Abstract
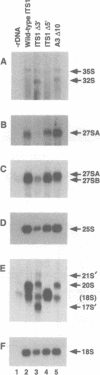
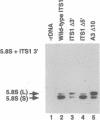
We have developed techniques for the detailed analysis of cis-acting sequences in the pre-rRNA of Saccharomyces cerevisiae and used these to study the processing of internal transcribed spacer 1 (ITS1) leading to the synthesis of 5.8S rRNA. As is the case for many eukaryotes, the 5' end of yeast 5.8S rRNA is heterogeneous; we designate the major, short form 5.8S(S), and the minor form (which is seven or eight nucleotides longer) 5.8S(L). These RNAs do not have a precursor/product relationship, but result from the use of alternative processing pathways. In the major pathway, a previously unidentified processing site in ITS1, designated A3, is cleaved. A 10 nucleotide deletion at site A3 strongly inhibits processing of A3 and the synthesis of 5.8S(S); processing is predominantly transferred to the alternative 5.8S(L) pathway. Site A3 lies 76 nucleotides 5' to the end of 5.8S(S), and acts as an entry site for 5'-->3' exonuclease digestion which generates the 5' end of 5.8S(S). This pathway is inhibited in strains mutant for XRN1p and RAT1p. Both of these proteins have been reported to have 5'-->3' exonuclease activity in vitro. Formation of 5.8S(L) is increased by mutations at A3 in cis or in RAT1p and XRN1p in trans, and is kinetically faster than 5.8S(S) synthesis.
Full text
PDF
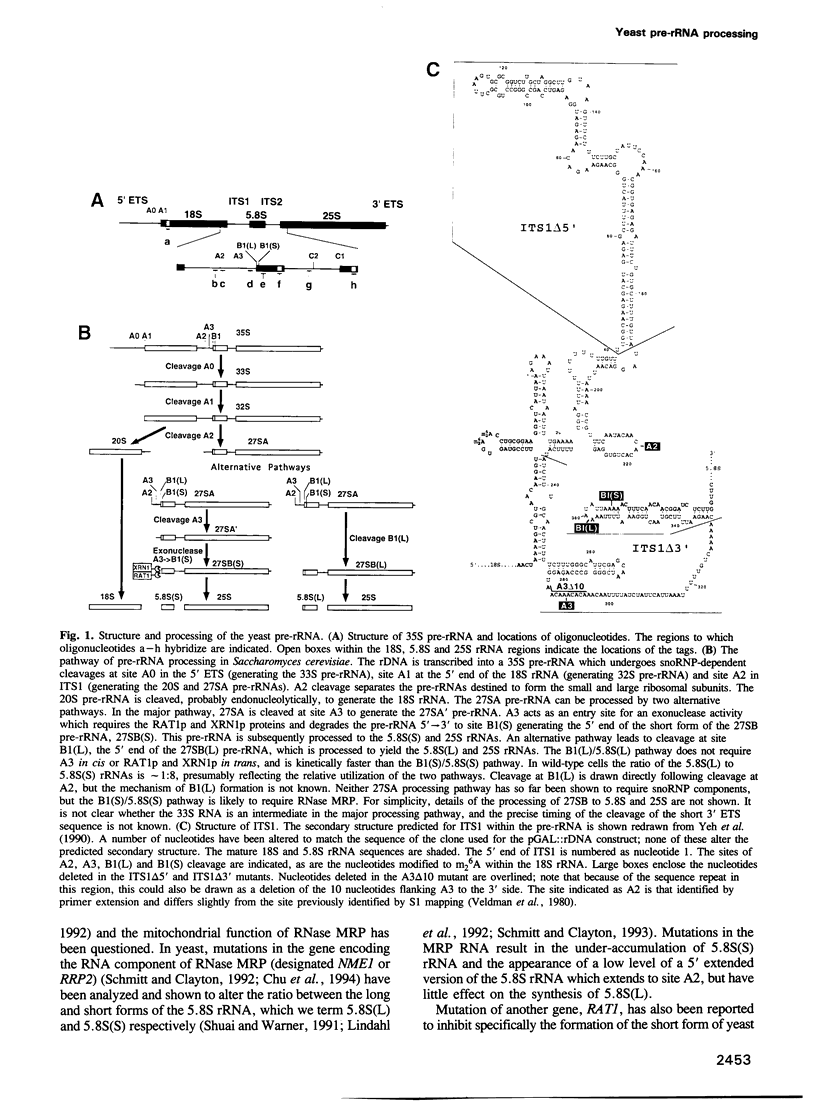










Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Aldrich T. L., Di Segni G., McConaughy B. L., Keen N. J., Whelen S., Hall B. D. Structure of the yeast TAP1 protein: dependence of transcription activation on the DNA context of the target gene. Mol Cell Biol. 1993 Jun;13(6):3434–3444. doi: 10.1128/mcb.13.6.3434. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beltrame M., Tollervey D. Identification and functional analysis of two U3 binding sites on yeast pre-ribosomal RNA. EMBO J. 1992 Apr;11(4):1531–1542. doi: 10.1002/j.1460-2075.1992.tb05198.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brand R. C., Klootwijk J., Van Steenbergen T. J., De Kok A. J., Planta R. J. Secondary methylation of yeast ribosomal precursor RNA. Eur J Biochem. 1977 May 2;75(1):311–318. doi: 10.1111/j.1432-1033.1977.tb11531.x. [DOI] [PubMed] [Google Scholar]
- Chang D. D., Clayton D. A. A mammalian mitochondrial RNA processing activity contains nucleus-encoded RNA. Science. 1987 Mar 6;235(4793):1178–1184. doi: 10.1126/science.2434997. [DOI] [PubMed] [Google Scholar]
- Chang D. D., Clayton D. A. A novel endoribonuclease cleaves at a priming site of mouse mitochondrial DNA replication. EMBO J. 1987 Feb;6(2):409–417. doi: 10.1002/j.1460-2075.1987.tb04770.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chu S., Archer R. H., Zengel J. M., Lindahl L. The RNA of RNase MRP is required for normal processing of ribosomal RNA. Proc Natl Acad Sci U S A. 1994 Jan 18;91(2):659–663. doi: 10.1073/pnas.91.2.659. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Di Segni G., McConaughy B. L., Shapiro R. A., Aldrich T. L., Hall B. D. TAP1, a yeast gene that activates the expression of a tRNA gene with a defective internal promoter. Mol Cell Biol. 1993 Jun;13(6):3424–3433. doi: 10.1128/mcb.13.6.3424. