Abstract
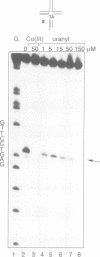
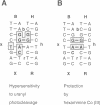
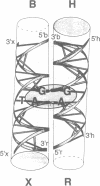
Metal ions are very important in mediating the folding of nucleic acids, as exemplified by the folding of the four-way DNA junction into the stacked X-conformation. Uranyl ion-mediated photocleavage provides a method for the localization of high-affinity ion binding sites in nucleic acids, and we have applied this to the four-way DNA junction. We have made the following observations. (i) Uranyl ions (UO2(2+)) suppressed the reactivity of junction thymine bases against attack by osmium tetroxide, indicating that the uranyl ion induces folding of the junction into a stacked conformation. (ii) DNA located immediately at the point of strand exchange on the two exchanging strands was hypersensitive to uranyl photocleavage. The relative hypersensitivity was considerably accentuated when the photocleavage was carried out in the presence of citrate ions. This suggests the presence of a tight binding site for the uranyl ion in the junction. (iii) The same positions were significantly protected from uranyl cleavage by the presence of hexamminecobalt (III) or spermidine. These ions are known to induce the folded conformation of the four-way junction with high efficiency, suggesting a direct competition between the ions. By contrast, magnesium ions failed to generate a similar protection against photocleavage. These results suggest that the uranyl, hexamminecobalt (III) and spermidine ions compete for the same high affinity binding site on the junction. This site is located at the centre of the junction, at the point where the exchanging strands pass between the stacked helices. We believe that we have observed the first known example of a metal ion 'footprint' on a folded nucleic acid structure.
Full text
PDF




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Behe M., Felsenfeld G. Effects of methylation on a synthetic polynucleotide: the B--Z transition in poly(dG-m5dC).poly(dG-m5dC). Proc Natl Acad Sci U S A. 1981 Mar;78(3):1619–1623. doi: 10.1073/pnas.78.3.1619. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Celander D. W., Cech T. R. Visualizing the higher order folding of a catalytic RNA molecule. Science. 1991 Jan 25;251(4992):401–407. doi: 10.1126/science.1989074. [DOI] [PubMed] [Google Scholar]
- Churchill M. E., Tullius T. D., Kallenbach N. R., Seeman N. C. A Holliday recombination intermediate is twofold symmetric. Proc Natl Acad Sci U S A. 1988 Jul;85(13):4653–4656. doi: 10.1073/pnas.85.13.4653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clegg R. M., Murchie A. I., Lilley D. M. The solution structure of the four-way DNA junction at low-salt conditions: a fluorescence resonance energy transfer analysis. Biophys J. 1994 Jan;66(1):99–109. doi: 10.1016/S0006-3495(94)80765-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clegg R. M., Murchie A. I., Zechel A., Carlberg C., Diekmann S., Lilley D. M. Fluorescence resonance energy transfer analysis of the structure of the four-way DNA junction. Biochemistry. 1992 May 26;31(20):4846–4856. doi: 10.1021/bi00135a016. [DOI] [PubMed] [Google Scholar]
- Cooper J. P., Hagerman P. J. Gel electrophoretic analysis of the geometry of a DNA four-way junction. J Mol Biol. 1987 Dec 20;198(4):711–719. doi: 10.1016/0022-2836(87)90212-9. [DOI] [PubMed] [Google Scholar]
- Cooper J. P., Hagerman P. J. Geometry of a branched DNA structure in solution. Proc Natl Acad Sci U S A. 1989 Oct;86(19):7336–7340. doi: 10.1073/pnas.86.19.7336. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duckett D. R., Lilley D. M. Effects of base mismatches on the structure of the four-way DNA junction. J Mol Biol. 1991 Sep 5;221(1):147–161. doi: 10.1016/0022-2836(91)80211-c. [DOI] [PubMed] [Google Scholar]
- Duckett D. R., Murchie A. I., Diekmann S., von Kitzing E., Kemper B., Lilley D. M. The structure of the Holliday junction, and its resolution. Cell. 1988 Oct 7;55(1):79–89. doi: 10.1016/0092-8674(88)90011-6. [DOI] [PubMed] [Google Scholar]
- Duckett D. R., Murchie A. I., Lilley D. M. The role of metal ions in the conformation of the four-way DNA junction. EMBO J. 1990 Feb;9(2):583–590. doi: 10.1002/j.1460-2075.1990.tb08146.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gough G. W., Lilley D. M. DNA bending induced by cruciform formation. Nature. 1985 Jan 10;313(5998):154–156. doi: 10.1038/313154a0. [DOI] [PubMed] [Google Scholar]
