Abstract
Background
The human left and right atria have different susceptibilities to develop atrial fibrillation (AF). However, the molecular events related to structural and functional changes that enhance AF susceptibility are still poorly understood.
Objective
To characterize gene expression and genetic variation in human atria.
Methods
We studied the gene expression profiles and genetic variations in 53 left atrial and 52 right atrial tissue samples collected from the Myocardial Applied Genomics Network (MAGNet) repository. The tissues were collected from heart failure patients undergoing transplantation and from unused organ donor hearts with normal ventricular function. Gene expression was profiled using the Affymetrix GeneChip Human Genome U133A Array. Genetic variation was profiled using the Affymetrix Genome-Wide Human SNP Array 6.0.
Results
We found that 109 genes were differentially expressed between left and right atrial tissues. A total of 187 and 259 significant cis-associations between transcript levels and genetic variants were identified in left and right atrial tissues, respectively. We also found that a SNP at a known AF locus, rs3740293, was associated with the expression of MYOZ1 in both left and right atrial tissues.
Conclusion
We found a distinct transcriptional profile between the right and left atrium, and extensive cis-associations between atrial transcripts and common genetic variants. Our results implicate MYOZ1 as the causative gene at the chromosome 10q22 locus for AF.
Keywords: genetics, eQTL, gene expression, atrial tissues
The human left and right atria have distinct electrophysiological and pathophysiological differences. Atrial fibrillation (AF) is characterized predominantly by left atrial (LA) structural and contractile remodeling.1 Left atrial enlargement is also associated with many complex diseases such as stroke,2 obesity,3 and cardiovascular diseases.4, 5 In contrast, typical atrial flutter is known to be an arrhythmia arising from the right atrium.
Thousands of genes are expressed in the human heart,6 and many genes are well-known to be differentially regulated between the atria and ventricles7, 8 as well as in disease states including AF,9 heart failure,6, 10 and hypertrophy.11, 12 Earlier studies on mice and human atria have found several genes, including BMP10 and PITX2, are differentially expressed between left and right atria.13, 14 However, the sample sizes of these studies were usually small (<15), and the relation between atrial gene expression and genetic variants has not been systematically investigated.
In the past few years, genome wide association studies (GWAS) have been used successfully to identify thousands of genetic loci associated with a variety of diseases and phenotypic traits.15 Unfortunately, many novel candidate loci do not have defined functions, and the mechanisms to confer disease susceptibility remain largely unknown. Expression quantitative trait loci (eQTL) analyses can be used to correlate the relation between single nucleotide polymorphisms (SNPs) and gene expression. Such eQTL analysis is considered as an intermediate phenotype between genetic variations and diseases.16, 17 It would thus be interesting to determine if any genetic variations were associated with gene expression in atrial tissues.
Our objectives were three-fold: 1) to characterize the expression profiles of the right and left atria; 2) to perform an eQTL analysis in atrial tissue, and 3) using AF as an example, to determine if GWAS disease variants correlate with atrial gene expression.
Methods
Study Samples
We studied samples collected from 64 genetically-inferred European ancestry participants in the Myocardial Applied Genomics Network (MAGNet) repository. The tissues were collected from discarded hearts of heart failure patients undergoing transplantation and from unused organ donor hearts with normal ventricular function. Twelve had only LA, 11 had only right atrial (RA) tissue, and 41 individuals had both atrial tissues collected. The sample collection was approved by the Institutional Review Board at the University of Pennsylvania.
Genotyping
Genomic DNA was extracted using the Gentra Puregene Tissue Kit (Qiagen), which was then hybridized to the Affymetrix Genome-Wide Human SNP Array 6.0 in accordance with the manufacturer’s standard recommendations. The single nucleotide polymorphism (SNP) calling from raw CEL files were performed using Birdsuite software package.18 We filtered out SNPs with missing rate higher than 20%, minor allele frequency less than 5%, or Hardy–Weinberg Equilibrium P value less than 1×10−3 (Fisher’s exact test). At the end, 637,607 SNPs were used for downstream analyses. The quality control was performed using PLINK software package.19 All the participants were of European ancestry, which was inferred using multi-dimension scaling of analysis Affymetrix 6.0 genotypes with similar data from a larger cohort of know ancestry (N=340).