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Girard J. P., Lehtonen H., Caizergues-Ferrer M., Amalric F., Tollervey D., Lapeyre B. GAR1 is an essential small nucleolar RNP protein required for pre-rRNA processing in yeast. EMBO J. 1992 Feb;11(2):673–682. doi: 10.1002/j.1460-2075.1992.tb05099.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsu C. L., Stevens A. Yeast cells lacking 5'-->3' exoribonuclease 1 contain mRNA species that are poly(A) deficient and partially lack the 5' cap structure. Mol Cell Biol. 1993 Aug;13(8):4826–4835. doi: 10.1128/mcb.13.8.4826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hughes J. M., Ares M., Jr Depletion of U3 small nucleolar RNA inhibits cleavage in the 5' external transcribed spacer of yeast pre-ribosomal RNA and impairs formation of 18S ribosomal RNA. EMBO J. 1991 Dec;10(13):4231–4239. doi: 10.1002/j.1460-2075.1991.tb05001.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jansen R., Tollervey D., Hurt E. C. A U3 snoRNP protein with homology to splicing factor PRP4 and G beta domains is required for ribosomal RNA processing. EMBO J. 1993 Jun;12(6):2549–2558. doi: 10.1002/j.1460-2075.1993.tb05910.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kearsey S., Kipling D. Recombination and RNA processing: a common strand? Trends Cell Biol. 1991 Nov;1(5):110–112. doi: 10.1016/0962-8924(91)90101-e. [DOI] [PubMed] [Google Scholar]
- Kenna M., Stevens A., McCammon M., Douglas M. G. An essential yeast gene with homology to the exonuclease-encoding XRN1/KEM1 gene also encodes a protein with exoribonuclease activity. Mol Cell Biol. 1993 Jan;13(1):341–350. doi: 10.1128/mcb.13.1.341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kipling D., Tambini C., Kearsey S. E. rar mutations which increase artificial chromosome stability in Saccharomyces cerevisiae identify transcription and recombination proteins. Nucleic Acids Res. 1991 Apr 11;19(7):1385–1391. doi: 10.1093/nar/19.7.1385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kiss T., Marshallsay C., Filipowicz W. 7-2/MRP RNAs in plant and mammalian cells: association with higher order structures in the nucleolus. EMBO J. 1992 Oct;11(10):3737–3746. doi: 10.1002/j.1460-2075.1992.tb05459.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klootwijk J., Planta R. J. Isolation and characterization of yeast ribosomal RNA precursors and preribosomes. Methods Enzymol. 1989;180:96–109. doi: 10.1016/0076-6879(89)80095-3. [DOI] [PubMed] [Google Scholar]
- Lamm G. M., Blencowe B. J., Sproat B. S., Iribarren A. M., Ryder U., Lamond A. I. Antisense probes containing 2-aminoadenosine allow efficient depletion of U5 snRNP from HeLa splicing extracts. Nucleic Acids Res. 1991 Jun 25;19(12):3193–3198. doi: 10.1093/nar/19.12.3193. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lamond A. I., Sproat B. S. Antisense oligonucleotides made of 2'-O-alkylRNA: their properties and applications in RNA biochemistry. FEBS Lett. 1993 Jun 28;325(1-2):123–127. doi: 10.1016/0014-5793(93)81427-2. [DOI] [PubMed] [Google Scholar]
- Larimer F. W., Hsu C. L., Maupin M. K., Stevens A. Characterization of the XRN1 gene encoding a 5'-->3' exoribonuclease: sequence data and analysis of disparate protein and mRNA levels of gene-disrupted yeast cells. Gene. 1992 Oct 12;120(1):51–57. doi: 10.1016/0378-1119(92)90008-d. [DOI] [PubMed] [Google Scholar]
