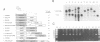Abstract
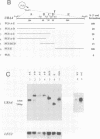
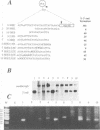
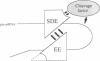
We have defined sequences in the 3' non-coding region of the Schizosaccharomyces pombe ura4 gene that are required for efficient mRNA 3' end formation. Three separate sequence elements have been identified. Two of these are site determining elements which specify alternative sites of polyadenylation [the major poly(A) site and a minor downstream poly(A) site]. The third sequence, located downstream of both poly(A) sites, functions as an efficiency element that enhances utilization of either polyadenylation site. By employing sensitive RT-PCR analysis, we demonstrate that although low levels of transcripts are detected up to the efficiency element, none is detected beyond this point. The downstream site determining element and efficiency element have both been delineated to specific 16 nt sequences which we show are together sufficient for ura4 mRNA 3' end formation. We have further characterized the interaction between these two elements and show that the efficiency element behaves in a position-independent, orientation-dependent manner, but cannot form 3' ends independently of the site determining element. Surprisingly, we find that the efficiency element can be functionally replaced by a second copy of either site determining element. We present a model for the mechanism of RNA 3' end formation of the ura4 gene and note that this bipartite structure for a poly(A) signal in S.pombe may be related to the AAUAAA and downstream GU-rich sequences of poly(A) signals in mammalian genes.
Full text
PDF

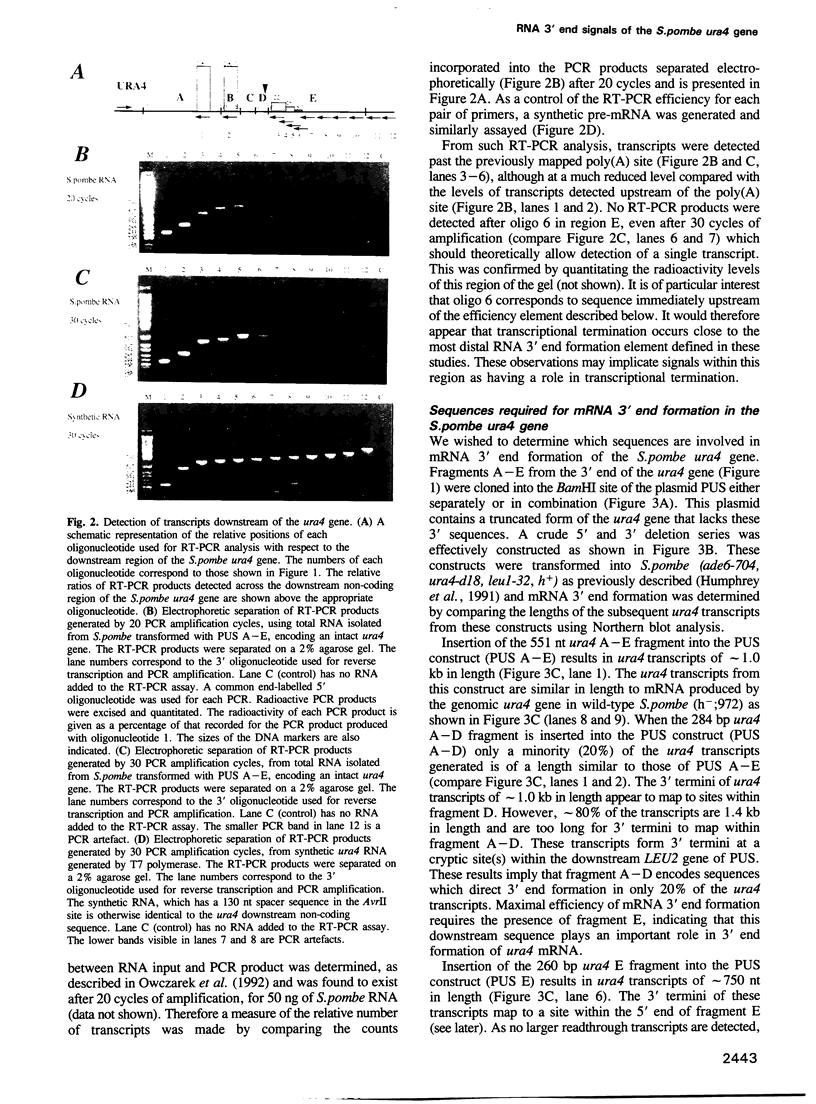

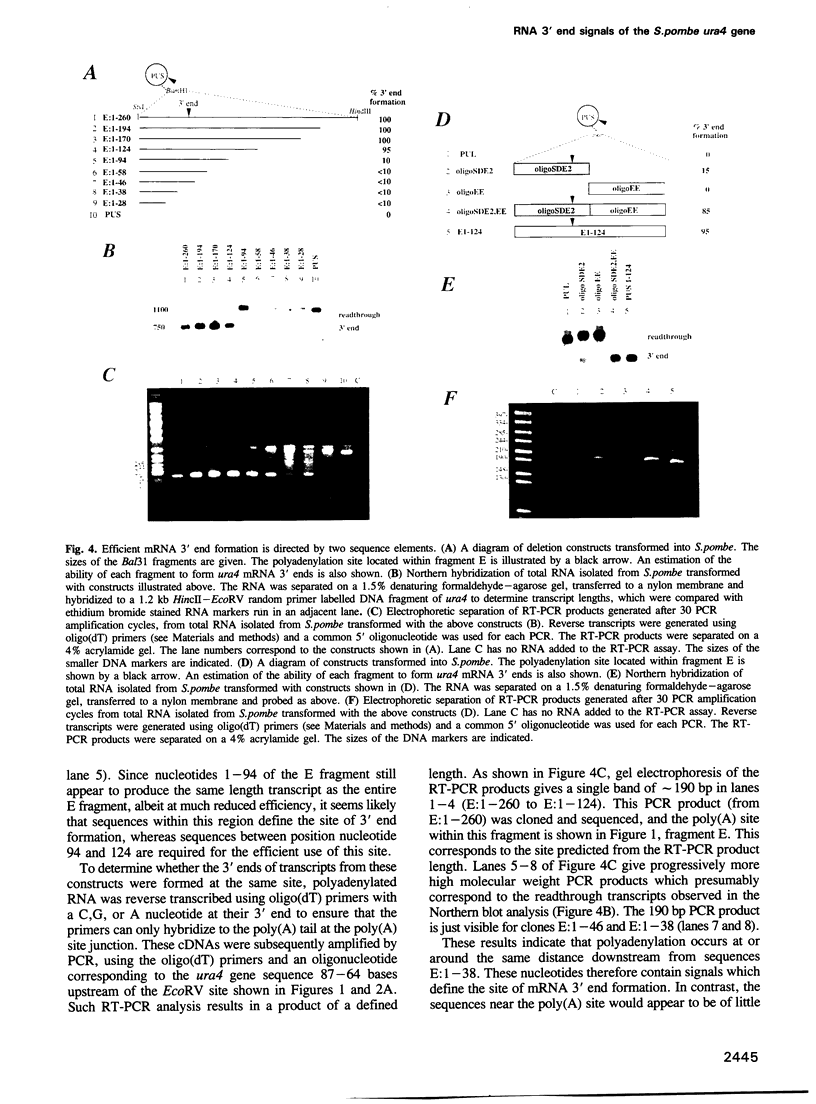






Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Abe A., Hiraoka Y., Fukasawa T. Signal sequence for generation of mRNA 3' end in the Saccharomyces cerevisiae GAL7 gene. EMBO J. 1990 Nov;9(11):3691–3697. doi: 10.1002/j.1460-2075.1990.tb07581.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Butler J. S., Platt T. RNA processing generates the mature 3' end of yeast CYC1 messenger RNA in vitro. Science. 1988 Dec 2;242(4883):1270–1274. doi: 10.1126/science.2848317. [DOI] [PubMed] [Google Scholar]
