Abstract
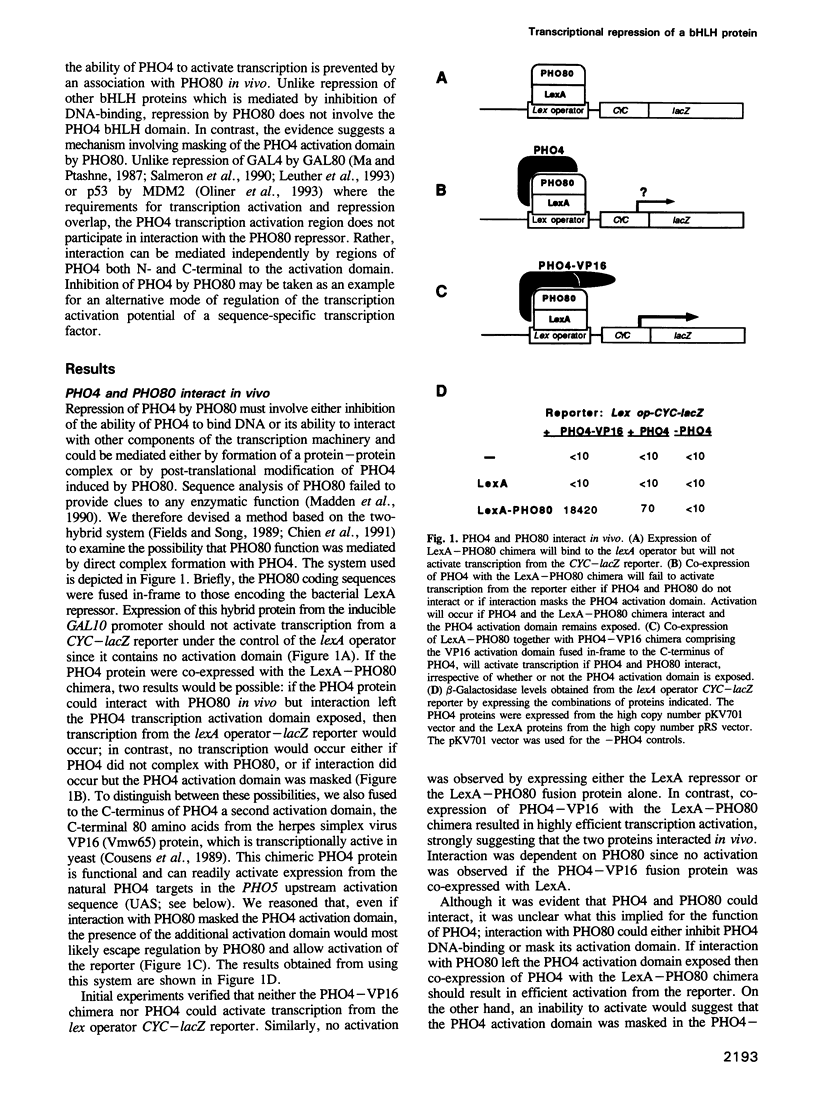
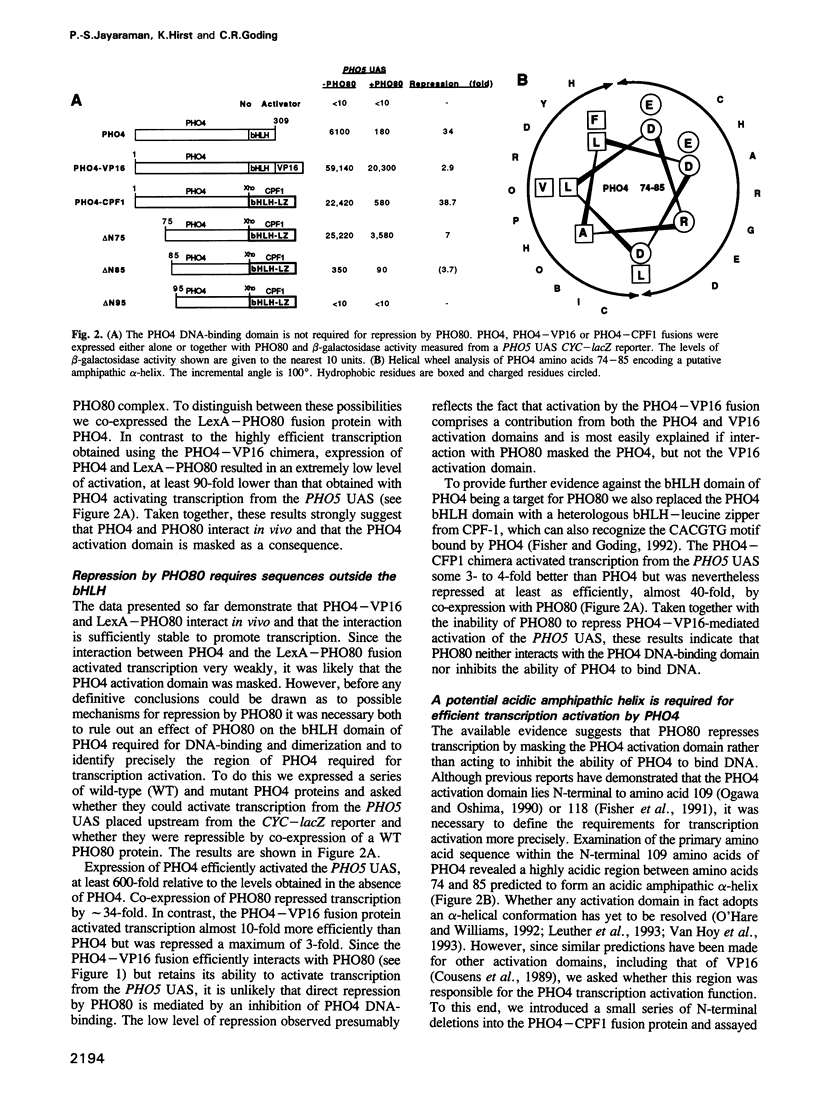
While there are many examples of protein-protein interactions modulating the DNA-binding activity of transcription factors, little is known of the molecular mechanisms underlying the regulation of the transcription activation function. Using a two-hybrid system we show here that transcription repression of the basic domain/helix-loop-helix factor PHO4 is mediated by complex formation with the PHO80 repressor. In contrast to other systems, such as inhibition of GAL4 by GAL80 or of p53 by MDM2, where repression is mediated by direct interaction at regions overlapping the transcription activation domain, interaction with PHO80 involves two regions of PHO4 distinct from those involved in transcription activation or DNA-binding and dimerization. The possibility that repression of PHO4 by PHO80 may represent a general mechanism of transcription control, including regulation of the cell-type-specific transcription activation domain of c-Jun, is discussed.