Transcriptional Profiling
The RNA was extracted using the Trizol reagent,20 and cDNA was hybridized to the Affymetrix Human Genome U133A Array according to the manufacturer’s instructions. The raw CEL files were pre-processed using Bioconductor R package.21 The data was quantile-normalized and log2 transformed, followed by summarization using Robust Multi-array Average.22 Potential batch effects were corrected by ComBat, an Empirical Bayes based approach.23 The gene annotation was downloaded from Affymetrix NetAffxTM Analysis Center (version 32). We excluded transcripts that were not aligned uniquely to the reference genome, and those that were not expressed in any of samples using MAS 5.0 algorithm.24, 25 Many genes were represented by multiple probesets, corresponding to different transcripts. We treated each transcript equally, and reported the transcript-based analysis, but the result was interpreted at the gene level. A total of 11,818 transcripts, corresponding to 8,644 genes, were used for the downstream analyses after adjusting for age and sex.
Differential Gene Expression
Differential expression between LA and RA tissues was assessed using unpaired Student’s t-test. To adjust for multiple testing, we used the highly conservative Bonferroni correction method. Significance was claimed if the P value was less than 4.2×10−6 (0.05/11,818 transcripts). The differential analysis was performed using R software packages (www.r-project.org/). We also used GOrilla web tool26 to test the enrichment of differentially expressed genes in Gene Ontology categories. We limited the enrichment analysis on the biological processes, and the enrichment was claimed if the FDR was less than 5%.27 Using a similar approach, we also compared the differential expression between participants with and without heart failure or AF.
Expression Quantitative Trait Loci (eQTL) Analysis
We used a linear regression model to test the association between genetic variations and gene expression. The significance was indicated by the regression P value. We defined cis-eQTLs as transcripts that were associated with SNPs within 1Mb from the gene boundary, and trans-eQTLs as transcripts that were associated with SNPs at least 1Mb far away or on different chromosomes.28 A total of 7,535,239,526 associations (637,607 SNP x 11,818 Transcripts) were tested. Empirical false discovery rate (FDR) was estimated by the permutation test (Supplemental Materials).
qPCR Confirmation of Significant Association
A total of 70 human LA samples were used for the replication studies. Surgical samples were obtained at Massachusetts General Hospital during cardiac surgery for valvular heart disease (n=32) or cardiac transplantation (n=27). Normal LA tissue was obtained from the National Disease Research Interchange repository (n=11). The sample collection was approved by the Institutional Review Board at Massachusetts General Hospital. Details of DNA and RNA extraction are available at the Supplemental Materials.
Results
Differential Gene Expression between the Left and Right Atrium
The clinical characteristics of the study sample from MAGNet are provided in Table 1. We observed that 109 transcripts, corresponding to 106 unique genes, were differentially expressed between left and right atria (P<4.2×10−6 by unpaired t-test). Thirty-three genes were expressed higher in LA tissue (Supplemental Table 1a) and 76 transcripts were expressed higher in RA tissue (Supplemental Table 1b). The most significantly over-expressed gene in LA tissue was AKR1B1 (P=6.0×10−13), which encodes a reductase that catalyzes the reduction of aldehydes. The most significantly over-expressed gene in RA tissue is SMAD6 (P=2.3×10−17), a transcriptional repressor critical for cardiac development.29 An analysis by GOrilla26 found that 218 biological processes were enriched with differentially expressed genes, and the top three were negative regulation of biological processes (P=4.1×10−9), anatomical structure development (P=5.4×10−9), and single-organism developmental processes (P=7.8×10−9).
Table 1.
Clinical characteristics of the study samples*
| Characteristics | Left Atrium (n=53) | Right Atrium (n=52) |
|---|---|---|
| Women, n (%) | 10 (18.9%) | 5 (9.6%) |
| Age, year ± SD | 55.9 ± 12.5 | 55.8 ± 13.3 |
| Advanced heart failure, n (%) | 47 (88.7%) | 48 (92.3%) |
| History of atrial fibrillation, n (%) | 9 (17.0%) | 9 (17.3%) |
Forty-one individuals had both left and right atrial tissue collected.