- Butler J. S., Sadhale P. P., Platt T. RNA processing in vitro produces mature 3' ends of a variety of Saccharomyces cerevisiae mRNAs. Mol Cell Biol. 1990 Jun;10(6):2599–2605. doi: 10.1128/mcb.10.6.2599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen J., Moore C. Separation of factors required for cleavage and polyadenylation of yeast pre-mRNA. Mol Cell Biol. 1992 Aug;12(8):3470–3481. doi: 10.1128/mcb.12.8.3470. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feinberg A. P., Vogelstein B. "A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity". Addendum. Anal Biochem. 1984 Feb;137(1):266–267. doi: 10.1016/0003-2697(84)90381-6. [DOI] [PubMed] [Google Scholar]
- Girard J. P., Feliu J., Caizergues-Ferrer M., Lapeyre B. Study of multiple fibrillarin mRNAs reveals that 3' end formation in Schizosaccharomyces pombe is sensitive to cold shock. Nucleic Acids Res. 1993 Apr 25;21(8):1881–1887. doi: 10.1093/nar/21.8.1881. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grimm C., Kohli J., Murray J., Maundrell K. Genetic engineering of Schizosaccharomyces pombe: a system for gene disruption and replacement using the ura4 gene as a selectable marker. Mol Gen Genet. 1988 Dec;215(1):81–86. doi: 10.1007/BF00331307. [DOI] [PubMed] [Google Scholar]
- Heidmann S., Obermaier B., Vogel K., Domdey H. Identification of pre-mRNA polyadenylation sites in Saccharomyces cerevisiae. Mol Cell Biol. 1992 Sep;12(9):4215–4229. doi: 10.1128/mcb.12.9.4215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henikoff S., Kelly J. D., Cohen E. H. Transcription terminates in yeast distal to a control sequence. Cell. 1983 Jun;33(2):607–614. doi: 10.1016/0092-8674(83)90441-5. [DOI] [PubMed] [Google Scholar]
- Humphrey T., Sadhale P., Platt T., Proudfoot N. Homologous mRNA 3' end formation in fission and budding yeast. EMBO J. 1991 Nov;10(11):3503–3511. doi: 10.1002/j.1460-2075.1991.tb04914.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hyman L. E., Moore C. L. Termination and pausing of RNA polymerase II downstream of yeast polyadenylation sites. Mol Cell Biol. 1993 Sep;13(9):5159–5167. doi: 10.1128/mcb.13.9.5159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hyman L. E., Seiler S. H., Whoriskey J., Moore C. L. Point mutations upstream of the yeast ADH2 poly(A) site significantly reduce the efficiency of 3'-end formation. Mol Cell Biol. 1991 Apr;11(4):2004–2012. doi: 10.1128/mcb.11.4.