Full text
PDF



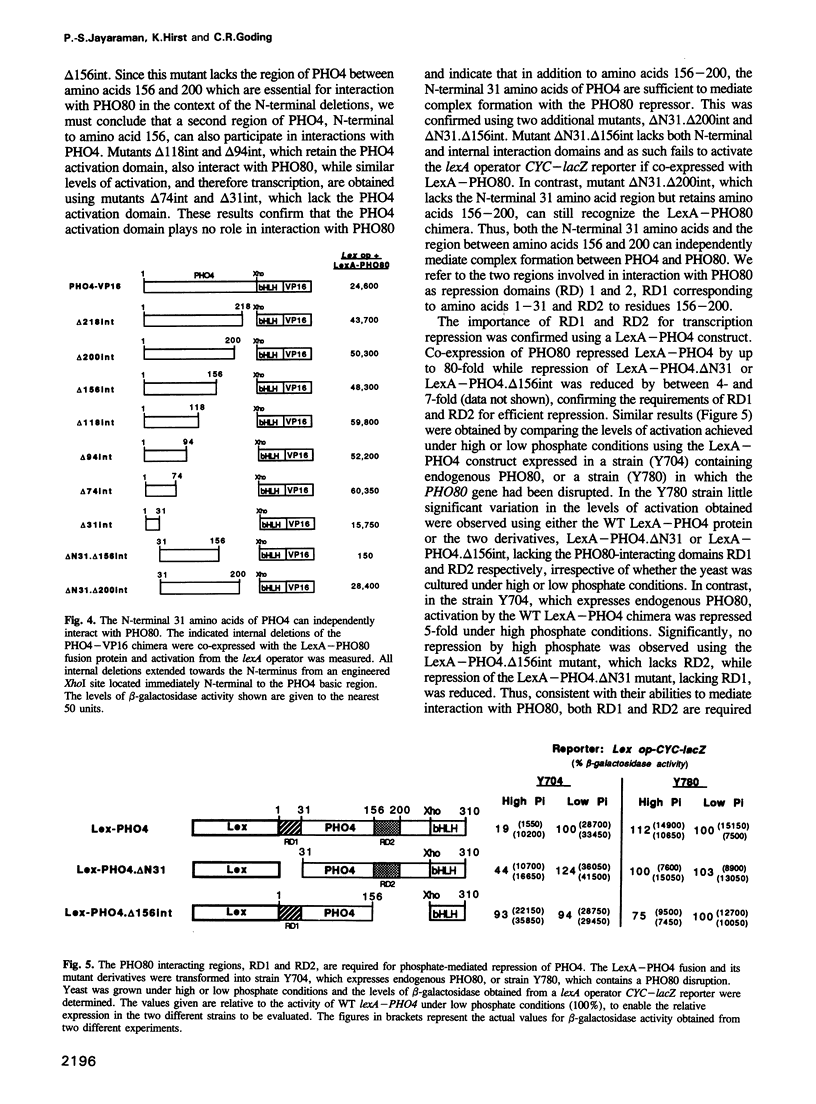



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Baichwal V. R., Park A., Tjian R. The cell-type-specific activator region of c-Jun juxtaposes constitutive and negatively regulated domains. Genes Dev. 1992 Aug;6(8):1493–1502. doi: 10.1101/gad.6.8.1493. [DOI] [PubMed] [Google Scholar]
- Baker R. E., Masison D. C. Isolation of the gene encoding the Saccharomyces cerevisiae centromere-binding protein CP1. Mol Cell Biol. 1990 Jun;10(6):2458–2467. doi: 10.1128/mcb.10.6.2458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cai M., Davis R. W. Yeast centromere binding protein CBF1, of the helix-loop-helix protein family, is required for chromosome stability and methionine prototrophy. Cell. 1990 May 4;61(3):437–446. doi: 10.1016/0092-8674(90)90525-j. [DOI] [PubMed] [Google Scholar]
- Chien C. T., Bartel P. L., Sternglanz R., Fields S. The two-hybrid system: a method to identify and clone genes for proteins that interact with a protein of interest. Proc Natl Acad Sci U S A. 1991 Nov 1;88(21):9578–9582. doi: 10.1073/pnas.88.21.9578. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cousens D. J., Greaves R., Goding C. R., O'Hare P. The C-terminal 79 amino acids of the herpes simplex virus regulatory protein, Vmw65, efficiently activate transcription in yeast and mammalian cells in chimeric DNA-binding proteins. EMBO J. 1989 Aug;8(8):2337–2342. doi: 10.1002/j.1460-2075.1989.tb08361.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fields S., Song O. A novel genetic system to detect protein-protein interactions. Nature. 1989 Jul 20;340(6230):245–246. doi: 10.1038/340245a0. [DOI] [PubMed] [Google Scholar]
- Fisher F., Goding C. R. Single amino acid substitutions alter helix-loop-helix protein specificity for bases flanking the core CANNTG motif. EMBO J. 1992 Nov;11(11):4103–4109. doi: 10.1002/j.1460-2075.1992.tb05503.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fisher F., Jayaraman P. S., Goding C. R. C-myc and the yeast transcription factor PHO4 share a common CACGTG-binding motif. Oncogene. 1991 Jul;6(7):1099–1104. [PubMed] [Google Scholar]
- Harrison S. C. A structural taxonomy of DNA-binding domains. Nature. 1991 Oct 24;353(6346):715–719. doi: 10.1038/353715a0. [DOI] [PubMed] [Google Scholar]
- Harshman K. D., Moye-Rowley W. S., Parker C. S. Transcriptional activation by the SV40 AP-1 recognition element in yeast is mediated by a factor similar to AP-1 that is distinct from GCN4. Cell. 1988 Apr 22;53(2):321–330. doi: 10.1016/0092-8674(88)90393-5. [DOI] [PubMed] [Google Scholar]
- Hinnen A., Hicks J. B., Fink G. R. Transformation of yeast. Proc Natl Acad Sci U S A. 1978 Apr;75(4):1929–1933. doi: 10.1073/pnas.75.4.1929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoshizaki D. K., Hill J. E., Henry S. A. The Saccharomyces cerevisiae INO4 gene encodes a small, highly basic protein required for derepression of phospholipid biosynthetic enzymes. J Biol Chem. 1990 Mar 15;265(8):4736–4745. [PubMed] [Google Scholar]
