Abstract
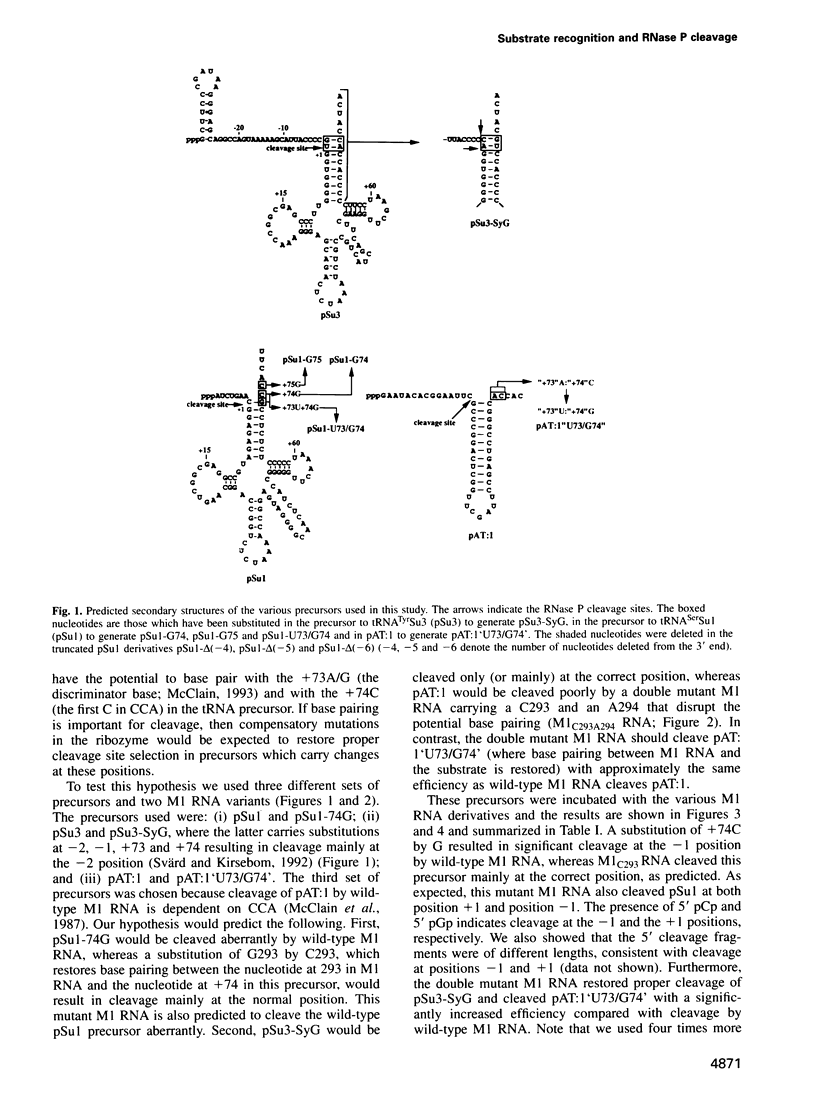
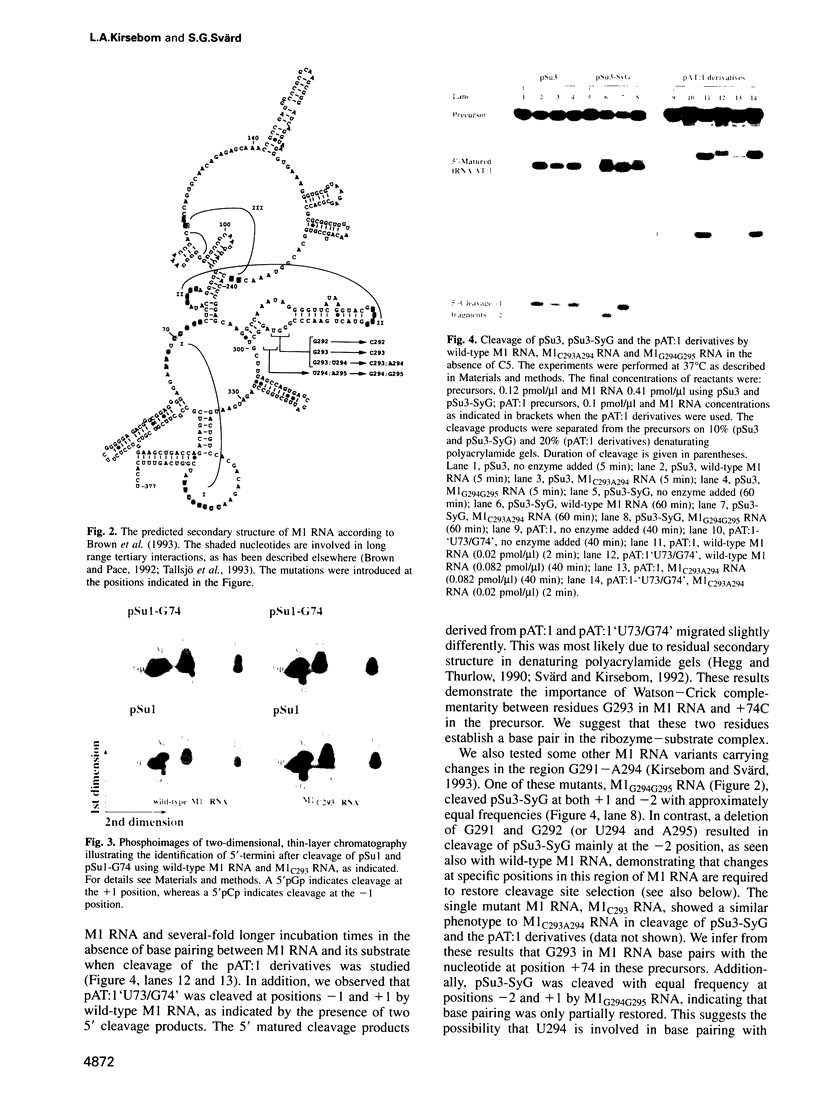
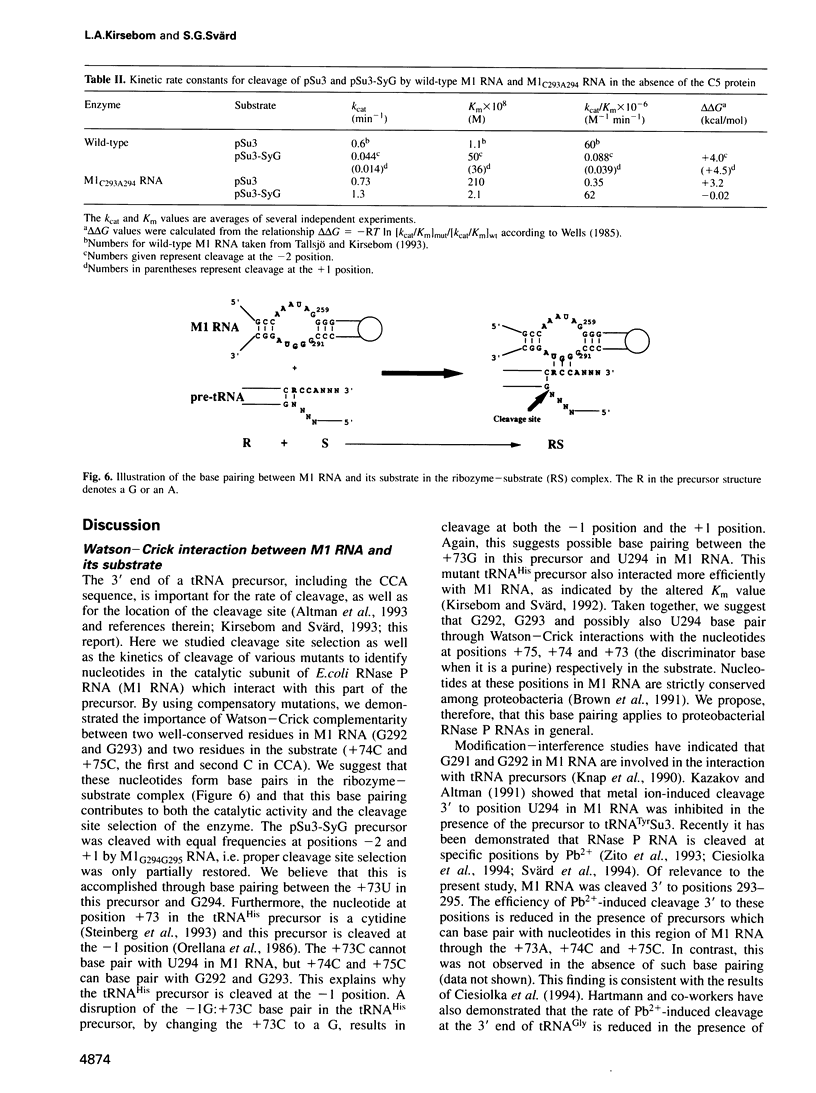
Base pairing between the substrate and the ribozyme has previously been shown to be essential for catalytic activity of most ribozymes, but not for RNase P RNA. By using compensatory mutations we have demonstrated the importance of Watson-Crick complementarity between two well-conserved residues in Escherichia coli RNase P RNA (M1 RNA), G292 and G293, and two residues in the substrate, +74C and +75C (the first and second C residues in CCA). We suggest that these nucleotides base pair (G292/+75C and G293/+74C) in the ribozyme-substrate complex and as a consequence the amino acid acceptor stem of the precursor is partly unfolded. Thus, a function of M1 RNA is to anchor the substrate through this base pairing, thereby exposing the cleavage site such that cleavage is accomplished at the correct position. Our data also suggest possible base pairing between U294 in M1 RNA and the discriminator base at position +73 of the precursor. Our findings are also discussed in terms of evolution.
