Abstract
The rhoptries are key secretory organelles from apicomplexan parasites that contain proteins involved in invasion and modulation of the host cell. Some rhoptry proteins are restricted to the posterior bulb (ROPs) and others to the anterior neck (RONs). As many rhoptry proteins have been shown to be key players in Toxoplasma invasion and virulence, it is important to identify, understand and characterize the biological function of the components of the rhoptries. In this report, we identified putative novel rhoptry candidate genes by identifying Toxoplasma genes with similar cyclical expression profiles as known rhoptry protein encoding genes across its cell cycle. Using this approach we identified two new rhoptry bulb (ROP47 and ROP48) and one new rhoptry neck protein (RON12). ROP47 is secreted and traffics to the host cell nucleus, RON12 was not detected at the moving junction during invasion. Deletion of ROP47 or ROP48 in a type II strain did not show major influence in in vitro growth or virulence in mice.
Keywords: Toxoplasma gondii, Rhoptry, Rhoptry neck, Host-pathogen interaction
1. Introduction
Toxoplasma gondii is a highly successful parasite infecting approximately 30% of people worldwide and is the second largest cause of death due to foodborne illness in the United States (Scallan et al., 2011). Toxoplasmosis can be fatal in the unborn fetus and in immunosuppressed individuals if the disease is not recognized and treated early, in contrast to immunocompetent patients in whom it is mainly self-limited (Montoya and Liesenfeld, 2004; Scallan et al., 2011). Toxoplasma resides within a non-fusogenic parasitophorous vacuole and has three apical secretory organelles: the dense granules, micronemes and rhoptries. Rhoptries are club-shaped organelles divided into two distinct compartments, the posterior bulb and the more anterior duct (neck) through which rhoptry proteins are secreted. Proteins derived from the rhoptry secretory organelles are crucial for the invasion and survival of apicomplexan parasites within host cells and thus rhoptry protein targeting is a vital process for Toxoplasma. Some rhoptry proteins are restricted to the bulb (ROPs) and others to the neck (RONs). The role of rhoptries in the invasion process, virulence and/or host cell modulation has been well documented, although the molecular mechanisms remain only partially understood. RONs 2, 4, 5 and 8 have been shown to be involved in parasite invasion of the host cell (Besteiro et al., 2009; Straub et al., 2009); ROP16 activates the transcription factors STAT3 and STAT6 (Saeij et al., 2007; Yamamoto et al., 2009; Ong et al., 2010); ROP38 downregulates Mitogen Activated Protein Kinase (MAPK) pathway activation (Peixoto et al., 2010); and ROP5 and ROP18 act jointly to block immunity-related GTPase (IRG) mediated clearance of the parasite by the host cell (Behnke et al., 2012; Fleckenstein et al., 2012; Niedelman et al., 2012). Once rhoptry proteins are secreted into the host cell cytosol they traffic to distinct cellular destinations. For example, ROP16 and the rhoptry protein phosphatase 2 C (PP2C-hn) carry a nuclear localization signal (NLS), which mediates their trafficking to the host nucleus (Gilbert et al., 2007; Saeij et al., 2007), ROP5 and ROP18, following secretion into the host cell, traffic back to the outside of the parasitophorous membrane vacuole (PVM) via an arginine-rich amphipathic helix domain that is essential for this localization (Reese and Boothroyd, 2009; Fentress et al., 2012). Other rhoptry proteins, such as Toxofilin, remain in the host cell cytosol upon secretion (Lodoen et al., 2010).
Rhoptry proteins contain a classic eukaryotic signal peptide for entrance into the secretory pathway and are trafficked from the endoplasmic reticulum through the Golgi by a conserved pathway before being packaged into the apically located secretory organelles (Sadak et al., 1988; Bradley and Boothroyd, 1999; Bradley et al., 2004; Carey et al., 2004; Hajj et al., 2006b, 2007; Turetzky et al., 2010). N-terminal pro-domains have been implicated in rhoptry protein sorting and indeed several rhoptry proteins exhibit N-terminal processing. However, the failure to remove the pro-domain does not seem to disrupt targeting (Bradley et al., 2002; Miller et al., 2003; Turetzky et al., 2010). Other rhoptry proteins do not appear to be processed and the mechanism by which they are targeted to the rhoptries is unknown. Additionally, any soluble protein recombinantly fused to a signal peptide is delivered to the dense granules by default (Joiner and Roos, 2002). Therefore, motifs that mediate trafficking of proteins to the rhoptry organelles are insufficiently defined, or insufficiently specific, to allow genome-wide identification of rhoptry proteins.
Proteomic and genomic approaches have been widely used to identify the contents of the rhoptries of apicomplexan parasites (Hoppe et al., 2000; Bradley et al., 2005; Peixoto et al., 2010; Marugán-Hernández et al., 2011; Reid et al., 2012; Oakes et al., 2013). A proteomic study of Toxoplasma rhoptry contents led to the identification of 38 rhoptry proteins (Bradley et al., 2005). Twenty of these proteins were shown to localize to the rhoptry organelles, 11 to the rhoptry bulb and nine to the rhoptry neck (Bradley et al., 2005; Taylor et al., 2006; Gilbert et al., 2007; Proellocks et al., 2009; Straub et al., 2009; Peixoto et al., 2010; Lamarque et al., 2012). As expected, several previously known rhoptry proteins were readily detected in this proteomic analysis. However, TgNHE2 and TgSUB2, previously characterized as rhoptry proteins, were missed. The absence of these known rhoptry proteins is likely due to limitations of the technique, such as size cut-off or low amounts of protein.
Because several rhoptry proteins, such as the aforementioned ROP5, ROP16, ROP18 and ROP38, contain kinase-like domains (Hajj et al., 2006a; Peixoto et al., 2010), another study exploited a phylogenomic approach to characterize the Toxoplasma kinome, defining a 44-member family of kinase-like rhoptry proteins based on sequence similarities, including all previously reported kinase-like rhoptry proteins (Peixoto et al., 2010). To evaluate the accuracy of these predictions, nine of the identified kinase-like rhoptry proteins were confirmed to localize to the rhoptries, whereas two (ROP21 and ROP22) did not but were still annotated as rhoptry proteins. Surprisingly, the overlap between these two studies is relatively small, with only 11 genes found in common, which emphasizes the complementarity of different methodologies and the likelihood that there are still rhoptry proteins yet to be identified.
Previous characterization of the cell cycle transcriptome of Toxoplasma covering 12 h post-synchronization and nearly two tachyzoite replication cycles showed that the mRNA levels of proteins secreted from the rhoptry organelles display a cyclical expression profile, reaching peak levels in late S phase/early mitosis followed by a rapid and dramatic decline of these transcripts in early G1, before peaking again in the next S phase (Behnke et al., 2010). Inner membrane complex (IMC) mRNAs presented a very similar cyclical expression profile. Microneme mRNAs were offset by 1–2 h from rhoptry mRNAs and defined a distinct temporal class. By contrast, dense granule mRNAs largely were not regulated in the tachyzoite cell cycle.
Here we combined in silico and in vivo methods to identify novel rhoptry proteins likely to be involved in parasite modulation of host cells and found two novel rhoptry bulb proteins and one novel rhoptry neck protein.
2. Materials and methods
2.1. Parasites and cell lines
Parasites were maintained in vitro by serial passage on monolayers of human foreskin fibroblasts (HFFs) at 37°C in 5% CO2. HFFs were grown in DMEM supplemented with 10% FBS. C57BL6/J mouse embryonic fibroblasts (MEFs) were a gift from A. Sinai (University of Kentucky College of Medicine, Lexington, KY, USA) and were grown in HFF media supplemented with 10 mM HEPES, 1 mM sodium pyruvate and 1X non-essential amino acids.
2.2. Identification of candidate gene
For identification of new rhoptry protein coding genes, cell cycle transcriptome data of the Toxoplasma gondii during synchronized growth in human foreskin fibroblasts (HFFs) deposited at Gene Expression Omnibus (GEO)with accession number GSE19092 were used (Behnke et al., 2010). The data were normalized using Robust Multi Array (RMA) algorithm using Affymetrix Expression Console Software version 1.3.1, and all background values less than 6.5 were set to 6.5. Genes with an expression value of less than 6.5 in 12 or more samples where removed from the analysis. K-means clustering of each duplicated sample (time-point) was performed using the default distance metric (Pearson correlation) and a maximum number of 50, 45, 40, 35, 30, 25, 20, 12 and 10 clusters in Multiple Array Viewer 4.6. The cluster with the largest number of previously annotated rhoptry proteins was chosen for further analysis. The candidate genes were chosen using the following criteria: i) contained a predicted signal peptide, ii) had unannotated function and iii) unknown subcellular localization.
2.3. Gene tagging
Genomic sequences were obtained from the ToxoDB database (Version 8.0, ToxoDB.org) (Kissinger et al., 2003). For endogenous gene tagging (Huynh and Carruthers, 2009), primers were designed to amplify 1 – 3 kb of the predicted 3’ ends of genes. For heterologous expression of ROP47, the coding regions of TGME49_261740, together with putative promoter (~1500 bp upstream of ATG start codon) were amplified by PCR. All forward primers contained the 5’-CACC-3’ sequence required to perform directional TOPO cloning in pENTR/D-TOPO (Invitrogen, USA) and all reverse primers contained the hemagglutinin (HA) tag sequence followed by a stop codon (Table 1). HA-tagged sequences were cloned in vector pTKOatt by Gateway Recombination Cloning Technology (Invitrogen, USA). For endogenous gene tagging, the resulting vectors were linearized using a restriction enzyme with a unique restriction site within the cloned fragment. Linearized vector was transfected into RHΔhxgprtΔku80 (a gift from V. Carruthers, University of Michigan, Ann Arbor, MI, USA) parasites by electroporation. For heterologous expression of ROP47, vector was not linearized and was transfected into RHΔhxgprt. Electroporation was done in a 2 mm cuvette (Bio-Rad Laboratories, USA) with 2 mM ATP (MP Biomedicals, USA) and 5 mM glutathione (EMD, Germany) in a Gene Pulser Xcell (Bio-Rad Laboratories), with the following settings: 25 µFD, 1.25 kV, ∞Ω. Stable integrants were selected in media with 50 µg/ml of mycophenolic acid (Axxora, USA) and 50 µg/ml of xanthine (Alfa Aesar, USA) and cloned by limiting dilution. The correct tagging of each gene was confirmed by PCR, using a primer upstream of the plasmid integration site and a primer specific for the HA tag and/or by immunofluorescence (IF) analysis.
Table 1.
Primers used in this study.
