Abstract
The evolutionary history of the Mycobacterium tuberculosis complex (MTBC) has previously been studied by analysis of sequence diversity in extant strains, but not addressed by direct examination of strain genotypes in archaeological remains. Here, we use ancient DNA sequencing to type 11 single nucleotide polymorphisms and two large sequence polymorphisms in the MTBC strains present in 10 archaeological samples from skeletons from Britain and Europe dating to the second–nineteenth centuries AD. The results enable us to assign the strains to groupings and lineages recognized in the extant MTBC. We show that at least during the eighteenth–nineteenth centuries AD, strains of M. tuberculosis belonging to different genetic groups were present in Britain at the same time, possibly even at a single location, and we present evidence for a mixed infection in at least one individual. Our study shows that ancient DNA typing applied to multiple samples can provide sufficiently detailed information to contribute to both archaeological and evolutionary knowledge of the history of tuberculosis.
Keywords: tuberculosis, ancient DNA, strain typing, single nucleotide polymorphisms, mixed infection
1. Introduction
Tuberculosis (TB) has caused millions of deaths throughout history and is still a major burden in many parts of the world. Especially in the seventeenth–nineteenth centuries AD, TB was highly prevalent throughout Europe, urbanization facilitating spread of the disease in overcrowded environments [1]. The improved living standards in the late nineteenth century AD led to a decline in incidence rates and a further drop was achieved by the use of antibiotics in the mid-twentieth century [1,2]. Since its re-emergence in the 1980s, however, TB has once again become one of the leading infectious diseases, causing morbidity and mortality in all parts of the world, with an estimated 1.4 million deaths in 2011 [3].
TB is caused by the members of the Mycobacterium tuberculosis complex (MTBC), with M. tuberculosis being the most common infecting species in humans. The MTBC also comprises the human pathogens M. africanum and M. canettii as well as the primarily animal infecting species M. bovis, M. microti, M. pinnipedii and M. caprae, all of which have been identified as causative agents of TB in humans [4–8]. The continuing appearance of antibiotic-resistant MTBC strains has stimulated interest in the evolutionary history of TB, in particular the possible coevolution between MTBC lineages and human populations [9]. Modern genetic data indicate that the MTBC may have coexisted with humans for at least 15 000 years [10–14], and archaeological evidence suggests that it has afflicted humankind since the Neolithic [15–18]. Throughout the past two decades, studies of ancient DNA (aDNA) in archaeological remains have begun to contribute to our understanding of the evolutionary history of the MTBC, several publications reporting the presence of MTBC aDNA in human remains, and some attempting to classify the infecting MTBC strains based on the identities of various genetic markers [17–28].
Several methods are used in clinical research to classify MTBC isolates into groups of related strains, including IS6110 restriction fragment length polymorphism (IS6110 RFLP) [29], mycobacterial interspersed repetitive unit-variable number tandem repeat (MIRU-VNTR) typing [30,31], spacer oligotyping (spoligotyping) [32], targeting of large sequence polymorphisms (LSPs) [33–35] and typing of single nucleotide polymorphisms (SNPs) [13,36–39]. RFLP analysis of IS6110, MIRU-VNTR typing and spoligotyping are less suitable for phylogenetic analyses as convergent evolution may result in homoplasy of the targeted loci [40,41]. Spoligotyping has been applied in studies of ancient TB [17–24] but its use has been questioned [42], not just because of its limited phylogenetic value but also because of its methodological problems when highly degraded aDNA is being analysed. LSPs and SNPs are considered to be the most suitable markers for strain identification and phylogenetic analysis, although they are not exempt from convergent evolution [43,44]. Early sets of SNPs were discovered by analysing selected genes or comparing genome sequences from only few strains, thereby not accessing a large part of the global variation [45]. More recently, de novo sequencing of 89 genes in 108 strains from all parts of the world identified 488 SNPs and resulted in a detailed phylogeny of the MTBC [46]. Whole genome comparison of 22 globally representative, mainly newly sequenced strains subsequently revealed 9037 unique SNPs and further resolved the phylogeny of the MTBC [47], and 34 167 SNPs identified from 259 strains have been used to study the coevolutionary history of M. tuberculosis and prehistoric human populations [14]. Targeting such large sets of loci is not feasible in aDNA studies if conventional PCR procedures are applied because the amount of extract available for analysis is usually very limited. These methodological constraints can be overcome by next generation sequencing (NGS), which is now being adopted in human aDNA studies and has recently been applied to historic strains of plague [48], TB [49] and leprosy [50]. However, NGS methods have their own limitations, requiring relatively large amounts of aDNA, being computationally intensive, and suffering from missing data owing to the absence of sequence reads covering particular SNPs, even ones that can be typed in the same sample by conventional PCR [49].
In this study, we show that an informative comparison of MTBC strains in archaeological human bone and dental samples is possible by conventional PCR of eleven SNPs and two LSPs (table 1). These markers enable MTBC strains to be classified into principal genetic groups (PGGs) 1–3 [36], lineages I–IV and M. bovis [38], SNP cluster groups (SCGs) 1–7 [39], ‘modern’ M. tuberculosis [33] and the Euro-American lineage of modern M. tuberculosis [34,35]. Five additional SNPs [51] allow a more precise classification of strains of the Euro-American lineage. We typed these markers in British and other European archaeological bone and dental samples from skeletons dated to the second–nineteenth centuries AD, revealing historic variations in genotype identities.
Table 1.
Loci targeted in this study in order to classify ancient MTBC strains.
| locus type | locusa | classification into | references |
|---|---|---|---|
| SNP | gyrA284 | PGGs 1–3 | [36] |
| katG1388 | |||
| oxyR37 | lineages I–IV and M. bovis | [38] | |
| oxyR285 | |||
| rpoB2646 | |||
| rpoB3243 | |||
| leuB (3352929) | SNP cluster groups 1–7 | [39] | |
| qcrB (2460626) | |||
| recN (1920118) | |||
| Rv0083 (92197) | |||
| Rv2802c (3111473) | |||
| LSP | TbD1 | ancient/modern M. tuberculosis | [33] |
| pks15/1 | Euro-American lineage | [34,35] |
aNumbers in parentheses denote the nucleotide position in H37Rv, as given in Filliol et al. [39].