- Hoess R., Wierzbicki A., Abremski K. Isolation and characterization of intermediates in site-specific recombination. Proc Natl Acad Sci U S A. 1987 Oct;84(19):6840–6844. doi: 10.1073/pnas.84.19.6840. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jayaram M., Crain K. L., Parsons R. L., Harshey R. M. Holliday junctions in FLP recombination: resolution by step-arrest mutants of FLP protein. Proc Natl Acad Sci U S A. 1988 Nov;85(21):7902–7906. doi: 10.1073/pnas.85.21.7902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jeppesen C., Nielsen P. E. Uranyl mediated photofootprinting reveals strong E. coli RNA polymerase--DNA backbone contacts in the +10 region of the DeoP1 promoter open complex. Nucleic Acids Res. 1989 Jul 11;17(13):4947–4956. doi: 10.1093/nar/17.13.4947. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karpel R. L., Miller N. S., Lesk A. M., Fresco J. R. Stabilization of the native tertiary stucture of yeast tRNALeu3 by cationic metal complexes. J Mol Biol. 1975 Oct 5;97(4):519–532. doi: 10.1016/s0022-2836(75)80056-8. [DOI] [PubMed] [Google Scholar]
- Kitts P. A., Nash H. A. Homology-dependent interactions in phage lambda site-specific recombination. Nature. 1987 Sep 24;329(6137):346–348. doi: 10.1038/329346a0. [DOI] [PubMed] [Google Scholar]
- Lilley D. M., Clegg R. M. The structure of the four-way junction in DNA. Annu Rev Biophys Biomol Struct. 1993;22:299–328. doi: 10.1146/annurev.bb.22.060193.001503. [DOI] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 1980;65(1):499–560. doi: 10.1016/s0076-6879(80)65059-9. [DOI] [PubMed] [Google Scholar]
- Meselson M. S., Radding C. M. A general model for genetic recombination. Proc Natl Acad Sci U S A. 1975 Jan;72(1):358–361. doi: 10.1073/pnas.72.1.358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murchie A. I., Carter W. A., Portugal J., Lilley D. M. The tertiary structure of the four-way DNA junction affords protection against DNase I cleavage. Nucleic Acids Res. 1990 May 11;18(9):2599–2606. doi: 10.1093/nar/18.9.2599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murchie A. I., Clegg R. M., von Kitzing E., Duckett D. R., Diekmann S., Lilley D. M. Fluorescence energy transfer shows that the four-way DNA junction is a right-handed cross of antiparallel molecules. Nature. 1989 Oct 26;341(6244):763–766. doi: 10.1038/341763a0. [DOI] [PubMed] [Google Scholar]
- Murchie A. I., Portugal J., Lilley D. M. Cleavage of a four-way DNA junction by a restriction enzyme spanning the point of strand exchange. EMBO J. 1991 Mar;10(3):713–718. doi: 10.1002/j.1460-2075.1991.tb08001.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nielsen P. E., Jeppesen C., Buchardt O. Uranyl salts as photochemical agents for cleavage of DNA and probing of protein-DNA contacts. FEBS Lett. 1988 Aug 1;235(1-2):122–124. doi: 10.1016/0014-5793(88)81245-6. [DOI] [PubMed] [Google Scholar]
- Nunes-Düby S. E., Matsumoto L., Landy A. Site-specific recombination intermediates trapped with suicide substrates. Cell. 1987 Aug 28;50(5):779–788. doi: 10.1016/0092-8674(87)90336-9. [DOI] [PubMed] [Google Scholar]
- Piccirilli J. A., Vyle J. S., Caruthers M. H., Cech T. R. Metal ion catalysis in the Tetrahymena ribozyme reaction. Nature. 1993 Jan 7;361(6407):85–88. doi: 10.1038/361085a0. [DOI] [PubMed] [Google Scholar]
- Pörschke D. Thermodynamic and kinetic parameters of ion condensation to polynucleotides. Outer sphere complex formed by Mg++ions. Biophys Chem. 1976 Jul;4(4):383–394. doi: 10.1016/0301-4622(76)80018-x. [DOI] [PubMed] [Google Scholar]
- Sinha N. D., Biernat J., McManus J., Köster H. Polymer support oligonucleotide synthesis XVIII: use of beta-cyanoethyl-N,N-dialkylamino-/N-morpholino phosphoramidite of deoxynucleosides for the synthesis of DNA fragments simplifying deprotection and isolation of the final product. Nucleic Acids Res. 1984 Jun 11;12(11):4539–4557. doi: 10.1093/nar/12.11.4539. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Uhlenbeck O. C. A small catalytic oligoribonucleotide. Nature. 1987 Aug 13;328(6131):596–600. doi: 10.1038/328596a0. [DOI] [PubMed] [Google Scholar]
- von Kitzing E., Lilley D. M., Diekmann S. The stereochemistry of a four-way DNA junction: a theoretical study. Nucleic Acids Res. 1990 May 11;18(9):2671–2683. doi: 10.1093/nar/18.9.2671. [DOI] [PMC free article] [PubMed] [Google Scholar]