Full text
PDF



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Altman S., Kirsebom L., Talbot S. Recent studies of ribonuclease P. FASEB J. 1993 Jan;7(1):7–14. doi: 10.1096/fasebj.7.1.7916700. [DOI] [PubMed] [Google Scholar]
- Baer M. F., Reilly R. M., McCorkle G. M., Hai T. Y., Altman S., RajBhandary U. L. The recognition by RNase P of precursor tRNAs. J Biol Chem. 1988 Feb 15;263(5):2344–2351. [PubMed] [Google Scholar]
- Brown J. W., Haas E. S., James B. D., Hunt D. A., Liu J. S., Pace N. R. Phylogenetic analysis and evolution of RNase P RNA in proteobacteria. J Bacteriol. 1991 Jun;173(12):3855–3863. doi: 10.1128/jb.173.12.3855-3863.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown J. W., Haas E. S., Pace N. R. Characterization of ribonuclease P RNAs from thermophilic bacteria. Nucleic Acids Res. 1993 Feb 11;21(3):671–679. doi: 10.1093/nar/21.3.671. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown J. W., Pace N. R. Ribonuclease P RNA and protein subunits from bacteria. Nucleic Acids Res. 1992 Apr 11;20(7):1451–1456. doi: 10.1093/nar/20.7.1451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ciesiolka J., Hardt W. D., Schlegl J., Erdmann V. A., Hartmann R. K. Lead-ion-induced cleavage of RNase P RNA. Eur J Biochem. 1994 Jan 15;219(1-2):49–56. doi: 10.1111/j.1432-1033.1994.tb19913.x. [DOI] [PubMed] [Google Scholar]
- Green C. J., Vold B. S. A cluster of nine tRNA genes between ribosomal gene operons in Bacillus subtilis. J Bacteriol. 1992 May;174(10):3147–3151. doi: 10.1128/jb.174.10.3147-3151.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Green C. J., Vold B. S. Staphylococcus aureus has clustered tRNA genes. J Bacteriol. 1993 Aug;175(16):5091–5096. doi: 10.1128/jb.175.16.5091-5096.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Green C. J., Vold B. S. Structural requirements for processing of synthetic tRNAHis precursors by the catalytic RNA component of RNase P. J Biol Chem. 1988 Jan 15;263(2):652–657. [PubMed] [Google Scholar]
- Gregory S. T., Lieberman K. R., Dahlberg A. E. Mutations in the peptidyl transferase region of E. coli 23S rRNA affecting translational accuracy. Nucleic Acids Res. 1994 Feb 11;22(3):279–284. doi: 10.1093/nar/22.3.279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guerrier-Takada C., Gardiner K., Marsh T., Pace N., Altman S. The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell. 1983 Dec;35(3 Pt 2):849–857. doi: 10.1016/0092-8674(83)90117-4. [DOI] [PubMed] [Google Scholar]
- Guerrier-Takada C., Lumelsky N., Altman S. Specific interactions in RNA enzyme-substrate complexes. Science. 1989 Dec 22;246(4937):1578–1584. doi: 10.1126/science.2480641. [DOI] [PubMed] [Google Scholar]
- Guerrier-Takada C., McClain W. H., Altman S. Cleavage of tRNA precursors by the RNA subunit of E. coli ribonuclease P (M1 RNA) is influenced by 3'-proximal CCA in the substrates. Cell. 1984 Aug;38(1):219–224. doi: 10.1016/0092-8674(84)90543-9. [DOI] [PubMed] [Google Scholar]
- Guerrier-Takada C., van Belkum A., Pleij C. W., Altman S. Novel reactions of RNAase P with a tRNA-like structure in turnip yellow mosaic virus RNA. Cell. 1988 Apr 22;53(2):267–272. doi: 10.1016/0092-8674(88)90388-1. [DOI] [PubMed] [Google Scholar]
- Hardt W. D., Schlegl J., Erdmann V. A., Hartmann R. K. Role of the D arm and the anticodon arm in tRNA recognition by eubacterial and eukaryotic RNase P enzymes. Biochemistry. 1993 Dec 7;32(48):13046–13053. doi: 10.1021/bi00211a014. [DOI] [PubMed] [Google Scholar]