2. Material and methods
Thirty-four bone and dental samples were selected based on the positive outcomes of PCRs directed at the IS6110 and IS1081 insertion sequences [52], which are looked on as specific for the MTBC group of bacteria. The samples come from skeletons dated to the second–nineteenth centuries AD and most but not all displayed lesions specific or non-specific for TB (electronic supplementary material, table S1). We have previously reported NGS and conventional SNP typing results for one of these samples, St George's Crypt 4006 [49].
Samples were taken under clean conditions by personnel wearing forensic suits, hair nets, face masks and sterile gloves, and stored in sterile plastic bags under dry conditions. Work was performed at the University of Manchester and the Complutense University of Madrid. The aDNA facility at the University of Manchester comprises independent, physically isolated laboratories for extraction and PCR set-up, each with an ultrafiltered air supply maintaining positive displacement pressure. DNA extractions were prepared in a Class II biological safety cabinet, and PCRs were set up in a laminar flow hood. Surfaces were sterilized by UV irradiation and regularly cleaned with 5% bleach and 70% ethanol. All equipment was treated with DNA-Away (Molecular Bioproducts) and tubes, pipettes and aqueous solutions were UV irradiated (254 nm, 120 000 µJ cm−2) for at least 10 min before use. Personnel wore protective clothing at all times, including forensic suits, face masks, hair nets, goggles and two pairs of sterile gloves. Work in Madrid was also carried out in physically separated laboratories for DNA extraction and PCR set-up, UV irradiated both before and after use. Surfaces and laboratory equipment were regularly cleaned with bleach. Personnel wore disposable forensic suits, face masks, caps, glasses, shoe covers and gloves. All reagents and consumables were DNase and RNase free. All procedures were carried out in a laminar flow cabinet, UV irradiated and cleaned with bleach before use. DNA extractions were accompanied by two blanks (extraction without skeletal material) per five samples (Manchester) or one blank per seven samples (Madrid). A set of 5–7 PCRs was always accompanied by at least two PCR blanks (set up with water instead of DNA extract).
Samples were prepared by removing approximately 1–2 mm of the outer surface of the bone mechanically, followed by UV irradiation (254 nm, 120 000 µJ cm–2) for 2 × 5 min, with 180° rotation between the two exposures, and subsequent crushing of the bone into powder. Each tooth was cleaned externally by placing it for 5 min in a small beaker containing 5% bleach, without allowing bleach to enter the root canal. The tooth was then dried with a paper towel, placed in a second beaker and rinsed in Millipore water, again avoiding entry of water into the root canal. After drying, a 37% phosphoric acid etching solution was applied to the tooth surface, left for 1 min and wiped off. The tooth was rinsed in Millipore water, dried for 10 min, and dentine powder collected using a dental pick.
At least two extractions were prepared for each sample, using 250 mg of bone or 50–100 mg of dentine powder. Extractions at Manchester used the method described by Bouwman & Brown [53] and/or a protocol based on Rohland & Hofreiter [54] and Rohland et al. [55], previously described by Bouwman et al. [49]. A subset of samples was re-extracted at Madrid using the latter method only. PCRs were directed at up to 16 SNPs and two LSPs (table 1; electronic supplementary material). PCRs were set up in a final volume of 30 µl, comprising 3–5 µl of DNA extract, 1× AmpliTaq Gold PCR Master Mix (Applied Biosystems), 400 nM each primer and 1% BSA. Cycling conditions were: 7 min at 95°C; followed by 45 cycles of 1 min at x°C and 1 min at 94°C; followed by 10 min at 72°C, where x°C is the primer-specific annealing temperature (electronic supplementary material, table S2). For primers with an annealing temperature less than or equal to 60°C, a three-temperature PCR was set up, with each annealing step at x°C followed by a 1 min extension at 72°C. All PCR products were examined by electrophoresis in 2% agarose gels, purified either from the gel or directly using Qiaquick columns (Qiagen) and subsequently cloned into Escherichia coli XL1-Blue competent cells (Agilent) using the CloneJet PCR cloning kit (Fermentas). Colony PCR was performed in 20 μl comprising 1× Taq buffer (New England Biolabs), 200 nM each primer, 200 μM dNTPs and 0.625 units Taq DNA polymerase (New England Biolabs), with cycling at: 95°C for 3 min; 30 cycles of 30 s at 94°C, 30 s at 60°C, 1 min at 72°C; 10 min at 72°C. PCR products were then sequenced (GATC Biotech, Cologne) and sequences aligned with the M. tuberculosis H37Rv reference sequence using Geneious v. 6.0.3 (http://www.geneious.com/).
To compare our results with 21 extant MTBC strains, targeted regions available from the six samples that provided unambiguous typing results were concatenated and aligned with the equivalent concatenated regions of the extant MTBC strains. For Auldhame 43, this procedure was repeated for all available sequences, including the four additional SNPs taken from Abadia et al. [51]. Neighbour-joining trees were created using MEGA v. 5.2.1 [56] and visualized with Dendroscope v. 3.2.4 [57].
3. Results
We analysed 34 samples from 31 individuals from 22 archaeological sites and one pathological reference collection (electronic supplementary material, table S1) for 11 SNPs and two LSPs (table 1). We obtained data for ten samples from seven archaeological sites (electronic supplementary material, table S3), enabling us to classify the infecting MTBC strains according to their SNP identities, the presence/absence of the TbD1 locus, and/or the presence of deletions within pks15/1 (table 2). Six of the samples (Auldhame 43, Saint Amé 20, St George's Crypt 4006, St Peter's Church 1390, St Peter's Collegiate Church 62, Whitefriars 10466) provided sufficient information, reproducible with two independent extracts, to assign the infecting strains to both their PGGs [36] and SCGs [39]. Two other samples, St Peter's Collegiate Church 28 and St George's Crypt 5003, gave incomplete results that enabled a PGG and/or SCG to be assigned from one extract but not confirmed with the second extract. Ambiguous results were obtained with two additional samples. Whilst Whitefriars 657 was identified as containing a SCG 6 strain, the gyrA284 SNP was typed as a C in one extract and G in the other, suggesting either PGG 2 or 3. With Ashchurch 705, clone sequences from the first extract gave a mixture of C and G at this position, and the qcrB SNP, which distinguishes SCG 3 from SCG 4, could be typed with only one of the two extracts (electronic supplementary material, table S3).
