Abstract
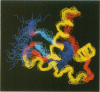
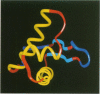
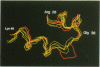
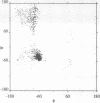
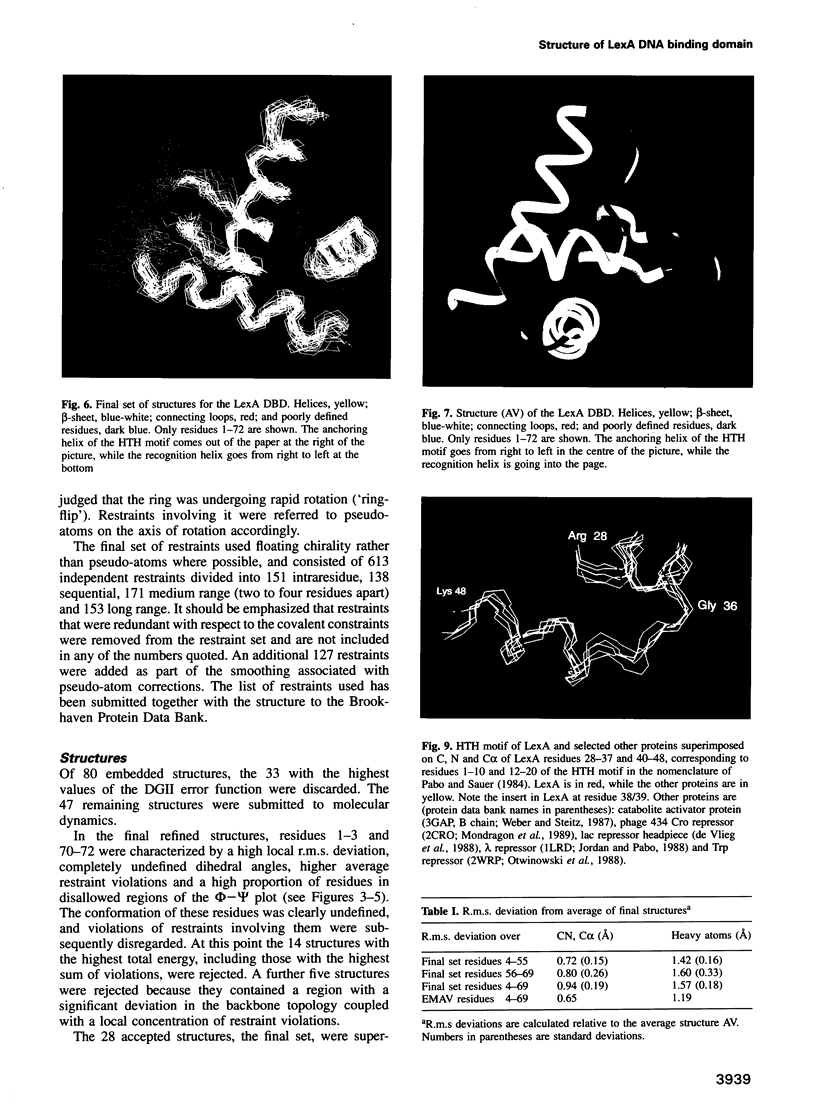
The structure of the 84 residue DNA binding domain of the Escherichia coli LexA repressor has been determined from NMR data using distance geometry and restrained molecular dynamics. The assignment of the 1H NMR spectrum of the molecule, derived from 2- and 3-D homonuclear experiments, is also reported. A total of 613 non-redundant distance restraints were used to give a final family of 28 structures. The structured region of the molecule consisted of residues 4-69 and yielded a r.m.s. deviation from an average of 0.9 A for backbone and 1.6 A for all heavy atoms. The structure contains three regular alpha-helices at residues 6-21 (I), 28-35 (II) and 41-52 (III), and an antiparallel beta-sheet at residues 56-58 and 66-68. Helices II and III form a variant helix-turn-helix DNA binding motif, with an unusual one residue insert at residue 38. The topology of the LexA DNA binding domain is found to be the same as for the DNA binding domains of the catabolic activator protein, human histone 5, the HNF-3/fork head protein and the Kluyveromyces lactis heat shock transcription factor.