| Gene ID | Forward primer | Reverse primer | ||
|---|---|---|---|---|
| Primers used for gene tagging (5'–3') | ||||
| TGME49_201860 | CACCTCGCGACCACCTGCGTTTTGA | TTACGCGTAGTCCGGGACGTCGTACGGGTAGAGTTCAGCGAAAGCACCTGC | ||
| TGME49_210370 | CACCACAGCCCGTTACGTCTTGCGAC | TTACGCGTAGTCCGGGACGTCGTACGGGTAAACGGAGGGAAGAAACGGGG | ||
| TGME49_218270 | CACCGGCTGGAGATTTTTCCCCGGAACT | TTACGCGTAGTCCGGGACGTCGTACGGGTAAGCCGGACTTGCAGAAGGCAC | ||
| TGME49_225160 | CACCCCATGCTTCTGTTCGCAGAAAT | TTACGCGTAGTCCGGGACGTCGTACGGGTATGAATCCTTGAAACTGCGAATC | ||
| TGME49_232020 | CACCTGGCTCTCCAGCCGCGGCGATT | TTACGCGTAGTCCGGGACGTCGTACGGGTATCGTCGCCGGCGCCTTCCGC | ||
| TGME49_237180 | CACCAGAAACCGCTGCTGAGGAATGG | TTACGCGTAGTCCGGGACGTCGTACGGGTACATAAGAAATTTTATTTTATGGAGGCG | ||
| TGME49_258360 | CACCGCGGTGTTCGTCAGGATCTGCT | TTACGCGTAGTCCGGGACGTCGTACGGGTACCCGTTAAGATGCGCAACGAC | ||
| TGME49_261740 | CACCACAAAACGGGGAGCA | TTACGCGTAGTCCGGGACGTCGTACGGGTACGGTCTTTTTCCACCTTTCACACG | ||
| Primers used for gene knock-out (5'–3') | ||||
| TGME49_201860 | GGGGACAAGTTTGTACAAAAAAGCAGGCTTATGCTTCGTACGACACCAACG | GGGGACAACTTTGTATAGAAAAGTTGTGGCTAATGTATCCACGTGG | ||
| TGME49_201860 | GGGGACAACTTTGTATAATAAAGTTGCTGAGGTCCAAGCAGCACCGAA | GGGGACCACTTTGTACAAGAAAGCTGGGTAGAGAGAGTCGGGTTCGTTCG | ||
| TGME49_218270 | GGGGACAAGTTTGTACAAAAAAGCAGGCTTAAAGACCTCTCGCCTCGGATT | GGGGACAACTTTGTATAGAAAAGTTGCTT AGGCTAAGTGTTGGAGC | ||
| TGME49_218270 | GGGGACAACTTTGTATAATAAAGTTGCTCTAGTCGTCTCGGATGATGG | GGGGACCACTTTGTACAAGAAAGCTGGGTATTGTCGCTGGAGGATTCATGG | ||
| TGME49_261740 | GGGGACAAGTTTGTACAAAAAAGCAGGCTGCTTAGTAACCTGCGGATACACTTC | GGGGACAACTTTGTATAGAAAAGTTGGGTGGACCGCCGAACATCATTGTTTC | ||
| TGME49_261740 | GGGGACAACTTTGTATAGAAAAGTTGGGTGGACCGCCGAACATCATTGTTTC | GGGGACCACTTTCTTGTACAAAGTGGTGGTAAAGGGCATGTTTTGACACGGG | ||
| P1 | P2 | P3 | P4 | |
| TGME49_201860 | CGTGACTTGAAATGCAGTCC | GATCCAGACGTCTTCAATGC | CACCTCGCGACCACCTGCGTTTTGA | TTACGCGTAGTCCGGGACGTCGTACGGGTAGAGTTCAGCGAAAGCACCTGC |
| TGME49_218270 | CACTTCGACTGACATCTCAG | GATCCAGACGTCTTCAATGC | CACCGGCTGGAGATTTTTCCCCGGAACT | TTACGCGTAGTCCGGGACGTCGTACGGGTAAGCCGGACTTGCAGAAGGCAC |
| Primers used for generation of recombinant ROP47 peptide antigen (5'–3') | ||||
| TGME49_261740 | CACCATTTGTCTCGCCGC | CTTTCTTTTTCGACCTTTCACACG | ||
2.4. IF analysis
Parasites were allowed to invade cells on coverslips and incubated for 16 – 24 h. The cells were then fixed with 3% (vol/vol) formaldehyde in PBS for 20 min at room temperature and blocked in PBS with 3% (wt/vol) BSA and 5% (vol/vol) goat serum. Coverslips were incubated with primary antibody for 1 h at room temperature or overnight at 4°C, and fluorescent secondary antibodies and Hoechst dye were used for antigen and DNA visualization, respectively. Coverslips were mounted on a glass slide with Vectashield (Vector Laboratories, USA), and photographs were taken using NIS-Elements software (Nikon, Japan) and a digital camera (CoolSNAP EZ; Roper Industries, USA) connected to an inverted fluorescence microscope (model eclipse Ti-S; Nikon). To determine seroconversion, peripheral blood serum was used as a primary antibody. For co-localization experiments, GFP-expressing parasites were fixed with methanol or 3% (vol/vol) formaldehyde. For moving junction staining, this standard IF protocol was modified slightly. Parasites were added to HFFs on coverslips, spun down to bring them into contact with host cells, and allowed to attach to and invade host cells for 5 min at 37°C. Unattached parasites were washed off with PBS, and cells were fixed with 3% (vol/vol) formaldehyde in PBS for 20 min at room temperature, blocked in PBS with 5% (vol/vol) FBS and 5% (vol/vol) normal goat serum for 1 – 2 h at room temperature, and permeabilized by incubation in PBS with 0.2% (wt/vol) saponin at 37°C for 20 min.
2.5. Antibodies
Recombinant ROP47 peptide antigen was generated by PCR amplification of sequences immediately downstream of the predicted ROP47 signal peptide and immediately upstream of the ROP47 stop codon from PruΔhxgprt genomic DNA. The amplicon was cloned in frame into pET102/D-TOPO (Invitrogen), creating a Thioredoxin-ROP47-V5-His6 construct. This protein was expressed in BL21 Star (Invitrogen) cells and purified on a Ni-NTA column (Invitrogen) eluted with 250 mM imidazole and dialyzed under native conditions according to the manufacturer’s directions. Rabbit polyclonal antibodies (Covance, USA) were raised against the purified peptide antigen. Antibodies were affinity purified from the ROP47 antiserum with Affi-Gel 10 resin (Bio-Rad) covalently coupled to the antigen peptide and eluted with 100 mM Glycine, pH 2.5, then immediately neutralized with 1 M Tris pH 8.0. The eluate was subsequently incubated with Affi-Gel 10 resin covalently coupled to a Thioredoxin-V5-His6 peptide that was expressed and purified as above to remove antibodies in the serum reacting against thioredoxin and the epitope tags. The flow-through was collected, tested for specificity and used for immunogenic assays.
Antibodies against HA (Roche), Toxoplasma surface antigen (SAG)-1 (DG52, (Burg et al., 1988)), Toxoplasma rhoptry protein ROP1 (Tg49; (Ossorio et al., 1992)), Toxoplasma dense granule protein GRA7 (Dunn et al., 2008), Toxoplasma rhoptry neck protein RON4 (generously provided by P. Bradley, University of California, Los Angeles, CA, USA), Toxoplasma inner membrane complex proteins IMC1 and MLP1 (generously provided by M.J. Gubbels, Boston College, Boston, MA, USA) and mouse TGTP (A-20; Santa Cruz Biotechnology, USA) were used in the IF assay. IF secondary antibodies were coupled with Alexa Fluor 488 or Alexa Fluor 594 (Invitrogen).
2.6. Western blot
Parasites were syringe lysed from infected HFFs with lysis buffer, boiled for 5 min and subjected to 10% SDS-PAGE. Proteins were transferred to a polyvinylidene difluoride membrane, which was blocked in PBS/0.1% Tween-20/5% non-fat dry milk and incubated with primary and secondary antibodies. The blot was incubated with a luminal-based substrate (Immun-Star WesternC; Bio-Rad Laboratories) and chemiluminescence was detected using a charge-coupled device camera (Chemidoc XRS; Bio-Rad Laboratories). The bands were visualized using Quantity One 1-D analysis software.
2.7. Generation of gene knockouts (KOs)
The 5’ and 3’ flanking regions of the genes to be knocked out were cloned in pTKO2 (Rosowski et al., 2011) around the hypoxanthine-xanthine-guanine ribosyl transferase (HXGPRT) selectable marker using Multisite Gateway Pro 3-Fragment Recombination (Invitrogen). Flanking regions (5’ and 3’) of TGME49_201860, TGME49_218270 and TGME49_261740 were cloned from type II genomic DNA. Primers contained att recombination sites (denoted in primer sequence with italics, Table 1) and amplified ≈2 kb upstream of the start codon and downstream of the stop codon. These flanking regions were then cloned around the HXGPRT selectable marker flanked by 5’ and 3’ untranslated region (UTRs) from dihydrofolate reductase (DHFR), as previously described (Rosowski et al., 2011). Before transfection, the KO vector was linearized. PruΔhxgprtΔku80 (a gift from D. Bzik, Dartmouth Medical School, Lebanon, NH, USA) parasites were transfected with the KO construct by electroporation, and stable integrants were selected and cloned by limiting dilution, as described in Section 2.3. PCR with a forward primer upstream of the 5’ flanking region (P1) and a reverse primer within the HXGPRT cassette (P2) confirmed the disruption in the desired loci (Supplementary Fig. S1A). Additionally, PCR was performed to confirm the inability to amplify the target genes (P3 and P4) (Supplementary Fig S1A).
2.8. Plaque assays
For the plaque assays, 100–500 parasites per well were added to monolayers of MEFs seeded the day before and either previously stimulated with 1000 U/mL of mouse IFNγ or left unstimulated for 24 h before infection in a 24 well plate in MEF media. Infections were then incubated for 5 days at 37°C and the number of plaques was counted using a microscope.
2.9. Animal infections
Six to 10 week old female C57BL/6J mice (The Jackson Laboratory, USA) were used in all experiments. For i.p. infection, tachyzoites were grown in vitro and extracted from host cells by passage through a 30-gauge needle, washed twice in PBS and quantified with a hemocytometer. Parasites were diluted in PBS and mice were inoculated i.p. with 500 tachyzoites of each strain (in 100 µL) using a 28-gauge needle. For oral infection, brain homogenate of chronically infected mice was stained with dolichos biflorus-FITC (Vector Laboratories) and cysts were enumerated by microscopy. The mice were orally gavaged with 1000 cysts. All of the animals were monitored daily and weighed three times per week. Peripheral blood serum was collected on days 7 and 30 of the experiment and the levels of IFNγ were determined using commercially available ELISA kits, according to the manufacturer’s protocol (eBioscience, USA). The Massachusetts Institute for Technology, USA, Committee on Animal Care approved all protocols. All mice were maintained under specific pathogen-free conditions, in accordance with institutional and federal regulations.
3. Results and Discussion
3.1. Identification of novel rhoptry proteins
Previous characterization of the Toxoplasma cell cycle transcriptome showed that the mRNA levels of rhoptry encoding genes display the same cyclical expression profile (Behnke et al., 2010). We hypothesized that unidentified rhoptry encoding genes would show a similar expression profile. Genes were identified that showed the same expression pattern as established rhoptry encoding genes by performing K-means clustering of 13 duplicate samples spanning 12 h post-thymidine release into the tachyzoite cell cycle. The cluster with the largest number of previously annotated rhoptry protein encoding genes (67 out of 75 annotated rhoptry encoding genes were expressed above background and were present on the Toxoplasma array; Supplementary Table S1) was chosen for further analysis (Table 2, Supplementary Fig. S2). This cluster contained 190 unique genes, of which 45 encoded proteins annotated as rhoptries (out of 67 possible genes) and four encoded IMC proteins. Thirty-one out of these 45 have been previously confirmed to localize to the rhoptry organelles.
Table 2.
Toxoplasma genes in the rhoptry cluster.
