Abstract
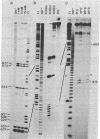
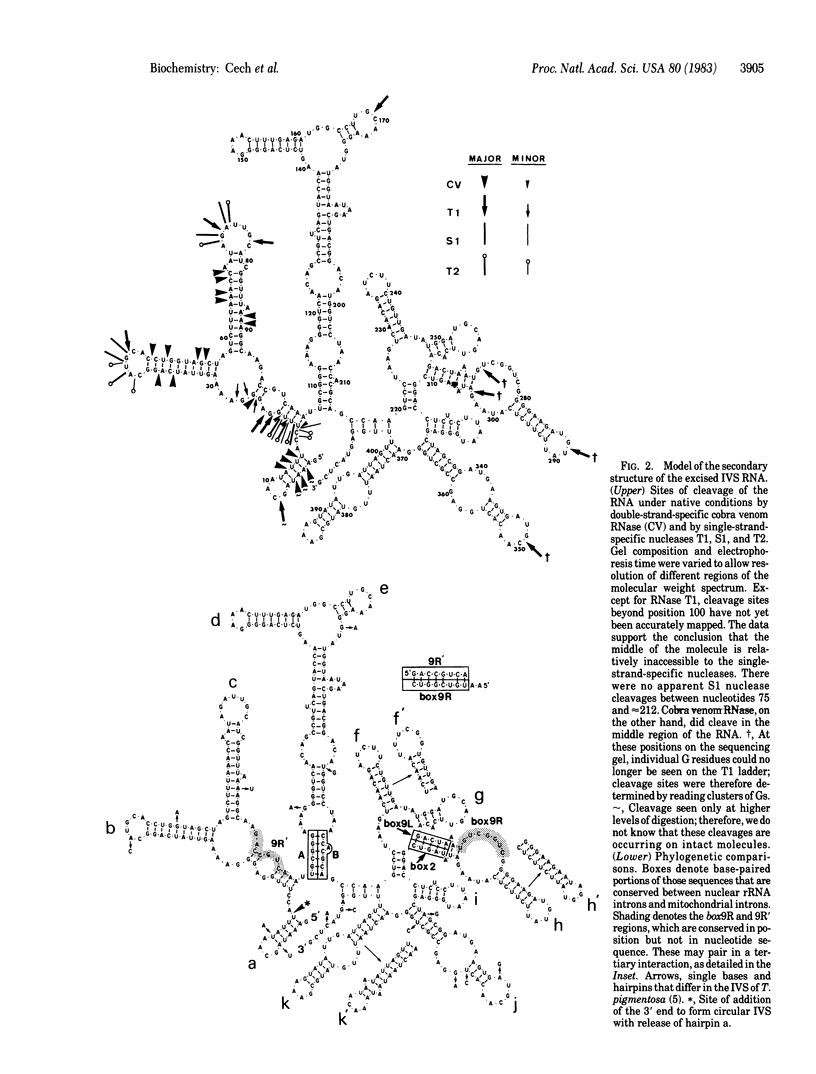
Splicing of the ribosomal RNA precursor of Tetrahymena is an autocatalytic reaction, requiring no enzyme or other protein in vitro. The structure of the intervening sequence (IVS) appears to direct the cleavage/ligation reactions involved in pre-rRNA splicing and IVS cyclization. We have probed this structure by treating the linear excised IVS RNA under nondenaturing conditions with various single- and double-strand-specific nucleases and then mapping the cleavage sites by using sequencing gel electrophoresis. A computer program was then used to predict the lowest-free-energy secondary structure consistent with the nuclease cleavage data. The resulting structure is appealing in that the ends of the IVS are in proximity; thus, the IVS can help align the adjacent coding regions (exons) for ligation, and IVS cyclization can occur. The Tetrahymena IVS has several sequences in common with those of fungal mitochondrial mRNA and rRNA IVSs, sequences that by genetic analysis are known to be important cis-acting elements for splicing of the mitochondrial RNAs. In the predicted structure of the Tetrahymena IVS, these sequences interact in a pairwise manner similar to that postulated for the mitochondrial IVSs. These findings suggest a common origin of some nuclear and mitochondrial introns and common elements in the mechanism of their splicing.
