Abstract
Antimicrobial susceptibility data on Escherichia coli F4, Pasteurella multocida, and Streptococcus suis isolates from Ontario swine (January 1998 to October 2010) were acquired from a comprehensive diagnostic veterinary laboratory in Ontario, Canada. In relation to the possible development of a surveillance system for antimicrobial resistance, data were assessed for ease of management, completeness, consistency, and applicability for temporal and spatial statistical analyses. Limited farm location data precluded spatial analyses and missing demographic data limited their use as predictors within multivariable statistical models. Changes in the standard panel of antimicrobials used for susceptibility testing reduced the number of antimicrobials available for temporal analyses. Data consistency and quality could improve over time in this and similar diagnostic laboratory settings by encouraging complete reporting with sample submission and by modifying database systems to limit free-text data entry. These changes could make more statistical methods available for disease surveillance and cluster detection.
Résumé
Sensibilité antimicrobienne des isolats d’Escherichia coli F4, de Pasteurella multocida et de Streptococcus suis transmise par un laboratoire de diagnostic vétérinaire et recommandations pour un système de surveillance. Les données de sensibilité antimicrobienne sur les isolats d’Escherichia coli F4, de Pasteurella multocida et de Streptococcus suis provenant des porcs de l’Ontario (de janvier 1998 à octobre 2010) ont été acquises auprès d’un laboratoire de diagnostic vétérinaire complet situé en Ontario, au Canada. En relation avec la création éventuelle d’un système de surveillance pour l’antibiorésistance, des données ont été évaluées pour déterminer la facilité de gestion, l’intégralité, la cohérence et l’applicabilité des analyses temporelles et spatiales. Des données limitées sur l’emplacement de la ferme empêchaient des analyses spatiales et des données démographiques manquantes limitaient leur utilisation comme prédicteurs au sein de modèles statistiques multivariables. Les changements du groupe standard d’antimicrobiens utilisés pour les tests de sensibilité ont réduit le nombre d’antimicrobiens disponibles pour des analyses temporelles. La cohérence et la qualité des données pourraient être améliorées au fil du temps dans ce laboratoire de diagnostic et d’autres installations semblables en encourageant la production de rapports complets avec la soumission d’échantillons et en modifiant les systèmes des bases de données afin de limiter l’entrée de données en forme libre. Ces changements pourraient rendre d’autres méthodes statistiques disponibles pour la surveillance des maladies et la détection de grappes.
(Traduit par Isabelle Vallières)
Introduction
Surveillance of antimicrobial resistance (AMR) in bacterial pathogens is becoming increasingly important in the face of rising resistance in many pathogens and reduced availability of effective antimicrobial drugs. Surveillance of AMR may include detection of temporal trends and identification of opportunities for interventions to mitigate dissemination and increases in the prevalence of resistance. These interventions may include efforts to reduce the use of specific antimicrobials, and/or replacing the use of drugs of concern with alternatives showing high in-vitro or in-vivo effectiveness. Furthermore, AMR surveillance provides information about the prevalence of resistance in relevant pathogens, which may be useful to practitioners making treatment decisions for animals affected by these pathogens.
There are AMR surveillance programs in several countries that focus on human enteric pathogens and foodborne commensal bacteria from various sources (1–6). These systems have emerged in response to increasing human health concerns about antimicrobial use in the agri-food sector and its possible impacts on resistance in zoonotic and enteric pathogens of humans. Other AMR surveillance systems focus on non-enteric (e.g., nosocomial, respiratory, and sexually transmitted) human pathogens (7–9). However, programs that focus on AMR in target pathogens of animals are rare. This is an important gap in AMR surveillance, as bacterial species that routinely affect animals clinically have developed resistance (10). Resistant animal pathogens are potential reservoirs of resistance genes for human pathogens, can serve as indicators of antimicrobial use in a population, and may be drivers of antimicrobial use as resistance to 1 drug begets increased use of second and third line options.
The need for surveillance of AMR in animal pathogens was expressed by the European Parliament in 2011 (11). Earlier, the Advisory Committee on Animal Uses of Antimicrobials and Impact on Resistance and Human Health included the design and implementation of AMR surveillance in food animal production as a key recommendation in their 2009 report (12). A surveillance system for AMR in isolates associated with clinical disease in food-animals would help to identify whether resistance is a problem in important animal pathogens, trends in resistance prevalence, and potentially provide veterinarians and farmers with information they can use to increase the responsible use of antimicrobials in farm animal production.
Surveillance systems may be active or passive (13). In the former, samples are obtained for the express purpose of surveillance; the surveillance team actively seeks out samples and the required supporting data. In contrast, a passive system obtains data on cases or submissions indirectly from sources such as laboratories or practitioners upon diagnosis or at regular intervals, or from databases storing information for other purposes (13). Passively collected data are typically limited to the information collected for their original purpose such as clinical diagnosis. Passive methods can require less labor and financial input to implement and maintain than active methods and therefore may be favored over active methods when a suitable passive data source is available. However, passively collected data may have more missing epidemiological information than that collected actively, may not be available in a format appropriate for surveillance analysis, and may be limited in the availability of epidemiological information by the data requirements of the original user. Therefore, careful scrutiny of the quality and applicability of the available passive data is warranted before the data are used for a surveillance system.
The Animal Health Laboratory (AHL) of the University of Guelph is a source of passive data that may be suitable for the development of a surveillance system for AMR in agricultural animal pathogens in Ontario.