Full text
PDF

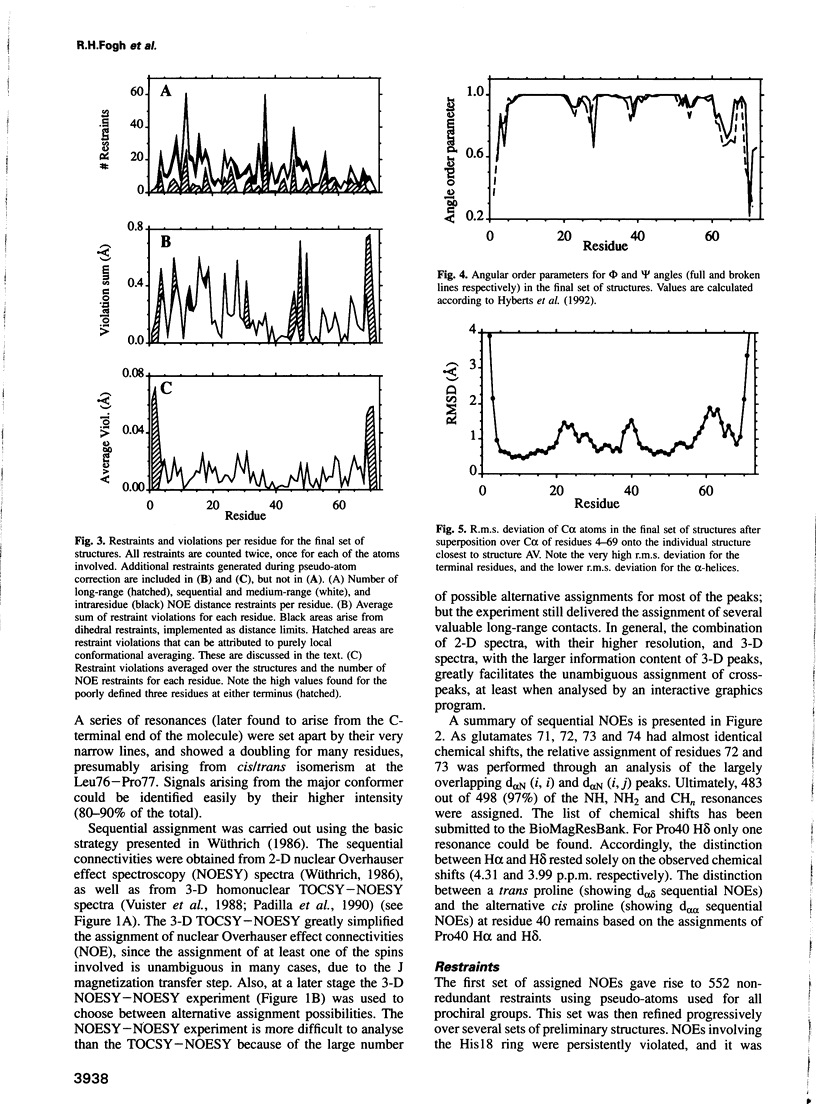

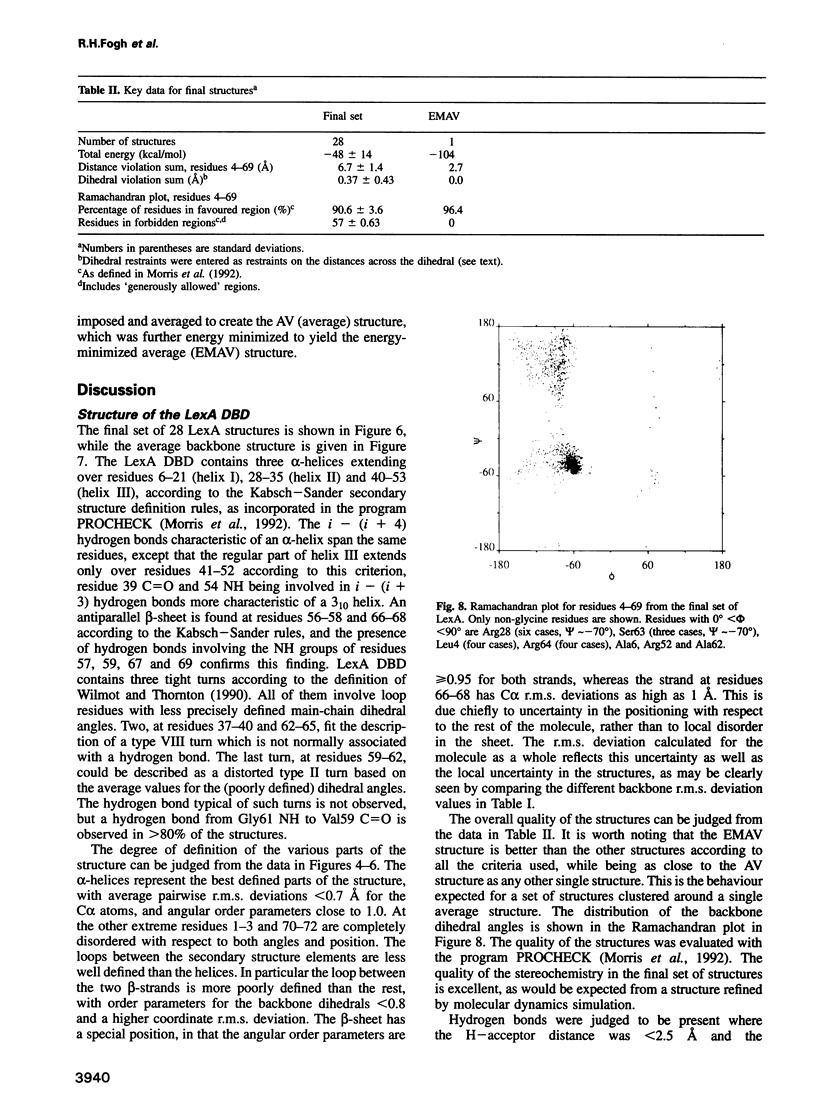




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Baker E. N., Hubbard R. E. Hydrogen bonding in globular proteins. Prog Biophys Mol Biol. 1984;44(2):97–179. doi: 10.1016/0079-6107(84)90007-5. [DOI] [PubMed] [Google Scholar]
- Bertrand-Burggraf E., Hurstel S., Daune M., Schnarr M. Promoter properties and negative regulation of the uvrA gene by the LexA repressor and its amino-terminal DNA binding domain. J Mol Biol. 1987 Jan 20;193(2):293–302. doi: 10.1016/0022-2836(87)90220-8. [DOI] [PubMed] [Google Scholar]
- Clark K. L., Halay E. D., Lai E., Burley S. K. Co-crystal structure of the HNF-3/fork head DNA-recognition motif resembles histone H5. Nature. 1993 Jul 29;364(6436):412–420. doi: 10.1038/364412a0. [DOI] [PubMed] [Google Scholar]
- Dodd I. B., Egan J. B. Improved detection of helix-turn-helix DNA-binding motifs in protein sequences. Nucleic Acids Res. 1990 Sep 11;18(17):5019–5026. doi: 10.1093/nar/18.17.5019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dodd I. B., Egan J. B. Systematic method for the detection of potential lambda Cro-like DNA-binding regions in proteins. J Mol Biol. 1987 Apr 5;194(3):557–564. doi: 10.1016/0022-2836(87)90681-4. [DOI] [PubMed] [Google Scholar]
- Dumoulin P., Oertel-Buchheit P., Granger-Schnarr M., Schnarr M. Orientation of the LexA DNA-binding motif on operator DNA as inferred from cysteine-mediated phenyl azide crosslinking. Proc Natl Acad Sci U S A. 1993 Mar 1;90(5):2030–2034. doi: 10.1073/pnas.90.5.2030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fogh R. H., Kem W. R., Norton R. S. Solution structure of neurotoxin I from the sea anemone Stichodactyla helianthus. A nuclear magnetic resonance, distance geometry, and restrained molecular dynamics study. J Biol Chem. 1990 Aug 5;265(22):13016–13028. doi: 10.2210/pdb2sh1/pdb. [DOI] [PubMed] [Google Scholar]
- Harrison C. J., Bohm A. A., Nelson H. C. Crystal structure of the DNA binding domain of the heat shock transcription factor. Science. 1994 Jan 14;263(5144):224–227. doi: 10.1126/science.8284672. [DOI] [PubMed] [Google Scholar]
- Havel T. F. An evaluation of computational strategies for use in the determination of protein structure from distance constraints obtained by nuclear magnetic resonance. Prog Biophys Mol Biol. 1991;56(1):43–78. doi: 10.1016/0079-6107(91)90007-f. [DOI] [PubMed] [Google Scholar]
