer stem-like cells of pancreas and prostate. Mol Ther. 19: 188–195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Wu L, Noyan Ashraf MH, Facci M, Wang R, Paterson PG, et al. (2004) Dietary approach to attenuate oxidative stress, hypertension, and inflammation in the cardiovascular system. Proc Natl Acad Sci U S A. 101: 7094–7099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Bourdon V, Naef F, Rao PH, Reuter V, Mok SC, et al. (2002) Genomic and expression analysis of the 12p11-p12 amplicon using EST arrays identifies two novel amplified and overexpressed genes. Cancer Res. 62: 6218–6223. [PubMed] [Google Scholar]
- 19. Cheung F, Win J, Lang JM, Hamilton J, Vuong H, et al. (2008) Analysis of the Pythium ultimum transcriptome using Sanger and pyrosequencing approaches. BMC Genomics 9: 542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Ruyter-Spira CP, de Koning DJ, van der Poel JJ, Crooijmans RP, Dijkhof RJ, et al. (1998) Developing microsatellite markers from cDNA: A tool for adding expressed sequence tags to the genetic linkage map of the chicken. Anim Genet 29: 85–90. [DOI] [PubMed] [Google Scholar]
- 21. Gonzalo MJ, Oliver M, Garcia-Mas J, Monfort A, Dolcet-Sanjuan R, et al. (2005) Simple-sequence repeat markers used in merging linkage maps of melon (Cucumis melo L.). Theor Appl Genet 110: 802–811. [DOI] [PubMed] [Google Scholar]
- 22. Seki M, Narusaka M, Kamiya A, Ishida J, Satou M, et al. (2002) Functional annotation of a full-length Arabidopsis cDNA collection. Science 296: 141–145. [DOI] [PubMed] [Google Scholar]
- 23. Fei Z, Tang X, Alba RM, White JA, Ronning CM, et al. (2004) Comprehensive EST analysis of tomato and comparative genomics of fruit ripening. Plant J 40: 47–59. [DOI] [PubMed] [Google Scholar]
- 24.USDA National Agricultural Statistics Service.
- 25. Adams MD, Kelley JM, Gocayne JD, Dubnick M, Polymeropoulos MH, et al. (1991) Complementary DNA sequencing: expressed sequence tags and human genome project. Science 252: 1651–1656. [DOI] [PubMed] [Google Scholar]
- 26. Metzker ML (2010) Sequencing technologies - the next generation. Nat Rev Genet 11: 31–46. [DOI] [PubMed] [Google Scholar]
- 27. Jiang L, Schlesinger F, Davis CA, Zhang Y, Li R, et al. (2011) Synthetic spike-in standards for RNA-seq experiments. Genome Res 21: 1543–1551. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, et al. (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. 25: 25–29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Powell S, Szklarczyk D, Trachana K, Roth A, Kuhn M, et al. (2012) eggNOG v3.0: orthologous groups covering 1133 organisms at 41 different taxonomic ranges. Nucleic Acids Res 40(Database issue): D284–289. [DOI] [PMC free article] [PubMed]
- 30. Tatusov RL, Koonin EV, Lipman DJ (1997) A genomic perspective on protein families. Science 278: 631–637. [DOI] [PubMed] [Google Scholar]
- 31. Fan H, Xiao Y, Yang Y, Xia W, Mason AS, et al. (2013) RNA-Seq analysis of Cocos nucifera: transcriptome sequencing and de novo assembly for subsequent functional genomics approaches. PLoS One 8: e59997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Zhang XM, Zhao L, Larson-Rabin Z, Li DZ, Guo ZH (2012) De novo sequencing and characterization of the floral transcriptome of Dendrocalamus latiflorus (Poaceae: Bambusoideae). PLoS One 7: e42082. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Li D, Deng Z, Qin B, Liu X, Men Z (2012) De novo assembly and characterization of bark transcriptome using Illumina sequencing and development of EST-SSR markers in rubber tree (Hevea brasiliensis Muell. Arg.). BMC Genomics 13: 192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Kanehisa M, Goto S (2000) KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28: 27–30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Qing DJ, Lu HF, Li N, Dong HT, Dong DF, et al. (2009) Comparative profiles of gene expression in leaves and roots of maize seedlings under conditions of salt stress and the removal of salt stress. Plant Cell Physiol 50: 889–903. [DOI] [PubMed] [Google Scholar]