The objective of this research was to assess the quantity, consistency, and quality of antimicrobial susceptibility data available from selected swine pathogens isolated at the AHL and to evaluate their potential for analyses in a semi-automated surveillance system. The pathogens included were: Escherichia coli F4 (previously K88), Streptococcus suis, and Pasteurella multocida. These pathogens were chosen through a pre-study review of available data. They were the 3 most frequently isolated bacterial pathogens from swine in the AHL data, and monthly isolation counts for these pathogens remained > 0 throughout the study period. Furthermore, these pathogens are important in Ontario swine production as causes of post-weaning diarrhea, neurological and respiratory infections, and secondary bacterial respiratory infections following viral infection, respectively (14). The data characteristics assessed for these pathogens included ease of data management, completeness of reporting, and availability of consistent antimicrobial susceptibility testing over time, with respect to statistical analyses such as cluster detection or multivariable logistic regression.
Materials and methods
Data detailing sample submissions, resulting isolations, and susceptibility tests were obtained from the AHL from January 1998 to October 2010 for E. coli F4, S. suis, and P. multocida. Isolation and antimicrobial susceptibility testing were performed according to the standard operating procedures of the AHL. All data pertaining to isolates were requested, including location and name for the clinic and farm submitting the sample, case demographic information, and all antimicrobial susceptibility results. During the study period, the AHL used one laboratory information system (LIMS) from January 1998 to May 2007 [Veterinary Animal Disease Diagnostic System (VADDS); Advanced Technology Corp., Ramsey, New Jersey, USA] and another LIMS from May 2007 to October 2010, which continues to be used in 2013 (Sapphire; LabVantage Solutions Inc., Bridgewater, New Jersey, USA). Data fields to be assessed were determined from the AHL swine submission form: case type (diagnostic, research, or monitoring), number of animals at risk, number sick, number dead, weight and age of animal, submitting clinic telephone number, name, and postal code; and owner contact information including farm/owner name, unique identification number, and postal code (15). A confidentiality agreement was signed regarding farm and veterinary clinic information such that individuals are not identified in results, and that data will be destroyed upon project completion.
To avoid potential biases, data from ongoing research programs and duplicate samples were excluded before any assessments or analyses were performed. Data pertaining to isolates not subjected to susceptibility testing were also removed. When the numbers of animals at risk, sick, and dead were all reported as zero, we assumed that the data were missing. Furthermore, when number at risk was not reported and number sick and number dead were reported as zero, we assumed that the number sick and number dead values were missing. Recording consistency for the new LIMS data was compared with all data using the Z-test for proportions. Similarly, recording consistencies for the top 4, 8, and 10 submitting practices were compared with the recording consistency for all data using the Z-test for proportions.
As it was not deemed feasible to assess susceptibility to every tested antimicrobial for each pathogen, an a priori decision was made to identify suitable pathogen/antimicrobial combinations for surveillance; antimicrobials were identified as suitable for continued surveillance for each pathogen if they were on the 2010 testing panel, displayed ≥ 2 years of historical data, and ≥ 1 susceptibility test was performed per month on average.
Potential sentinel practices were identified by enumerating the number of observations acquired from each practice after cleaning the veterinary practice field. Three groups of potential sentinel practices were identified; those with > 400, > 200, and > 150 observations, which reflect major groupings in the practice list.
All data merging, cleaning, protocol development, graphical methods, and analyses were performed using Stata/MP 11.0 (StataCorp 2009, College Station, Texas, USA).
Results
Ease of data management
For both LIMS, demographic and isolate information was provided in separate datasets for all pathogens. A third dataset was acquired detailing E. coli F4 results, as these results could not be extracted with the isolate information. Each case and sample in these datasets was provided with a submission identification number unique to the farm and date of submission, but not to the isolate. As such, multiple isolates from a single case may have shared submission identification numbers. Demographic variables (e.g., farm and practice data) were merged to the isolate data by the submission identification number. A single dataset was developed by merging all common data fields from the 2 LIMSs, and a semi-automated protocol was developed for future merging. Similarly, a semi-automated protocol was developed to append new data from the current LIMS on a prospective basis.
Semi-automated protocols for identifying the specific E. coli isolates positive for the F4 antigen could not be used because serotyping information was linked to the submission identification without an isolate-specific identifier. Therefore, in cases in which multiple E. coli isolates were obtained from a single submission, it was not possible to identify the specific isolate (and associated susceptibility results) that was tested for F4 reactivity. In these instances, the raw diagnostic laboratory reports for the submissions were reviewed to determine the isolate of interest.
Due to free-text entry of veterinary clinic name, and farm/owner name data fields, a considerable amount of cleaning was required to ensure that all data were in the same format. For example, variations of veterinary clinic name could appear due to spelling or abridgement inconsistencies. Similarly, the recording of antimicrobial susceptibilities as uppercase or lowercase text was variable from the VADDS LIMS, which disrupts statistical analyses. In order to address these issues, an automated protocol was developed to ensure that uppercase text was used for all susceptibility recordings. This approach was not sufficient for editing the clinic or farm/owner names, however, as there is a potentially infinite number of combinations of recording inconsistencies in these fields. To address this issue, changes were recorded manually and added to an automatic protocol to correct these errors in subsequent runs; however, as isolates are entered into the LIMS and accessed for surveillance, unique recording inconsistencies could be added to the database at any time.
