Abstract
Two conserved elements direct the 3' end processing of histone messenger RNA: a stem-loop structure immediately upstream of the site of cleavage and the histone downstream element (HDE), located 12-19 nucleotides downstream of the stem-loop in the premessenger RNA. We studied the role of these two elements by systematically inserting up to 10 C residues between them in the mouse H2A-614 histone pre-mRNA. 3' End mapping of RNAs processed in vitro demonstrated that as the HDE is move downstream, the site of cleavage correspondingly moves 3'. In addition, the efficiency of processing declines. In the wild-type substrate, cleavage occurs 3' of an A residue; modest increases in the efficiency of processing of the insertion mutants were observed when an A residue was placed at the new cleavage site. The results of psoralen cross-linking studies and immunoprecipitations using anti-trimethylguanosine antibodies indicated that the decreased processing efficiency of the insertion mutants is not due to impaired binding of the U7 small nuclear ribonucleoprotein (snRNP). We conclude that the mammalian U7 snRNP acts as a molecular ruler, targeting enzymatic components of cleave histone pre-mRNAs a fixed distance from its binding site, the HDE.
Full text
PDF

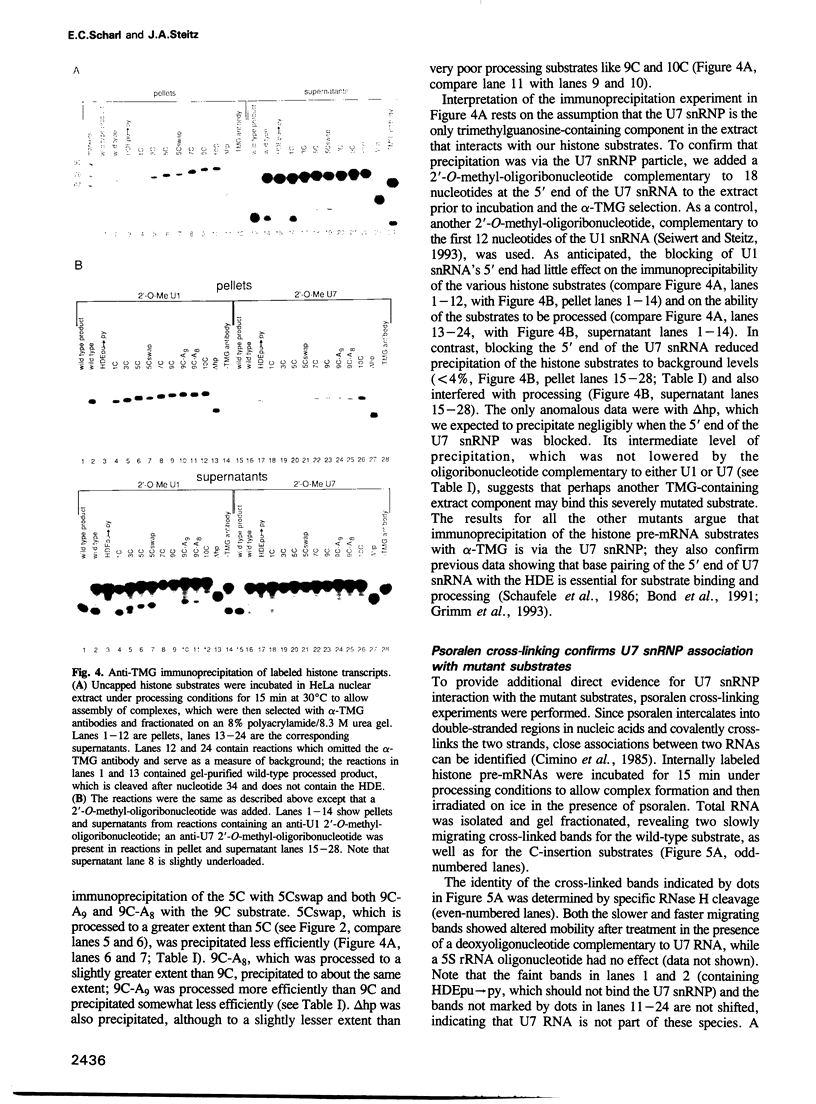
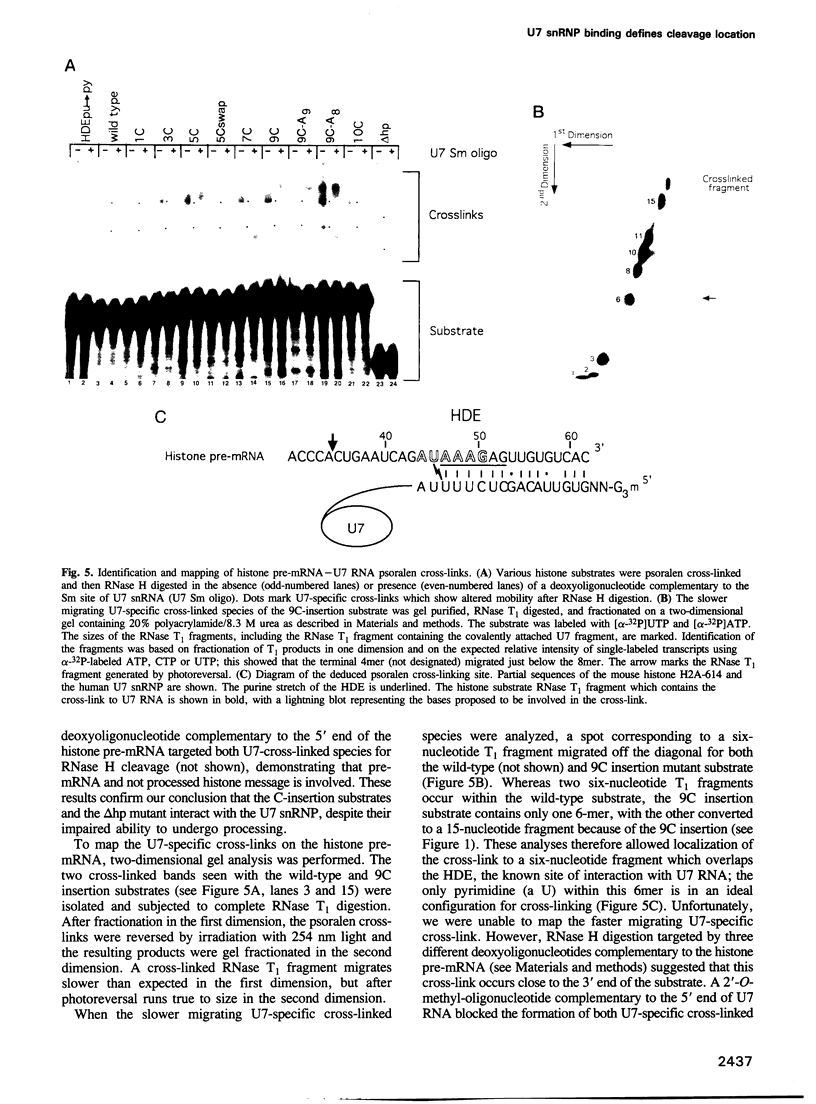



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Baldi M. I., Mattoccia E., Bufardeci E., Fabbri S., Tocchini-Valentini G. P. Participation of the intron in the reaction catalyzed by the Xenopus tRNA splicing endonuclease. Science. 1992 Mar 13;255(5050):1404–1408. doi: 10.1126/science.1542788. [DOI] [PubMed] [Google Scholar]
- Bond U. M., Yario T. A., Steitz J. A. Multiple processing-defective mutations in a mammalian histone pre-mRNA are suppressed by compensatory changes in U7 RNA both in vivo and in vitro. Genes Dev. 1991 Sep;5(9):1709–1722. doi: 10.1101/gad.5.9.1709. [DOI] [PubMed] [Google Scholar]
- Cimino G. D., Gamper H. B., Isaacs S. T., Hearst J. E. Psoralens as photoactive probes of nucleic acid structure and function: organic chemistry, photochemistry, and biochemistry. Annu Rev Biochem. 1985;54:1151–1193. doi: 10.1146/annurev.bi.54.070185.005443. [DOI] [PubMed] [Google Scholar]