Full text
PDF



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Anziano P. Q., Hanson D. K., Mahler H. R., Perlman P. S. Functional domains in introns: trans-acting and cis-acting regions of intron 4 of the cob gene. Cell. 1982 Oct;30(3):925–932. doi: 10.1016/0092-8674(82)90297-5. [DOI] [PubMed] [Google Scholar]
- Bonitz S. G., Coruzzi G., Thalenfeld B. E., Tzagoloff A., Macino G. Assembly of the mitochondrial membrane system. Structure and nucleotide sequence of the gene coding for subunit 1 of yeast cytochrme oxidase. J Biol Chem. 1980 Dec 25;255(24):11927–11941. [PubMed] [Google Scholar]
- Burke J. M., RajBhandary U. L. Intron within the large rRNA gene of N. crassa mitochondria: a long open reading frame and a consensus sequence possibly important in splicing. Cell. 1982 Dec;31(3 Pt 2):509–520. doi: 10.1016/0092-8674(82)90307-5. [DOI] [PubMed] [Google Scholar]
- Davies R. W., Waring R. B., Ray J. A., Brown T. A., Scazzocchio C. Making ends meet: a model for RNA splicing in fungal mitochondria. Nature. 1982 Dec 23;300(5894):719–724. doi: 10.1038/300719a0. [DOI] [PubMed] [Google Scholar]
- De La Salle H., Jacq C., Slonimski P. P. Critical sequences within mitochondrial introns: pleiotropic mRNA maturase and cis-dominant signals of the box intron controlling reductase and oxidase. Cell. 1982 Apr;28(4):721–732. doi: 10.1016/0092-8674(82)90051-4. [DOI] [PubMed] [Google Scholar]
- Dieckmann C. L., Pape L. K., Tzagoloff A. Identification and cloning of a yeast nuclear gene (CBP1) involved in expression of mitochondrial cytochrome b. Proc Natl Acad Sci U S A. 1982 Mar;79(6):1805–1809. doi: 10.1073/pnas.79.6.1805. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Donis-Keller H., Maxam A. M., Gilbert W. Mapping adenines, guanines, and pyrimidines in RNA. Nucleic Acids Res. 1977 Aug;4(8):2527–2538. doi: 10.1093/nar/4.8.2527. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dujon B. Sequence of the intron and flanking exons of the mitochondrial 21S rRNA gene of yeast strains having different alleles at the omega and rib-1 loci. Cell. 1980 May;20(1):185–197. doi: 10.1016/0092-8674(80)90246-9. [DOI] [PubMed] [Google Scholar]
- Farrelly F., Butow R. A. Rearranged mitochondrial genes in the yeast nuclear genome. Nature. 1983 Jan 27;301(5898):296–301. doi: 10.1038/301296a0. [DOI] [PubMed] [Google Scholar]
- Fox G. E., Woese C. R. 5S RNA secondary structure. Nature. 1975 Aug 7;256(5517):505–507. doi: 10.1038/256505a0. [DOI] [PubMed] [Google Scholar]
- Grabowski P. J., Zaug A. J., Cech T. R. The intervening sequence of the ribosomal RNA precursor is converted to a circular RNA in isolated nuclei of Tetrahymena. Cell. 1981 Feb;23(2):467–476. doi: 10.1016/0092-8674(81)90142-2. [DOI] [PubMed] [Google Scholar]
- Kan N. C., Gall J. G. The intervening sequence of the ribosomal RNA gene is highly conserved between two Tetrahymena species. Nucleic Acids Res. 1982 May 11;10(9):2809–2822. doi: 10.1093/nar/10.9.2809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kruger K., Grabowski P. J., Zaug A. J., Sands J., Gottschling D. E., Cech T. R. Self-splicing RNA: autoexcision and autocyclization of the ribosomal RNA intervening sequence of Tetrahymena. Cell. 1982 Nov;31(1):147–157. doi: 10.1016/0092-8674(82)90414-7. [DOI] [PubMed] [Google Scholar]
- Lazowska J., Jacq C., Slonimski P. P. Sequence of introns and flanking exons in wild-type and box3 mutants of cytochrome b reveals an interlaced splicing protein coded by an intron. Cell. 1980 Nov;22(2 Pt 2):333–348. doi: 10.1016/0092-8674(80)90344-x. [DOI] [PubMed] [Google Scholar]
- Mannella C. A., Collins R. A., Green M. R., Lambowitz A. M. Defective splicing of mitochondrial rRNA in cytochrome-deficient nuclear mutants of Neurospora crassa. Proc Natl Acad Sci U S A. 1979 Jun;76(6):2635–2639. doi: 10.1073/pnas.76.6.2635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Michel F., Dujon B. Conservation of RNA secondary structures in two intron families including mitochondrial-, chloroplast- and nuclear-encoded members. EMBO J. 1983;2(1):33–38. doi: 10.1002/j.1460-2075.1983.tb01376.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Michel F., Jacquier A., Dujon B. Comparison of fungal mitochondrial introns reveals extensive homologies in RNA secondary structure. Biochimie. 1982 Oct;64(10):867–881. doi: 10.1016/s0300-9084(82)80349-0. [DOI] [PubMed] [Google Scholar]