Consistency of recording
Following merging of all data into a common working database, they were assessed for consistency of recording and missing values. The recording consistency of each variable is displayed in Table 1 for all data (both LIMSs), and for the current LIMS alone. Data regarding the veterinary clinic was more consistently available in the new LIMS compared with all data. The clinic identification number was recorded at a significantly higher rate (P < 0.001) in the new system than all data. However, the identification number was not consistent within a clinic itself; multiple identification numbers were associated with each clinic, both within and between LIMSs. Moreover, farm demographic and herd health information was recorded in a significantly lower proportion of data from the current system than in all the data (P < 0.001) (Table 1).
Table 1.
Recording consistency (percentage of submissions with values recorded) for data fields from clinical swine submissions to the Animal Health Laboratory, displayed for all data (January 1998 to October 2010) and data from the current LIMS (May 2007 to October 2010)
| Escherichia coli F4 | Streptococcus suis | Pasteurella multocida | ||||
|---|---|---|---|---|---|---|
|
|
|
|
||||
| Data field | All data | Current LIMS | All data | Current LIMS | All data | Current LIMS |
| Submission number | 100 | 100 | 100 | 100 | 100 | 100 |
| Submission date | 100 | 100 | 100 | 100 | 100 | 100 |
| Clinic name | 100 | 100 | 100 | 100 | 100 | 100 |
| Clinic postal code | 100 | 100 | 99.96 | 100 | 99.93 | 100 |
| Clinic number | 77.40 | 100a | 15.81 | 100a | 74.45 | 100a |
| Case type | 99.92 | 100 | 98.94 | 100a | 98.09 | 100 |
| Breed | 78.91 | 69.96 | 79.95 | 72.21b | 81.28 | 77.95 |
| Owner or farm name | 15.95 | 80.23a | 12.79 | 80.89a | 10.66 | 80.00a |
| Owner unique id | 57.52 | 2.66b | 57.71 | 1.49b | 61.34 | 1.54b |
| Owner postal code | 54.35 | 0b | 54.57 | 0b | 57.31 | 0b |
| Number at risk | 63.30 | 37.71b | 59.51 | 42.93b | 61.61 | 45.96b |
| Number sick | 44.62 | 26.27b | 41.52 | 29.78b | 42.26 | 30.81b |
| Number dead | 44.62 | 26.27b | 41.52 | 29.78b | 42.50 | 32.32b |
Recording consistency significantly higher from the new system than all data, P < 0.001 by Z-test.
Recording consistency significantly lower from the new system than all data, P < 0.001 by Z-test.
Completeness of recording (based upon the sample submission form) was further assessed by veterinary clinic to determine whether data from sentinel practices may be better suited for use in a surveillance system than all available data. Approximately 150 practices were represented within the dataset; however, a small number of practices accounted for a disproportionate amount of data within the system. Ten practices had > 150 observations (> 71% of the total), 8 practices had > 200 observations (~65%), and 4 practices had > 400 observations (~45%). Recording consistency for the top 4, 8, and 10 submitting practices did not differ significantly from the recording consistency for all variables (P > 0.5; Table 2).
Table 2.
Percent of isolates, maximum distance between practices, and demographic variable recording consistency for the highest 4, 8, and 10 submitting practices within swine antimicrobial susceptibility testing data from the Animal Health Laboratory (January 1998 to October 2010) compared with the overall data
| Top 4 | Top 8 | Top 10 | Overall | |
|---|---|---|---|---|
| Percent of isolates | 44.94 | 65.01 | 71.38 | 100 |
| Approximate maximum distance between practices (km) | 57.5 | 196.0 | 196.0 | 675.0 |
| Recording consistency (%)a: | ||||
| Owner or farm name | 12.80 | 13.75 | 12.94 | 12.99 |
| Unique identification number | 54.59 | 55.90 | 57.71 | 58.66 |
| Number sick | 37.36 | 35.66 | 33.71 | 35.59 |
| Number dead | 37.53 | 35.72 | 33.79 | 35.64 |
| Number at risk | 52.21 | 52.26 | 59.57 | 51.12 |
No significant differences in the recording consistency between the top 4, 8, or 10 submitting practices and all data.
Consistency of susceptibility testing over time
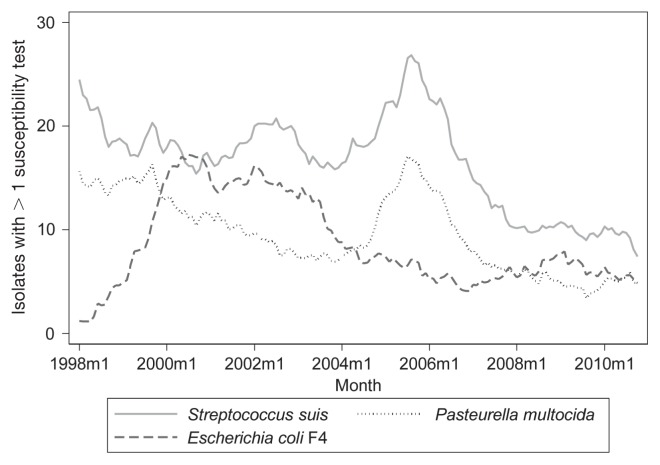
Over the study time frame, there were 1323 E. coli F4, 2549 S. suis, and 1464 P. multocida isolates with ≥ 1 antimicrobial susceptibility test result. Among the pathogens, the number of isolates with susceptibility results was variable over time (Figure 1); S. suis isolates had the highest number of susceptibility tests performed each year, while E. coli F4 and P. multocida varied in their relative ranking. From 2008 on, more susceptibility tests were performed on E. coli F4 isolates than P. multocida (Figure 1). Among all 3 pathogens, the monthly number of isolates tested stabilized during the 2008 to 2010 time period at approximately 8 to 10 isolates per pathogen per month.