- Jones N. Transcriptional regulation by dimerization: two sides to an incestuous relationship. Cell. 1990 Apr 6;61(1):9–11. doi: 10.1016/0092-8674(90)90207-u. [DOI] [PubMed] [Google Scholar]
- Karin M. Signal transduction and gene control. Curr Opin Cell Biol. 1991 Jun;3(3):467–473. doi: 10.1016/0955-0674(91)90075-a. [DOI] [PubMed] [Google Scholar]
- Karin M. Too many transcription factors: positive and negative interactions. New Biol. 1990 Feb;2(2):126–131. [PubMed] [Google Scholar]
- Koren R., LeVitre J., Bostian K. A. Isolation of the positive-acting regulatory gene PHO4 from Saccharomyces cerevisiae. Gene. 1986;41(2-3):271–280. doi: 10.1016/0378-1119(86)90107-1. [DOI] [PubMed] [Google Scholar]
- Legrain M., De Wilde M., Hilger F. Isolation, physical characterization and expression analysis of the Saccharomyces cerevisiae positive regulatory gene PHO4. Nucleic Acids Res. 1986 Apr 11;14(7):3059–3073. doi: 10.1093/nar/14.7.3059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lemire J. M., Willcocks T., Halvorson H. O., Bostian K. A. Regulation of repressible acid phosphatase gene transcription in Saccharomyces cerevisiae. Mol Cell Biol. 1985 Aug;5(8):2131–2141. doi: 10.1128/mcb.5.8.2131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leuther K. K., Johnston S. A. Nondissociation of GAL4 and GAL80 in vivo after galactose induction. Science. 1992 May 29;256(5061):1333–1335. doi: 10.1126/science.1598579. [DOI] [PubMed] [Google Scholar]
- Leuther K. K., Salmeron J. M., Johnston S. A. Genetic evidence that an activation domain of GAL4 does not require acidity and may form a beta sheet. Cell. 1993 Feb 26;72(4):575–585. doi: 10.1016/0092-8674(93)90076-3. [DOI] [PubMed] [Google Scholar]
- Ma J., Ptashne M. The carboxy-terminal 30 amino acids of GAL4 are recognized by GAL80. Cell. 1987 Jul 3;50(1):137–142. doi: 10.1016/0092-8674(87)90670-2. [DOI] [PubMed] [Google Scholar]
- Madden S. L., Johnson D. L., Bergman L. W. Molecular and expression analysis of the negative regulators involved in the transcriptional regulation of acid phosphatase production in Saccharomyces cerevisiae. Mol Cell Biol. 1990 Nov;10(11):5950–5957. doi: 10.1128/mcb.10.11.5950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mellor J., Jiang W., Funk M., Rathjen J., Barnes C. A., Hinz T., Hegemann J. H., Philippsen P. CPF1, a yeast protein which functions in centromeres and promoters. EMBO J. 1990 Dec;9(12):4017–4026. doi: 10.1002/j.1460-2075.1990.tb07623.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mitchell P. J., Tjian R. Transcriptional regulation in mammalian cells by sequence-specific DNA binding proteins. Science. 1989 Jul 28;245(4916):371–378. doi: 10.1126/science.2667136. [DOI] [PubMed] [Google Scholar]