We also compared our results with the top 20 differentially expressed genes reported in a recently published article.14 As shown in Supplemental Table 2, 11 out of the 20 top genes were identified in the current study. Despite the difference in platforms and sample size, 9 of 11 shared genes also showed differential expression in current study (P<0.05). Only two of the previously reported genes, SALL1 (P=0.23) and KRT7 (P=0.13), were not significant, but the direction of over-expression for these two genes was consistent with the previous study.
eQTL Analyses: Association of Gene Expression with Genetic Variations
A total of 5,000,634 cis-associations were tested. We found 187 significant cis-associations in LA with FDR less than 5%, which was equivalent to P<1.3×10−6. The associations involved 181 SNPs and 46 transcripts (referred as eSNPs and eQTLs respectively). The most significant cis-association in LA tissues was POMZP3 with rs2110465 (P=2.2×10−13, R2=0.81). The SNP is located at 102kb away from POMZP3, a fusion of genes POM121 and ZP3. We also found 259 cis-associations in RA tissues with FDR less than 5%, which was equivalent to P<2.4×10−6. The associations involved 233 eSNPs and 67 eQTLs. The most significant cis-association in RA was RPS26 with rs11171739 (P=3.4×10−14, R2=0.83). The SNP is located at 33kb downstream of RPS26, which encodes a key component of the ribosome 40S subunit. We also compared the top cis-associations with those reported in other tissue types (http://eqtl.uchicago.edu/cgi-bin/gbrowse/eqtl/). About 45% of the top cis-associations for LA and 57% of the top cis-associations for RA already have been reported in other tissues. The full list of top cis-associations in LA and RA is provided in Supplemental Tables 3a and 3b, respectively.
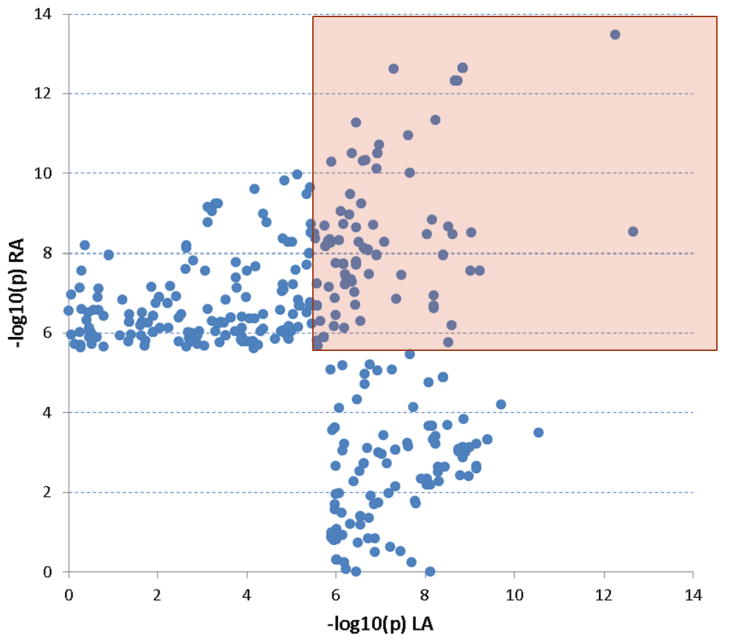
Figure 1 shows the comparison of top cis-associations between LA and RA tissues. Seventy-six cis-associations were significant in both LA and RA (FDR<5%), and 52 of them (68.4%) have been reported in other tissue types. Among the remaining 111 LA-specific cis-associations, 83 of them were also nominally significant in RA. Similarly, among the remaining 183 RA-specific cis-associations, 155 were nominally significant in LA. Our results suggest that the majority of cis-genetic determinants on gene expression are shared between LA and RA.
Figure 1. Comparison of top cis-associations between LA and RA.