Table 2.
Summary of SNP and LSP data.a —, no result obtained; n.d., not done; TbD1−, deletion of TbD1 has occurred; Δ7 bp, 7 bp deletion in pks15/1 has occurred.
| sample ID | sample date | TbD1 | pks15/1 | PGG | lineage | SCG |
|---|---|---|---|---|---|---|
| Ashchurch 705b | 129–317 calAD | TbD1−/TbD1− | Δ7 bp/Δ7 bp | 2/3/2 | I or II/– | 3/3 or 4 |
| Auldhame 43b | 1280–1394 calAD | TbD1−/n.d. | Δ7 bp/Δ7 bp | 2/2 | II/– | 5/5 |
| Saint Amé 20 | 16th–18th centuries AD | –/n.d. | –/Δ7 bp | 3/3 | not IV/not III | 6/6 |
| St George's Crypt 4006b | mid-19th century AD | TbD1−/TbD1− | Δ7 bp/Δ7 bp | 3/3 | II/II | 6/6 |
| St George's Crypt 5003 | mid-19th century AD | –/TbD1− | Δ7 bp/– | –/– | not IV/– | 6/– |
| St Peter's Church 1390b | 1016–1155 calAD | TbD1−/n.d. | Δ7 bp/Δ7bp | 2/2 | I or II/– | 3/3 |
| St Peter's Collegiate Church 28 | 19th century AD | TbD1−/n.d. | Δ7 bp/– | 3/– | I or II/– | 6/– |
| St Peter's Collegiate Church 62b | 19th century AD | TbD1−/TbD1− | Δ7 bp/Δ7 bp | 2/2 | I or II/II | 4/4 |
| Whitefriars 657 | 18th–19th centuries AD | –/n.d. | Δ7 bp/– | 2/3 | I or II/– | 6/6 |
| Whitefriars 10466b | 18th–19th centuries AD | TbD1−/n.d. | –/Δ7 bp | 2/2 | I or II/– | 4/4 |
aResults listed as first/second extraction.
bSecond extraction performed in Madrid, except for TbD1.
The oxyR and rpoB PCRs used to assign strains to lineages I–IV [38] were less successful owing to lack of reproducibility and amplification of non-specific rpoB3243 targets. Nine of the 10 samples could be assigned to lineages I or II, but distinction between these lineages, which requires rpoB3243, was only possible for Auldhame 43, St Peter's Collegiate Church 62 and St George's Crypt 4006, each of which was identified as lineage II based on results with one or both extracts. St George's Crypt 5003 could only be identified as not lineage IV. The pks15/1 PCR revealed a 7 bp deletion for each of the 10 samples, suggesting that the strains belong to the Euro-American lineage, and amplification of the region flanking TbD1 was achieved for eight samples, again indicating that these are modern M. tuberculosis strains.
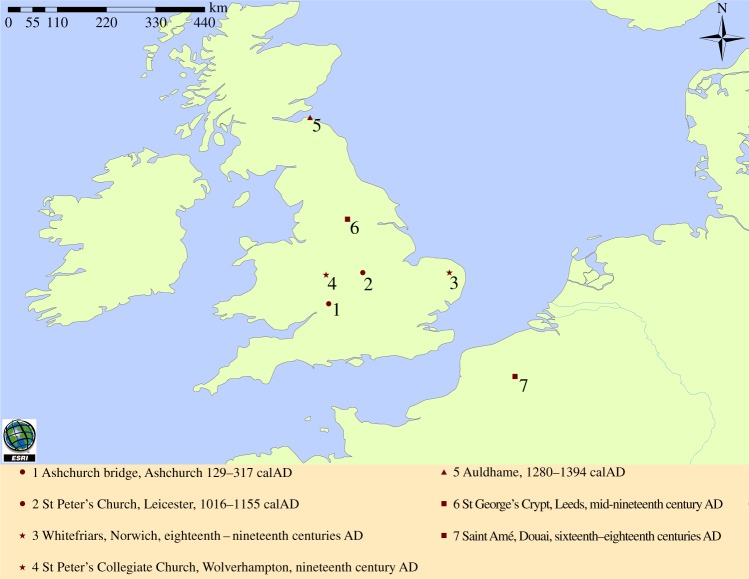
All but one of the 10 samples for which we obtained strain data came from British excavation sites spanning the second–nineteenth centuries AD, the one exception coming from sixteenth–eighteenth centuries AD Douai in northern France (figure 1). The relationships between these strains and modern ones are depicted as a neighbour-joining tree in figure 2a. Additional targeting of five of the SNPs reported by Abadia et al. [51] was attempted with one extract of Auldhame 43, a member of SCG 5. Four SNPs were typed, enabling more accurate resolution of the position of Auldhame 43 compared to the extant MTBC strains (figure 2b).
Figure 1.
Location of the sites for the 10 samples for which we report strain data. Circles, location of a strain belonging to PGG 2 and SCG 3; triangle, PGG 2 and SCG 5 strains; squares, PGG 3 and SCG 6 strains; stars, sites for which both PGG 2 and PGG 3 as well as SCG 4 and SCG 6 strains were identified. Note that evidence for an additional second strain belonging to PGG 3 was obtained for the sample from the individual from Ashchurch. (Online version in colour.)
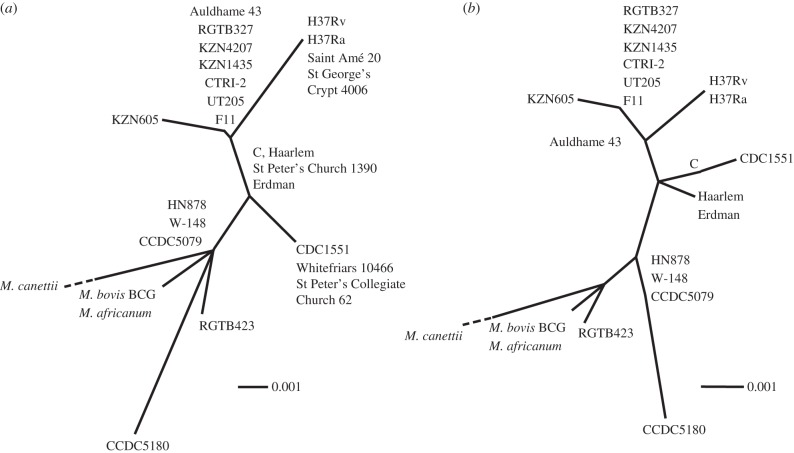
Figure 2.