- 36. Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5(7): 621–628. [DOI] [PubMed] [Google Scholar]
- 37. Anders S, Huber W (2010) Dierential expression analysis for sequence count data. Genome Biol 11: R106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Jiao Y, Yang H, Ma L, Sun N (2003) A genome-wide analysis of blue-light regulation of Arabidopsis transcription factor gene expression during seedling development. Plant Physiol 133: 1480–1493. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Pérez-Rodríguez P, Riaño-Pachón DM, Corrêa LG, Rensing SA, Kersten B, et al. (2010) PlnTFDB: updated content and new features of the plant transcription factor database. Nucleic Acids Res 38(Database issue): D822–827. [DOI] [PMC free article] [PubMed]
- 40. Saleh Abdelaty, Pagés Montserrat (2003) Plant AP2/ERF transcription factors. Genetika 35: 37–50. [Google Scholar]
- 41. Riechmann JL, Meyerowitz EM (1998) The AP2/EREBP family of plant transcription factors. Biol Chem 379: 633–646. [DOI] [PubMed] [Google Scholar]
- 42. Jofuku KD, Omidyar PK, Gee Z, Okamuro JK (2005) Control of seed mass and seed yield by the floral homeotic gene APETALA2. Proc Natl Acad Sci U S A 102: 3117–3122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Ohto MA, Fischer RL, Goldberg RB, Nakamura K, Harada JJ (2005) Control of seed mass by APETALA2. Proc Natl Acad Sci U S A 102: 3123–3128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Hongtao Zhao, Xia Li, Ligeng Ma (2012) Basic helix-loop-helix transcription factors andepidermal cell fate determination in Arabidopsis. Plant Signal Behav 7: 1556–1560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Duek PD, Fankhauser C (2005) bHLH class transcription factors take centre stage in phytochrome signalling. Trends Plant Sci 10: 51–54. [DOI] [PubMed] [Google Scholar]
- 46. Khanna R, Huq E, Kikis EA, Al-Sady B, Lanzatella C, et al. (2004) A novel molecular recognition motif necessary for targeting photoactivated phytochrome signaling to specific basic helix–loop–helix transcription factors. Plant Cell 16: 3033–3044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Ni M, Tepperman JM, Quail PH (1999) Binding of phytochrome B to its nuclear signalling partner PIF3 is reversibly induced by light. Nature 400: 781–784. [DOI] [PubMed] [Google Scholar]
- 48. Toledo-Ortiz G, Huq E, Quail PH (2003) The Arabidopsis basic/helix–loop–helix transcription factor family. Plant Cell 15: 1749–1770. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Ichihashi Y, Horiguchi G, Gleissberg S, Tsukaya H (2010) The bHLH transcription factor SPATULA controls final leaf size in Arabidopsis thaliana. Plant Cell Physiol 51: 252–261. [DOI] [PubMed] [Google Scholar]
- 50. Dubos C, Stracke R, Grotewold E, Weisshaar B, Martin C, et al. (2010) MYB transcription factors in Arabidopsis. Trends Plant Sci. 15: 573–581. [DOI] [PubMed] [Google Scholar]
- 51. Wang X, Niu QW, Teng C, Li C, Mu J, et al. (2009) Overexpression of PGA37/MYB118 and MYB115 promotes vegetative-to-embryonic transition in Arabidopsis. Cell Res. 19: 224–235. [DOI] [PubMed] [Google Scholar]