- Hurstel S., Granger-Schnarr M., Daune M., Schnarr M. In vitro binding of LexA repressor to DNA: evidence for the involvement of the amino-terminal domain. EMBO J. 1986 Apr;5(4):793–798. doi: 10.1002/j.1460-2075.1986.tb04283.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hurstel S., Granger-Schnarr M., Schnarr M. Contacts between the LexA repressor--or its DNA-binding domain--and the backbone of the recA operator DNA. EMBO J. 1988 Jan;7(1):269–275. doi: 10.1002/j.1460-2075.1988.tb02809.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hyberts S. G., Goldberg M. S., Havel T. F., Wagner G. The solution structure of eglin c based on measurements of many NOEs and coupling constants and its comparison with X-ray structures. Protein Sci. 1992 Jun;1(6):736–751. doi: 10.1002/pro.5560010606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jordan S. R., Pabo C. O. Structure of the lambda complex at 2.5 A resolution: details of the repressor-operator interactions. Science. 1988 Nov 11;242(4880):893–899. doi: 10.1126/science.3187530. [DOI] [PubMed] [Google Scholar]
- Lamerichs R. M., Padilla A., Boelens R., Kaptein R., Ottleben G., Rüterjans H., Granger-Schnarr M., Oertel P., Schnarr M. The amino-terminal domain of LexA repressor is alpha-helical but differs from canonical helix-turn-helix proteins: a two-dimensional 1H NMR study. Proc Natl Acad Sci U S A. 1989 Sep;86(18):6863–6867. doi: 10.1073/pnas.86.18.6863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Little J. W. Autodigestion of lexA and phage lambda repressors. Proc Natl Acad Sci U S A. 1984 Mar;81(5):1375–1379. doi: 10.1073/pnas.81.5.1375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Little J. W., Hill S. A. Deletions within a hinge region of a specific DNA-binding protein. Proc Natl Acad Sci U S A. 1985 Apr;82(8):2301–2305. doi: 10.1073/pnas.82.8.2301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Little J. W. Mechanism of specific LexA cleavage: autodigestion and the role of RecA coprotease. Biochimie. 1991 Apr;73(4):411–421. doi: 10.1016/0300-9084(91)90108-d. [DOI] [PubMed] [Google Scholar]
- Little J. W., Mount D. W. The SOS regulatory system of Escherichia coli. Cell. 1982 May;29(1):11–22. doi: 10.1016/0092-8674(82)90085-x. [DOI] [PubMed] [Google Scholar]
- Mondragón A., Wolberger C., Harrison S. C. Structure of phage 434 Cro protein at 2.35 A resolution. J Mol Biol. 1989 Jan 5;205(1):179–188. doi: 10.1016/0022-2836(89)90374-4. [DOI] [PubMed] [Google Scholar]
- Morris A. L., MacArthur M. W., Hutchinson E. G., Thornton J. M. Stereochemical quality of protein structure coordinates. Proteins. 1992 Apr;12(4):345–364. doi: 10.1002/prot.340120407. [DOI] [PubMed] [Google Scholar]
- Oertel-Buchheit P., Lamerichs R. M., Schnarr M., Granger-Schnarr M. Genetic analysis of the LexA repressor: isolation and characterization of LexA(Def) mutant proteins. Mol Gen Genet. 1990 Aug;223(1):40–48. doi: 10.1007/BF00315795. [DOI] [PubMed] [Google Scholar]