- Cotten M., Gick O., Vasserot A., Schaffner G., Birnstiel M. L. Specific contacts between mammalian U7 snRNA and histone precursor RNA are indispensable for the in vitro 3' RNA processing reaction. EMBO J. 1988 Mar;7(3):801–808. doi: 10.1002/j.1460-2075.1988.tb02878.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Lorenzi M., Rohrer U., Birnstiel M. L. Analysis of a sea urchin gene cluster coding for the small nuclear U7 RNA, a rare RNA species implicated in the 3' editing of histone precursor mRNAs. Proc Natl Acad Sci U S A. 1986 May;83(10):3243–3247. doi: 10.1073/pnas.83.10.3243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dignam J. D., Lebovitz R. M., Roeder R. G. Accurate transcription initiation by RNA polymerase II in a soluble extract from isolated mammalian nuclei. Nucleic Acids Res. 1983 Mar 11;11(5):1475–1489. doi: 10.1093/nar/11.5.1475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Galli G., Hofstetter H., Stunnenberg H. G., Birnstiel M. L. Biochemical complementation with RNA in the Xenopus oocyte: a small RNA is required for the generation of 3' histone mRNA termini. Cell. 1983 Oct;34(3):823–828. doi: 10.1016/0092-8674(83)90539-1. [DOI] [PubMed] [Google Scholar]
- Georgiev O., Birnstiel M. L. The conserved CAAGAAAGA spacer sequence is an essential element for the formation of 3' termini of the sea urchin H3 histone mRNA by RNA processing. EMBO J. 1985 Feb;4(2):481–489. doi: 10.1002/j.1460-2075.1985.tb03654.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gick O., Krämer A., Vasserot A., Birnstiel M. L. Heat-labile regulatory factor is required for 3' processing of histone precursor mRNAs. Proc Natl Acad Sci U S A. 1987 Dec;84(24):8937–8940. doi: 10.1073/pnas.84.24.8937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gilmartin G. M., Schaufele F., Schaffner G., Birnstiel M. L. Functional analysis of the sea urchin U7 small nuclear RNA. Mol Cell Biol. 1988 Mar;8(3):1076–1084. doi: 10.1128/mcb.8.3.1076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Green M. R. Biochemical mechanisms of constitutive and regulated pre-mRNA splicing. Annu Rev Cell Biol. 1991;7:559–599. doi: 10.1146/annurev.cb.07.110191.003015. [DOI] [PubMed] [Google Scholar]
- Grimm C., Stefanovic B., Schümperli D. The low abundance of U7 snRNA is partly determined by its Sm binding site. EMBO J. 1993 Mar;12(3):1229–1238. doi: 10.1002/j.1460-2075.1993.tb05764.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gruber A., Soldati D., Burri M., Schümperli D. Isolation of an active gene and of two pseudogenes for mouse U7 small nuclear RNA. Biochim Biophys Acta. 1991 Jan 17;1088(1):151–154. doi: 10.1016/0167-4781(91)90167-k. [DOI] [PubMed] [Google Scholar]