Figure 1.
Smootheda number of isolates per month that underwent ≥ 1 antimicrobial susceptibility test at the Animal Health Laboratory from January 1998 to October 2010.
a Smoothing was performed by using the current observation, 6 lagged, and 5 forward observations.
Twenty-eight antimicrobials were used for susceptibility testing over the study period. The numbers of isolates tested for susceptibility to each antimicrobial per year are displayed in Tables 3a–c. Changes in the most commonly used susceptibility panel occurred with the introduction or withdrawal of drugs over time (details available from the author).
Table 3a.
Number of susceptibility tests performed on Escherichia coli F4 isolates at the Animal Health Laboratory by year (January 1998 to October 2010)
| 1998 | 1999 | 2000 | 2001 | 2002 | 2003 | 2004 | 2005 | 2006 | 2007 | 2008 | 2009 | 2010 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Ampicillina | 27 | 91 | 207 | 176 | 170 | 153 | 81 | 79 | 60 | 64 | 72 | 80 | 60 |
| Apramycina | — | — | — | — | — | — | — | — | — | 51 | 72 | 79 | 56 |
| Ceftiofura | 2 | — | 2 | 9 | 41 | 128 | 80 | 79 | 59 | 64 | 72 | 80 | 60 |
| Cephalothin | 27 | 92 | 204 | 173 | 165 | 143 | 3 | — | — | — | — | — | — |
| Gentamicina | 27 | 92 | 207 | 179 | 170 | 153 | 79 | 79 | 60 | 64 | 72 | 79 | 57 |
| Kanamycina | — | — | — | — | — | — | — | 8 | 59 | 64 | 72 | 79 | 60 |
| Neomycin | 27 | 91 | 205 | 176 | 169 | 153 | 61 | 61 | — | — | — | — | — |
| Spectinomycina | 25 | 88 | 207 | 177 | 169 | 153 | 74 | 71 | 52 | 64 | 72 | 80 | 60 |
| Sulfisoxazolea | 26 | 87 | 203 | 173 | 164 | 142 | 80 | 79 | 60 | 64 | 70 | 80 | 60 |
| Tetracyclinea | 27 | 92 | 207 | 177 | 170 | 153 | 81 | 79 | 60 | 64 | 72 | 80 | 60 |
| Tobramycin | 25 | 87 | 95 | — | — | — | — | — | — | — | — | — | — |
| Trimethoprim/ sulfamethoxazolea | 27 | 92 | 207 | 177 | 170 | 153 | 81 | 79 | 60 | 64 | 72 | 80 | 60 |
Chosen for inclusion in a potential AMR surveillance system, — = no data available.
Table 3c.
Number of susceptibility tests performed on Streptococcus suis isolates at the Animal Health Laboratory by year (January 1998 to October 2010)
| 1998 | 1999 | 2000 | 2001 | 2002 | 2003 | 2004 | 2005 | 2006 | 2007 | 2008 | 2009 | 2010 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Amikacin | 175 | 168 | 69 | — | — | — | — | — | — | — | — | — | — |
| Ampicillina | 249 | 226 | 194 | 204 | 249 | 194 | 216 | 319 | 219 | 149 | 119 | 112 | 97 |
| Ceftiofura | 184 | 203 | 193 | 196 | 247 | 194 | 211 | 318 | 219 | 149 | 118 | 112 | 97 |
| Cephalothin | 249 | 227 | 82 | 2 | 1 | 1 | — | — | — | — | — | — | — |
| Clindamycin | 246 | 222 | 195 | 199 | 248 | 193 | 43 | — | — | — | — | — | — |
| Erythromycin | 249 | 227 | 82 | 4 | — | — | — | — | — | — | — | — | — |
| Florfenicol | 148 | 196 | 77 | — | — | — | — | — | — | — | — | 1 | — |
| Gentamicin | 246 | 227 | 195 | 203 | 249 | 194 | 216 | 319 | 219 | 66 | 1 | — | — |
| Kanamycin | — | — | — | — | — | — | — | 88 | 218 | 66 | 1 | — | — |
| Neomycin | 247 | 225 | 194 | 200 | 244 | 194 | 216 | 230 | — | — | — | — | — |
| Oxacillin | 243 | 221 | 84 | 3 | — | — | — | — | — | — | — | — | — |
| Penicillin Ga | 249 | 226 | 194 | 204 | 248 | 193 | 207 | 315 | 217 | 148 | 117 | 112 | 97 |
| Spectinomycina | 151 | 168 | 179 | 201 | 219 | 194 | 216 | 319 | 217 | 149 | 118 | 111 | 97 |
| Sulfisoxazolea | 4 | 2 | 3 | 2 | — | — | 183 | 319 | 219 | 149 | 112 | 112 | 95 |
| Tetracyclinea | 241 | 225 | 195 | 203 | 249 | 194 | 216 | 319 | 219 | 149 | 118 | 112 | 97 |
| Tiamulina | 146 | 163 | 174 | 29 | — | 2 | 187 | 319 | 131 | 146 | 117 | 112 | 97 |
| Tilmicosin | 147 | 163 | 163 | 104 | 233 | 193 | 215 | 319 | 218 | 68 | — | — | — |
| Trimethoprim/sulfamethoxazolea | 248 | 226 | 195 | 202 | 249 | 194 | 215 | 318 | 219 | 149 | 118 | 112 | 97 |
| Tylosin | 146 | 163 | 66 | — | — | — | — | — | — | — | — | — | — |
Chosen for inclusion in a potential AMR surveillance system, — = no data available.