- Oertel-Buchheit P., Porte D., Schnarr M., Granger-Schnarr M. Isolation and characterization of LexA mutant repressors with enhanced DNA binding affinity. J Mol Biol. 1992 Jun 5;225(3):609–620. doi: 10.1016/0022-2836(92)90389-2. [DOI] [PubMed] [Google Scholar]
- Otwinowski Z., Schevitz R. W., Zhang R. G., Lawson C. L., Joachimiak A., Marmorstein R. Q., Luisi B. F., Sigler P. B. Crystal structure of trp repressor/operator complex at atomic resolution. Nature. 1988 Sep 22;335(6188):321–329. doi: 10.1038/335321a0. [DOI] [PubMed] [Google Scholar]
- Pabo C. O., Sauer R. T. Protein-DNA recognition. Annu Rev Biochem. 1984;53:293–321. doi: 10.1146/annurev.bi.53.070184.001453. [DOI] [PubMed] [Google Scholar]
- Pabo C. O., Sauer R. T. Transcription factors: structural families and principles of DNA recognition. Annu Rev Biochem. 1992;61:1053–1095. doi: 10.1146/annurev.bi.61.070192.005201. [DOI] [PubMed] [Google Scholar]
- Presta L. G., Rose G. D. Helix signals in proteins. Science. 1988 Jun 17;240(4859):1632–1641. doi: 10.1126/science.2837824. [DOI] [PubMed] [Google Scholar]
- Ramakrishnan V., Finch J. T., Graziano V., Lee P. L., Sweet R. M. Crystal structure of globular domain of histone H5 and its implications for nucleosome binding. Nature. 1993 Mar 18;362(6417):219–223. doi: 10.1038/362219a0. [DOI] [PubMed] [Google Scholar]
- Richardson J. S., Richardson D. C. Amino acid preferences for specific locations at the ends of alpha helices. Science. 1988 Jun 17;240(4859):1648–1652. doi: 10.1126/science.3381086. [DOI] [PubMed] [Google Scholar]
- Schnarr M., Granger-Schnarr M., Hurstel S., Pouyet J. The carboxy-terminal domain of the LexA repressor oligomerises essentially as the entire protein. FEBS Lett. 1988 Jul 4;234(1):56–60. doi: 10.1016/0014-5793(88)81302-4. [DOI] [PubMed] [Google Scholar]
- Schnarr M., Oertel-Buchheit P., Kazmaier M., Granger-Schnarr M. DNA binding properties of the LexA repressor. Biochimie. 1991 Apr;73(4):423–431. doi: 10.1016/0300-9084(91)90109-e. [DOI] [PubMed] [Google Scholar]
- Schnarr M., Pouyet J., Granger-Schnarr M., Daune M. Large-scale purification, oligomerization equilibria, and specific interaction of the LexA repressor of Escherichia coli. Biochemistry. 1985 May 21;24(11):2812–2818. doi: 10.1021/bi00332a032. [DOI] [PubMed] [Google Scholar]
- Schultz S. C., Shields G. C., Steitz T. A. Crystal structure of a CAP-DNA complex: the DNA is bent by 90 degrees. Science. 1991 Aug 30;253(5023):1001–1007. doi: 10.1126/science.1653449. [DOI] [PubMed] [Google Scholar]