- Guthrie C. Messenger RNA splicing in yeast: clues to why the spliceosome is a ribonucleoprotein. Science. 1991 Jul 12;253(5016):157–163. doi: 10.1126/science.1853200. [DOI] [PubMed] [Google Scholar]
- Hausner T. P., Giglio L. M., Weiner A. M. Evidence for base-pairing between mammalian U2 and U6 small nuclear ribonucleoprotein particles. Genes Dev. 1990 Dec;4(12A):2146–2156. doi: 10.1101/gad.4.12a.2146. [DOI] [PubMed] [Google Scholar]
- Heintz N., Roeder R. G. Transcription of human histone genes in extracts from synchronized HeLa cells. Proc Natl Acad Sci U S A. 1984 May;81(9):2713–2717. doi: 10.1073/pnas.81.9.2713. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hurt M. M., Chodchoy N., Marzluff W. F. The mouse histone H2a.2 gene from chromosome 3. Nucleic Acids Res. 1989 Nov 11;17(21):8876–8876. doi: 10.1093/nar/17.21.8876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krinke L., Wulff D. L. The cleavage specificity of RNase III. Nucleic Acids Res. 1990 Aug 25;18(16):4809–4815. doi: 10.1093/nar/18.16.4809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kunkel T. A., Roberts J. D., Zakour R. A. Rapid and efficient site-specific mutagenesis without phenotypic selection. Methods Enzymol. 1987;154:367–382. doi: 10.1016/0076-6879(87)54085-x. [DOI] [PubMed] [Google Scholar]
- Lüscher B., Schümperli D. RNA 3' processing regulates histone mRNA levels in a mammalian cell cycle mutant. A processing factor becomes limiting in G1-arrested cells. EMBO J. 1987 Jun;6(6):1721–1726. doi: 10.1002/j.1460-2075.1987.tb02423.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marzluff W. F. Histone 3' ends: essential and regulatory functions. Gene Expr. 1992;2(2):93–97. [PMC free article] [PubMed] [Google Scholar]
- Melin L., Soldati D., Mital R., Streit A., Schümperli D. Biochemical demonstration of complex formation of histone pre-mRNA with U7 small nuclear ribonucleoprotein and hairpin binding factors. EMBO J. 1992 Feb;11(2):691–697. doi: 10.1002/j.1460-2075.1992.tb05101.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mital R., Albrecht U., Schümperli D. Detection of UV-induced RNA:protein crosslinks in snRNPs by oligonucleotides complementary to the snRNA. Nucleic Acids Res. 1993 Feb 25;21(4):1049–1050. doi: 10.1093/nar/21.4.1049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mowry K. L., Oh R., Steitz J. A. Each of the conserved sequence elements flanking the cleavage site of mammalian histone pre-mRNAs has a distinct