| Probeset_id | ToxoDB V8_ID | Product Description | Subcellular localization |
Reference | Confirmed | Exons | Transmembrane Domains |
Predicted Signal Peptide |
Cell Cycle Microarray RMA |
|
|---|---|---|---|---|---|---|---|---|---|---|
| Max | Min | |||||||||
| 20.m08222 | TGME49_203990 | rhoptry protein ROP12 (ROP12) | RHOPTRY | Bradley et al., 2005 | Confirmed | 3 | 1 | Yes | 12.0 | 7.7 |
| 20.m03896 | TGME49_205250 | rhoptry protein ROP18 (ROP18) | RHOPTRY | Taylor et al., 2006 | Confirmed | 1 | 1 | Yes | 13.8 | 10.6 |
| 27.m00091 | TGME49_211290 | rhoptry protein ROP15 (ROP15) | RHOPTRY | Bradley et al., 2005 | Confirmed | 6 | 0 | Yes | 12.7 | 8.9 |
| 33.m02185 | TGME49_214080 | toxofilin | RHOPTRY | Bradley et al., 2005 | Confirmed | 1 | 1 | Yes | 12.2 | 8.9 |
| 33.m01398 | TGME49_215775 | rhoptry protein ROP8 (ROP8) | RHOPTRY | Beckers et al., 1997 | Confirmed | 1 | 1 | Yes | 13.0 | 9.4 |
| 42.m00026 | TGME49_223920 | rhoptry neck protein RON3 (RON3) | RHOPTRY | Bradley et al., 2005 | Confirmed | 21 | 1 | Yes | 12.3 | 7.9 |
| 42.m03584 | TGME49_227810 | rhoptry kinase family protein ROP11 (incomplete catalytic triad) (ROP11) |
RHOPTRY | Bradley et al., 2005 | Confirmed | 1 | 0 | Yes | 13.1 | 9.0 |
| 44.m06355 | TGME49_229010 | rhoptry neck protein RON4 (RON4) | RHOPTRY | Bradley et al., 2005 | Confirmed | 20 | 0 | Yes | 11.8 | 7.7 |
| 44.m00026 | TGME49_230350 | rhoptry neck protein RON11 (RON11) | RHOPTRY | Beck et al., 2013 | Confirmed | 16 | 4 | 10.4 | 6.5 | |
| 49.m03277 | TGME49_242240 | rhoptry kinase family protein ROP19A (ROP19A) |
RHOPTRY | Peixoto et al., 2010 | Confirmed | 1 | 0 | Yes | 10.7 | 7.4 |
| 52.m01543 | TGME49_252360 | rhoptry kinase family protein ROP24 (incomplete catalytic triad) (ROP24) |
RHOPTRY | Peixoto et al., 2010 | Confirmed | 1 | 0 | Yes | 11.8 | 8.0 |
| 55.m04748 | TGME49_258230 | rhoptry kinase family protein ROP20 (ROP20) |
RHOPTRY | Peixoto et al., 2010 | Confirmed | 1 | 1 | Yes | 10.1 | 6.5 |
| 55.m08191 | TGME49_258580 | rhoptry protein ROP17 (ROP17) | RHOPTRY | Peixoto et al., 2010 | Confirmed | 1 | 0 | Yes | 13.1 | 9.8 |
| 55.m00092 | TGME49_258660 | rhoptry protein ROP6 (ROP6) | RHOPTRY | Sohn et al., 1999 | Confirmed | 5 | 1 | Yes | 13.3 | 9.5 |
| 55.m00167 | TGME49_261750 | rhoptry neck protein RON10 (RON10) | RHOPTRY | Lamarque et al., 2012 | Confirmed | 7 | 0 | 11.3 | 6.7 | |
| 55.m08219 | TGME49_262730 | rhoptry protein ROP16 (ROP16) | RHOPTRY | Bradley et al., 2005 | Confirmed | 1 | 0 | Yes | 11.9 | 8.5 |
| 59.m03479 | TGME49_269885 | rhoptry metalloprotease toxolysin TLN1 (TLN1) |
Hajagos et al., 2011 | Confirmed | 12 | 1 | Yes | 11.6 | 6.9 | |
| 74.m00767 | TGME49_282055 | protein phosphatase PP2C–hn (PP2CHN) |
RHOPTRY | Gilbert et al., 2007 | Confirmed | 13 | 0 | 11.5 | 6.9 | |
| 80.m02343 | TGME49_291960 | rhoptry kinase family protein ROP40 (incomplete catalytic triad) (ROP40) |
RHOPTRY | Peixoto et al., 2010 | Confirmed | 3 | 0 | Yes | 13.1 | 9.2 |
| 83.m02145 | TGME49_295110 | rhoptry protein ROP7 (ROP7) | RHOPTRY | Hajj et al., 2006 | Confirmed | 1 | 1 | Yes | 12.7 | 8.6 |
| 113.m00009 | TGME49_297960 | rhoptry neck protein RON6 (RON6) | RHOPTRY | Proellocks et al., 2006 | Confirmed | 10 | 1 | Yes | 11.5 | 7.0 |
| 129.m00252 | TGME49_299060 | sodium/hydrogen exchanger NHE2 | RHOPTRY | Karasov et al., 2005 | Confirmed | 12 | 13 | 10.4 | 6.5 | |
| 145.m00331 | TGME49_300100 | rhoptry neck protein RON2 (RON2) | RHOPTRY | Bradley et al., 2005 | Confirmed | 10 | 3 | Yes | 11.4 | 6.5 |
| 541.m00141 | TGME49_306060 | rhoptry neck protein RON8 (RON8) | RHOPTRY | Straub et al., 2009 | Confirmed | 15 | 1 | Yes | 11.9 | 7.0 |
| 551.m00238 | TGME49_308090 | rhoptry protein ROP5 (ROP5) | RHOPTRY | Bradley et al., 2005 | Confirmed | 1 | 0 | Yes | 13.5 | 10.2 |
| 583.m09207 | TGME49_308810 | rhoptry neck protein RON9 (RON9) | RHOPTRY | Lamarque et al., 2012 | Confirmed | 9 | 1 | 10.8 | 6.5 | |
| 583.m00003 | TGME49_309590 | rhoptry protein ROP1 (ROP1) | RHOPTRY | Ossorio et al., 1992 | Confirmed | 1 | 0 | Yes | 12.6 | 9.4 |
| 583.m00597 | TGME49_310010 | rhoptry neck protein RON1 (RON1) | RHOPTRY | Bradley et al., 2005 | Confirmed | 5 | 1 | Yes | 11.3 | 6.8 |
| 583.m00636 | TGME49_311470 | rhoptry neck protein RON5 (RON5) | RHOPTRY | Straub et al., 2009 | Confirmed | 38 | 0 | Yes | 12.3 | 7.6 |
| 583.m00692 | TGME49_315220 | rhoptry protein ROP14 (ROP14) | RHOPTRY | Bradley et al., 2005 | Confirmed | 11 | 11 | 10.9 | 6.5 | |
| 583.m05686 | TGME49_315490 | rhoptry protein ROP10 (ROP10) | RHOPTRY | Bradley et al., 2005 | Confirmed | 1 | 0 | Yes | 11.1 | 6.6 |
| 49.m05689 | TGME49_242118 | myosin-light-chain kinase | RHOPTRY | Peixoto et al., 2010 | 1 | 1 | 10.8 | 6.5 | ||
| 20.m00331 | TGME49_202200 | hypothetical protein | RHOPTRY | Bradley et al., 2005 | 7 | 0 | 12.0 | 7.9 | ||
| 27.m00846 | TGME49_211260 | rhoptry kinase family protein ROP26 (incomplete catalytic triad) (ROP26) |
RHOPTRY | Peixoto et al, 2010 | 2 | 0 | 12.7 | 9.8 | ||
| 41.m01337 | TGME49_222100 | hypothetical protein | RHOPTRY | Bradley et al., 2005 | 1 | 0 | Yes | 11.0 | 6.5 | |
| 49.m03276 | TGME49_242230 | rhoptry kinase family protein ROP29 (ROP29) |
RHOPTRY | Peixoto et al, 2010 | 1 | 1 | 10.8 | 6.9 | ||
| 49.m03399 | TGME49_244250 | hypothetical protein | RHOPTRY | Bradley et al., 2005 | 4 | 0 | 11.2 | 6.9 | ||
| 52.m01529 | TGME49_252200 |
Toxoplasma palmitoyl acyltransferase TgDHHC7 |
RHOPTRY | Beck et al, 2013 | 8 | 4 | 10.1 | 6.5 | ||
| 52.m01582 | TGME49_253370 | hypothetical protein (RON4L1) | RHOPTRY | Boothroyd and Dubremetz, 2008 |
30 | 0 | Yes | 10.0 | 6.5 | |
| 55.m04788 | TGME49_258800 | rhoptry kinase family protein ROP31 (ROP31) |
RHOPTRY | Peixoto et al., 2010 | 1 | 1 | Yes | 10.0 | 6.9 | |
| 55.m05020 | TGME49_262920 | trypsin domain-containing protein | RHOPTRY | Bradley et al., 2005 | 11 | 1 | Yes | 9.8 | 6.5 | |
| 83.m01271 | TGME49_294560 | rhoptry kinase family protein ROP37 (incomplete catalytic triad) (ROP37) |
RHOPTRY | Peixoto et al., 2010 | 1 | 0 | Yes | 10.6 | 6.5 | |
| 83.m01285 | TGME49_294790 | hypothetical protein | RHOPTRY | Bradley et al., 2005 | 1 | 0 | 13.0 | 9.7 | ||
| 113.m00755 | TGME49_297070 | hypothetical protein | RHOPTRY | Bradley et al., 2005 | 2 | 0 | Yes | 12.6 | 8.2 | |
| 583.m00694 | TGME49_315210 | rhoptry protein, putative (ROP14B) | RHOPTRY | Reid et al., 2012 | 11 | 7 | 10.7 | 6.5 | ||
| 20.m00355 | TGME49_201520 | protein phosphatase 2C domain-containing protein | 9 | 0 | Yes | 11.0 | 6.6 | |||
| 20.m05880 | TGME49_201520 | protein phosphatase 2C domain-containing protein | 9 | 0 | Yes | 11.0 | 6.5 | |||
| 20.m03673 | TGME49_201760 | hypothetical protein | 4 | 0 | 10.5 | 6.5 | ||||
| 20.m03682 | TGME49_201860 | hypothetical protein | 2 | 1 | Yes | 13.3 | 9.8 | |||
| 20.m03718 | TGME49_202420 | hypothetical protein | 2 | 1 | 9.2 | 6.5 | ||||
| 20.m00003 | TGME49_202500 | GAPM1a | 4 | 6 | 13.7 | 10.2 | ||||
| 20.m03760 | TGME49_203010 | aurora kinase | 1 | 0 | 9.9 | 6.5 | ||||
| 20.m05981 | TGME49_203930 | hypothetical protein | 6 | 5 | 10.6 | 6.6 | ||||
| 20.m03880 | TGME49_204880 | hypothetical protein | 1 | 0 | Yes | 9.3 | 6.5 | |||
| 20.m03902 | TGME49_205330 | hypothetical protein | 2 | 8 | 12.7 | 8.6 | ||||
| 20.m03905 | TGME49_205360 | hypothetical protein | 20 | 2 | Yes | 9.7 | 6.5 | |||
| 20.m03977 | TGME49_206710 | hypothetical protein | 5 | 0 | 8.5 | 6.5 | ||||
| 25.m01833 | TGME49_208910 | hypothetical protein | 3 | 0 | 9.2 | 6.5 | ||||
| 25.m01852 | TGME49_209170 | hypothetical protein | 1 | 0 | Yes | 9.6 | 6.5 | |||
| 25.m01855 | TGME49_209200 | hypothetical protein | 12 | 0 | 9.0 | 6.5 | ||||
| 25.m01859 | TGME49_209250 | hypothetical protein | 7 | 1 | 10.8 | 7.8 | ||||
| 26.m00235 | TGME49_210270 | hypothetical protein | 3 | 0 | 10.1 | 7.6 | ||||
| 26.m00242 | TGME49_210370 | hypothetical protein | 1 | 0 | Yes | 12.2 | 8.2 | |||
| 26.m00370 | TGME49_210420 | hypothetical protein | 1 | 1 | 11.3 | 7.1 | ||||
| 27.m00828 | TGME49_210820 | hypothetical protein | 3 | 10 | 9.3 | 6.5 | ||||
| 28.m00429 | TGME49_211850 | hypothetical protein | 3 | 0 | 11.8 | 9.1 | ||||
| 31.m00934 | TGME49_212980 | hypothetical protein | 2 | 1 | 11.8 | 8.6 | ||||
| 33.m02670 | TGME49_214400 | hypothetical protein | 2 | 0 | 10.7 | 6.7 | ||||
| 35.m00004 | TGME49_216080 | apical complex lysine methyltransferase |
4 | 0 | 11.3 | 8.3 | ||||
| 35.m00895 | TGME49_216620 | EF hand domain-containing protein | 39 | 7 | 9.0 | 6.5 | ||||
| 37.m00748 | TGME49_217520 | hypothetical protein | 2 | 0 | Yes | 12.5 | 8.8 | |||
| 38.m01037 | TGME49_218240 | hypothetical protein | 2 | 1 | Yes | 9.6 | 6.5 | |||
| 38.m01040 | TGME49_218270 | hypothetical protein | 1 | 8 | Yes | 12.7 | 8.4 | |||
| 41.m01283 | TGME49_221250 | hypothetical protein | 8 | 0 | 9.8 | 6.5 | ||||
| 41.m00036 | TGME49_221620 | beta-tubulin, putative | 4 | 0 | 13.1 | 9.7 | ||||
| 41.m01316 | TGME49_221675 | hypothetical protein | 9 | 1 | Yes | 8.9 | 6.5 | |||
| 42.m03399 | TGME49_225020 | hypothetical protein | 4 | 0 | 9.5 | 6.5 | ||||
| 42.m03409 | TGME49_225160 | hypothetical protein | 1 | 2 | Yes | 11.5 | 7.8 | |||
| 42.m07438 | TGME49_225200 | hypothetical protein | 33 | 0 | Yes | 9.2 | 6.5 | |||
| 42.m00061 | TGME49_225320 | hypothetical protein | 5 | 0 | Yes | 10.8 | 6.5 | |||
| 42.m00060 | TGME49_225330 | hypothetical protein | 6 | 0 | Yes | 9.8 | 6.5 | |||
| 42.m03456 | TGME49_225860 | hypothetical protein | 4 | 0 | 9.8 | 6.5 | ||||