Each point represents one cis-association. The x-axis is the −log10(p) of the association in LA, whereas the y-axis is the −log10(p) in RA. Cis-associations that were not significant (FDR<5%) in either LA or RA were not shown (lower left). Points within the upper right rectangle represent cis-associations that were significant in both LA and RA.
We also studied the association of genetic variations with trans-gene expression, which included evaluating 7,530,238,892 potential trans-associations. The most significant trans-association in RA was rs2419490 with PSPH (P=1.7×10−14, R2=0.84). In fact it was the only trans-association that reached the FDR cutoff of 5%. It was also the most significant trans-association in LA (P=1.6×10−11, R2=0.77) but did not reach the FDR cutoff in LA. The SNP is located in an intergenic region between LOC441242 and VKORC1L1, and it was 9.2 Mb upstream of PSPH, a gene encoding the phosphoserine phosphatase. The SNP is in complete linkage disequilibrium with another SNP (rs2419481), which is located in a DNAse hypersensitive region30 and could change the binding motifs of multiple transcription factors.31
Application of eQTL Analyses to AF-Related Genetic Loci
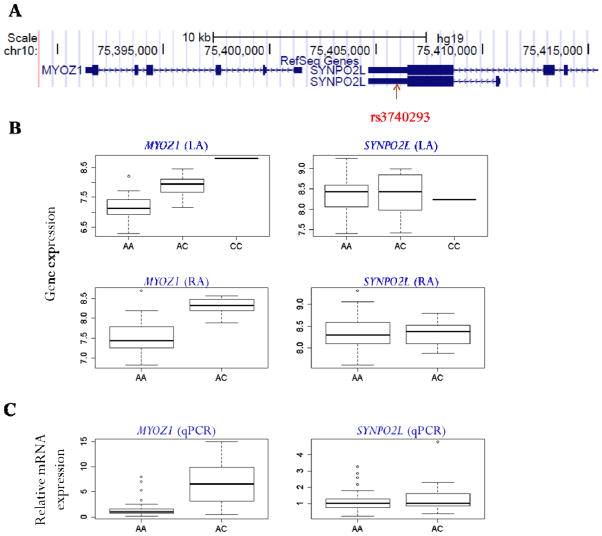
As an example of the potential utility of the transcriptional profiling and eQTL analyses generated in this study, we tested if the gene expression in atrial tissue was associated with 9 genome-wide AF-related loci.32 For each of the AF loci, we included the top SNP together with the neighboring SNPs within 500kb. We then tested the association of these SNPs with all the transcripts in the genome. The most significant association for both LA and RA was rs3740293 with MYOZ1 (P=4.0×10−9 for LA and 1.1×10−8 for RA). The SNP is located at the 3′ UTR of SYNPO2L, about 5kb upstream of MYOZ1 (Figure 2). However, it was not associated with SYNPO2L in either LA (P=0.90) or RA (P=0.97). The SNP is in high linkage disequilibrium (R2=0.80) with the top SNP at this locus (rs10824026), and it was strongly associated with AF (P=7.2×10−7).32
Figure 2. Cis-association between rs3740293 and MYOZ1.
A) The genomic location of rs3740293 from the UCSC Genome Browser (http://genome.ucsc.edu/cgi-bin/hgGateway). The SNP is located at the 3′UTR of SYNPO2L, approximately 5kb upstream of MYOZ1.
B) The association of rs3740293 with two MYOZ1 and SYNPO2L in LA and RA. The x-axis represents the genotype, and the y-axis is the boxplot of gene expression. The expression of MYOZ1 increases with the increasing copies of C allele of rs3740293 in both LA (upper left) and RA (lower left). No association was found between rs3740293 and SYNPO2L in either LA (upper right) or RA (lower right).
C) qPCR validation showing a significantly higher MYOZ1 expression in the left atrium of the samples with AC genotype as compared to AA genotype, whereas SYNPO2L expression did not differ between two groups.