(a) Neighbour-joining tree comparing the concatenated sequences of eight regions (478 bp—gyrA, katG, leuB, oxyR37, pks15/1, qcrB, rpoB2646 and Rv0083) obtained from a total of six samples (Auldhame 43, Saint Amé 20, St Peter's Church 1390, St Peter's Collegiate Church 62, St George's Crypt 4006 and Whitefriars 10466) with the equivalent regions of 21 extant MTBC strains. These extant MTBC strains are M. tuberculosis strains H37Rv (National Center for Biotechnology Information reference sequence NC_000962.2), H37Ra (NC_009525.1), ATCC35801 str. Erdman (AP012340.1), CCDC5079 (NC_017523.1), CCDC5180 (NC_017522.1), CDC1551 (NC_002755.2), CTRI2 (NC_017524.1), F11 (NC_009565.1), KZN605 (NC_018078.1), KZN1435 (NC_012943.1), KZN4207 (NC_016768.1), RGTB327 (NC_017026.1), RGTB423 (NC_017528.1), UT205 (NC_016934.1) and HN878 (CM001043.1) as well as M. bovis bacillus Calmette–Guérin str. Mexico, (NC_016804.1), M. africanum GM041182 (NC_015758.1) and M. canettii CIPT140010059 (NC_015848.1). Further strain data from whole genome shotgun sequencing projects was available from the Broad Institute (M. tuberculosis comparative sequencing project, Broad Institute of Harvard and MIT (http://www.broadinstitute.org/)) for: M. tuberculosis C (GenBank accession number AAKR00000000), M. tuberculosis Haarlem (AASN00000000) and M. tuberculosis W-148 (ACSX00000000.1). (b) Neighbour-joining tree comparing the concatenated sequences of 16 regions (1054 bp) obtained from Auldhame 43 with the equivalent regions of the 21 extant MTBC strains listed in (a). Bootstrap values were weak for both trees, as expected due to the small character set.
4. Discussion
We obtained sufficient SNP and/or LSP data to classify the M. tuberculosis strains present in 10 of the 34 bone and dental samples that we studied. Our results clearly show that, at least during the eighteenth–nineteenth centuries AD, strains of M. tuberculosis belonging to different genetic groups were present in Britain at the same time, possibly even at a single location. Within this time period, we discovered PGG 2/SCG 4 strains in individuals at sites in Norwich, eastern England (Whitefriars 10466), and Wolverhampton, central England (St Peter's Collegiate Church 62), and a PGG 3/SCG 6 strain at Leeds, northern England (St George's Crypt 4006). Strains of both types might even have coexisted in the same local areas, as a second Wolverhampton individual (St Peter's Collegiate Church 28) had a PGG 3/SCG 6 strain, and a second individual from Norwich (Whitefriars 657) had a PGG 2/3/SCG 6 strain. Both of these identifications are, however, tentative (at least according to the strict technical regime that we adopted) because they were not fully reproducible. Only one extract from St Peter's Collegiate Church 28 gave results, and the PGG classification for Whitefriars 657 was ambiguous, PGG 2 being identified with the first extraction and PGG 3 with the second.
One explanation of the ambiguous Whitefriars 657 result is that this individual was co-infected with two strains of M. tuberculosis, one strain belonging to PGG 2 and the other to PGG 3. Based on the presence of a T at Rv0083, in both extracts, the infecting strains could further be assigned to SCG 6. However, SNP typing success with the second extract was limited (table 2; electronic supplementary material, table S3), and for this reason we look on the individual from Whitefriars (657) as having possible but not definite mixed infection, i.e. TB caused by two different strains. A second, more convincing example of mixed infection was presented by the Roman sample Ashchurch 705. One of the extracts of this sample gave both possible nucleotides for gyrA284 (C and G), indicating an infection with both PGG 2 and PGG 3 strains. Owing to the type of polymorphism, we exclude the disparity as resulting from a miscoding lesion [58–60], and there was no evidence of contamination (electronic supplementary material). Mixed infection has been reported with patients today, either as concurrent infections with multiple MTBC strains or as exogenous re-infections [61]. The presence of a mixed infection in Ashchurch 705 also indicates that both PGG 2 and PGG 3 strains coexisted in southwest Britain during the second–fourth centuries AD. Since the second extract for Ashchurch 705 revealed a PGG 2 strain only, we assume that the SCG genotype obtained from both extracts derived from this PGG 2 strain. However, we cannot establish if the PGG 3 strain displayed the same SCG genotype as the PGG 2 strain.
Filliol et al. [39] have suggested that, of the four SCGs 3–6, SCG 3 preceded SCG 5 with the latter followed by SCG 4 and SCG 6 appearing lastly. Although our sample size is small, the results are consistent with this succession of SCG types. The three oldest samples that yielded genotype data were from skeleton 705 from Ashchurch (129–317 calAD) and St Peter's Church 1390 (1016–1155 calAD), both of whom had strains belonging to PGG 2/SCG 3, and skeleton 43 from Auldhame (1280–1394 calAD), which was identified as PGG 2/SCG 5. The remaining six British samples, dated to the eighteenth–nineteenth centuries AD, were from individuals with either PGG 2/SCG 4 or PGG 3/SCG 6, with skeleton 657 from Whitefriars possibly also harbouring a PGG 2 strain. Additionally, a sixteenth–eighteenth century AD individual from Douai, northern France (Saint Amé 20), was shown to contain a PGG 3/SCG 6 strain.
The markers we typed do not distinguish the a and b subgroups of SCG 6. However, we have previously shown by NGS genotyping that skeleton 4006 from St George's Crypt had a SCG 6b strain [49]. This is the same SCG as the MTBC reference strain H37Rv [39], which was first isolated at the beginning of the twentieth century from a North American patient [62]. Our results therefore indicate that strains similar to H37Rv might have been present in continental Europe in the sixteenth–eighteenth centuries AD and geographically dispersed in eighteenth–nineteenth centuries AD Britain.