- 52. Hong SH, Kim HJ, Ryu JS, Choi H, Jeong S, et al. (2008) CRY1 inhibits COP1-mediated degradation of BIT1, a MYB transcription factor, to activate blue light-dependent gene expression in Arabidopsis. Plant J 55: 361–371. [DOI] [PubMed] [Google Scholar]
- 53. Yang SW, Jang IC, Henriques R, Chua NH (2009) FAR-RED ELONGATED HYPOCOTYL1 and FHY1-LIKE associate with the Arabidopsis transcription factors LAF1 and HFR1 to transmit phytochrome A signals for inhibition of hypocotyl elongation. Plant Cell 21: 1341–1359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Zhang Y, Cao G, Qu LJ, Gu H (2009) Characterization of Arabidopsis MYB transcription factor gene AtMYB17 and its possible regulation by LEAFY and AGL15. J. Genet. Genomics 36: 99–107. [DOI] [PubMed] [Google Scholar]
- 55. Zhou J, Lee C, Zhong R, Ye ZH (2009) MYB58 and MYB63 are transcriptional activators of the lignin biosynthetic pathway during secondary cell wall formation in Arabidopsis. Plant Cell 21: 248–266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Zhong R, Lee C, Zhou J, McCarthy RL, Ye ZH (2008) A battery of transcription factors involved in the regulation of secondary cell wall biosynthesis in Arabidopsis. Plant Cell 20: 2763–2782. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Feng C, Andreasson E, Maslak A, Mock HP, Mattsson O, et al. (2004) Arabidopsis MYB68 in development and responses to environmental cues. Plant Sci. 167: 1099–1107. [Google Scholar]
- 58. Zhong R, Richardson EA, Ye ZH (2007) The MYB46 transcription factor is a direct target of SND1 and regulates secondary wall biosynthesis in Arabidopsis. Plant Cell 19: 2776–2792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59. Shamimuzzaman M, Vodkin L (2013) Genome-wide identification of binding sites for NAC and YABBY transcription factors and co-regulated genes during soybean seedling development by ChIP-Seq and RNA-Seq. BMC Genomics 14: 477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. Olsen AN, Ernst HA, Leggio LL, Skriver K (2005) NAC transcription factors: structurally sidtinct, functionally diverse. Trends Plant Sci 10: 79–87. [DOI] [PubMed] [Google Scholar]
- 61. Razin SV, Borunova VV, Maksimenko OG, Kantidze OL (2012) Cys2His2 zinc finger protein family: classification, functions, and major members. Biochemistry (Mosc) 77: 217–226. [DOI] [PubMed] [Google Scholar]
- 62. Knill T, Schuster J, Reichelt M, Gershenzon J, Binder S (2008) Arabidopsis branched-chain aminotransferase 3 functions in both amino acid and glucosinolate biosynthesis. Plant Physiol 146: 1028–1039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63. Nozawa A, Takano J, Miwa K, Nakagawa Y, Fujiwara T (2005) Cloning of cDNAs encoding isopropylmalate dehydrogenase from Arabidopsis thaliana and accumulation patterns of their transcripts. Biosci Biotechnol Biochem 69: 806–810. [DOI] [PubMed] [Google Scholar]
- 64. Tantikanjana T, Mikkelsen MD, Hussain M, Halkier BA, Sundaresan V (2004) Functional analysis of the tandem-duplicated P450 genes SPS/BUS/CYP79F1 and CYP79F2 in glucosinolate biosynthesis and plant development by Ds transposition-generated double mutants. Plant Physiol 135: 840–848. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65. Hansen BG, Kliebenstein DJ, Halkier BA (2007) Identification of a flavin-monooxygenase as the S-oxygenating enzyme in aliphatic glucosinolate biosynthesis in Arabidopsis. Plant J 50: 902–910. [DOI] [PubMed] [Google Scholar]