In order to further confirm the association of rs3740293 with MYOZ1 expression, we performed qPCR validation in additional 70 samples collected from Massachusetts General Hospital (see Methods). Samples were divided into two groups based on their genotype: 57 samples with “AA” allele and 13 samples with “AC” allele. As shown in Figure 2c, the expression of MYOZ1 was significantly higher in samples with “AC” allele than those with “AA” allele (P=1.2×10−3). In contrast, no difference was observed for the expression of SYNPO2L (P=0.38).
Discussion
The overall goal of our study was to examine transcriptional profile differences between the LA and RA, to perform eQTL analyses in the atria, and then to apply our findings to AF. Our study is among the largest atrial expression analyses performed to date, and among the earliest attempts to investigate the impact of genome-wide genetic variations on atrial transcription. We have identified a distinct transcriptional profile between the left and right atrium, created a genome-wide map relating genetic variants to atrial gene expression, and applied these results to implicate the MYOZ1 gene in AF.
We began our study by creating a transcriptional map of the right versus the left atrium. It is expected that different tissues would have distinct transcriptional profiles that contribute to their unique anatomical structure and functional activity. Furthermore, gene expression is a quantitative and heritable phenotype that is mediated by the interplay of environmental and genetic factors.17, 33 We therefore used a repository with both right and left atrial tissue samples that were largely obtained at the time of cardiac surgery to define the transcriptional signature for each atrium. We found that over 100 genes were differentially expressed between the right and left atrium. As anticipated, many of the differentially expressed genes are involved in the anatomical structure development (P=5.4×10−9). Our findings also provide a transcriptional basis for the electrophysiological and pathophysiological differences between the atria. Genes that are preferentially expressed in the right versus the left atrium could be considered novel candidate genes for atrial flutter and atrial fibrillation, respectively.
We then genotyped each individual using a high-density SNP array of nearly a million SNPs. By combining our transcriptional and genetic data, we created a genome-wide map that correlates SNPs with the expression of genes either locally (cis) or at distance (trans). Such eQTL maps are powerful tools to determine which SNPs regulate gene expression and can identify functional SNPs and candidate genes within GWAS loci. Initial eQTL maps were derived from blood or lymphoblastoid cell lines;28, 33, 34 however, except for housekeeping genes, the genetic variants that control gene expression could be quite unique to a given tissue.28, 35 Thus, analysis of tissue-specific gene expression is essential.
We systemically studied the association of genetic variants with gene expression in atrial tissue, and identified hundreds of cis-associations. It is interesting to note that all but one of the significant associations that we observed were in cis, suggesting that the majority of genetic determinants of gene expression are likely to exert their effects on neighboring genes. Given that genome-wide association studies typically report the closest genes to the most significant SNPs, our result suggests that such practice is generally reasonable. The lack of additional significant trans-associations could be also attributable to an insufficient sample size to detect distant associations with small effects.
Finally, as an example of the potential utility of an atrial eQTL map, we sought to apply our results to AF. We systematically examined the relationship between the nine recently reported AF loci and expression of genes in the local region.32 Our results have strongly implicated MYOZ1 rather than SYNPO2L as the gene related to AF at the chromosome 10q22 locus. The MYOZ1 gene encodes calsarcin-2 or myozenin which is localized to the Z line and has been shown to interact with alpha-actinin, gamma-filamin and calcineurin.36, 37 Calsarcin-2 is predominately expressed in skeletal muscle,36 but is present at a lower level in cardiac tissue.36 Calsarcin-2 knockout mice had improved exercise performance and a switch to slow-twitch oxidative fibers in skeletal muscle.38 Although the relation between calsarcin-2 and AF is currently unknown, our findings prioritize MYOZ1 as gene worth further study as a potential mediator of AF.