Strains belonging to SCG 3b, 3c, 4 and 5 fall into PGG 2 as they harbour a polymorphism at katG1388 but not gyrA284 [39]. By contrast, SCG 6 strains display polymorphisms at both katG1388 and gyrA284 and are therefore classified as PGG 3 [39]. Further analyses will be necessary in order to identify whether skeletons from Ashchurch (705) and St Peter's Church (1390) indeed had strains belonging to SCG 3b or 3c as suggested by the PGG 2 determination or if they had strains belonging to subgroup SCG 3a which is primarily found within PGG 1 [39], with possible exceptions [63]. PGGs 2 and 3 (and likewise their associated SCGs) are also known to comprise lineage II strains, as identified by a polymorphism at rpoB3243 [38], and to lack the M. tuberculosis-specific deletion TbD1 [33] as well as a 7 bp region within pks15/1 [34]. Deletion of the 7 bp region was revealed for all 10 of our samples, with eight of them also shown to lack TbD1, classifying them as strains belonging to the Euro-American clade of modern M. tuberculosis. While deletion of TbD1 has previously been reported in an individual from Britain dating to approximately 2200 years BP [25], our results for Ashchurch 705 now additionally disclose that the deletion of the 7 bp region within pks15/1 had already taken place by the second–fourth centuries AD in Britain.
By typing four of the SNPs reported by Abadia et al. [51], we showed that the infecting SCG 5 strain found in the skeleton of Auldhame 43 is a member of the group that is ancestral to extant strains of the Latin-American Mediterranean clade within the Euro-American lineage. The detection of a SCG 5 strain at Auldhame (a coastal settlement east of Edinburgh, Scotland) but a SCG 3 strain at Leicester only about 100–200 years earlier raises the possibility that the Auldhame strain was not introduced into the human population in Scotland from southern parts of Britain but from Scandinavia instead. The first archaeological evidence of TB in Scotland [64] pre-dates Viking invasions in the late eighth century AD and Scandinavian contact in subsequent centuries [65], and stable isotope analysis of samples from individuals from the Auldhame site has indicated that the skeletal population itself is most likely composed of local individuals [66]. Nevertheless, introduction of (new) TB strains via this route is a possible scenario as the osteological evidence suggests the presence of TB in Scandinavia during the Iron Age (fifth–first centuries BC) as well as the medieval period (1050–1536 AD) [1,67–69].
Acknowledgements
We thank Darlene Weston (University of British Columbia) for collection of samples and Abigail Bouwman (University of Zürich) for sampling and extraction of a subset of samples at Manchester. We further thank Eva Fernández Domínguez (John Moores University of Liverpool) for extraction of samples and helping R.M. in Madrid, Cristina Gamba (University College Dublin) for supporting R.M. and Eduardo Arroyo-Pardo (Complutense University of Madrid) for the invitation to analyse our samples in Madrid. We also thank the following people and/or institutions for providing skeletal samples: Bedford Museum (Kempston); Wessex Archaeology (West Thurrock, Purfleet); Gloucester County Archaeology Service (Ashchurch Bridge, Ashchurch); Division of Archaeological and Environmental Sciences, University of Bradford (Kingsholm; St Peter's Collegiate Church, Wolverhampton); Cotswold Archaeology (Wheatpieces, Tewkesbury); Museum of London Archaeology Services (St Benet Sherehog, London); Department of Archaeology, Durham University (Manchester Hanging Ditch); North Hertfordshire Museums Resource Centre (Baldock); University of Leicester Archaeological Services (St Peter's Church, Leicester); English Heritage (Ancaster); Lindsey Archaeological Services (Horncastle); Cambridgeshire Archaeology and Norfolk Museums and Archaeology Service (Whitefriars, Norwich); Peterborough Archaeology (Ashton); Northamptonshire Archaeology (Water Lane, Towcester); Oxfordshire Museum Service (Queensford Mill); MAP Archaeological Consultancy Ltd (St George's Crypt, Leeds); Prof. D. Rimantas Jankauskas, Department of Anatomy, Histology, and Anthropology, University of Vilnius, Lithuania (Obelia); Dr Inna Potekhina and Dr Aleksandra Kozak, Institute of Archaeology, Ukrainian Academy of Sciences, Kiev, Ukraine (Shchekavitsa; Naberezhno-Kreschatitskaya, Kiev); Dr William Devriendt, Direction de l'Archéologie Préventive, Communauté d'Agglomération du Douaisis (Saint Amé, Douai); Prof. Bernd Herrmann and Dr Birgit Großkopf, Historical Anthropology und Human Ecology, Georg-August University Göttingen (Göttingen, Anthropology Pathology Collections); Jaime Jennings (Durham University), who sampled the Auldhame skeleton that was excavated by AOC Archaeology on behalf of Historic Scotland.
Funding statement
This work was supported by NERC grants NE/E015697/1 and NE/E018564/1 awarded to C.A.R. and T.A.B. R.M. acknowledges support of a Natural Environment Research Council studentship.