- Li H. D., Zagorski J., Fournier M. J. Depletion of U14 small nuclear RNA (snR128) disrupts production of 18S rRNA in Saccharomyces cerevisiae. Mol Cell Biol. 1990 Mar;10(3):1145–1152. doi: 10.1128/mcb.10.3.1145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lindahl L., Archer R. H., Zengel J. M. A new rRNA processing mutant of Saccharomyces cerevisiae. Nucleic Acids Res. 1992 Jan 25;20(2):295–301. doi: 10.1093/nar/20.2.295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maden B. E. The numerous modified nucleotides in eukaryotic ribosomal RNA. Prog Nucleic Acid Res Mol Biol. 1990;39:241–303. doi: 10.1016/s0079-6603(08)60629-7. [DOI] [PubMed] [Google Scholar]
- Morrissey J. P., Tollervey D. Yeast snR30 is a small nucleolar RNA required for 18S rRNA synthesis. Mol Cell Biol. 1993 Apr;13(4):2469–2477. doi: 10.1128/mcb.13.4.2469. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Musters W., Boon K., van der Sande C. A., van Heerikhuizen H., Planta R. J. Functional analysis of transcribed spacers of yeast ribosomal DNA. EMBO J. 1990 Dec;9(12):3989–3996. doi: 10.1002/j.1460-2075.1990.tb07620.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Musters W., Venema J., van der Linden G., van Heerikhuizen H., Klootwijk J., Planta R. J. A system for the analysis of yeast ribosomal DNA mutations. Mol Cell Biol. 1989 Feb;9(2):551–559. doi: 10.1128/mcb.9.2.551. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nogi Y., Yano R., Dodd J., Carles C., Nomura M. Gene RRN4 in Saccharomyces cerevisiae encodes the A12.2 subunit of RNA polymerase I and is essential only at high temperatures. Mol Cell Biol. 1993 Jan;13(1):114–122. doi: 10.1128/mcb.13.1.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nogi Y., Yano R., Nomura M. Synthesis of large rRNAs by RNA polymerase II in mutants of Saccharomyces cerevisiae defective in RNA polymerase I. Proc Natl Acad Sci U S A. 1991 May 1;88(9):3962–3966. doi: 10.1073/pnas.88.9.3962. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reddy R., Henning D., Busch H. Primary and secondary structure of U8 small nuclear RNA. J Biol Chem. 1985 Sep 15;260(20):10930–10935. [PubMed] [Google Scholar]
- Reddy R., Li W. Y., Henning D., Choi Y. C., Nohga K., Busch H. Characterization and subcellular localization of 7-8 S RNAs of Novikoff hepatoma. J Biol Chem. 1981 Aug 25;256(16):8452–8457. [PubMed] [Google Scholar]
- Schmitt M. E., Clayton D. A. Nuclear RNase MRP is required for correct processing of pre-5.8S rRNA in Saccharomyces cerevisiae. Mol Cell Biol. 1993 Dec;13(12):7935–7941. doi: 10.1128/mcb.13.12.7935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmitt M. E., Clayton D. A. Yeast site-specific ribonucleoprotein endoribonuclease MRP contains an RNA component homologous to mammalian RNase MRP RNA and essential for cell viability. Genes Dev. 1992 Oct;6(10):1975–1985. doi: 10.1101/gad.6.10.1975. [DOI] [PubMed] [Google Scholar]