| 42.m03481 | TGME49_226220 | alveolin domain containing intermediate filament IMC9 (ALV6/IMC9) |
9 | 0 | 9.7 | 6.5 | ||||
| 42.m00093 | TGME49_227000 | hypothetical protein | 16 | 0 | 9.9 | 6.5 | ||||
| 44.m02549 | TGME49_229500 | hypothetical protein | 2 | 3 | 9.8 | 6.6 | ||||
| 44.m02600 | TGME49_230480 | hypothetical protein | 1 | 2 | 11.4 | 7.5 | ||||
| 44.m02630 | TGME49_231000 | START domain-containing protein | 18 | 1 | 9.0 | 6.5 | ||||
| 44.m02636 | TGME49_231070 | protein kinase | 1 | 0 | 10.8 | 6.8 | ||||
| 44.m02644 | TGME49_231160 | hypothetical protein | 3 | 0 | 12.7 | 9.5 | ||||
| 44.m02678 | TGME49_231840 | hypothetical protein | 6 | 0 | 9.7 | 6.5 | ||||
| 44.m02696 | TGME49_232020 | hypothetical protein | 2 | 1 | 8.8 | 6.5 | ||||
| 44.m02714 | TGME49_232260 | hypothetical protein | 3 | 3 | 10.0 | 6.5 | ||||
| 44.m02750 | TGME49_232780 | hypothetical protein | 8 | 0 | 9.5 | 6.5 | ||||
| 46.m01616 | TGME49_234540 | hypothetical protein | 7 | 9 | 10.2 | 6.5 | ||||
| 46.m02875 | TGME49_235130 | transmembrane protein | 4 | 0 | Yes | 9.3 | 6.5 | |||
| 46.m01643 | TGME49_235380 | hypothetical protein | 11 | 0 | 8.9 | 6.5 | ||||
| 46.m01716 | TGME49_236860 | haloacid dehalogenase family hydrolase domain-containing protein |
2 | 12 | 9.8 | 6.5 | ||||
| 46.m01722 | TGME49_236960 | transporter, major facilitator family protein |
10 | 10 | 10.8 | 6.8 | ||||
| 46.m01740 | TGME49_237180 | hypothetical protein | 1 | 0 | Yes | 12.1 | 7.8 | |||
| 46.m01741 | TGME49_237190 | hypothetical protein | 4 | 0 | 9.7 | 6.5 | ||||
| 46.m01743 | TGME49_237210 | Tyrosine kinase-like (TKL) protein | 4 | 0 | 10.2 | 6.5 | ||||
| 49.m03087 | TGME49_238150 | hypothetical protein | 1 | 8 | 12.7 | 10.2 | ||||
| 49.m03179 | TGME49_239830 | TBC domain-containing protein | 13 | 0 | 10.7 | 6.5 | ||||
| 49.m03213 | TGME49_240460 | AP2 domain transcription factor AP2VI-1 (AP2VI1) |
2 | 0 | 10.9 | 6.8 | ||||
| 49.m07198 | TGME49_240730 | hypothetical protein | 2 | 4 | 9.0 | 6.5 | ||||
| 49.m07245 | TGME49_241000 | hypothetical protein | 1 | 4 | 11.0 | 7.0 | ||||
| 49.m03332 | TGME49_243200 | hypothetical protein | 5 | 0 | 12.2 | 6.6 | ||||
| 49.m00056 | TGME49_243690 | hypothetical protein | 4 | 1 | 12.1 | 8.0 | ||||
| 49.m03388 | TGME49_244080 | hypothetical protein | 3 | 1 | Yes | 10.7 | 6.5 | |||
| 49.m03412 | TGME49_244470 | hypothetical protein | 8 | 0 | 10.0 | 7.0 | ||||
| 50.m03074 | TGME49_245550 | hypothetical protein | 1 | 4 | Yes | 10.0 | 6.5 | |||
| 50.m03107 | TGME49_246182 | hypothetical protein | 10.1 | 7.2 | ||||||
| 50.m03131 | TGME49_246710 | hypothetical protein | 1 | 3 | 9.0 | 6.5 | ||||
| 50.m03132 | TGME49_246720 | hypothetical protein | 1 | 0 | 10.2 | 7.2 | ||||
| 50.m03154 | TGME49_247195 | microneme protein MIC15 (MIC15) | 42 | 1 | Yes | 9.8 | 6.5 | |||
| 50.m03253 | TGME49_248690 | hypothetical protein | 7 | 0 | 9.4 | 6.8 | ||||
| 50.m03302 | TGME49_249440 | hypothetical protein | 2 | 0 | 9.9 | 6.5 | ||||
| 50.m03315 | TGME49_249570 | hypothetical protein | 20 | 1 | Yes | 11.4 | 6.8 | |||
| 52.m01567 | TGME49_253140 | hypothetical protein | 3 | 0 | 9.3 | 6.5 | ||||
| 52.m01583 | TGME49_253380 | AP2 domain transcription factor AP2III-2 (AP2III2) |
1 | 0 | 10.5 | 6.8 | ||||
| 52.m01598 | TGME49_253600 | hypothetical protein | 11 | 0 | 9.6 | 6.5 | ||||
| 52.m01644 | TGME49_254290 | hypothetical protein | 1 | 0 | 9.0 | 6.5 | ||||
| 55.m04629 | TGME49_255700 | hypothetical protein | 6 | 0 | 9.7 | 6.5 | ||||
| 55.m08188 | TGME49_256030 | hypothetical protein | 6 | 0 | 12.1 | 7.6 | ||||
| 55.m04752 | TGME49_258360 | hypothetical protein | 7 | 0 | Yes | 11.6 | 7.0 | |||
| 55.m00096 | TGME49_258700 | transporter, major facilitator family protein |
12 | 11 | 9.3 | 6.5 | ||||
| 55.m04796 | TGME49_258900 | hypothetical protein | 0 | 9.3 | 6.5 | |||||
| 55.m04843 | TGME49_259700 | hypothetical protein | 2 | 3 | 11.1 | 6.9 | ||||
| 55.m00144 | TGME49_260820 | IMC sub-compartment protein ISP1 (ISP1) |
3 | 0 | 12.0 | 8.5 | ||||
| 55.m04955 | TGME49_261740 | hypothetical protein | 1 | 1 | Yes | 14.2 | 11.9 | |||
| 57.m01689 | TGME49_264600 | hypothetical protein | 2 | 4 | 12.2 | 7.5 | ||||
| 57.m01720 | TGME49_265080 | Tubulin-tyrosine ligase family protein | 12 | 0 | 9.6 | 6.5 | ||||
| 57.m01755 | TGME49_265650 | protein phosphatase 2C domain-containing protein | 1 | 0 | Yes | 9.7 | 6.5 | |||
| 57.m01783 | TGME49_266300 | hypothetical protein | 2 | 0 | 12.3 | 8.0 | ||||
| 57.m01792 | TGME49_266435 | hypothetical protein | 34 | 0 | 9.4 | 6.5 | ||||
| 57.m01834 | TGME49_267070 | aquaporin 2 | 1 | 11 | 11.6 | 7.7 | ||||
| 59.m03403 | TGME49_268760 | hypothetical protein | 12.2 | 8.6 | ||||||
| 59.m00087 | TGME49_268870 | tetratricopeptide repeat-containing protein |
10 | 0 | 10.3 | 6.5 | ||||
| 59.m00092 | TGME49_269330 | hypothetical protein | 10 | 0 | 10.1 | 6.6 | ||||
| 59.m00029 | TGME49_269340 | hypothetical protein | 4 | 1 | 11.4 | 6.9 | ||||
| 59.m03542 | TGME49_270890 | hypothetical protein | 3 | 0 | 9.4 | 6.5 | ||||
| 59.m00038 | TGME49_271270 | hypothetical protein | 10 | 1 | 11.2 | 7.0 | ||||
| 59.m03707 | TGME49_273860 | hypothetical protein | 1 | 0 | 11.0 | 8.2 | ||||
| 64.m00327 | TGME49_275670 | alveolin domain containing intermediate filament IMC15 (ALV5/IMC15) |
9 | 0 | 10.8 | 6.5 | ||||
| 64.m00582 | TGME49_276130 | cathepsin CPC2 (CPC2) | 10 | 0 | Yes | 9.7 | 6.5 | |||
| 65.m01152 | TGME49_278130 | hypothetical protein | 14 | 0 | 9.3 | 6.5 | ||||
| 65.m01964 | TGME49_278510 | protein phosphatase 2C domain-containing protein | 10 | 1 | Yes | 11.1 | 7.3 | |||
| 65.m02537 | TGME49_278920 | hypothetical protein | 5 | 6 | 9.5 | 6.5 | ||||
| 69.m00143 | TGME49_279420 | hypothetical protein | 1 | 0 | 11.3 | 6.5 | ||||
| 72.m00385 | TGME49_280480 | EF hand domain-containing protein | 1 | 4 | 11.5 | 6.8 | ||||
| 72.m00685 | TGME49_280670 | hypothetical protein | 7 | 10 | Yes | 11.3 | 6.7 | |||
| 74.m00455 | TGME49_282070 | hypothetical protein | 9 | 0 | 10.8 | 6.5 | ||||
| 74.m00465 | TGME49_282210 | AP2 domain transcription factor AP2VIIa-8 (AP2VIIA8) |
2 | 0 | 9.3 | 6.5 | ||||
| 76.m01597 | TGME49_285290 | hypothetical protein | 1 | 3 | Yes | 10.3 | 6.5 | |||
| 76.m01608 | TGME49_285650 | hypothetical protein | 2 | 0 | 8.9 | 6.5 | ||||
| 76.m01626 | TGME49_285870 | SAG-related sequence SRS20A (SRS20A) |
2 | 1 | Yes | 12.8 | 8.7 | |||
| 76.m01662 | TGME49_286500 | hypothetical protein | 1 | 10 | 9.9 | 6.5 | ||||
| 80.m02122 | TGME49_287970 | hypothetical protein | 3 | 0 | 10.4 | 6.5 | ||||
| 80.m02181 | TGME49_288950 | AP2 domain transcription factor AP2IX-4 (AP2IX4) |
1 | 0 | 9.7 | 6.5 | ||||
| 80.m03982 | TGME49_289150 | hypothetical protein | 5 | 1 | 9.4 | 6.5 | ||||
| 80.m03946 | TGME49_289970 | hypothetical protein | 3 | 0 | 11.7 | 9.3 | ||||
| 83.m01220 | TGME49_293540 | hypothetical protein | 6 | 0 | 10.3 | 6.5 | ||||
| 83.m01311 | TGME49_295100 | hypothetical protein | 4 | 0 | 9.5 | 6.9 | ||||
| 86.m00370 | TGME49_295420 | hypothetical protein | 9 | 0 | 10.4 | 7.0 | ||||
| 86.m00377 | TGME49_295620 | hypothetical protein | 4 | 0 | 9.7 | 6.5 | ||||
| 113.m01286 | TGME49_298010 | hypothetical protein | 52 | 0 | 9.1 | 6.5 | ||||
| 145.m00603 | TGME49_300220 | hypothetical protein | 15 | 4 | Yes | 9.4 | 6.5 | |||
| 145.m00607 | TGME49_300360 | ADP/ATP translocase | 6 | 2 | 9.5 | 6.5 | ||||
| 162.m00326 | TGME49_301420 | hypothetical protein | 7 | 0 | 11.8 | 7.5 | ||||
| 541.m00127 | TGME49_305250 | hypothetical protein | 6 | 6 | 10.5 | 7.1 | ||||
| 541.m01166 | TGME49_305270 | hypothetical protein | 6 | 0 | Yes | 10.2 | 6.5 | |||
| 541.m00131 | TGME49_305510 | hypothetical protein | 7 | 0 | Yes | 11.9 | 7.9 | |||
| 541.m01185 | TGME49_305590 | ABC transporter transmembrane region domain-containing protein |
9 | 9 | 9.5 | 6.5 | ||||
| 542.m00226 | TGME49_307020 | hypothetical protein | 3 | 0 | 9.4 | 6.5 | ||||
| 551.m00232 | TGME49_308010 | hypothetical protein | 1 | 0 | 9.0 | 6.5 | ||||
| 583.m05275 | TGME49_309160 | IgA-specific metalloendopeptidase | 1 | 0 | Yes | 10.4 | 6.5 | |||
| 583.m09102 | TGME49_310240 | hypothetical protein | 1 | 2 | 10.3 | 6.5 | ||||
| 583.m11449 | TGME49_310740 | hypothetical protein | 2 | 7 | 9.3 | 6.5 | ||||
| 583.m00659 | TGME49_311800 | endonuclease/exonuclease/phosphatase family protein |
6 | 0 | 9.1 | 6.6 | ||||
| 583.m00645 | TGME49_312150 | hypothetical protein | 6 | 1 | 11.7 | 7.6 | ||||
| 583.m09217 | TGME49_313780 | hypothetical protein | 1 | 0 | 11.1 | 7.3 | ||||
| 583.m09134 | TGME49_314260 | hypothetical protein | 1 | 1 | 12.1 | 8.2 | ||||
| 583.m05709 | TGME49_315780 | myosin regulatory light chain, putative | 7 | 0 | 11.5 | 8.6 | ||||
| 583.m05738 | TGME49_316260 | hypothetical protein | 2 | 7 | 12.3 | 8.1 | ||||
| 583.m09158 | TGME49_316280 | transporter, major facilitator family protein |
4 | 8 | 11.7 | 7.3 | ||||
| 583.m05758 | TGME49_316540 | IMC sub-compartment protein ISP3 (ISP3) |
1 | 0 | 11.1 | 8.0 | ||||
| 611.m00052 | TGME49_317705 | enoyl-CoA hydratase/isomerase family protein |
8 | 0 | 10.5 | 7.2 | ||||
| 641.m01483 | TGME49_318470 | AP2 domain transcription factor AP2IV-4 (AP2IV4) |
1 | 0 | 9.3 | 6.5 | ||||
| 641.m00181 | TGME49_320740 | hypothetical protein | 5 | 1 | Yes | 11.7 | 8.6 | |||
| 645.m00324 | TGME49_321600 | hypothetical protein | 7 | 0 | 9.6 | 6.5 | ||||
Genes highlighted in grey were further characterized in this study.