The antimicrobials deemed suitable for surveillance in E. coli F4 were: ampicillin, apramycin, ceftiofur, gentamicin, kanamycin, spectinomycin, sulfisoxazole, tetracycline, and trimethoprim/sulfa (TMS) (Table 3a). The antimicrobials deemed suitable for surveillance in P. multocida were: ampicillin, ceftiofur, florfenicol, penicillin G, spectinomycin, sulfisoxazole, tetracycline, tiamulin, TMS, and tulathromycin (Table 3b). The antimicrobials deemed suitable for surveillance in S. suis were: ampicillin, ceftiofur, penicillin G, spectinomycin, sulfisoxazole, tetracycline, tiamulin, and TMS (Table 3c).
Table 3b.
Number of susceptibility tests performed on Pasteurella multocida isolates at the Animal Health Laboratory by year (January 1998 to October 2010)
| 1998 | 1999 | 2000 | 2001 | 2002 | 2003 | 2004 | 2005 | 2006 | 2007 | 2008 | 2009 | 2010 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Amikacin | 130 | 124 | 39 | — | — | — | — | — | — | — | — | — | — |
| Ampicillina | 174 | 183 | 134 | 122 | 102 | 89 | 99 | 205 | 126 | 79 | 48 | 53 | 49 |
| Ceftiofura | 142 | 172 | 132 | 120 | 101 | 88 | 96 | 205 | 126 | 79 | 49 | 53 | 49 |
| Cephalothin | 172 | 183 | 58 | — | — | — | — | — | — | — | — | — | — |
| Clindamycin | 103 | 127 | 116 | 122 | 102 | 89 | 19 | — | — | — | — | — | — |
| Erythromycin | 168 | 181 | 57 | 1 | — | — | — | — | 1 | — | — | — | — |
| Florfenicola | 106 | 168 | 48 | — | — | — | — | — | — | 43 | 49 | 51 | 49 |
| Gentamicin | 171 | 183 | 134 | 122 | 102 | 89 | 99 | 205 | 126 | 31 | — | — | — |
| Kanamycin | — | — | — | — | — | — | — | 51 | 125 | 31 | — | — | — |
| Neomycin | 113 | 131 | 116 | 121 | 102 | 89 | 99 | 154 | — | — | — | — | — |
| Oxacillin | 101 | 127 | 40 | 2 | — | — | — | — | — | — | — | — | — |
| Penicillin Ga | 172 | 183 | 131 | 122 | 101 | 88 | 78 | 42 | 10 | 38 | 39 | 44 | 49 |
| Spectinomycina | 102 | 124 | 116 | 121 | 100 | 89 | 99 | 205 | 126 | 79 | 49 | 53 | 49 |
| Sulfisoxazolea | 2 | 2 | — | — | — | — | 80 | 205 | 126 | 79 | 47 | 53 | 48 |
| Tetracyclinea | 174 | 183 | 134 | 122 | 102 | 89 | 99 | 205 | 126 | 79 | 49 | 53 | 49 |
| Tiamulina | 98 | 120 | 112 | 16 | — | — | 82 | 205 | 68 | 77 | 49 | 53 | 49 |
| Tilmicosin | 100 | 123 | 100 | 72 | 94 | 89 | 99 | 205 | 125 | 32 | — | — | — |
| Trimethoprim/ sulfamethoxazolea | 172 | 182 | 134 | 122 | 102 | 88 | 99 | 204 | 126 | 79 | 49 | 53 | 49 |
| Tulathromycina | — | — | — | — | — | — | — | — | — | 43 | 49 | 53 | 49 |
Chosen for inclusion in a potential AMR surveillance system, — = no data available.
Discussion
Antimicrobial resistance data for selected swine pathogens isolated at the AHL at the University of Guelph have potential for use in a prospective surveillance system. It is believed that the AHL handles a larger portion of the clinical submissions from Ontario livestock than other laboratories. Therefore, data from the AHL are expected to be fairly representative of those requested by veterinarians serving the Ontario livestock industry as a whole. However, swine submissions received at the AHL are affected by factors other than disease incidence, including the presence of disease outbreaks and economic factors such as the value of the Canadian dollar and auction prices (16,17). Furthermore, this is a tertiary data source, and as such, is inherently subject to reporting bias. Although the data have features, such as missing covariate data and some recording inconsistency that limit analyses to temporal options, knowledge of the prevalence of resistance in isolates from submitted clinical samples may be informative to veterinary practitioners facing decisions for treatment of clinically ill animals.
The availability of representative data, the high training and expertise of AHL personnel and ease of data acquisition make a passive surveillance system for antimicrobial resistance in Ontario livestock using AHL data an attractive option. Through automated protocols for data analysis and online reporting of results, an efficient system could be developed to provide useful information for veterinary practitioners in the province. Semi-automated protocols have been generated for S. suis and P. multocida isolates, and could be implemented into a surveillance program with current AHL protocols. Manual evaluation of the E. coli diagnostic reports was an extensive task, however. It was confirmed with laboratory personnel that susceptibility reports for E. coli are primarily made for the F4-positive isolates, unless otherwise requested by the referring veterinarian. Therefore, with a small decrease in the specificity of F4 results, F4 E. coli data are also suitable for semi-automated AMR surveillance using the AHL data. However, isolate specific identifier data are suggested for future LIMS development.