References
- 1.Roberts CA, Buikstra JE. 2003. The bioarchaeology of tuberculosis. A global view on a reemerging disease. Gainesville, FL: University Press of Florida [Google Scholar]
- 2.Lönnroth K, Jaramillo E, Williams BG, Dye C, Raviglione M. 2009. Drivers of tuberculosis epidemics: the role of risk factors and social determinants. Soc. Sci. Med. 68, 2240–2246 (doi:10.1016/j.socscimed.2009.03.041) [DOI] [PubMed] [Google Scholar]
- 3.World Health Organisation 2012. Global tuberculosis report 2012. Geneva, Switzerland: World Health Organisation [Google Scholar]
- 4.Gutiérrez M, Samper S, Jiménez MS, van Embden JD, Marín JF, Martín C. 1997. Identification by spoligotyping of a caprine genotype in Mycobacterium bovis strains causing human tuberculosis . J. Clin. Microbiol. 35, 3328–3330 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Van Soolingen D, et al. 1997. A novel pathogenic taxon of the Mycobacterium tuberculosis complex, canetti: characterization of an exceptional isolate from Africa. Int. J. Syst. Bacteriol. 47, 1236–1245 (doi:10.1099/00207713-47-4-1236) [DOI] [PubMed] [Google Scholar]
- 6.Van Soolingen D, et al. 1998. Diagnosis of Mycobacterium microti infections among humans by using novel genetic markers. J. Clin. Microbiol. 36, 1840–1845 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Aranaz A, Cousins D, Mateos A, Domínguez L. 2003. Elevation of Mycobacterium tuberculosis subsp. caprae Aranaz et al. 1999 to species rank as Mycobacterium caprae comb. nov., sp. nov. Int. J. Syst. Evol. Microbiol. 53, 1785–1789 (doi:10.1099/ijs.0.02532-0) [DOI] [PubMed] [Google Scholar]
- 8.Kiers A, Klarenbeek A, Mendelts B, Van Soolingen D, Koëter G. 2008. Transmission of Mycobacterium pinnipedii to humans in a zoo with marine mammals. Int. J. Tuberc. Lung Dis. 12, 1469–1473 [PubMed] [Google Scholar]
- 9.Gagneux S. 2012. Host–pathogen coevolution in human tuberculosis . Phil. Trans. R. Soc. B 367, 850–859 (doi:10.1098/rstb.2011.0316) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Kapur V, Whittam TS, Musser JM. 1994. Is Mycobacterium tuberculosis 15,000 years old? J. Infect. Dis. 170, 1348–1349 (doi:10.1093/infdis/170.5.1348) [DOI] [PubMed] [Google Scholar]
- 11.Hughes AL, Friedman R, Murray M. 2002. Genomewide pattern of synonymous nucleotide substitution in two complete genomes of Mycobacterium tuberculosis. Emerg. Infect. Dis. 8, 1342–1346 (doi:10.3201/eid0811.020064) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Wirth T, et al. 2008. Origin, spread and demography of the Mycobacterium tuberculosis complex. PLoS Pathog. 4, e1000160 (doi:10.1371/journal.ppat.1000160) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Gutierrez MC, Brisse S, Brosch R, Fabre M, Omaïs B, Marmiesse M, Supply P, Vincent V. 2005. Ancient origin and gene mosaicism of the progenitor of Mycobacterium tuberculosis. PLoS Pathog. 1, e5 (doi:10.1371/journal.ppat.0010005) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Comas I, et al. 2013. Out-of-Africa migration and Neolithic coexpansion of Mycobacterium tuberculosis with modern humans. Nat. Genet. 45, 1176–1182 (doi:10.1038/ng.2744) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Formicola V, Milanesi Q, Scarsini C. 1987. Evidence of spinal tuberculosis at the beginning of the fourth millenium BC from Arene Candide Cave (Liguria, Italy). Am. J. Phys. Anthropol. 72, 1–6 (doi:10.1002/ajpa.1330720102) [DOI] [PubMed] [Google Scholar]
- 16.Canci A, Minozzi S, Borgognini Tarli SM. 1996. New evidence of tuberculous spondylitis from Neolithic Liguria (Italy). Int. J. Osteoarchaeol. 6, 497–501 (doi:10.1002/(SICI)1099-1212(199612)6:5<497::AID-OA291>3.0.CO;2-O) [Google Scholar]
- 17.Nicklisch N, Maixner F, Ganslmeier R, Friederich S, Dresely V, Meller H, Zink A, Alt KW. 2012. Rib lesions in skeletons from Early Neolithic sites in Central Germany: on the trail of tuberculosis at the onset of agriculture. Am. J. Phys. Anthropol. 149, 391–404 (doi:10.1002/ajpa.22137) [DOI] [PubMed] [Google Scholar]
- 18.Hershkovitz I, et al. 2008. Detection and molecular characterization of 9000-year-old Mycobacterium tuberculosis from a Neolithic settlement in the Eastern Mediterranean. PLoS ONE 3, e3426 (doi:10.1371/journal.pone.0003426) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Fletcher HA, Donoghues HD, Taylor GM, van der Zanden AGM, Spigelman M. 2003. Molecular analysis of Mycobacterium tuberculosis DNA from a family of 18th century Hungarians. Microbiology 149, 143–151 (doi:10.1099/mic.0.25961-0) [DOI] [PubMed] [Google Scholar]
- 20.Taylor GM, Goyal M, Legge AJ, Shaw RJ, Young D. 1999. Genotypic analysis of Mycobacterium tuberculosis from medieval human remains. Microbiology 145, 899–904 (doi:10.1099/13500872-145-4-899) [DOI] [PubMed] [Google Scholar]
- 21.Zink AR, Sola C, Reischl U, Grabner W, Rastogi N, Wolf H, Nerlich AG. 2004. Molecular identification and characterization of Mycobacterium tuberculosis complex in ancient Egyptian mummies. Int. J. Osteoarcheol. 14, 404–413 (doi:10.1002/oa.724) [Google Scholar]
- 22.Zink AR, Sola C, Reischl U, Grabner W, Rastogi N, Wolf H, Nerlich AG. 2003. Characterization of Mycobacterium tuberculosis complex DNAs from Egyptian mummies by spoligotyping. J. Clin. Microbiol. 41, 359–367 (doi:10.1128/JCM.41.1.359-367.2003) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Zink AR, Molnár E, Motamedi N, Pálfy G, Marcsik A, Nerlich AG. 2007. Molecular history of tuberculosis from ancient mummies and skeletons. Int. J. Osteoarchaeol. 17, 380–391 (doi:10.1002/oa.909) [Google Scholar]