RMA, Robust Multi-array Average
The Toxoplasma genome is predicted to encode 8127 genes, 1920 of which have a signal peptide (23.4%). Our analysis imposed no explicit selection for genes coding for proteins with signal peptide sequences; however, the resulting list is enriched (65 genes with a predicted signal peptide out of 190, P = 0.0003, Hypergeometric distribution) in signal-peptide-containing proteins (34%).
Interestingly, 22 out of the 75 tachyzoite rhoptry proteins encoding genes never clustered with the remainder of the rhoptries (Supplementary Table S1). Two of these genes (RON2L1 and BRP1 (Schwarz et al., 2005; Fritz et al., 2012)) encode bradyzoite- or sporozoite-specific rhoptry proteins and six encode confirmed tachyzoite rhoptries (ROP9, ROP38, ROP39, TgARO, Toxolysin 1 and Subtilisin 2). According to the cell cycle expression data (Behnke et al., 2010), ROP9, ROP38 and ROP39 do not display an obvious cyclical expression profile. In addition, ROP9 was shown to be secreted with micronemal proteins, in a calcium-dependent manner (Kawase et al., 2007). The cellular localization of the proteins encoded by the other 14 genes annotated as rhoptry protein genes (Supplementary Table S1) was never confirmed by IF and it is therefore possible that these are not localized to the rhoptries.
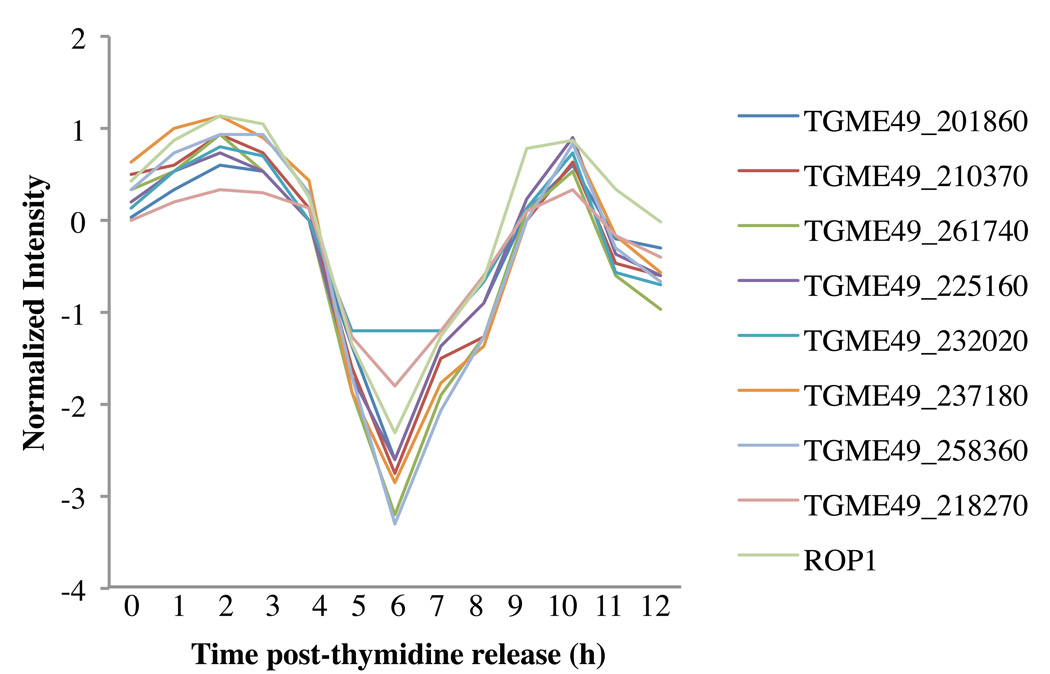
These results underscore the relevance of our approach and strongly suggest that our rhoptry protein cluster might contain many of the remainder of the unidentified rhoptry protein-encoding genes of tachyzoites. To identify new putative rhoptry protein encoding genes that could modulate host cell functions, we chose eight genes that encoded a protein with a predicted signal peptide, unannotated function and unknown subcellular localization from within our rhoptry cluster. The eight candidate genes are described in Table 2, highlighted in grey, and all display the cyclic expression profile described above (Fig. 1).
Fig. 1.
Identification of new Toxoplasma rhoptry proteins. The gene expression profile of the entire genome of Toxoplasma was compared with the cyclical gene expression profile of rhoptry genes throughout the cell cycle by performing clustering of duplicate samples spanning 12 h post-synchronization (Supplementary Fig. S2). Shown are eight genes that displayed an expression profile signature similar to the rhoptry pattern and that were chosen for further analysis. The expression profile of ROP1 is also shown.
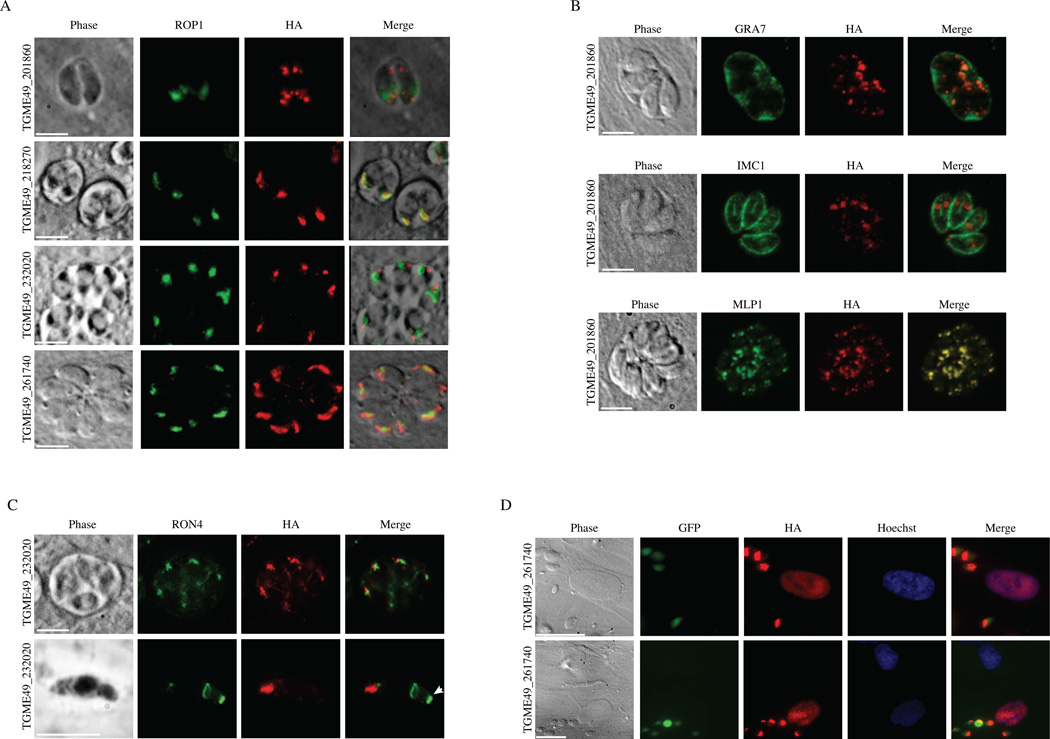
To determine the localization of the candidate gene products within the parasite, we genetically engineered Toxoplasma strains expressing an HA-tagged version of the candidate genes at their endogenous loci. We were unable to endogenously tag TGME49_261740 and therefore determined its localization by heterologous expression of a C-terminal HA-tagged copy of the candidate gene, including at least 1500 bp of the putative endogenous promoter. The correct tagging of each endogenously tagged gene was confirmed by PCR (Supplementary Fig. S3). IF analysis was performed on HFFs infected with each of the HA-tagged parasites. The staining pattern of four out of eight candidate gene products was unlike that of the secretory organelles and appeared to label the whole parasite (Supplementary Fig. S4). The genes whose labeling appeared consistent with that of the secretory organelles were TGME49_201860, TGME49_218270, TGME49_232020 and TGME49_261740 (Fig. 2A). Some characteristics of the proteins encoded by these genes are described in Table 3.
Fig. 2.
Cellular localization of C-terminally tagged genes. Human foreskin fibroblasts (HFFs) were infected with RH expressing hemagglutinin (HA)-tagged TGME49_201860, TGME49_218270 (rop48), TGME49_232020 (ron12) or TGME49_261740 (rop47), fixed and stained with (A) α-HA and rhoptry protein marker α–ROP1, (B) α-HA and dense granule protein marker α-GRA7, α-HA and inner membrane complex marker α-IMC1, α-HA and inner membrane complex marker α-MLP1. (C) α-HA and rhoptry neck/moving junction marker, α-RON4 and (D) α–HA, α-ROP47 and Hoechst dye. (A–C) Scale bars represent 5 µm. (D) Scale bars represent 20 µm.
Table 3.
Characteristics of new putative rhoptry protein encoding genes identified in this study
| Candidate gene ToxoDB V8 ID |
Alias | Signal peptide | Transmembrane domains |
Conserved domains |
Paralogue (% protein identity) |
Paralogue present in rhoptry cluster |
Homologue in Neospora caninum (% protein identity) |
|---|---|---|---|---|---|---|---|
| TGME49_201860 | - | Yes | 1 | - | TGME49_301390 (31%) | No | NCLIV_022900 (75%) |
| TGME49_218270 | ROP48 | Yes | 8 | - | TGME49_209810 (29%) | No | NCLIV_061950 (71%) |
| TGME49_232020 | RON12 | Yes, in GT1 and CEP | 1 | - | TGME49_244726 (27%) | No | NCLIV_032020 (59%) NCLIV_025740 (43%) |
| TGME49_261740 | ROP47 | Yes | 1 | - | - |
The product of TGME49_201860-HA appears to have a punctate distribution, present at both the posterior and the apical end of the parasite (Fig. 2A). TGME49_218270-HA seems to label the rhoptry bulb. We were unable to detect these two proteins using an anti-HA western blot, and therefore do not know whether they undergo post-transcriptional processing.
TGME49_232020-HA localizes to the apical pole of the parasite, a labeling consistent with the rhoptry neck (Fig. 2A). This gene is predicted to encode a protein with no signal peptide in type II parasites. However, as first exon prediction is notoriously difficult, in type I and type III parasites, this protein is predicted have a signal peptide. Moreover, TGME49_232020 encodes a protein with a predicted molecular weight of 135 kDa. However, western blot analysis of the TGME49_232020-HA strain detected a band of approximately 40 kDa, instead of the predicted 135 kDa (Supplementary Fig. S5). Indeed, several rhoptry proteins exhibit N-terminal processing and that is also probably the case for TGME49_232020. The rhoptry subtilisin TgSUB2 recognizes and cleaves after the consensus sequence SΦXE (Miller et al., 2003). There is a putative SUB2 cleavage site (SPQE) between amino acids 968 and 971 of TGME49_232020. Cleavage at this site would generate a ~32 kDa tagged product, which could be consistent with the band observed. Longer exposures did not reveal a 135 kDa pro-protein, suggesting that the half-life of the pro-protein is very short.
TGME49_261740-HA appears to label the entire rhoptry organelle (Fig. 2A). Interestingly, TGME49_261740 is in the top five of the most highly expressed genes across the Toxoplasma cell cycle (Behnke et al., 2010) and is one of the most polymorphic Toxoplasma genes (Minot et al., 2012). Anti-HA western blotting of this protein detected a band of approximately 15 kDa, consistent with the predicted size of 14 kDa (Supplementary Fig. S1B).