- Nikoloff D. M., McGraw P., Henry S. A. The INO2 gene of Saccharomyces cerevisiae encodes a helix-loop-helix protein that is required for activation of phospholipid synthesis. Nucleic Acids Res. 1992 Jun 25;20(12):3253–3253. doi: 10.1093/nar/20.12.3253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Hare P., Williams G. Structural studies of the acidic transactivation domain of the Vmw65 protein of herpes simplex virus using 1H NMR. Biochemistry. 1992 Apr 28;31(16):4150–4156. doi: 10.1021/bi00131a035. [DOI] [PubMed] [Google Scholar]
- Ogawa N., Oshima Y. Functional domains of a positive regulatory protein, PHO4, for transcriptional control of the phosphatase regulon in Saccharomyces cerevisiae. Mol Cell Biol. 1990 May;10(5):2224–2236. doi: 10.1128/mcb.10.5.2224. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Okada H., Toh-e A. A novel mutation occurring in the PHO80 gene suppresses the PHO4c mutations of Saccharomyces cerevisiae. Curr Genet. 1992 Feb;21(2):95–99. doi: 10.1007/BF00318466. [DOI] [PubMed] [Google Scholar]
- Oliner J. D., Pietenpol J. A., Thiagalingam S., Gyuris J., Kinzler K. W., Vogelstein B. Oncoprotein MDM2 conceals the activation domain of tumour suppressor p53. Nature. 1993 Apr 29;362(6423):857–860. doi: 10.1038/362857a0. [DOI] [PubMed] [Google Scholar]
- Salmeron J. M., Jr, Leuther K. K., Johnston S. A. GAL4 mutations that separate the transcriptional activation and GAL80-interactive functions of the yeast GAL4 protein. Genetics. 1990 May;125(1):21–27. doi: 10.1093/genetics/125.1.21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Uesono Y., Tanaka K., Toh-e A. Negative regulators of the PHO system in Saccharomyces cerevisiae: isolation and structural characterization of PHO85. Nucleic Acids Res. 1987 Dec 23;15(24):10299–10309. doi: 10.1093/nar/15.24.10299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Hoy M., Leuther K. K., Kodadek T., Johnston S. A. The acidic activation domains of the GCN4 and GAL4 proteins are not alpha helical but form beta sheets. Cell. 1993 Feb 26;72(4):587–594. doi: 10.1016/0092-8674(93)90077-4. [DOI] [PubMed] [Google Scholar]
- Vogel K., Hörz W., Hinnen A. The two positively acting regulatory proteins PHO2 and PHO4 physically interact with PHO5 upstream activation regions. Mol Cell Biol. 1989 May;9(5):2050–2057. doi: 10.1128/mcb.9.5.2050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yoshida K., Kuromitsu Z., Ogawa N., Oshima Y. Mode of expression of the positive regulatory genes PHO2 and PHO4 of the phosphatase regulon in Saccharomyces cerevisiae. Mol Gen Genet. 1989 May;217(1):31–39. doi: 10.1007/BF00330939. [DOI] [PubMed] [Google Scholar]
- Yoshida K., Ogawa N., Oshima Y. Function of the PHO regulatory genes for repressible acid phosphatase synthesis in Saccharomyces cerevisiae. Mol Gen Genet. 1989 May;217(1):40–46. doi: 10.1007/BF00330940. [DOI] [PubMed] [Google Scholar]