- 24.Mays S, Taylor GM, Legge AJ, Young DB, Turner-Walker GT. 2001. Paleopathological and biomolecular study of tuberculosis in a medieval collection from England . Am. J. Phys. Anthropol. 114, 298–311 (doi:10.1002/ajpa.1042) [DOI] [PubMed] [Google Scholar]
- 25.Taylor GM, Young DB, Mays SA. 2005. Genotypic analysis of the earliest known prehistoric case of tuberculosis in Britain. J. Clin. Microbiol. 43, 2236–2240 (doi:10.1128/JCM.43.5.2236-2240.2005) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Mays S, Taylor GM. 2002. Osteological and biomolecular study of two possible cases of hypertrophic osteoarthropathy from Mediaeval England. J. Archaeol. Sci. 29, 1267–1276 (doi:10.1006/jasc.2001.0769) [Google Scholar]
- 27.Taylor GM, Murphy E, Hopkins R, Rutland P, Chistov Y. 2007. First report of Mycobacterium bovis DNA in human remains from the Iron Age. Microbiology 153, 1243–1249 (doi:10.1099/mic.0.2006/002154-0) [DOI] [PubMed] [Google Scholar]
- 28.Fletcher HA, Donoghue HD, Holton H, Pap I, Spigelman M. 2003. Widespread occurrence of Mycobacterium tuberculosis DNA from 18th–19th century Hungarians. Am. J. Phys. Anthropol. 120, 144–152 (doi:10.1002/ajpa.10114) [DOI] [PubMed] [Google Scholar]
- 29.Van Embden JDA, et al. 1993. Strain identification of Mycobacterium tuberculosis by DNA fingerprinting: recommendations for a standardized methodology. J. Clin. Microbiol. 31, 406–409 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Supply P, Mazars E, Lesjean S, Vincent V, Gicquel B, Locht C. 2000. Variable human minisatellite-like regions in the Mycobacterium tuberculosis genome. Mol. Microbiol. 36, 762–771 (doi:10.1046/j.1365-2958.2000.01905.x) [DOI] [PubMed] [Google Scholar]
- 31.Supply P, et al. 2006. Proposal for standardization of mycobacterial interspersed repetitive unit–variable-number tandem repeat typing of Mycobacterium tuberculosis. J. Clin. Microbiol. 44, 4498–4510 (doi:10.1128/JCM.01392-06) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Kamerbeek J, et al. 1997. Simultaneous detection and strain differentiation of Mycobacterium tuberculosis for diagnosis and epidemiology. J. Clin. Microbiol. 35, 907–914 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Brosch R, et al. 2002. A new evolutionary scenario for the Mycobacterium tuberculosis complex. Proc. Natl Acad. Sci. USA 99, 3684–3689 (doi:10.1073/pnas.052548299) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Marmiesse M, Brodin P, Buchrieser C, Gutierrez C, Simoes N, Vincent V, Glaser P, Cole ST, Brosch R. 2004. Macro-array and bioinformatic analyses reveal mycobacterial ‘core’ genes, variation in the ESAT-6 gene family and new phylogenetic markers for the Mycobacterium tuberculosis complex. Microbiology 150, 483–496 (doi:10.1099/mic.0.26662-0) [DOI] [PubMed] [Google Scholar]
- 35.Gagneux S, et al. 2006. Variable host-pathogen compatibility in Mycobacterium tuberculosis. Proc. Natl Acad. Sci. USA 103, 2869–2873 (doi:10.1073/pnas.0511240103) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Sreevatsan S, Pan X, Stockbauer KE, Connell ND, Kreiswirth BN, Whittam TS, Musser JM. 1997. Restricted structural gene polymorphism in the Mycobacterium tuberculosis complex indicates evolutionarily recent global dissemination. Proc. Natl Acad. Sci. USA 94, 9869–9874 (doi:10.1073/pnas.94.18.9869) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Gutacker MM, et al. 2002. Genome-wide analysis of synonymous single nucleotide polymorphisms in Mycobacterium tuberculosis complex organisms: resolution of genetic relationships among closely related microbial strains. Genetics 162, 1533–1543 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Baker L, Brown T, Maiden MC, Drobniewski F. 2004. Silent nucleotide polymorphisms and a phylogeny for Mycobacterium tuberculosis. Emerg. Infect. Dis. 10, 1568–1577 (doi:10.3201/eid1009.040046) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Filliol I, et al. 2006. Global phylogeny of Mycobacterium tuberculosis based on single nucleotide polymorphism (SNP) analysis: insights into tuberculosis evolution, phylogenetic accuracy of other DNA fingerprinting systems, and recommendations for a minimal standard SNP set. J. Bacteriol. 188, 759–772 (doi:10.1128/JB.188.2.759-772.2006) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Warren RM, Streicher EM, Sampson SL, van der Spuy GD, Richardson M, Nguyen D, Behr MA, Victor TC, van Helden PD. 2002. Microevolution of the direct repeat region of Mycobacterium tuberculosis: implications for interpretation of spoligotyping data. J. Clin. Microbiol. 40, 4457–4465 (doi:10.1128/JCM.40.12.4457-4465.2002) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Comas I, Homolka S, Niemann S, Gagneux S. 2009. Genotyping of genetically monomorphic bacteria: DNA sequencing in Mycobacterium tuberculosis highlights the limitations of current methodologies . PloS ONE 4, e7815 (doi:10.1371/journal.pone.0007815) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Wilbur AK, Bouwman AS, Stone AC, Roberts CA, Pfister L-A, Buikstra JE, Brown TA. 2009. Deficiencies and challenges in the study of ancient tuberculosis DNA. J. Archaeol. Sci. 36, 1990–1997 (doi:10.1016/j.jas.2009.05.020) [Google Scholar]
- 43.Faksri K, Drobniewski F, Nikolayevskyy V, Brown T, Prammananan T, Palittapongarnpim P, Prayoonwiwat N, Chaiprasert A. 2011. Genetic diversity of the Mycobacterium tuberculosis Beijing family based on IS6110, SNP, LSP and VNTR profiles from Thailand. Infect. Genet. Evol. 11, 1142–1149 (doi:10.1016/j.meegid.2011.04.007) [DOI] [PubMed] [Google Scholar]
- 44.Nakanishi N, Wada T, Arikawa K, Millet J, Rastogi N, Iwamoto T. 2013. Evolutionary robust SNPs reveal the misclassification of Mycobacterium tuberculosis Beijing family strains into sublineages. Infect. Genet. Evol. 16, 174–177 (doi:10.1016/j.meegid.2013.02.007) [DOI] [PubMed] [Google Scholar]