These four proteins are conserved between Toxoplasma and Neospora (Table 3), but protein BLAST analysis did not reveal the presence of close (30% or more protein identity) homologues in other apicomplexans. Additionally, no known conserved domains were detected, giving no indication about the potential function of these proteins.
3.2. TGME49_218270, TGME49_232020 and TGME49_261740 encode new rhoptry proteins
To further confirm the localization of TGME49_201860, TGME49_218270, TGME49_232020 and TGME49_261740 within the parasite, we performed co-staining of each protein with a rhoptry bulb marker, ROP1, on intracellular parasites (Fig. 2A). TGME49_218270-HA shows a perfect co-localization with ROP1.
TGME49_201860-HA does not overlap with this rhoptry marker. To further investigate the cellular localization of this protein, we co-stained the TGME49_201860-HA expressing parasites with dense granule (GRA7) and inner membrane complex (IMC1 and MIP1-like protein-1 (MLP1)) markers (Fig. 2B). TGME49_201860-HA appears to partially overlap with GRA7. In addition, while it does not co-localize with IMC1, it co-localizes with MLP1, a protein that is present at the apical cap and basal complex of mature as well as budding daughter parasites (MJ Gubbels, personal communication).
TGME49_232020-HA showed labeling of the apical pole anterior to the rhoptry bulb protein ROP1, as well as co-localization with RON4 staining (Fig. 2C), confirming rhoptry neck localization in intracellular parasites. Interestingly, TGME49_232020-HA does not localize to the moving junction in invading parasites (Fig. 2C). It was recently shown that RON9, RON10 and RON11, in contrast to the other RONs described to date, do not relocalize from the rhoptry neck to the moving junction during host invasion (Lamarque et al., 2012; Beck et al., 2013). TGME49_232020 localization also seems to be independent of the moving junction. Similar to TGME49_232020, RON10 has a predicted signal peptide in its N-terminus but no known domains or motifs have been identified. In contrast, RON9 harbors several protein-protein interaction domains. The disruption of either RON9 or RON10 led to the retention of either protein in the endoplasmic reticulum (ER), suggesting an interaction between RON9 and RON10 during their trafficking through the secretory pathway on the way to the rhoptries. Whether TGME49_232020 can interact with the RON9/RON10 complex or any other rhoptry neck protein in a similar fashion remains to be determined. TGME49_261740 co-localizes with ROP1 (Fig. 2B) and is also found in the nucleus of infected host cells (Fig. 2D). NLS Mapper (Kosugi et al., 2009) predicts that TGME49_261740 (ROP47) encodes a bipartite NLS with a score of 3.4. Moreover, this protein is predicted to have a size of 15 kDa and is probably able to diffuse through nuclear pores.
Altogether, these results suggest that TGME49_261740 and TGME49_218270 encode new rhoptry bulb proteins, hereafter referred to as ROP47 and ROP48, respectively. TGME49_232020 codes for a new rhoptry neck protein, from now on referred to as RON12.
3.3. TGME49_201860 and ROP48 are not implicated in evasion of the IFNγ response
To investigate the role of TGME49_201860, ROP47 and ROP48 in parasite biology, we removed these genes in the PruΔhxgprtΔku80 strain using double homologous recombination. PCR confirmed both the absence of the target gene coding sequences and the insertion of the hxgprt gene in the parasite genome. The deletion of ROP47 was confirmed by western blot (Supplementary Fig. S1B). Our attempts to generate a KO of RON12 in the PruΔhxgprtΔku80 strain have to date been unsuccessful.
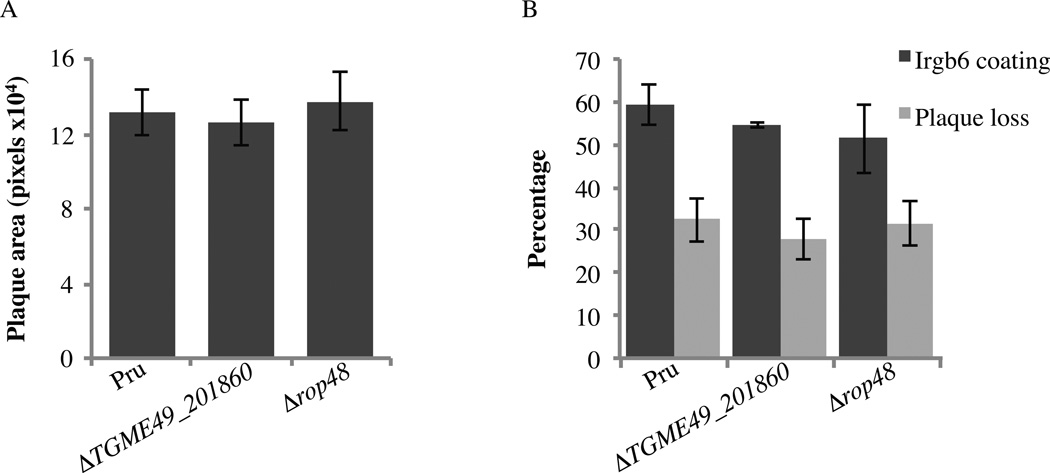
To determine a potential growth phenotype, monolayers of MEF were infected with PruΔhxgprtΔku80, Δtgme49_201860 or Δrop48 parasites, the parasites were allowed to grow for 5 days, and then the areas of the plaques formed on the monolayers were quantified (Fig. 3A). PruΔhxgprtΔku80, Δtgme49_201860 and Δrop48 strains did not form significantly different sized plaques. These data indicate that deletion of TGME49_201860 or ROP48 has no major influence on in vitro Toxoplasma growth.
Fig. 3.
Deletion of TGME49_201860 and ROP48 does not affect plaque loss in response to IFNγ. (A) Monolayers of mouse embryonic fibroblasts (MEF) were infected with PruΔhxgprtΔku80, ΔTGME49_201860 or Δrop48 parasites. The plaque area was quantified after 5 days. Mean ± S.D.; n ≥ 30 plaques. (B) Quantification of the localization of IFNγ-inducible immunity-related GTPase B6 (Irgb6) on the parasite containing vacuole and of the percentage of plaque loss after 5 days on MEF stimulated with IFNγ compared with unstimulated MEF. Mean ± S.D.; n = 4 experiments.
It is well established that IFNγ is the main mediator of resistance against Toxoplasma (Suzuki et al., 1988). An important class of downstream effectors of this immune activation is the IFNγ-inducible immunity-related GTPases (IRGs), which belong to the dynamin family of GTPases and can cooperatively oligomerize to vesiculate membranes. The IRGs are able to disrupt the PVM and kill the parasite (Butcher et al., 2005). Recently, two rhoptry proteins were shown to mediate Toxoplasma evasion of the IFNγ–induced IRGs in murine cells (Fentress et al., 2010; Steinfeldt et al., 2010; Niedelman et al., 2012). To study a potential role of ROP48 and TGME49_201860 in Toxoplasma resistance to IFNγ and the IRGs, we measured the percentage of vacuoles coated with Irgb6 by IF in IFNγ-stimulated MEFs infected with PruΔhxgprtΔku80, Δtgme49_201860 or Δrop48. The percentage of coated vacuoles was approximately 50% in both Δtgme49_201860 and Δrop48 and was not significantly different from that of the parental strain, PruΔhxgprtΔku80 (Fig. 3B). Although it is generally assumed that once the PVM is coated, it will eventually lead to killing of the parasite inside, it has also been shown that Toxoplasma can escape a coated vacuole and invade a new cell (Zhao et al., 2009). Therefore, to measure killing of Toxoplasma, a plaque loss assay was performed, as described in Niedelman et al., (2012). Briefly, 100 parasites were seeded on a monolayer of MEF, either previously stimulated for 24 h with IFN γ or left untreated, and the number of plaques that form after 5 days of growth was determined. The three strains had an average of ~30% plaque loss when comparing plaques formed on IFNγ-stimulated MEFs with unstimulated MEFs. This percentage of plaque loss was lower than the percentage of vacuoles coated with Irgb6, suggesting that some coated vacuoles can escape destruction by the IRGs. Overall, the results suggest that TGME49_201860 and ROP48 are not implicated in evading the IFNγ response in MEFs. Indeed, ROP5 and ROP18 were recently reported to mediate Toxoplasma evasion of the murine IFNγ response (Niedelman et al., 2012). These two proteins seem to determine the majority of strain differences in mouse IRG evasion, even for non-clonal strains for which virulence determinants have not been studied. However, neither ROP18 nor ROP5 markedly affect survival in IFNγ -activated human cells and it is reasonable to speculate that one or more still unidentified, secreted, potentially a rhoptry, protein could be involved in escaping IFNγ -mediated killing in other host cell types. Whether this (these) protein(s) can be found within the rhoptry cluster will be the object of future studies.
3.4. TGME49_201860, ROP47 and ROP48 are not implicated in Toxoplasma virulence in mice
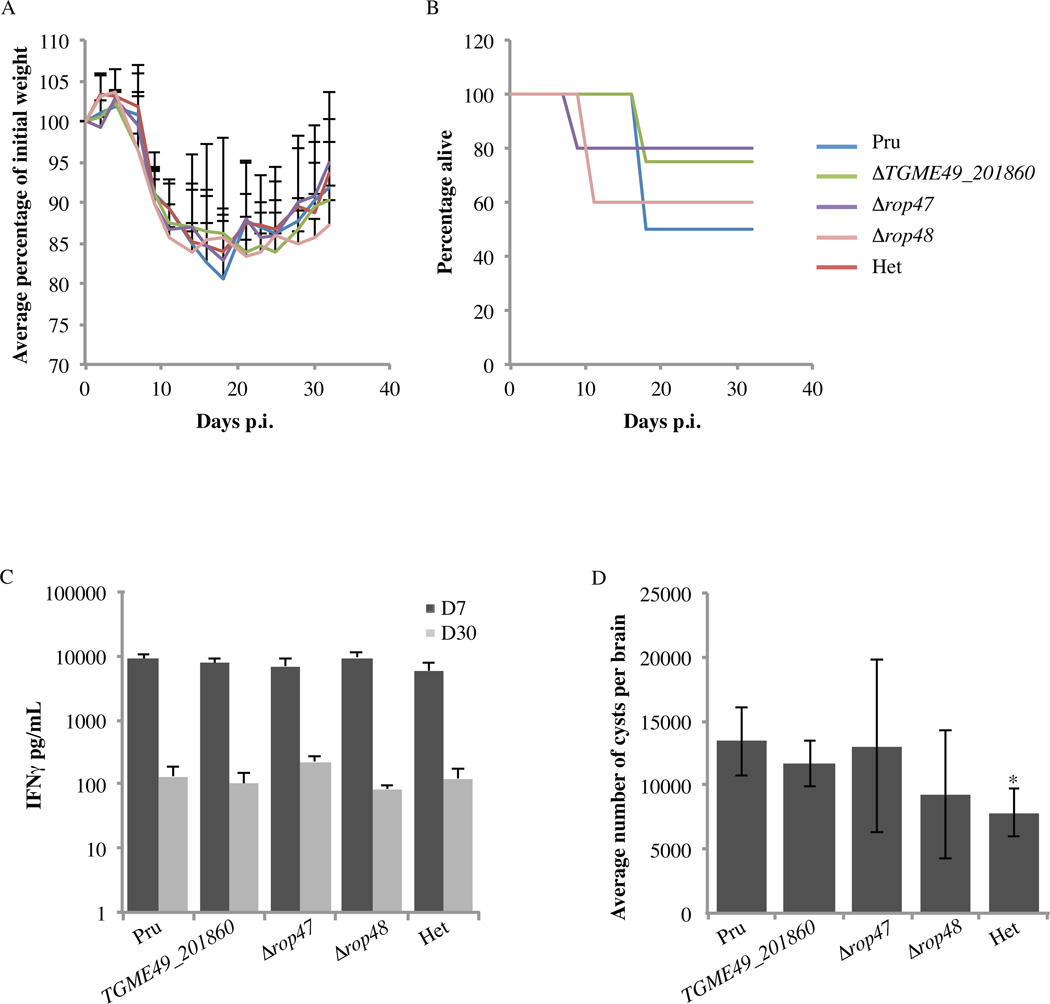
To examine the role of TGME49_201860, ROP47 and ROP48 on parasite virulence, we infected C57BL/6 mice by i.p. injection with 500 tachyzoites of PruΔhxgprtΔku80, Δtgme49_201860, Δrop47, Δrop48 or a heterologous control strain (Het) and assessed mouse morbidity (through monitoring of weight loss) and survival during the initial phase of infection (days 0–32) (Fig. 4A, B). Weight loss started at day 4 p.i. The mice reached their lowest weight between days 14 and 18 (~80% of their initial body weight) and did not regain their original weight. The survival of C57BL/6 mice infected with either strain did not show significant differences compared with PruΔhxgprtΔku80 infected mice (P = 0.9060, Log-rank Mantel-Cox test). Seroconversion was examined at day 30 p.i..and all the animals tested seropositive (data not shown). The level of IFNγ in peripheral blood serum was measured at day 7 and day 30 p.i. (Fig. 4D). IFNγ levels were higher at day 7 than day 30 and similar for all the strains. Prior to infection, animals did not display detectable levels of IFNγ. At day 35 p.i., the surviving mice were sacrificed and the number of brain cysts per mouse was determined (Fig. 4C). The numbers of cysts generated by the Δtgme49_201860, Δrop47 and Δrop48 parasites were not significantly different from that of their parental strain. The number of brain cysts observed in the mice injected i.p. with the heterologous control strain was significantly lower (~1.6 fold, P = 0.0011, Student’s t-test) than that observed in mice infected with PruΔhxgprtΔku80 parasites. Accordingly, it was previously reported that PruΔku80::hxgprt exhibited lower cyst burdens than PruΔhxgprtΔku80 (Fox et al., 2011), which suggests the presence of HXGPRT might affect generation or viability of brain cysts. Additionally, we infected C57BL/6 mice by oral gavage with 1000 brain cysts of the same strains. We found no difference in mouse morbidity and survival (data not shown).