Missing data were commonly encountered, reflecting incomplete form submission by the veterinarian. Missing farm location limits the application of spatial statistical methods to the level of the veterinary clinic. Veterinary clinic level data are not the ideal resolution for swine data in Ontario, as most swine herds are served by a limited number of veterinarians who cover large territories. Amezcua et al (18) found that 7 Ontario practitioners visited 23.6% of Ontario swine herds in 2006, and that these farms were located in 4 of the 5 Ontario agricultural regions (all but Northern Ontario, which was estimated to represent only 0.5% of all herds in 2006). For this reason, a clinic may serve herds separated by large distances (> 100 km), which makes clinic location an epidemiologically unimportant and potentially spurious predictor variable in a statistical model. Therefore, in the current state of data management and collection, temporal analyses are more appropriate for these AMR data than spatial or temporal-spatial analyses. For example, cluster detection methods may be used to indicate periods in time when the proportion of resistance is statistically higher (or lower) than expected, while logistic regression models with year and season as predictor variables or the application of time-series modeling may allow the visualization of statistically significant trends. The consistency of recording of herd health information (herd size, number sick, number dead) was also found to be lacking, limiting the use of these factors as predictors or offset values within statistical models. Missing information for these potential covariates (e.g., production system or age of animal) may be an issue when considering the confounding effects of uncontrolled variables (19), and changes in background populations may result in biased P-values if an incorrect assumption of stable background populations is made (20). As infectious agents and their resistance profiles may be particularly influenced by demographic factors such as animal age/stage of production, the statistical models produced with these data may be limited in their predictive abilities without these parameters. The availability of these predictors would allow surveillance personnel to determine whether an increase in resistance is true in the general swine population, or if it reflects an increase in submissions from (or growth within) a given level of the production process compared with other production levels. Stage of production information should be added to the submission form and veterinarians and producers submitting samples should be encouraged to be diligent in providing complete information in order to develop a complete database and support the use of AHL data for surveillance purposes. Emphasis must be placed on completeness of recording of data on intake forms, which may include continuing education for submitting veterinarians/owners about the importance of surveillance data and how they may benefit from these results. Alternatively, as AHL livestock and poultry testing are subsidized, it may be possible to make the preferential food animal pricing contingent on completion of submission forms with the required demographic data. Furthermore, simply informing practitioners of the data requirements may influence response as a form of stewardship to the industry (21), that is, practitioners accessing data may be more likely to submit samples with the knowledge that results may be improved with more samples.
Observations from high-submitting practices were evaluated in comparison to the average in regards to recording consistency, as these practices could potentially be used as sentinels for the province. Although there were no significant differences in completeness of records for these most frequently submitting practices compared with all practices, previous research has shown that Ontario swine clinics were willing to participate/comply with pilot testing of surveillance initiatives (18). Therefore, consideration should be given to requesting that these clinics provide all submission details on a prospective basis, providing an avenue for increasing data quality and consistency. With high compliance to this request, the consistency of the demographic data might be improved for sentinel practices compared with all Ontario practitioners. Alternatively, future work could assess whether resistance data from the top submitting practices differs significantly from all Ontario data. If resistance patterns from potential sentinel clinics are representative of all resistance patterns, sentinel clinic data may provide an option for temporal AMR surveillance.
Another common occurrence that may hinder data analysis in a semi-automated surveillance system is recording inconsistency. In many instances, a single clinic or farm name was recorded in multiple formats within the data acquired. Thus, one clinic might be treated as multiple clinics in a statistical model, purely due to formatting problems (either at data entry when specimens arrive at the lab or by the individuals submitting specimens). A standardized recording method should be used for clinic and farm names such as a unique identification code or by use of a drop-down menu approach in place of current free-text fields. Although the unique identification code field has a high compliance for recording, it currently does not fulfill the requirements for use in a surveillance system because each clinic has multiple “unique” codes recorded. It may be suitable to develop a system such that clinics are registered into the LIMS database using a standardized format, which would provide an opportunity to check new registrations against those that already exist within the data.
The choice of drugs to include on the panel of antimicrobials should be based on defined criteria that include use in therapy of swine diseases and importance for human medicine. The AHL data fulfill these criteria. As the AHL aims to provide susceptibility information that is practical to the practitioner, susceptibility is tested to antimicrobials used in swine to treat the organism in question. The most commonly employed panel of antimicrobials for each of the pathogens assessed in this study matches with the Canadian Veterinary Medical Association’s (CVMA) antimicrobial prudent use guidelines (22), and includes antimicrobials in each category of the Veterinary Drug Directorate (VDD) Categorization of Antimicrobial Drugs Based on Importance in Human Medicine (23). Accordingly, these antimicrobials are also those used in Canadian swine production (24).