Genome wide association studies have identified over thousands of disease or trait related loci in recent years and the vast majority of these loci are located in the intergenic or intronic regions.15 Similar to the analysis that we performed for AF, our atrial eQTL map could also be used to explore other atrial or cardiovascular traits including the PR or RR interval among others.39, 40
Our study has a number of potential limitations. The samples were from individuals of European ancestry, so our findings may not generalize to other races/ethnicities. They were collected from a mixture of heart failure or healthy donors. We had a modest number of samples, and a small number in the pre-specified subgroups with and without heart failure, and AF. Hence, our power for all analyses was modest (Supplemental Results). Given the similar transcriptional profiles in samples from both groups (Supplemental Figure 1), sample pooling would increase the statistical power comparing to treat each group separately (Supplemental Tables 4 and 5). The cross hybridization in microarray-based platforms might result in non-specific signals for some genes (e.g., HLA regions), thus further validation is warranted.41 Advances in RNA sequencing technologies is anticipated to be more accurate for detecting low-abundance or wide-dynamic-range transcripts.42 Given that we performed thousands or even millions of statistical tests, potential false findings might be a concern for our study. We thus applied a very stringent Bonferroni threshold or performed empirical imputation to define significant associations; however, such stringent thresholds might overlook subthreshold but nonetheless important associations.
In conclusion, we have characterized the transcriptional signature of the left and right atrium, created an eQTL map of the atria, and identified MYOZ1 as a candidate gene for AF. We anticipate that future increases in sample size and sequencing efforts will help to identify additional gene signatures underlying atrial structure and function.
Supplementary Material
Acknowledgments
This work is supported by NIH grants R01HL092577 (Ellinor and Benjamin), 1R01 HL102214 (Benjamin), R01HL104156 and K24HL105780 (Ellinor), R01HL105993 (Margulies and Cappola), KL2RR031981 and 1U01HL105268-01 (McManus), and American Heart Association Awards 13EIA14220013 (Ellinor) and 09FTF2190028 (Magnani).
Footnotes
Disclosures: The authors declare no commercial conflicts of interest.
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Allessie M, Ausma J, Schotten U. Electrical, contractile and structural remodeling during atrial fibrillation. Cardiovasc Res. 2002 May;54:230–246. doi: 10.1016/s0008-6363(02)00258-4. [DOI] [PubMed] [Google Scholar]
- 2.Benjamin EJ, D’Agostino RB, Belanger AJ, Wolf PA, Levy D. Left atrial size and the risk of stroke and death. The Framingham Heart Study. Circulation. 1995 Aug 15;92:835–841. doi: 10.1161/01.cir.92.4.835. [DOI] [PubMed] [Google Scholar]
- 3.Lavie CJ, Amodeo C, Ventura HO, Messerli FH. Left atrial abnormalities indicating diastolic ventricular dysfunction in cardiopathy of obesity. Chest. 1987 Dec;92:1042–1046. doi: 10.1378/chest.92.6.1042. [DOI] [PubMed] [Google Scholar]
- 4.Tsang TS, Barnes ME, Gersh BJ, Bailey KR, Seward JB. Left atrial volume as a morphophysiologic expression of left ventricular diastolic dysfunction and relation to cardiovascular risk burden. Am J Cardiol. 2002 Dec 15;90:1284–1289. doi: 10.1016/s0002-9149(02)02864-3. [DOI] [PubMed] [Google Scholar]
- 5.Miller JT, O’Rourke RA, Crawford MH. Left atrial enlargement: an early sign of hypertensive heart disease. Am Heart J. 1988 Oct;116:1048–1051. doi: 10.1016/0002-8703(88)90158-5. [DOI] [PubMed] [Google Scholar]
- 6.Putt ME, Hannenhalli S, Lu Y, et al. Evidence for coregulation of myocardial gene expression by MEF2 and NFAT in human heart failure. Circulation Cardiovascular genetics. 2009 Jun;2:212–219. doi: 10.1161/CIRCGENETICS.108.816686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Barth AS, Merk S, Arnoldi E, et al. Functional profiling of human atrial and ventricular gene expression. Pflugers Archiv : European journal of physiology. 2005 Jul;450:201–208. doi: 10.1007/s00424-005-1404-8. [DOI] [PubMed] [Google Scholar]