- 66. Li J, Hansen BG, Ober JA, Kliebenstein DJ, Halkier BA (2008) Subclade of flavin-monooxygenases involved in aliphatic glucosinolate biosynthesis. Plant Physiol 148: 1721–1733. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67. Pfalz M, Mikkelsen MD, Bednarek P, Olsen CE, Halkier BA, et al. (2011) Metabolic engineering in Nicotiana benthamiana reveals key enzyme functions in Arabidopsis indole glucosinolate modification. Plant Cell 23: 716–729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68. Zhao Y, Hull AK, Gupta NR, Goss KA, Alonso J, et al. (2002) Trp-dependent auxin biosynthesis in Arabidopsis: involvement of cytochrome P450s CYP79B2 and CYP79B3. Genes Dev 16: 3100–3112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69. Cobbett CS, May MJ, Howden R, Rolls B (1998) The glutathione-deficient, cadmiumsensitive mutant, cad2–1, of Arabidopsis thaliana is deficient in gamma-glutamylcysteine synthetase. Plant J 16: 73–78. [DOI] [PubMed] [Google Scholar]
- 70. Schlaeppi K, Bodenhausen N, Buchala A, Mauch F, Reymond P (2008) The glutathione-deficient mutant pad2–1 accumulates lower amounts of glucosinolates and is more susceptible to the insect herbivore Spodoptera littoralis. Plant J 55: 774–786. [DOI] [PubMed] [Google Scholar]
- 71. Gigolashvili T, Yatusevich R, Berger B, Müller C, Flügge UI (2007) The R2R3-MYB transcription factor HAG1/MYB28 is a regulator of methionine-derived glucosinolate biosynthesis in Arabidopsis thaliana . Plant J 51: 247–261. [DOI] [PubMed] [Google Scholar]
- 72. Gigolashvili T, Engqvist M, Yatusevich R, Müller C, Flügge UI (2008) HAG2/MYB76 and HAG3/MYB29 exert a specific and coordinated control on the regulation of aliphatic glucosinolate biosynthesis in Arabidopsis thaliana . New Phytol 177: 627–642. [DOI] [PubMed] [Google Scholar]
- 73. Hirai MY, Sugiyama K, Sawada Y, Tohge T, Obayashi T, et al. (2007) Omics-based identification of Arabidopsis Myb transcription factors regulating aliphatic glucosinolate biosynthesis. Proc Natl Acad Sci U S A 104: 6478–6483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74. Celenza JL, Quiel JA, Smolen GA, Merrikh H, Silvestro AR, et al. (2005) The Arabidopsis ATR1Myb transcription factor controls indolic glucosinolate homeostasis. Plant Physiol. 2005, 137, 253–262. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75. Gigolashvili T, Berger B, Mock HP, Müller C, Weisshaar B, et al. (2007) The transcription factor HIG1/MYB51 regulates indolic glucosinolate biosynthesis in Arabidopsis thaliana. Plant J. 50: 886–901. [DOI] [PubMed] [Google Scholar]
- 76. Bednarek P, Pislewska-Bednarek M, Svatos A, Schneider B, Doubsky J, et al. (2009) A glucosinolate metabolism pathway in living plant cells mediates broad-spectrum antifungal defense. Science 323: 101–106. [DOI] [PubMed] [Google Scholar]
- 77. Barth C, Jander G (2006) Arabidopsis myrosinases TGG1 and TGG2 have redundant function in glucosinolate breakdown and insect defense. Plant J 46: 549–562. [DOI] [PubMed] [Google Scholar]
- 78. Clay NK, Adio AM, Denoux C, Jander G, Ausubel FM (2009) Glucosinolate metabolites required for an Arabidopsis innate immune response. Science 323: 95–101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79. Husebye H, Chadchawan S, Winge P, Thangstad OP, Bones AM (2002) Guard cell- and phloem idioblast-specific expression of thioglucoside glucohydrolase 1 (myrosinase) in Arabidopsis . Plant Physiol 128: 1180–1188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80. Barth C, Jander G (2006) Arabidopsis myrosinases TGG1 and TGG2 have redundant function in glucosinolate breakdown and insect defense. Plant J 46: 549–562. [DOI] [PubMed] [Google Scholar]
- 81. Burow M, Rice M, Hause B, Gershenzon J, Wittstock U (2007) Cell- and tissue-specific localization and regulation of the epithiospecifier protein in Arabidopsis thaliana. Plant Mol Biol 64: 173–185. [DOI] [PubMed] [Google Scholar]