Addition or removal of antimicrobials from the commonly tested panels, and changes in interpretive criteria present a challenge for an ongoing surveillance system. Trends in resistance (including multidrug and multiclass resistance) cannot be reliably tracked for a pathogen/antimicrobial combination if susceptibility testing for the given antimicrobial changes over time. When these changes occur, documentation must be provided to stakeholders, along with the impact of the change on interpretation of the data. An example of suitable documentation is provided online by CIPARS (25). If possible, a standard minimum panel for each pathogen should be implemented such that a set of antimicrobials is available for which resistance rates could be followed over time. Changes to these panels may be appropriate when new drugs become available for agricultural use, or changes to antimicrobial labels including times when E. coli, P. multocida, or S. suis are added as targeted pathogens. However, removal of drugs from agricultural use may not warrant the removal of these drugs from the panel used for surveillance. Depending on the circumstances and the potential for reinstatement over time, such changes would require discussion at regular meetings between AHL staff and surveillance personnel. Similarly, changes in breakpoints for resistance that affect R/I/S interpretive criteria may be conveyed at these meetings. This highlights a key conflict between the uses of laboratory data; within a laboratory, the focus is on identifying the susceptibility results for the individual case/veterinarian/producer. Therefore, testing susceptibility of an antimicrobial no longer used in practice is illogical, and logical changes in breakpoints may occur over time as pathogens evolve. However, continued monitoring of a product that has been removed from the market provides key surveillance information; particularly about the rate at which AMR prevalence declines or persists upon the removal of a particular antimicrobial drug or class. Furthermore, surveillance personnel need to have breakpoint changes documented, as these changes affect the comparability of data.
The data management system used at the AHL requires significant technological expertise to retrieve data in a suitable format for surveillance. The system performs well when generating individual reports for farm owners or practitioners; however, retrieval of large datasets is difficult. The current system requires multiple searches and extraction/merging steps to develop a database that lists submission demographic information and results in formatting with 1 row per isolate. A system or subroutine that allows extraction/export in this format, in a single file, would increase efficiency with regards to speed, number of files saved, and possible points of error.
With knowledge of the current trends in resistance and predictors for specific animal pathogens, veterinarians may be able to make more informed decisions regarding their use of antimicrobials and its potential to increase selection pressure for AMR. Furthermore, knowledge of the current trends in AMR prevalence may allow for routine treatment options to be assessed in order to reduce the occurrence of treatment failure, to guide the development of provincial regulations and empirical treatment guidelines for antimicrobial use in livestock, and to examine the impacts of interventions to alter antimicrobial use in the province. Moreover, 2 of the 3 pathogens assessed in this paper (P. multocida and S. suis) had increases in isolation from 2004 to 2006, concurrent with an increase in submissions to the AHL and the Ontario porcine circovirus type 2 outbreak (16). These results suggest that a system for AMR surveillance may also allow for a form of syndromic surveillance for emerging diseases whereby increased submissions to the AHL, requests for antimicrobial susceptibility testing, or isolation of other pathogens affecting the anatomical system of interest may indicate times when a novel disease is circulating in the source population.
Therefore, the development of an AMR surveillance system for clinical isolates from Ontario swine will be an asset for local veterinarians and researchers. This surveillance system may be used to promote the health of swine herds in Ontario, to improve and monitor antimicrobial stewardship efforts in the province, and potentially identify unique or novel infections. Current data from the AHL will support a system using temporal statistical techniques, and with the adoption of improved recording practices, adjustment for covariates and spatial or temporal-spatial analyses may be supported in the future.
Acknowledgments
The authors express their appreciation to Dr. Jane Parmley, Dr. David Léger, and Dr. Agnes Agunos for their contributions to the conceptual framework for this project, and for their guidance in the preparation of this manuscript. CVJ
Footnotes
Support for the Pearl laboratory and the work of S.K. Glass-Kaastra were provided by the Animal Health Strategic Investment, Canada Foundation for Innovation, and the Ontario Research Fund.
Use of this article is limited to a single copy for personal study. Anyone interested in obtaining reprints should contact the CVMA office (hbroughton@cvma-acmv.org) for additional copies or permission to use this material elsewhere.
References
- 1.Public Health Agency of Canada. Canadian Integrated Program for Antimicrobial Resistance Surveillance (CIPARS) Final Report, 2007. [Last accessed January 15, 2014]. Available from: http://www.phac-aspc.gc.ca/cipars-picra/2007-eng.php.
- 2.National Antimicrobial Resistance Monitoring System for Enteric Bacteria (NARMS): Human Isolates Final Report, 2008. Centers for Disease Control and Prevention; [Last accessed January 15, 2013]. Available from: http://www.cdc.gov/narms/annual/2008/narms_2008_annual_report.pdf. [Google Scholar]
- 3.National Antimicrobial Resistance Monitoring System for Enteric Bacteria (NARMS): Animal Arm Annual Report, 2007. Atlanta, Georgia: United States Department of Health and Human Services; 2010. [Google Scholar]
- 4.National Antimicrobial Resistance Monitoring System for Enteric Bacteria (NARMS): Retail Meat Annual Report, 2007. Food and Drug Administration; [Last accessed January 15, 2013]. Available from: http://www.fda.gov/downloads/AnimalVeterinary/SafetyHealth/AntimicrobialResistance/NationalAntimicrobialResistanceMonitoringSystem/UCM166132.pdf. [Google Scholar]
- 5.Danish Integrated Antimicrobial Resistance Monitoring and Research Programme 2011 — Use of antimicrobial agents and occurrence of antimicrobial resistance in bacteria from food animals, foods and humans in Denmark. Copenhagen, Denmark: Statens Serum Institut, Danish Veterinary and Food Administration, Danish Medicines Agency, and Danish Institute for Food and Veterinary Research; 2012. [Google Scholar]
- 6.Government of Japan. A report on the Japanese veterinary antimicrobial resistance monitoring system 2000 to 2007. Tokyo, Japan: National Veterinary Assay Laboratory, Ministry of Agriculture, Forestry and Fisheries; 2009. [Google Scholar]
- 7.Gravel D, Matlow A, Ofner-Agostini M, et al. Canadian Nosocomial Infection Surveillance Program (CNISP) A point prevalence survey of healthcare associated infections in pediatric populations in major Canadian acute-care hospitals. Am J Infect Control. 2007;35:157–162. doi: 10.1016/j.ajic.2006.06.006. [DOI] [PubMed] [Google Scholar]
- 8.Respiratory virus detection surveillance system [homepage on the Internet] Public Health Agency of Canada; [Last accessed January 15, 2014]. [updated 2013 May 30]. Available from: http://www.phac-aspc.gc.ca/bid-bmi/dsd-dsm/rvdi-divr/index-eng.php. [Google Scholar]
- 9.Sexually transmitted infections surveillance and epidemiology section [homepage on the Internet] Public Health Agency of Canada; [Last accessed January 15, 2014]. [updated 2009 October 19]. Available from: http://www.phac-aspc.gc.ca/sti-its-surv-epi/about-eng.php. [Google Scholar]
- 10.Catry B, Laevens H, Devriese LA, Opsomer G, de Kruif A. Antimicrobial resistance in livestock. J Vet Pharmacol Ther. 2003;26:81–93. doi: 10.1046/j.1365-2885.2003.00463.x. [DOI] [PubMed] [Google Scholar]
- 11.De Castro P [report on the Internet] European Parliament resolution on antibiotic resistance. 2011. [Last accessed January 15, 2014]. Available from: http://www.europarl.europa.eu/sides/getDoc.do?type=MOTION&reference=B7-2011-0295&language=EN.