- Hegg L. A., Thurlow D. L. Residual tRNA secondary structure in 'denaturing' 8M urea/TBE polyacrylamide gels: effects on electrophoretic mobility and dependency on prior chemical modification of the tRNA. Nucleic Acids Res. 1990 May 25;18(10):2993–3000. doi: 10.1093/nar/18.10.2993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kazakov S., Altman S. Site-specific cleavage by metal ion cofactors and inhibitors of M1 RNA, the catalytic subunit of RNase P from Escherichia coli. Proc Natl Acad Sci U S A. 1991 Oct 15;88(20):9193–9197. doi: 10.1073/pnas.88.20.9193. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kirsebom L. A., Altman S. Reaction in vitro of some mutants of RNase P with wild-type and temperature-sensitive substrates. J Mol Biol. 1989 Jun 20;207(4):837–840. doi: 10.1016/0022-2836(89)90250-7. [DOI] [PubMed] [Google Scholar]
- Kirsebom L. A., Svärd S. G. Identification of a region within M1 RNA of Escherichia coli RNase P important for the location of the cleavage site on a wild-type tRNA precursor. J Mol Biol. 1993 Jun 5;231(3):594–604. doi: 10.1006/jmbi.1993.1312. [DOI] [PubMed] [Google Scholar]
- Kirsebom L. A., Svärd S. G. The kinetics and specificity of cleavage by RNase P is mainly dependent on the structure of the amino acid acceptor stem. Nucleic Acids Res. 1992 Feb 11;20(3):425–432. doi: 10.1093/nar/20.3.425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Knap A. K., Wesolowski D., Altman S. Protection from chemical modification of nucleotides in complexes of M1 RNA, the catalytic subunit of RNase P from E coli, and tRNA precursors. Biochimie. 1990 Nov;72(11):779–790. doi: 10.1016/0300-9084(90)90187-l. [DOI] [PubMed] [Google Scholar]
- Komine Y., Adachi T., Inokuchi H., Ozeki H. Genomic organization and physical mapping of the transfer RNA genes in Escherichia coli K12. J Mol Biol. 1990 Apr 20;212(4):579–598. doi: 10.1016/0022-2836(90)90224-A. [DOI] [PubMed] [Google Scholar]
- Krupp G., Kahle D., Vogt T., Char S. Sequence changes in both flanking sequences of a pre-tRNA influence the cleavage specificity of RNase P. J Mol Biol. 1991 Feb 20;217(4):637–648. doi: 10.1016/0022-2836(91)90522-8. [DOI] [PubMed] [Google Scholar]
- McClain W. H., Guerrier-Takada C., Altman S. Model substrates for an RNA enzyme. Science. 1987 Oct 23;238(4826):527–530. doi: 10.1126/science.2443980. [DOI] [PubMed] [Google Scholar]
- McClain W. H. Rules that govern tRNA identity in protein synthesis. J Mol Biol. 1993 Nov 20;234(2):257–280. doi: 10.1006/jmbi.1993.1582. [DOI] [PubMed] [Google Scholar]
- Milligan J. F., Groebe D. R., Witherell G. W., Uhlenbeck O. C. Oligoribonucleotide synthesis using T7 RNA polymerase and synthetic DNA templates. Nucleic Acids Res. 1987 Nov 11;15(21):8783–8798. doi: 10.1093/nar/15.21.8783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moazed D., Noller H. F. Sites of interaction of the CCA end of peptidyl-tRNA with 23S rRNA. Proc Natl Acad Sci U S A. 1991 May 1;88(9):3725–3728. doi: 10.1073/pnas.88.9.3725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Orellana O., Cooley L., Söll D. The additional guanylate at the 5' terminus of Escherichia coli tRNAHis is the result of unusual processing by RNase P. Mol Cell Biol. 1986 Feb;6(2):525–529. doi: 10.1128/mcb.6.2.525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perreault J. P., Altman S. Important 2'-hydroxyl groups in model substrates for M1 RNA, the catalytic RNA subunit of RNase P from Escherichia coli. J Mol Biol. 1992 Jul 20;226(2):399–409. doi: 10.1016/0022-2836(92)90955-j. [DOI] [PubMed] [Google Scholar]