- 45.Stucki D, Gagneux S. 2012. Single nucleotide polymorphisms in Mycobacterium tuberculosis and the need for a curated database. Tuberculosis 93, 30–39 (doi:10.1016/j.tube.2012.11.002) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Hershberg R, et al. 2008. High functional diversity in Mycobacterium tuberculosis driven by genetic drift and human demography . PloS Biol. 6, e311 (doi:10.1371/journal.pbio.0060311) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Comas I, Chakravartti J, Small PM, Galagan J, Niemann S, Kremer K, Ernst JD, Gagneux S. 2010. Human T cell epitopes of Mycobacterium tuberculosis are evolutionarily hyperconserved. Nat. Genet. 42, 498–503 (doi:10.1038/ng.590) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Bos KI, et al. 2011. A draft genome of Yersinia pestis from victims of the Black Death. Nature 478, 506–510 (doi:10.1038/nature10549) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Bouwman AS, Kennedy SL, Müller R, Stephens RH, Holst M, Caffell AC, Roberts CA, Brown TA. 2012. Genotype of a historic strain of Mycobacterium tuberculosis . Proc. Natl Acad. Sci. USA 109, 18 511–18 516 (doi:10.1073/pnas.1209444109) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Schuenemann VJ, et al. 2013. Genome-wide comparison of medieval and modern Mycobacterium leprae. Science 341, 179–183 (doi:10.1126/science.1238286) [DOI] [PubMed] [Google Scholar]
- 51.Abadia E, et al. 2010. Resolving lineage assignation on Mycobacterium tuberculosis clinical isolates classified by spoligotyping with a new high-throughput 3R SNPs based method. Infect. Genet. Evol. 10, 1066–1074 (doi:10.1016/j.meegid.2010.07.006) [DOI] [PubMed] [Google Scholar]
- 52.Müller R, Roberts CA, Brown TA. 2013. Biomolecular identification of ancient Mycobacterium tuberculosis complex DNA in human remains from Britain and Continental Europe. Am. J. Phys. Anthropol. 153, 178–189 (doi:10.1002/ajpa.22417) [DOI] [PubMed] [Google Scholar]
- 53.Bouwman AS, Brown TA. 2005. The limits of biomolecular palaeopathology: ancient DNA cannot be used to study venereal syphilis. J. Archaeol. Sci. 32, 703–713 (doi:10.1016/j.jas.2004.11.014) [Google Scholar]
- 54.Rohland N, Hofreiter M. 2007. Ancient DNA extraction from bones and teeth. Nat. Prot. 2, 1756–1762 (doi:10.1038/nprot.2007.247) [DOI] [PubMed] [Google Scholar]
- 55.Rohland N, Siedel H, Hofreiter M. 2010. A rapid column-based ancient DNA extraction method for increased sample throughput. Mol. Ecol. Resour. 10, 677–683 (doi:10.1111/j.1755-0998.2009.02824.x) [DOI] [PubMed] [Google Scholar]
- 56.Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. 2011. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (doi:10.1093/molbev/msr121) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Huson DH, Scornavacca C. 2012. Dendroscope 3: an interactive viewer for rooted phylogenetic trees and networks. Syst. Biol. 61, 1061–1067 (doi:10.1093/sysbio/sys062) [DOI] [PubMed] [Google Scholar]
- 58.Gilbert MT, Hansen AJ, Willerslev E, Rudbeck L, Barnes I, Lynnerup N, Cooper A. 2003. Characterization of genetic miscoding lesions caused by postmortem damage. Am. J. Hum. Genet. 72, 48–61 (doi:10.1086/345379) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Gilbert MTP, Binladen J, Miller W, Wiuf C, Willerslev E, Poinar H, Carlson JE, Leebens-Mack JH, Schuster SC. 2007. Recharacterization of ancient DNA miscoding lesions: insights in the era of sequencing-by-synthesis. Nucleic Acids Res. 35, 1–10 (doi:10.1093/nar/gkl483) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Brotherton P, Endicott P, Sanchez JJ, Beaumont M, Barnett R, Austin J, Cooper A. 2007. Novel high-resolution characterization of ancient DNA reveals C>U-type base modification events as the sole cause of post mortem miscoding lesions. Nucleic Acids Res. 35, 5717–5728 (doi:10.1093/nar/gkm588) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Chiang C-Y, Riley LW. 2005. Exogenous reinfection in tuberculosis. Lancet Infect. Dis. 5, 629–636 (doi:10.1016/S1473-3099(05)70240-1) [DOI] [PubMed] [Google Scholar]
- 62.Steenken W, Jr, Gardner LU. 1946. History of H37 strain of tubercle bacillus . Am. Rev. Tuberc. 54, 62–66 [DOI] [PubMed] [Google Scholar]
- 63.Ilina EN, et al. 2013. Comparative genomic analysis of Mycobacterium tuberculosis drug resistant strains from Russia. PLoS ONE 8, e56577 (doi:10.1371/journal.pone.0056577) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Cardy A. 1997. The environmental material: the human bones. In Whithorn and St Ninian. The excavation of a monastic town (ed. Hill P.), pp. 519–562 Stroud, UK: Sutton Publishing [Google Scholar]
- 65.Barrell ADM. 2000. Medieval Scotland, pp. 5–10 Cambridge, UK: Cambridge University Press [Google Scholar]
- 66.Lamb AL, Melikian M, Ives R, Evans J. 2012. Multi-isotope analysis of the population of the lost medieval village of Auldhame, East Lothian, Scotland. J. Anal. At. Spectrom. 27, 765–777 (doi:10.1039/C2JA10363J) [Google Scholar]
- 67.Bennike P. 1985. Palaeopathology of Danish skeletons: a comparative study of demography, disease and injury, pp. 183–194 Copenhagen, Denmark: Akademisk Forlag [Google Scholar]
- 68.Bennike P. 1999. Facts or myths? A re-evaluation of cases of diagnosed tuberculosis in the past in Denmark. In Tuberculosis past and present (eds Pálfi G, Dutour O, Deák J, Hutás I.), pp. 561–573 Budapest, Hungary: Golden Book Publisher [Google Scholar]
- 69.Kjellström A. 2012. Possible cases of leprosy and tuberculosis in medieval Sigtuna, Sweden. Int. J. Osteoarchaeol. 22, 261–283 (doi:10.1002/oa.1204) [Google Scholar]