Abstract
In multicellular organisms, transcription regulation is one of the central mechanisms modelling lineage differentiation and cell-fate determination1. Transcription requires dynamic chromatin configurations between promoters and their corresponding distal regulatory elements2. It is believed that their communication occurs within large discrete foci of aggregated RNA polymerases termed transcription factories in three-dimensional nuclear space3. However, the dynamic nature of chromatin connectivity has not been characterized at the genome-wide level. Here, through a chromatin interaction analysis with paired-end tagging approach3–5 using an antibody that primarily recognizes the pre-initiation complexes of RNA polymerase II6, we explore the transcriptional interactomes of three mouse cells of progressive lineage commitment, including pluripotent embryonic stem cells7, neural stem cells8 and neuro-sphere stem/progenitor cells9. Our global chromatin connectivity maps reveal approximately 40,000 long-range interactions, suggest precise enhancer–promoter associations and delineate cell-type-specific chromatin structures. Analysis of the complex regulatory repertoire shows that there are extensive colocalizations among promoters and distal-acting enhancers. Most of the enhancers associate with promoters located beyond their nearest active genes, indicating that the linear juxtaposition is not the only guiding principle driving enhancer target selection. Although promoter–enhancer interactions exhibit high cell-type specificity, promoters involved in interactions are found to be generally common and mostly active among different cells. Chromatin connectivity networks reveal that the pivotal genes of reprogramming functions are transcribed within physical proximity to each other in embryonic stem cells, linking chromatin architecture to coordinated gene expression. Our study sets the stage for the full-scale dissection of spatial and temporal genome structures and their roles in orchestrating development.
Gene regulatory networks are organized by spatial connectivity between distal regulatory elements (DREs) and their corresponding promoters3. Many of these DREs, including cell-specific enhancers, were characterized for their vital function in development and differentiation2. Increasing evidence has shown that DREs can function over long distances3, even on a different chromosome10 from their target genes. However, much of our current knowledge of cell-specific transcription regulation is based on extensive survey of DREs in the linear genome2,11,12. The direct delineation of genome-wide DRE–promoter interactions is still very limited, and how chromatin structure regulates transcription is largely unknown.
To explore the promoter-associated chromatin interactomes, we used the RNA polymerase II (RNAPII) chromatin interaction analysis with paired-end tagging (ChIA-PET) approach3 (see Methods) in three murine cell types: embryonic stem cells (ESCs), neural stem cells (NSCs) and neurosphere stem/progenitorcells (NPCs). NSCs and NPCs are two widely used neural development models representing different commitment steps. NSCs are clonally derived early neural stem cells obtained by in vitro differentiation of ESCs8, whereas NPCs are neural progenitor cells derived ex vivo from the forebrain telencephalic region9. Further examination of their transcription profiles confirms their expected cellular origins (Supplementary Information section 1). Chromatin immunoprecipitation (ChIP) efficiency and ChIA-PET library quality were evaluated (Extended Data Fig. 1 and Supplementary Information section 2) and the data were processed further to estimate reproducibility, noise and coverage (Supplementary Information sections 3, 4). Non-chimaeric, uniquely mapped PETs were used to define three classes of genome-wide information: the RNAPII-associated binding sites and long-range intra- and interchromosomal interaction clusters (Extended Data Fig. 2 and Supplementary Table 1). Using two independent approaches—quantitative PCR analysis of chromosome conformation capture (3C) and DNA fluorescent in situ hybridization (FISH)—we were able to validate the defined intra- and interchromosomal interactions (Supplementary Information section 5). In all we identified 40,000 RNAPII-bound interaction pairs present from a total of three cell types (Supplementary Table 2).
Extended Data Figure 1. Enrichments of RNAPII ChIP by ChIP-qPCR in three cell lines.
a–c, Fold enrichments (y axes) of RNAPII ChIP in selected regions (x axes) from three different cell lines (mouse ESCs (a), NSCs (b) and NPCs (c)) are shown. Two replicates of ChIP were tested via ChIP-qPCR and are represented as different colours (red and blue).
Extended Data Figure 2. Schematic overview of ChIA-PET analysis.
RNAPII binding sites, intra- and interchromosomal interactions identified from each cell type are shown.