- Rould M. A., Perona J. J., Söll D., Steitz T. A. Structure of E. coli glutaminyl-tRNA synthetase complexed with tRNA(Gln) and ATP at 2.8 A resolution. Science. 1989 Dec 1;246(4934):1135–1142. doi: 10.1126/science.2479982. [DOI] [PubMed] [Google Scholar]
- Saiki R. K., Gelfand D. H., Stoffel S., Scharf S. J., Higuchi R., Horn G. T., Mullis K. B., Erlich H. A. Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science. 1988 Jan 29;239(4839):487–491. doi: 10.1126/science.2448875. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seidman J. G., McClain W. H. Three steps in conversion of large precursor RNA into serine and proline transfer RNAs. Proc Natl Acad Sci U S A. 1975 Apr;72(4):1491–1495. doi: 10.1073/pnas.72.4.1491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steinberg S., Misch A., Sprinzl M. Compilation of tRNA sequences and sequences of tRNA genes. Nucleic Acids Res. 1993 Jul 1;21(13):3011–3015. doi: 10.1093/nar/21.13.3011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Svärd S. G., Kirsebom L. A. Determinants of Escherichia coli RNase P cleavage site selection: a detailed in vitro and in vivo analysis. Nucleic Acids Res. 1993 Feb 11;21(3):427–434. doi: 10.1093/nar/21.3.427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Svärd S. G., Kirsebom L. A. Several regions of a tRNA precursor determine the Escherichia coli RNase P cleavage site. J Mol Biol. 1992 Oct 20;227(4):1019–1031. doi: 10.1016/0022-2836(92)90518-o. [DOI] [PubMed] [Google Scholar]
- Svärd S. G., Mattsson J. G., Johansson K. E., Kirsebom L. A. Cloning and characterization of the RNase P RNA genes from two porcine mycoplasmas. Mol Microbiol. 1994 Mar;11(5):849–859. doi: 10.1111/j.1365-2958.1994.tb00363.x. [DOI] [PubMed] [Google Scholar]
- Tallsjö A., Kirsebom L. A. Product release is a rate-limiting step during cleavage by the catalytic RNA subunit of Escherichia coli RNase P. Nucleic Acids Res. 1993 Jan 11;21(1):51–57. doi: 10.1093/nar/21.1.51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tallsjö A., Svärd S. G., Kufel J., Kirsebom L. A. A novel tertiary interaction in M1 RNA, the catalytic subunit of Escherichia coli RNase P. Nucleic Acids Res. 1993 Aug 25;21(17):3927–3933. doi: 10.1093/nar/21.17.3927. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vioque A., Arnez J., Altman S. Protein-RNA interactions in the RNase P holoenzyme from Escherichia coli. J Mol Biol. 1988 Aug 20;202(4):835–848. doi: 10.1016/0022-2836(88)90562-1. [DOI] [PubMed] [Google Scholar]
- Vold B. S. Structure and organization of genes for transfer ribonucleic acid in Bacillus subtilis. Microbiol Rev. 1985 Mar;49(1):71–80. doi: 10.1128/mr.49.1.71-80.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wells J. A. Additivity of mutational effects in proteins. Biochemistry. 1990 Sep 18;29(37):8509–8517. doi: 10.1021/bi00489a001. [DOI] [PubMed] [Google Scholar]
- Zito K., Hüttenhofer A., Pace N. R. Lead-catalyzed cleavage of ribonuclease P RNA as a probe for integrity of tertiary structure. Nucleic Acids Res. 1993 Dec 25;21(25):5916–5920. doi: 10.1093/nar/21.25.5916. [DOI] [PMC free article] [PubMed] [Google Scholar]