- 8.Skopek P, Hynie S, Chottova-Dvorakova M, et al. Effects of acute stressors on the expression of oxytocin receptor mRNA in hearts of rats with different activity of HPA axis. Neuro endocrinology letters. 2012;33:124–132. [PubMed] [Google Scholar]
- 9.Dupont E, Ko Y, Rothery S, et al. The gap-junctional protein connexin40 is elevated in patients susceptible to postoperative atrial fibrillation. Circulation. 2001 Feb 13;103:842–849. doi: 10.1161/01.cir.103.6.842. [DOI] [PubMed] [Google Scholar]
- 10.Jones SP, Greer JJ, van Haperen R, Duncker DJ, de Crom R, Lefer DJ. Endothelial nitric oxide synthase overexpression attenuates congestive heart failure in mice. Proc Natl Acad Sci U S A. 2003 Apr 15;100:4891–4896. doi: 10.1073/pnas.0837428100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Taniyama Y, Ito M, Sato K, et al. Akt3 overexpression in the heart results in progression from adaptive to maladaptive hypertrophy. J Mol Cell Cardiol. 2005 Feb;38:375–385. doi: 10.1016/j.yjmcc.2004.12.002. [DOI] [PubMed] [Google Scholar]
- 12.Matsui T, Li L, Wu JC, et al. Phenotypic spectrum caused by transgenic overexpression of activated Akt in the heart. J Biol Chem. 2002 Jun 21;277:22896–22901. doi: 10.1074/jbc.M200347200. [DOI] [PubMed] [Google Scholar]
- 13.Kahr PC, Piccini I, Fabritz L, et al. Systematic analysis of gene expression differences between left and right atria in different mouse strains and in human atrial tissue. PLoS ONE. 2011;6:e26389. doi: 10.1371/journal.pone.0026389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Hsu J, Hanna P, Van Wagoner DR, et al. Whole genome expression differences in human left and right atria ascertained by RNA sequencing. Circ Cardiovasc Genet. 2012 Jun;5:327–335. doi: 10.1161/CIRCGENETICS.111.961631. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Hindorff LA, Sethupathy P, Junkins HA, et al. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc Natl Acad Sci U S A. 2009 Jun 9;106:9362–9367. doi: 10.1073/pnas.0903103106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Chen Y, Zhu J, Lum PY, et al. Variations in DNA elucidate molecular networks that cause disease. Nature. 2008 Mar 27;452:429–435. doi: 10.1038/nature06757. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Emilsson V, Thorleifsson G, Zhang B, et al. Genetics of gene expression and its effect on disease. Nature. 2008 Mar 27;452:423–428. doi: 10.1038/nature06758. [DOI] [PubMed] [Google Scholar]
- 18.Korn JM, Kuruvilla FG, McCarroll SA, et al. Integrated genotype calling and association analysis of SNPs, common copy number polymorphisms and rare CNVs. Nat Genet. 2008 Oct;40:1253–1260. doi: 10.1038/ng.237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Purcell S, Neale B, Todd-Brown K, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. American journal of human genetics. 2007 Sep;81:559–575. doi: 10.1086/519795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Margulies KB, Matiwala S, Cornejo C, Olsen H, Craven WA, Bednarik D. Mixed messages: transcription patterns in failing and recovering human myocardium. Circ Res. 2005 Mar 18;96:592–599. doi: 10.1161/01.RES.0000159390.03503.c3. [DOI] [PubMed] [Google Scholar]
- 21.Gentleman RC, Carey VJ, Bates DM, et al. Bioconductor: open software development for computational biology and bioinformatics. Genome biology. 2004;5:R80. doi: 10.1186/gb-2004-5-10-r80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Irizarry RA, Hobbs B, Collin F, et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics (Oxford, England) 2003 Apr;4:249–264. doi: 10.1093/biostatistics/4.2.249. [DOI] [PubMed] [Google Scholar]
- 23.Johnson WE, Li C, Rabinovic A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics (Oxford, England) 2007 Jan;8:118–127. doi: 10.1093/biostatistics/kxj037. [DOI] [PubMed] [Google Scholar]
- 24.Hubbell E, Liu WM, Mei R. Robust estimators for expression analysis. Bioinformatics. 2002 Dec;18:1585–1592. doi: 10.1093/bioinformatics/18.12.1585. [DOI] [PubMed] [Google Scholar]