- 82. Wittstock U, Burow M (2010) Glucosinolate breakdown in Arabidopsis: mechanism, regulation and biological significance. Arabidopsis Book 8: e0134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83. Zhao Z, Zhang W, Stanley BA, Assmann SM (2008) Functional proteomics of Arabidopsis thaliana guard cells uncovers new stomatal signaling pathways. Plant Cell 20: 3210–3226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84. Islam MM, Tani C, Watanabe-Sugimoto M (2009) Myrosinases, TGG1 and TGG2, redundantly function in ABA and MeJA signaling in Arabidopsis guard cells. Plant Cell Physiol 50: 1171–1175. [DOI] [PubMed] [Google Scholar]
- 85. Falk KL, Tokuhisa JG, Gershenzon J (2007) The effect of sulfur nutrition on plant glucosinolate content: physiology and molecular mechanisms. Plant Biol (Stuttg) 9: 573–581. [DOI] [PubMed] [Google Scholar]
- 86. Zerbino DR, Birney E (2008) Velvet: algorithms for de novo short read assembly using de Bruijn graphs. D.R. Zerbino and E. Birney. Genome Res 18: 821–829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87. Schulz MH, Zerbino DR, Vingron M, Birney E (2012) Oases: Robust de novo RNA-seq assembly across the dynamic range of expression levels. Bioinformatics 28: 1086–1092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Langmead B. (2010) Aligning short sequencing reads with Bowtie. Curr Protoc Bioinformatics Chapter 11:Unit 11.7. [DOI] [PMC free article] [PubMed]
- 89. Rice P, Longden I, Bleasby A (2000) EMBOSS: the European Molecular Biology Open Software Suite. Trends Genet 16: 276–277. [DOI] [PubMed] [Google Scholar]
- 90. Conesa A, Götz S, García-Gómez JM, Terol J, Talón M, et al. (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21: 3674–3676. [DOI] [PubMed] [Google Scholar]
- 91. Conesa A, Götz S (2008) Blast2GO: A Comprehensive Suite for Functional Analysis in Plant Genomics. Int J Plant Genomics 2008: 619832. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92. Götz S, García-Gómez JM, Terol J, Williams TD, Nagaraj SH, et al. (2008) High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res 36: 3420–3435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93. Götz S, Arnold R, Sebastián-León P, Martín-Rodríguez S, Tischler P, et al. (2011) B2G-FAR, a species-centered GO annotation repository. Bioinformatics 27: 919–924. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94. Anders S, Huber W (2010) Differential expression analysis for sequence count data. Genome Biology 11: R106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95. Eisen MB, Spellman PT, Brown PO, Botstein D (1998) Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci U S A 95: 14863–14868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96. Howe EA, Sinha R, Schlauch D, Quackenbush J (2011) RNA-Seq analysis in MeV. Bioinformatics 27: 3209–3210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97. Saldanha AJ (2004) Java Treeview–extensible visualization of microarray data. Bioinformatics 20: 3246–3248. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Length distribution of contigs, transcripts and unigenes.
(TIF)
Matching percentage of broccoli unigenes with different lengths to entries in public databases.
(TIF)
EggNOG classification of the broccoli seed and sprout transcriptome.
(TIF)
Classification of unigenes based on KEGG categorization.
(TIF)
Characterization of raw data and trimmed data.
(PDF)
Transcription factor members of every family detected in the broccoli seeds and sprouts.
(PDF)