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Irniger S., Braus G. H. Saturation mutagenesis of a polyadenylation signal reveals a hexanucleotide element essential for mRNA 3' end formation in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 1994 Jan 4;91(1):257–261. doi: 10.1073/pnas.91.1.257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Irniger S., Egli C. M., Braus G. H. Different classes of polyadenylation sites in the yeast Saccharomyces cerevisiae. Mol Cell Biol. 1991 Jun;11(6):3060–3069. doi: 10.1128/mcb.11.6.3060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Köhrer K., Domdey H. Preparation of high molecular weight RNA. Methods Enzymol. 1991;194:398–405. doi: 10.1016/0076-6879(91)94030-g. [DOI] [PubMed] [Google Scholar]
- Lingner J., Kellermann J., Keller W. Cloning and expression of the essential gene for poly(A) polymerase from S. cerevisiae. Nature. 1991 Dec 12;354(6353):496–498. doi: 10.1038/354496a0. [DOI] [PubMed] [Google Scholar]
- Osborne B. I., Guarente L. Mutational analysis of a yeast transcriptional terminator. Proc Natl Acad Sci U S A. 1989 Jun;86(11):4097–4101. doi: 10.1073/pnas.86.11.4097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Osborne B. I., Guarente L. Transcription by RNA polymerase II induces changes of DNA topology in yeast. Genes Dev. 1988 Jun;2(6):766–772. doi: 10.1101/gad.2.6.766. [DOI] [PubMed] [Google Scholar]
- Owczarek C. M., Enriquez-Harris P., Proudfoot N. J. The primary transcription unit of the human alpha 2 globin gene defined by quantitative RT/PCR. Nucleic Acids Res. 1992 Feb 25;20(4):851–858. doi: 10.1093/nar/20.4.851. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Proudfoot N. J. How RNA polymerase II terminates transcription in higher eukaryotes. Trends Biochem Sci. 1989 Mar;14(3):105–110. doi: 10.1016/0968-0004(89)90132-1. [DOI] [PubMed] [Google Scholar]
- Proudfoot N. Poly(A) signals. Cell. 1991 Feb 22;64(4):671–674. doi: 10.1016/0092-8674(91)90495-k. [DOI] [PubMed] [Google Scholar]
- Rave N., Crkvenjakov R., Boedtker H. Identification of procollagen mRNAs transferred to diazobenzyloxymethyl paper from formaldehyde agarose gels. Nucleic Acids Res. 1979 Aug 10;6(11):3559–3567. doi: 10.1093/nar/6.11.3559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruohola H., Baker S. M., Parker R., Platt T. Orientation-dependent function of a short CYC1 DNA fragment in directing mRNA 3' end formation in yeast. Proc Natl Acad Sci U S A. 1988 Jul;85(14):5041–5045. doi: 10.1073/pnas.85.14.5041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Russo P., Li W. Z., Guo Z., Sherman F. Signals that produce 3' termini in CYC1 mRNA of the yeast Saccharomyces cerevisiae. Mol Cell Biol. 1993 Dec;13(12):7836–7849. doi: 10.1128/mcb.13.12.7836. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Russo P., Li W. Z., Hampsey D. M., Zaret K. S., Sherman F. Distinct cis-acting signals enhance 3' endpoint formation of CYC1 mRNA in the yeast Saccharomyces cerevisiae. EMBO J. 1991 Mar;10(3):563–571. doi: 10.1002/j.1460-2075.1991.tb07983.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Russo P., Sherman F. Transcription terminates near the poly(A) site in the CYC1 gene of the yeast Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 1989 Nov;86(21):8348–8352. doi: 10.1073/pnas.86.21.8348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sadhale P. P., Platt T. Unusual aspects of in vitro RNA processing in the 3' regions of the GAL1, GAL7, and GAL10 genes in Saccharomyces cerevisiae. Mol Cell Biol. 1992 Oct;12(10):4262–4270. doi: 10.1128/mcb.12.10.4262. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sadhale P. P., Sapolsky R., Davis R. W., Butler J. S., Platt T. Polymerase chain reaction mapping of yeast GAL7 mRNA polyadenylation sites demonstrates that 3' end processing in vitro faithfully reproduces the 3' ends observed in vivo. Nucleic Acids Res. 1991 Jul 11;19(13):3683–3688. doi: 10.1093/nar/19.13.3683. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wahle E., Keller W. The biochemistry of 3'-end cleavage and polyadenylation of messenger RNA precursors. Annu Rev Biochem. 1992;61:419–440. doi: 10.1146/annurev.bi.61.070192.002223. [DOI] [PubMed] [Google Scholar]
- Wu S. Y., Platt T. Transcriptional arrest of yeast RNA polymerase II by Escherichia coli rho protein in vitro. Proc Natl Acad Sci U S A. 1993 Jul 15;90(14):6606–6610. doi: 10.1073/pnas.90.14.6606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zaret K. S., Sherman F. DNA sequence required for efficient transcription termination in yeast. Cell. 1982 Mar;28(3):563–573. doi: 10.1016/0092-8674(82)90211-2. [DOI] [PubMed] [Google Scholar]
- Zaret K. S., Sherman F. Mutationally altered 3' ends of yeast CYC1 mRNA affect transcript stability and translational efficiency. J Mol Biol. 1984 Jul 25;177(1):107–135. doi: 10.1016/0022-2836(84)90060-3. [DOI] [PubMed] [Google Scholar]