- Steitz T. A. Structural studies of protein-nucleic acid interaction: the sources of sequence-specific binding. Q Rev Biophys. 1990 Aug;23(3):205–280. doi: 10.1017/s0033583500005552. [DOI] [PubMed] [Google Scholar]
- Thliveris A. T., Little J. W., Mount D. W. Repression of the E coli recA gene requires at least two LexA protein monomers. Biochimie. 1991 Apr;73(4):449–456. doi: 10.1016/0300-9084(91)90112-e. [DOI] [PubMed] [Google Scholar]
- Thliveris A. T., Mount D. W. Genetic identification of the DNA binding domain of Escherichia coli LexA protein. Proc Natl Acad Sci U S A. 1992 May 15;89(10):4500–4504. doi: 10.1073/pnas.89.10.4500. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walker G. C. Mutagenesis and inducible responses to deoxyribonucleic acid damage in Escherichia coli. Microbiol Rev. 1984 Mar;48(1):60–93. doi: 10.1128/mr.48.1.60-93.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weber I. T., Steitz T. A. Structure of a complex of catabolite gene activator protein and cyclic AMP refined at 2.5 A resolution. J Mol Biol. 1987 Nov 20;198(2):311–326. doi: 10.1016/0022-2836(87)90315-9. [DOI] [PubMed] [Google Scholar]
- Wertman K. F., Mount D. W. Nucleotide sequence binding specificity of the LexA repressor of Escherichia coli K-12. J Bacteriol. 1985 Jul;163(1):376–384. doi: 10.1128/jb.163.1.376-384.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilmot C. M., Thornton J. M. Beta-turns and their distortions: a proposed new nomenclature. Protein Eng. 1990 May;3(6):479–493. doi: 10.1093/protein/3.6.479. [DOI] [PubMed] [Google Scholar]
- Wilson K. P., Shewchuk L. M., Brennan R. G., Otsuka A. J., Matthews B. W. Escherichia coli biotin holoenzyme synthetase/bio repressor crystal structure delineates the biotin- and DNA-binding domains. Proc Natl Acad Sci U S A. 1992 Oct 1;89(19):9257–9261. doi: 10.1073/pnas.89.19.9257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wüthrich K., Billeter M., Braun W. Pseudo-structures for the 20 common amino acids for use in studies of protein conformations by measurements of intramolecular proton-proton distance constraints with nuclear magnetic resonance. J Mol Biol. 1983 Oct 5;169(4):949–961. doi: 10.1016/s0022-2836(83)80144-2. [DOI] [PubMed] [Google Scholar]
- de Vlieg J., Scheek R. M., van Gunsteren W. F., Berendsen H. J., Kaptein R., Thomason J. Combined procedure of distance geometry and restrained molecular dynamics techniques for protein structure determination from nuclear magnetic resonance data: application to the DNA binding domain of lac repressor from Escherichia coli. Proteins. 1988;3(4):209–218. doi: 10.1002/prot.340030402. [DOI] [PubMed] [Google Scholar]