- 25.Liu WM, Mei R, Di X, et al. Analysis of high density expression microarrays with signed-rank call algorithms. Bioinformatics. 2002 Dec;18:1593–1599. doi: 10.1093/bioinformatics/18.12.1593. [DOI] [PubMed] [Google Scholar]
- 26.Eden E, Navon R, Steinfeld I, Lipson D, Yakhini Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinformatics. 2009;10:48. doi: 10.1186/1471-2105-10-48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. Journal of the Royal Statistical Society, Series B (Methodological) 1995;57:289–300. [Google Scholar]
- 28.Schadt EE, Molony C, Chudin E, et al. Mapping the genetic architecture of gene expression in human liver. PLoS Biol. 2008 May 6;6:e107. doi: 10.1371/journal.pbio.0060107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Galvin KM, Donovan MJ, Lynch CA, et al. A role for smad6 in development and homeostasis of the cardiovascular system. Nat Genet. 2000 Feb;24:171–174. doi: 10.1038/72835. [DOI] [PubMed] [Google Scholar]
- 30.Dunham I, Kundaje A, et al. ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature. 2012 Sep 6;489:57–74. doi: 10.1038/nature11247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Ernst J, Kheradpour P, Mikkelsen TS, et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature. 2011 May 5;473:43–49. doi: 10.1038/nature09906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Ellinor PT, Lunetta KL, Albert CM, et al. Meta-analysis identifies six new susceptibility loci for atrial fibrillation. Nat Genet. 2012;44:670–675. doi: 10.1038/ng.2261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Dixon AL, Liang L, Moffatt MF, et al. A genome-wide association study of global gene expression. Nat Genet. 2007 Oct;39:1202–1207. doi: 10.1038/ng2109. [DOI] [PubMed] [Google Scholar]
- 34.Stranger BE, Nica AC, Forrest MS, et al. Population genomics of human gene expression. Nat Genet. 2007 Oct;39:1217–1224. doi: 10.1038/ng2142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Powell JE, Henders AK, McRae AF, et al. Genetic control of gene expression in whole blood and lymphoblastoid cell lines is largely independent. Genome Res Mar. 2012;22:456–466. doi: 10.1101/gr.126540.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Takada F, Vander Woude DL, Tong HQ, et al. Myozenin: an alpha-actinin- and gamma-filamin-binding protein of skeletal muscle Z lines. Proc Natl Acad Sci U S A. 2001 Feb 13;98:1595–1600. doi: 10.1073/pnas.041609698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Frey N, Richardson JA, Olson EN. Calsarcins, a novel family of sarcomeric calcineurin- binding proteins. Proc Natl Acad Sci U S A. 2000 Dec 19;97:14632–14637. doi: 10.1073/pnas.260501097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Frey N, Frank D, Lippl S, et al. Calsarcin-2 deficiency increases exercise capacity in mice through calcineurin/NFAT activation. J Clin Invest. 2008 Nov;118:3598–3608. doi: 10.1172/JCI36277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Pfeufer A, van Noord C, Marciante KD, et al. Genome-wide association study of PR interval. Nat Genet. 2010 Feb;42:153–159. doi: 10.1038/ng.517. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Marroni F, Pfeufer A, Aulchenko YS, et al. A genome-wide association scan of RR and QT interval duration in 3 European genetically isolated populations: the EUROSPAN project. Circ Cardiovasc Genet. 2009 Aug;2:322–328. doi: 10.1161/CIRCGENETICS.108.833806. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Adam O, Lavall D, Theobald K, et al. Rac1-induced connective tissue growth factor regulates connexin 43 and N-cadherin expression in atrial fibrillation. J Am Coll Cardiol. 2010 Feb 2;55:469–480. doi: 10.1016/j.jacc.2009.08.064. [DOI] [PubMed] [Google Scholar]
- 42.Matkovich SJ, Zhang Y, Van Booven DJ, Dorn GW., 2nd Deep mRNA sequencing for in vivo functional analysis of cardiac transcriptional regulators: application to Galphaq. Circ Res. 2010 May 14;106:1459–1467. doi: 10.1161/CIRCRESAHA.110.217513. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.