role in the 3'-end processing reaction. Mol Cell Biol. 1989 Jul;9(7):3105–3108. doi: 10.1128/mcb.9.7.3105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mowry K. L., Steitz J. A. Both conserved signals on mammalian histone pre-mRNAs associate with small nuclear ribonucleoproteins during 3' end formation in vitro. Mol Cell Biol. 1987 May;7(5):1663–1672. doi: 10.1128/mcb.7.5.1663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mowry K. L., Steitz J. A. Identification of the human U7 snRNP as one of several factors involved in the 3' end maturation of histone premessenger RNA's. Science. 1987 Dec 18;238(4834):1682–1687. doi: 10.1126/science.2825355. [DOI] [PubMed] [Google Scholar]
- Mowry K. L., Steitz J. A. snRNP mediators of 3' end processing: functional fossils? Trends Biochem Sci. 1988 Nov;13(11):447–451. doi: 10.1016/0968-0004(88)90220-4. [DOI] [PubMed] [Google Scholar]
- Phillips S. C., Birnstiel M. L. Analysis of a gene cluster coding for the Xenopus laevis U7 snRNA. Biochim Biophys Acta. 1992 May 7;1131(1):95–98. doi: 10.1016/0167-4781(92)90104-8. [DOI] [PubMed] [Google Scholar]
- Phillips S. C., Turner P. C. Nucleotide sequence of the mouse U7 snRNA gene. Nucleic Acids Res. 1991 Mar 25;19(6):1344–1344. doi: 10.1093/nar/19.6.1344. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reyes V. M., Abelson J. Substrate recognition and splice site determination in yeast tRNA splicing. Cell. 1988 Nov 18;55(4):719–730. doi: 10.1016/0092-8674(88)90230-9. [DOI] [PubMed] [Google Scholar]
- Schaufele F., Gilmartin G. M., Bannwarth W., Birnstiel M. L. Compensatory mutations suggest that base-pairing with a small nuclear RNA is required to form the 3' end of H3 messenger RNA. 1986 Oct 30-Nov 5Nature. 323(6091):777–781. doi: 10.1038/323777a0. [DOI] [PubMed] [Google Scholar]
- Seiwert S. D., Steitz J. A. Uncoupling two functions of the U1 small nuclear ribonucleoprotein particle during in vitro splicing. Mol Cell Biol. 1993 Jun;13(6):3135–3145. doi: 10.1128/mcb.13.6.3135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sheets M. D., Ogg S. C., Wickens M. P. Point mutations in AAUAAA and the poly (A) addition site: effects on the accuracy and efficiency of cleavage and polyadenylation in vitro. Nucleic Acids Res. 1990 Oct 11;18(19):5799–5805. doi: 10.1093/nar/18.19.5799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soldati D., Schümperli D. Structural and functional characterization of mouse U7 small nuclear RNA active in 3' processing of histone pre-mRNA. Mol Cell Biol. 1988 Apr;8(4):1518–1524. doi: 10.1128/mcb.8.4.1518. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Streit A., Koning T. W., Soldati D., Melin L., Schümperli D. Variable effects of the conserved RNA hairpin element upon 3' end processing of histone pre-mRNA in vitro. Nucleic Acids Res. 1993 Apr 11;21(7):1569–1575. doi: 10.1093/nar/21.7.1569. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strub K., Birnstiel M. L. Genetic complementation in the Xenopus oocyte: co-expression of sea urchin histone and U7 RNAs restores 3' processing of H3 pre-mRNA in the oocyte. EMBO J. 1986 Jul;5(7):1675–1682. doi: 10.1002/j.1460-2075.1986.tb04411.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strub K., Galli G., Busslinger M., Birnstiel M. L. The cDNA sequences of the sea urchin U7 small nuclear RNA suggest specific contacts between histone mRNA precursor and U7 RNA during RNA processing. EMBO J. 1984 Dec 1;3(12):2801–2807. doi: 10.1002/j.1460-2075.1984.tb02212.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vasserot A. P., Schaufele F. J., Birnstiel M. L. Conserved terminal hairpin sequences of histone mRNA precursors are not involved in duplex formation with the U7 RNA but act as a target site for a distinct processing factor. Proc Natl Acad Sci U S A. 1989 Jun;86(12):4345–4349. doi: 10.1073/pnas.86.12.4345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wahle E., Keller W. The biochemistry of 3'-end cleavage and polyadenylation of messenger RNA precursors. Annu Rev Biochem. 1992;61:419–440. doi: 10.1146/annurev.bi.61.070192.002223. [DOI] [PubMed] [Google Scholar]
- Wassarman D. A., Steitz J. A. Interactions of small nuclear RNA's with precursor messenger RNA during in vitro splicing. Science. 1992 Sep 25;257(5078):1918–1925. doi: 10.1126/science.1411506. [DOI] [PubMed] [Google Scholar]
- Wassarman K. M., Steitz J. A. Association with terminal exons in pre-mRNAs: a new role for the U1 snRNP? Genes Dev. 1993 Apr;7(4):647–659. doi: 10.1101/gad.7.4.647. [DOI] [PubMed] [Google Scholar]
- Watkins N. J., Phillips S. C., Turner P. C. The Xenopus U7 snRNA-encoding gene has an unusually compact structure. Gene. 1992 Oct 21;120(2):271–276. doi: 10.1016/0378-1119(92)90104-w. [DOI] [PubMed] [Google Scholar]
- Wells D. E. Compilation analysis of histones and histone genes. Nucleic Acids Res. 1986;14 (Suppl):r119–r149. doi: 10.1093/nar/14.suppl.r119. [DOI] [PMC free article] [PubMed] [Google Scholar]