- Shuai K., Warner J. R. A temperature sensitive mutant of Saccharomyces cerevisiae defective in pre-rRNA processing. Nucleic Acids Res. 1991 Sep 25;19(18):5059–5064. doi: 10.1093/nar/19.18.5059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stevens A., Hsu C. L., Isham K. R., Larimer F. W. Fragments of the internal transcribed spacer 1 of pre-rRNA accumulate in Saccharomyces cerevisiae lacking 5'----3' exoribonuclease 1. J Bacteriol. 1991 Nov;173(21):7024–7028. doi: 10.1128/jb.173.21.7024-7028.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stevens A. Purification and characterization of a Saccharomyces cerevisiae exoribonuclease which yields 5'-mononucleotides by a 5' leads to 3' mode of hydrolysis. J Biol Chem. 1980 Apr 10;255(7):3080–3085. [PubMed] [Google Scholar]
- Stohl L. L., Clayton D. A. Saccharomyces cerevisiae contains an RNase MRP that cleaves at a conserved mitochondrial RNA sequence implicated in replication priming. Mol Cell Biol. 1992 Jun;12(6):2561–2569. doi: 10.1128/mcb.12.6.2561. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tollervey D., Lehtonen H., Carmo-Fonseca M., Hurt E. C. The small nucleolar RNP protein NOP1 (fibrillarin) is required for pre-rRNA processing in yeast. EMBO J. 1991 Mar;10(3):573–583. doi: 10.1002/j.1460-2075.1991.tb07984.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tollervey D., Mattaj I. W. Fungal small nuclear ribonucleoproteins share properties with plant and vertebrate U-snRNPs. EMBO J. 1987 Feb;6(2):469–476. doi: 10.1002/j.1460-2075.1987.tb04777.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Udem S. A., Warner J. R. Ribosomal RNA synthesis in Saccharomyces cerevisiae. J Mol Biol. 1972 Mar 28;65(2):227–242. doi: 10.1016/0022-2836(72)90279-3. [DOI] [PubMed] [Google Scholar]
- Veldman G. M., Brand R. C., Klootwijk J., Planta R. Some characteristics of processing sites in ribosomal precursor RNA of yeast. Nucleic Acids Res. 1980 Jul 11;8(13):2907–2920. doi: 10.1093/nar/8.13.2907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Veldman G. M., Klootwijk J., van Heerikhuizen H., Planta R. J. The nucleotide sequence of the intergenic region between the 5.8S and 26S rRNA genes of the yeast ribosomal RNA operon. Possible implications for the interaction between 5.8S and 26S rRNA and the processing of the primary transcript. Nucleic Acids Res. 1981 Oct 10;9(19):4847–4862. doi: 10.1093/nar/9.19.4847. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wittekind M., Dodd J., Vu L., Kolb J. M., Buhler J. M., Sentenac A., Nomura M. Isolation and characterization of temperature-sensitive mutations in RPA190, the gene encoding the largest subunit of RNA polymerase I from Saccharomyces cerevisiae. Mol Cell Biol. 1988 Oct;8(10):3997–4008. doi: 10.1128/mcb.8.10.3997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yeh L. C., Thweatt R., Lee J. C. Internal transcribed spacer 1 of the yeast precursor ribosomal RNA. Higher order structure and common structural motifs. Biochemistry. 1990 Jun 26;29(25):5911–5918. doi: 10.1021/bi00477a005. [DOI] [PubMed] [Google Scholar]
- Yuan Y., Singh R., Reddy R. Rat nucleolar 7-2 RNA is homologous to mouse mitochondrial RNase mitochondrial RNA-processing RNA. J Biol Chem. 1989 Sep 5;264(25):14835–14839. [PubMed] [Google Scholar]
- van der Sande C. A., Kwa M., van Nues R. W., van Heerikhuizen H., Raué H. A., Planta R. J. Functional analysis of internal transcribed spacer 2 of Saccharomyces cerevisiae ribosomal DNA. J Mol Biol. 1992 Feb 20;223(4):899–910. doi: 10.1016/0022-2836(92)90251-e. [DOI] [PubMed] [Google Scholar]