- 12.Uses of antimicrobials in food animals in Canada: Impact on resistance and Human health [report on the Internet] Health Canada, 2002. Veterinary Drugs Directorate; [Last accessed January 15, 2013]. Available from: http://www.hc-sc.gc.ca/dhp-mps/alt_formats/hpfb-dgpsa/pdf/pubs/amr-ram_final_report-rapport_06-27-eng.pdf. [Google Scholar]
- 13.Lee LM, Teutsch SM, Thacker SB, St. Louis ME. Principles and Practice of Public Health Surveillance. 3rd ed. New York, New York: Oxford University Press; 2010. [Google Scholar]
- 14.Straw BE, Zimmerman JJ, D’Allaire S, Taylor DJ. Diseases of Swine. 9th ed. Oxford, United Kingdom: Blackwell publishing; 2006. [Google Scholar]
- 15.Animal Health Laboratory swine submission form. University of Guelph Laboratory Services [pdf on the internet, last updated 2010] [Last accessed January 15, 2014]. Available from: http://www.guelphlabservices.com/AHL/Submission_Forms.aspx.
- 16.O’Sullivan TL, Friendship R, Pearl DL, McEwen B, Ker A, Dewey C. The association between submission counts to a veterinary diagnostic laboratory and the economic and disease challenges of the Ontario swine industry from 1998 to 2009. Prev Vet Med. 2012;3–4:275–283. doi: 10.1016/j.prevetmed.2012.03.005. [DOI] [PubMed] [Google Scholar]
- 17.O’Sullivan TL. PhD thesis. Guelph, Ontario: University of Guelph; 2011. An investigation of various sources of health data for swine disease surveillance. [Google Scholar]
- 18.Amezcua M, Pearl DL, Friendship RM, McNab WB. Evaluation of a veterinary-based syndromic surveillance system implemented for swine. Can J Vet Res. 2010;74:241–251. [PMC free article] [PubMed] [Google Scholar]
- 19.Doohoo I, Martin W, Stryhn H. Veterinary Epidemiologic Research. Charlottetown, Prince Edward Island: AVC Inc; 2003. [Google Scholar]
- 20.Alton GD, Pearl DP, Bateman KG, McNab WB, Berke O. Factors associated with whole carcass condemnation rates in provincially-inspected abattoirs in Ontario 2001–2007: Implications for food animal syndromic surveillance. BMC Vet Res. 2010;6:42. doi: 10.1186/1746-6148-6-42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Silk BJ, Berkelman RL. A review of strategies for enhancing the completeness of notifiable disease reporting. JPHMP. 2005;11:191–200. doi: 10.1097/00124784-200505000-00003. [DOI] [PubMed] [Google Scholar]
- 22.Antimicrobial prudent use guidelines [pdf on the internet] Canadian Veterinary Medical Association; 2008. [Last accessed January 15, 2014]. Available from: http://www.canwestconference.ca/PDF/speakers/Food%20Animal%20Sessions/Dr.%20Warren%20Skippon/CVMA%20Prudent%20Use%20Guidelines.pdf. [Google Scholar]
- 23.Categorization of Antimicrobial Drugs Based on Importance in Human Medicine [homepage on the Internet] Government of Canada; [Last accessed January 15, 2014]. [Last updated April 2009]. Available from: http://www.hc-sc.gc.ca/dhp-mps/vet/antimicrob/amr_ram_hum-med-rev-eng.php. [Google Scholar]
- 24.Government of Canada. Canadian Integrated Program for Antimicrobial Resistance Surveillance (CIPARS) 2008 — Farm Surveillance in Pigs Preliminary Results: Antimicrobial Use. Public Health Agency of Canada; [Last accessed January 15, 2014]. Available from: http://publications.gc.ca/site/eng/371048/publication.html. [Google Scholar]
- 25.About CIPARS, Changes to CIPARS. [Website on the internet] Government of Canada; [Last accessed January 15, 2014]. [Last updated 2007 August 15] Available from: http://www.phac-aspc.gc.ca/cipars-picra/about_change-eng.php#ref. [Google Scholar]