- Morimoto R., Locker J., Synenki R. M., Rabinowitz M. Transcription, processing, and mapping of mitochondrial RNA from grande and petite yeast. J Biol Chem. 1979 Dec 25;254(24):12461–12470. [PubMed] [Google Scholar]
- Netter P., Jacq C., Carignani G., Slonimski P. P. Critical sequences within mitochondrial introns: cis-dominant mutations of the "cytochrome-b-like" intron of the oxidase gene. Cell. 1982 Apr;28(4):733–738. doi: 10.1016/0092-8674(82)90052-6. [DOI] [PubMed] [Google Scholar]
- Netzker R., Köchel H. G., Basak N., Küntzel H. Nucleotide sequence of Aspergillus nidulans mitochondrial genes coding for ATPase subunit 6, cytochrome oxidase subunit 3, seven unidentified proteins, four tRNAs and L-rRNA. Nucleic Acids Res. 1982 Aug 11;10(15):4783–4794. doi: 10.1093/nar/10.15.4783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ninio J. Prediction of pairing schemes in RNA molecules-loop contributions and energy of wobble and non-wobble pairs. Biochimie. 1979;61(10):1133–1150. doi: 10.1016/s0300-9084(80)80227-6. [DOI] [PubMed] [Google Scholar]
- Nobrega F. G., Tzagoloff A. Assembly of the mitochondrial membrane system. DNA sequence and organization of the cytochrome b gene in Saccharomyces cerevisiae D273-10B. J Biol Chem. 1980 Oct 25;255(20):9828–9837. [PubMed] [Google Scholar]
- Nomiyama H., Kuhara S., Kukita T., Otsuka T., Sakaki Y. Nucleotide sequence of the ribosomal RNA gene of Physarum polycephalum: intron 2 and its flanking regions of the 26S rRNA gene. Nucleic Acids Res. 1981 Nov 11;9(21):5507–5520. doi: 10.1093/nar/9.21.5507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nomiyama H., Sakaki Y., Takagi Y. Nucleotide sequence of a ribosomal RNA gene intron from slime mold Physarum polycephalum. Proc Natl Acad Sci U S A. 1981 Mar;78(3):1376–1380. doi: 10.1073/pnas.78.3.1376. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nussinov R., Tinoco I., Jr, Jacobson A. B. Small changes in free energy assignments for unpaired bases do not affect predicted secondary structures in single stranded RNA. Nucleic Acids Res. 1982 Jan 11;10(1):341–349. doi: 10.1093/nar/10.1.341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vasilenko S. K., Ryte V. C. [Isolation of highly purified ribonuclease from cobra (Naja oxiana) venom]. Biokhimiia. 1975 May-Jun;40(3):578–583. [PubMed] [Google Scholar]
- Waring R. B., Davies R. W., Scazzocchio C., Brown T. A. Internal structure of a mitochondrial intron of Aspergillus nidulans. Proc Natl Acad Sci U S A. 1982 Oct;79(20):6332–6336. doi: 10.1073/pnas.79.20.6332. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiss-Brummer B., Rödel G., Schweyen R. J., Kaudewitz F. Expression of the split gene cob in yeast: evidence for a precursor of a "maturase" protein translated from intron 4 and preceding exons. Cell. 1982 Jun;29(2):527–536. doi: 10.1016/0092-8674(82)90169-6. [DOI] [PubMed] [Google Scholar]
- Wickstrom E., Tinoco I., Jr The stability of RNA hairpin loops containing A-U-G: An-U-G-Um. Biopolymers. 1974 Nov;13(11):2367–2383. doi: 10.1002/bip.1974.360131116. [DOI] [PubMed] [Google Scholar]
- Wild M. A., Sommer R. Sequence of a ribosomal RNA gene intron from Tetrahymena. Nature. 1980 Feb 14;283(5748):693–694. doi: 10.1038/283693a0. [DOI] [PubMed] [Google Scholar]
- Wright R. M., Cummings D. J. Integration of mitochondrial gene sequences within the nuclear genome during senescence in a fungus. Nature. 1983 Mar 3;302(5903):86–88. doi: 10.1038/302086a0. [DOI] [PubMed] [Google Scholar]
- Wurst R. M., Vournakis J. N., Maxam A. M. Structure mapping of 5'-32P-labeled RNA with S1 nuclease. Biochemistry. 1978 Oct 17;17(21):4493–4499. doi: 10.1021/bi00614a021. [DOI] [PubMed] [Google Scholar]
- Yuan R. C., Steitz J. A., Moore P. B., Crothers D. M. The 3' terminus of 16S rRNA: secondary structure and interaction with ribosomal protein S1. Nucleic Acids Res. 1979 Dec 20;7(8):2399–2418. doi: 10.1093/nar/7.8.2399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zaug A. J., Grabowski P. J., Cech T. R. Autocatalytic cyclization of an excised intervening sequence RNA is a cleavage-ligation reaction. Nature. 1983 Feb 17;301(5901):578–583. doi: 10.1038/301578a0. [DOI] [PubMed] [Google Scholar]
- Zuker M., Stiegler P. Optimal computer folding of large RNA sequences using thermodynamics and auxiliary information. Nucleic Acids Res. 1981 Jan 10;9(1):133–148. doi: 10.1093/nar/9.1.133. [DOI] [PMC free article] [PubMed] [Google Scholar]