Fig. 4.
Deletion of the genes encoding TGME49_201860, ROP47 and ROP48 does not affect Toxoplasma gondii virulence in mice. C57BL/6 mice were infected with 500 tachyzoites and weight and survival of mice was monitored. (A) Average percentage change in weight over time for mice i.p. infected with the indicated strains; n ≥ 4 for each strain, mean + S.D. (B) Mouse survival after i.p. infection with indicated strains; n ≥ 4 for each strain. (C) IFNγ cytokine levels in peripheral blood serum of surviving animals (n ≥ 2 for each strain) was determined by ELISA at days 7 (D7) and 30 (D30) p.i., mean ± S.D. Pru, PruΔhxgprtΔku80. Het, Heterologous control. (D) Average number of brain cysts at day 35 following i.p. infection with the indicated strains; n ≥ 2 for each strain; mean ± S. D. Pru - PruΔhxgprtΔku80. Het – Heterologous control. * P = 0.0011, Student’s t-test.
Overall, the results suggest that TGME49_201860, ROP47 and ROP48 are not implicated in Toxoplasma virulence in mice. Interestingly, relatively few rhoptry proteins seem to have been investigated on their ability to affect virulence in the mouse model. Of the 10 proteins that were reported, ROP5, ROP16, ROP13, ROP18, RON8, RON9/10, TLN1, BRP1 and PP2C-hn (Saeij et al., 2006; Gilbert et al., 2007; Turetzky et al., 2010; Hajagos et al., 2011; Straub et al., 2011; Lamarque et al., 2012; Niedelman et al., 2012;), only four (RON8, ROP5, ROP16, ROP18) have an effect on mouse survival. However, mouse virulence is often tested in tachyzoites, the asexually reproducing form of the parasite, leaving out events that are restricted to the sexual life cycle of the parasite. Alternatively, the mouse model used may not be the optimal setting to reveal an essential role for such proteins in infection. For instance, a role in virulence could only be apparent in other intermediate hosts and/or when cysts are ingested naturally. Elucidation of this question will be extremely challenging due to the remarkable host range of Toxoplasma.
Supplementary Material
Highlights.
Three novel Toxoplasma gondii rhoptry proteins, ROP47, ROP48 and RON12, were identified.
ROP47 is secreted into the host cell and traffics to the host nucleus.
Deletion of ROP47 or ROP48 did not influence in vitro growth or virulence in mice.
Acknowledgements
The authors thank V. Carruthers for RHΔhxgprtΔku80, D. Bzik for PruΔhxgprtΔku80, P. Bradley for the anti-RON4 antibody, M.J. Gubbels for the anti-IMC1 and anti-MLP1 antibodies, and the members of the Saeij laboratory for helpful discussions. This work was supported by a postdoctoral fellowship from the American Heart Association to AC, a postdoctoral fellowship from the Knights Templar Eye Foundation, USA, to DAG, an A*STAR NSS, Singapore, graduate scholarship to NY, postdoctoral fellowships from the Cancer Research Institute, USA, and the Charles A. King Trust, USA, to KDCJ and National Institutes of Health, USA grant R01-AI080621 to JPJS.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- Beck JR, Fung C, Straub KW, Coppens I, Vashisht AA, Wohlschlegel JA, Bradley PJ. A Toxoplasma Palmitoyl Acyl Transferase and the Palmitoylated Armadillo Repeat Protein TgARO Govern Apical Rhoptry Tethering and Reveal a Critical Role for the Rhoptries in Host Cell Invasion but Not Egress. PLoS Pathog. 2013;9:e1003162. doi: 10.1371/journal.ppat.1003162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beckers CJ, Wakefield T, Joiner KA. The expression of Toxoplasma proteins in Neospora caninum and the identification of a gene encoding a novel rhoptry protein. Mol. Biochem. Parasitol. 1997;89:209–223. doi: 10.1016/s0166-6851(97)00120-5. [DOI] [PubMed] [Google Scholar]
- Behnke MS, Fentress SJ, Mashayekhi M, Li LX, Taylor GA, Sibley LD. The polymorphic pseudokinase ROP5 controls virulence in Toxoplasma gondii by regulating the active kinase ROP18. PLoS Pathog. 2012;8:e1002992. doi: 10.1371/journal.ppat.1002992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Behnke MS, Wootton JC, Lehmann MM, Radke JB, Lucas O, Nawas J, Sibley LD, White MW. Coordinated progression through two subtranscriptomes underlies the tachyzoite cycle of Toxoplasma gondii . PLoS ONE. 2010;5:e12354. doi: 10.1371/journal.pone.0012354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Besteiro S, Michelin A, Poncet J, Dubremetz J-F, Lebrun M. Export of a Toxoplasma gondii rhoptry neck protein complex at the host cell membrane to form the moving junction during invasion. PLoS Pathog. 2009;5:e1000309. doi: 10.1371/journal.ppat.1000309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boothroyd JC, Dubremetz J-F. Kiss and spit: the dual roles of Toxoplasma rhoptries. Nat. Rev. Microbiol. 2008;6:79–88. doi: 10.1038/nrmicro1800. [DOI] [PubMed] [Google Scholar]
- Bradley PJ, Boothroyd JC. Identification of the pro-mature processing site of Toxoplasma ROP1 by mass spectrometry. Mol. Biochem. Parasitol. 1999;100:103–109. doi: 10.1016/s0166-6851(99)00035-3. [DOI] [PubMed] [Google Scholar]
- Bradley PJ, Hsieh CL, Boothroyd JC. Unprocessed Toxoplasma ROP1 is effectively targeted and secreted into the nascent parasitophorous vacuole. Mol. Biochem. Parasitol. 2002;125:189–193. doi: 10.1016/s0166-6851(02)00162-7. [DOI] [PubMed] [Google Scholar]
- Bradley PJ, Li N, Boothroyd JC. A GFP-based motif-trap reveals a novel mechanism of targeting for the Toxoplasma ROP4 protein. Mol. Biochem. Parasitol. 2004;137:111–120. doi: 10.1016/j.molbiopara.2004.05.003. [DOI] [PubMed] [Google Scholar]
- Bradley PJ, Ward C, Cheng SJ, Alexander DL, Coller S, Coombs GH, Dunn JD, Ferguson DJ, Sanderson SJ, Wastling JM, Boothroyd JC. Proteomic analysis of rhoptry organelles reveals many novel constituents for host-parasite interactions in Toxoplasma gondii . J. Biol. Chem. 2005;280:34245–34258. doi: 10.1074/jbc.M504158200. [DOI] [PubMed] [Google Scholar]
- Burg JL, Perelman D, Kasper LH, Ware PL, Boothroyd JC. Molecular analysis of the gene encoding the major surface antigen of Toxoplasma gondii . J. Immunol. 1988;141:3584–3591. [PubMed] [Google Scholar]
- Butcher BA, Greene RI, Henry SC, Annecharico KL, Weinberg JB, Denkers EY, Sher A, Taylor GA. p47 GTPases regulate Toxoplasma gondii survival in activated macrophages. Infect. Immun. 2005;73:3278–3286. doi: 10.1128/IAI.73.6.3278-3286.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cabrera A, Herrmann S, Warszta D, Santos JM, John Peter AT, Kono M, Debrouver S, Jacobs T, Spielmann T, Ungermann C, et al. Dissection of Minimal Sequence Requirements for Rhoptry Membrane Targeting in the Malaria Parasite. Traffic. 2012;13:1335–1350. doi: 10.1111/j.1600-0854.2012.01394.x. [DOI] [PubMed] [Google Scholar]
- Carey KL, Jongco AM, Kim K, Ward GE. The Toxoplasma gondii rhoptry protein ROP4 is secreted into the parasitophorous vacuole and becomes phosphorylated in infected cells. Eukaryotic Cell. 2004;3:1320–1330. doi: 10.1128/EC.3.5.1320-1330.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dunn JD, Ravindran S, Kim S-K, Boothroyd JC. The Toxoplasma gondii dense granule protein GRA7 is phosphorylated upon invasion and forms an unexpected association with the rhoptry proteins ROP2 and ROP4 . Infect. Immun. 2008;76:5853–5861. doi: 10.1128/IAI.01667-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fentress SJ, Behnke MS, Dunay IR, Mashayekhi M, Rommereim LM, Fox BA, Bzik DJ, Taylor GA, Turk BE, Lichti CF, Townsend RR, Qiu W, Hui R, Beatty WL, Sibley LD. Phosphorylation of Immunity-Related GTPases by a Toxoplasma gondii-Secreted Kinase Promotes Macrophage Survival and Virulence. Cell Host Microbe. 2010;8:484–495. doi: 10.1016/j.chom.2010.11.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fentress SJ, Steinfeldt T, Howard JC, Sibley LD. The arginine-rich N-terminal domain of ROP18 is necessary for vacuole targeting and virulence of Toxoplasma gondii . Cell. Microbiol. 2012;14:1921–1933. doi: 10.1111/cmi.12022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fleckenstein MC, Reese ML, Könen-Waisman S, Boothroyd JC, Howard JC, Steinfeldt T. A Toxoplasma gondii pseudokinase inhibits host IRG resistance proteins. PLoS Biol. 2012;10:e1001358. doi: 10.1371/journal.pbio.1001358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fox BA, Falla A, Rommereim LM, Tomita T, Gigley JP, Mercier C, Cesbron-Delauw M-F, Weiss LM, Bzik DJ. Type II Toxoplasma gondii KU80 knockout strains enable functional analysis of genes required for cyst development and latent infection. Eukaryotic Cell. 2011;10:1193–1206. doi: 10.1128/EC.00297-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fritz HM, Buchholz KR, Chen X, Durbin-Johnson B, Rocke DM, Conrad PA, Boothroyd JC. Transcriptomic Analysis of Toxoplasma Development Reveals Many Novel Functions and Structures Specific to Sporozoites and Oocysts. PLoS ONE. 2012;7:e29998. doi: 10.1371/journal.pone.0029998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gilbert LA, Ravindran S, Turetzky JM, Boothroyd JC, Bradley PJ. Toxoplasma gondii targets a protein phosphatase 2C to the nuclei of infected host cells. Eukaryotic Cell. 2007;6:73–83. doi: 10.1128/EC.00309-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hajagos BE, Turetzky JM, Peng ED, Cheng SJ, Ryan CM, Souda P, Whitelegge JP, Lebrun M, Dubremetz J-F, Bradley PJ. Molecular Dissection of Novel Trafficking and Processing of the Toxoplasma gondii Rhoptry Metalloprotease Toxolysin-1. Traffic. 2011;13:292–304. doi: 10.1111/j.1600-0854.2011.01308.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hajj, El H, Demey E, Poncet J, Lebrun M, Wu B, Galéotti N, Fourmaux MN, Mercereau-Puijalon O, Vial H, Labesse G, Dubremetz J-F. The ROP2 family of Toxoplasma gondii rhoptry proteins: Proteomic and genomic characterization and molecular modeling. Proteomics. 2006a;6:5773–5784. doi: 10.1002/pmic.200600187. [DOI] [PubMed] [Google Scholar]
- Hajj, El H, Lebrun M, Fourmaux MN, Vial H, Dubremetz J-F. Characterization, biosynthesis and fate of ROP7, a ROP2 related rhoptry protein of Toxoplasma gondii . Mol. Biochem. Parasitol. 2006b;146:98–100. doi: 10.1016/j.molbiopara.2005.10.011. [DOI] [PubMed] [Google Scholar]
- Hajj, El H, Lebrun M, Arold ST, Vial H, Labesse G, Dubremetz J-F. ROP18 Is a Rhoptry Kinase Controlling the Intracellular Proliferation of Toxoplasma gondii . PLoS Pathog. 2007;3:e14. doi: 10.1371/journal.ppat.0030014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoppe HC, Ngô HM, Yang M, Joiner KA. Targeting to rhoptry organelles of Toxoplasma gondii involves evolutionarily conserved mechanisms. Nat. Cell Biol. 2000;2:449–456. doi: 10.1038/35017090. [DOI] [PubMed] [Google Scholar]
- Huynh M-H, Carruthers VB. Tagging of endogenous genes in a Toxoplasma gondii strain lacking Ku80. Eukaryotic Cell. 2009;8:530–539. doi: 10.1128/EC.00358-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Joiner KA, Roos DS. Secretory traffic in the eukaryotic parasite Toxoplasma gondii: less is more. J. Cell Biol. 2002;157:557–563. doi: 10.1083/jcb.200112144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karasov AO, Boothroyd JC, Arrizabalaga G. Identification and disruption of a rhoptry-localized homologue of sodium hydrogen exchangers in Toxoplasma gondii . Int. J. Parasitol. 2005;35:285–291. doi: 10.1016/j.ijpara.2004.11.015. [DOI] [PubMed] [Google Scholar]
- Kawase O, Nishikawa Y, Bannai H, Zhang H, Zhang G, Jin S, Lee E-G, Xuan X. Proteomic analysis of calcium-dependent secretion in Toxoplasma gondii . Proteomics. 2007;7:3718–3725. doi: 10.1002/pmic.200700362. [DOI] [PubMed] [Google Scholar]
- Kissinger JC, Gajria B, Li L, Paulsen IT, Roos DS. ToxoDB: accessing the Toxoplasma gondii genome. Nucleic Acids Res. 2003;31:234–236. doi: 10.1093/nar/gkg072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kosugi S, Hasebe M, Tomita M, Yanagawa H. Systematic identification of cell cycle-dependent yeast nucleocytoplasmic shuttling proteins by prediction of composite motifs. Proc. Natl. Acad. Sci. U.S.A. 2009;106:10171–10176. doi: 10.1073/pnas.0900604106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lamarque MH, Papoin J, Finizio A-L, Lentini G, Pfaff AW, Candolfi E, Dubremetz J-F, Lebrun M. Identification of a New Rhoptry Neck Complex RON9/RON10 in the Apicomplexa Parasite Toxoplasma gondii . PLoS ONE. 2012;7:e32457. doi: 10.1371/journal.pone.0032457. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lodoen MB, Gerke C, Boothroyd JC. A highly sensitive FRET-based approach reveals secretion of the actin-binding protein toxofilin during Toxoplasma gondii infection. Cell. Microbiol. 2010;12:55–66. doi: 10.1111/j.1462-5822.2009.01378.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marugán-Hernández V, Álvarez-García G, Tomley F, Hemphill A, Regidor-Cerrillo J, Ortega-Mora LM. Identification of novel rhoptry proteins in Neospora caninum by LC/MS-MS analysis of subcellular fractions. J. Proteomics. 2011;74:629–642. doi: 10.1016/j.jprot.2011.02.004. [DOI] [PubMed] [Google Scholar]
- Miller SA, Thathy V, Ajioka JW, Blackman MJ, Kim K. TgSUB2 is a Toxoplasma gondii rhoptry organelle processing proteinase. Molec. Microbiol. 2003;49:883–894. doi: 10.1046/j.1365-2958.2003.03604.x. [DOI] [PubMed] [Google Scholar]
- Minot S, Melo MB, Li F, Lu D, Niedelman W, Levine SS, Saeij JPJ. Admixture and recombination among Toxoplasma gondii lineages explain global genome diversity. Proc. Natl. Acad. Sci. U.S.A. 2012;109:13458–13463. doi: 10.1073/pnas.1117047109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Montoya JG, Liesenfeld O. Toxoplasmosis. Lancet. 2004;363:1965–1976. doi: 10.1016/S0140-6736(04)16412-X. [DOI] [PubMed] [Google Scholar]
- Niedelman W, Gold DA, Rosowski EE, Sprokholt JK, Lim D, Farid Arenas A, Melo MB, Spooner E, Yaffe MB, Saeij JPJ. The rhoptry proteins ROP18 and ROP5 mediate Toxoplasma gondii evasion of the murine, but not the human, interferon-gamma response. PLoS Pathog. 2012;8:e1002784. doi: 10.1371/journal.ppat.1002784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oakes RD, Kurian D, Bromley E, Ward C, Lal K, Blake DP, Reid AJ, Pain A, Sinden RE, Wastling JM, Tomley FM. The rhoptry proteome of Eimeria tenella sporozoites. Int. J. Parasitol. 2013;43:181–188. doi: 10.1016/j.ijpara.2012.10.024. [DOI] [PubMed] [Google Scholar]
- Ong Y-C, Reese ML, Boothroyd JC. Toxoplasma rhoptry protein 16 (ROP16) subverts host function by direct tyrosine phosphorylation of STAT6. J. Biol. Chem. 2010;285:28731–28740. doi: 10.1074/jbc.M110.112359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ossorio PN, Schwartzman JD, Boothroyd JC. A Toxoplasma gondii rhoptry protein associated with host cell penetration has unusual charge asymmetry. Mol. Biochem. Parasitol. 1992;50:1–15. doi: 10.1016/0166-6851(92)90239-g. [DOI] [PubMed] [Google Scholar]
- Peixoto L, Chen F, Harb OS, Davis PH, Beiting DP, Brownback CS, Ouloguem D, Roos DS. Integrative genomic approaches highlight a family of parasite-specific kinases that regulate host responses. Cell Host Microbe. 2010;8:208–218. doi: 10.1016/j.chom.2010.07.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Proellocks NI, Kats LM, Sheffield DA, Hanssen E, Black CG, Waller KL, Coppel RL. Characterisation of PfRON6, a Plasmodium falciparum rhoptry neck protein with a novel cysteine-rich domain. Int. J. Parasitol. 2009;39:683–692. doi: 10.1016/j.ijpara.2008.11.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reese ML, Boothroyd JC. A Helical Membrane-Binding Domain Targets the Toxoplasma ROP2 Family to the Parasitophorous Vacuole. Traffic. 2009;10:1458–1470. doi: 10.1111/j.1600-0854.2009.00958.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reichmann G, Długońska H, Fischer H-G. Characterization of TgROP9 p36, a novel rhoptry protein of Toxoplasma gondii tachyzoites identified by T cell clone. Mol. Biochem. Parasitol. 2002;119:43–54. doi: 10.1016/s0166-6851(01)00397-8. [DOI] [PubMed] [Google Scholar]
- Reid AJ, Vermont SJ, Cotton JA, Harris D, Hill-Cawthorne GA, Könen-Waisman S, Latham SM, Mourier T, Norton R, Quail MA, Sanders M, Shanmugam D, Sohal A, Wasmuth JD, Brunk B, Grigg ME, Howard JC, Parkinson J, Roos DS, Trees AJ, Berriman M, Pain A, Wastling JM. Comparative genomics of the apicomplexan parasites Toxoplasma gondii and Neospora caninum: Coccidia differing in host range and transmission strategy. PLoS Pathog. 2012;8:e1002567. doi: 10.1371/journal.ppat.1002567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rosowski EE, Lu D, Julien L, Rodda L, Gaiser RA, Jensen KDC, Saeij JPJ. Strain-specific activation of the NF-kappaB pathway by GRA15, a novel Toxoplasma gondii dense granule protein. J. Exp. Med. 2011;208:195–212. doi: 10.1084/jem.20100717. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sadak A, Taghy Z, Fortier B, Dubremetz JF. Characterization of a family of rhoptry proteins of Toxoplasma gondii . Mol. Biochem. Parasitol. 1988;29:203–211. doi: 10.1016/0166-6851(88)90075-8. [DOI] [PubMed] [Google Scholar]
- Saeij JPJ, Boyle JP, Coller S, Taylor S, Sibley LD, Brooke-Powell ET, Ajioka JW, Boothroyd JC. Polymorphic secreted kinases are key virulence factors in toxoplasmosis. Science. 2006;314:1780–1783. doi: 10.1126/science.1133690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saeij JPJ, Coller S, Boyle JP, Jerome ME, White MW, Boothroyd JC. Toxoplasma co-opts host gene expression by injection of a polymorphic kinase homologue. Nature. 2007;445:324–327. doi: 10.1038/nature05395. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scallan E, Hoekstra RM, Angulo FJ, Tauxe RV, Widdowson M-A, Roy SL, Jones JL, Griffin PM. Foodborne illness acquired in the United States--major pathogens. Emerging Infect. Dis. 2011;17:7–15. doi: 10.3201/eid1701.P11101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwarz JA, Fouts AE, Cummings CA, Ferguson DJP, Boothroyd JC. A novel rhoptry protein in Toxoplasma gondii bradyzoites and merozoites. Mol. Biochem. Parasitol. 2005;144:159–166. doi: 10.1016/j.molbiopara.2005.08.011. [DOI] [PubMed] [Google Scholar]
- Sohn WMW, Nam HWH. Western blot analysis of stray cat sera against Toxoplasma gondii and the diagnostic availability of monoclonal antibodies in sandwich-ELISA. Korean J Parasitol. 1999;37:249–256. doi: 10.3347/kjp.1999.37.4.249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steinfeldt T, Könen-Waisman S, Tong L, Pawlowski N, Lamkemeyer T, Sibley LD, Hunn JP, Howard JC. Phosphorylation of Mouse Immunity-Related GTPase (IRG) Resistance Proteins Is an Evasion Strategy for Virulent Toxoplasma gondii . PLoS Biol. 2010;8:e1000576. doi: 10.1371/journal.pbio.1000576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Straub KW, Cheng SJ, Sohn CS, Bradley PJ. Novel components of the Apicomplexan moving junction reveal conserved and coccidia-restricted elements. Cell. Microbiol. 2009;11:590–603. doi: 10.1111/j.1462-5822.2008.01276.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Straub KW, Peng ED, Hajagos BE, Tyler JS, Bradley PJ. The moving junction protein RON8 facilitates firm attachment and host cell invasion in Toxoplasma gondii . PLoS Pathog. 2011;7:e1002007. doi: 10.1371/journal.ppat.1002007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suzuki Y, Orellana M, Schreiber R, Remington J. Interferon-gamma: the major mediator of resistance against Toxoplasma gondii . Science. 1988;240:516–518. doi: 10.1126/science.3128869. [DOI] [PubMed] [Google Scholar]
- Taylor S, Barragan A, Su C, Fux B, Fentress SJ, Tang K, Beatty WL, Hajj HE, Jerome M, Behnke MS, White M, Wootton JC, Sibley LD. A secreted serine-threonine kinase determines virulence in the eukaryotic pathogen Toxoplasma gondii . Science. 2006;314:1776–1780. doi: 10.1126/science.1133643. [DOI] [PubMed] [Google Scholar]
- Turetzky JM, Chu DK, Hajagos BE, Bradley PJ. Processing and secretion of ROP13: A unique Toxoplasma effector protein. Int. J. Parasitol. 2010;40:1037–1044. doi: 10.1016/j.ijpara.2010.02.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamamoto M, Standley DM, Takashima S, Saiga H, Okuyama M, Kayama H, Kubo E, Ito H, Takaura M, Matsuda T, Soldati-Favre D, Takeda K. A single polymorphic amino acid on Toxoplasma gondii kinase ROP16 determines the direct and strain-specific activation of Stat3. J. Exp. Med. 2009;206:2747–2760. doi: 10.1084/jem.20091703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao YO, Rohde C, Lilue JT, Könen-Waisman S, Khaminets A, Hunn JP, Howard JC. Toxoplasma gondii and the Immunity-Related GTPase (IRG) resistance system in mice: a review. Mem. Inst. Oswaldo Cruz. 2009;104:234–240. doi: 10.1590/s0074-02762009000200016. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.