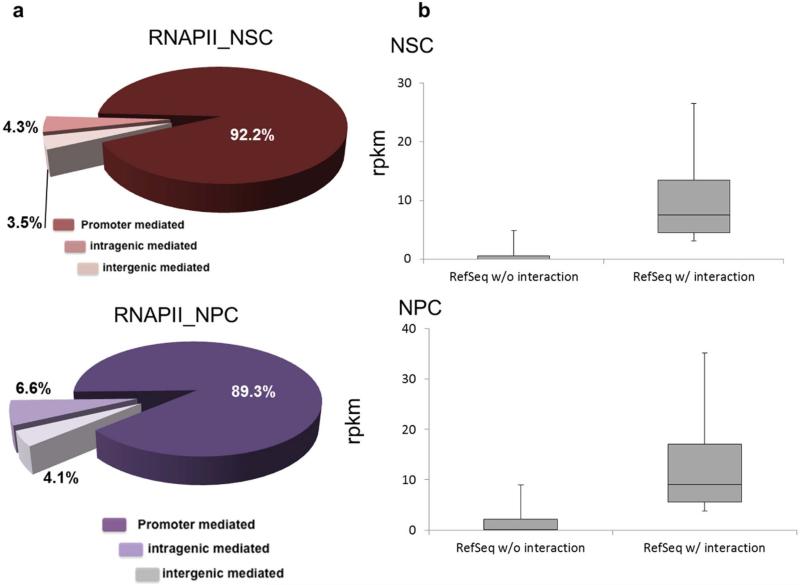
Consistent with its role in transcription initiation, this form of RNAPII, with a non-phosphorylated carboxy-terminal domain, tethers most (86–92%) of the chromatin interactions surrounding promoter regions (±2.5 kilobases (kb) of transcription start sites of UCSC known genes). In ESCs, roughly half of the promoter-tethered interactions connect two promoters (P–P), indicating the prevalence of promoter associations in the nucleus. The other half is distributed among promoters connecting to either intergenic (24%; P–inter) or intragenic (20%; P–intra) regions (Fig. 1a). Similar profiles are also found in NSCs and NPCs (Extended Data Fig. 3a). Through visual examination of these interactions, we identified many known enhancer–promoter (E–P) interactions, including the genic loci of Oct4, Nanog, Phc1 and Lefty1 (ref. 13) (Extended Data Fig. 4). Thus, these promoter-centric connectivity maps reveal large numbers of putative enhancers through both the inter- and intragenic interacting loci. In total, we identified 8,309, 4,463 and 3,649 putative DREs in ESCs, NSCs and NPCs, respectively. In ESCs, these distal loci exhibit enhancer characteristics; namely monomethylated histone H3 lysine 4 (H3K4me1) enrichment (P < 1 × 10–4), occupancy by pluripotent transcription factors and co-activator p300 (also known as Ep300), as well as sequence conservation14 (Fig. 1b, c and Extended Data Fig. 5). Furthermore, approximately half of the multiple-transcription-factor-binding loci (MTL) previously identified in ESCs12 overlap with these distal interaction sites. These regions also share significant overlap (P < 1 × 10–4) with occupancy of mediator, a protein complex known to be associated with enhancers13. Last, the expression levels of the genes involved in the RNAPII interactions are significantly higher than those with no detected interaction (P ≤ 2.2 × 10–16), suggesting that their promoters are transcriptionally more active (Fig. 1d and Extended Data Fig. 3b). Interestingly, among all the putative enhancers captured by RNAPII-bound interactions, we found 563 potential ‘poised enhancers’ connecting to the bivalent promoters15. Poised enhancers in ESCs are thought to ‘prime’ genes for subsequent for cell-type-dependent transcription activity during development16.
Figure 1. RNAPII tethers promoter–enhancer interactions.
a, Distribution of promoter-, intragenic- and intergenic-centred interactions in ESCs. The percentages of P–inter, P–intra and P–P interactions are listed.
b, Enrichment of H3K4me1 signal density28 (y axis: RPM, reads per million) along the non-promoter nodes (x axis). Signal from random control regions is shown in blue.
c, Enrichment of 12 transcription factors, Ctcf, p300 and MTL (multiple-transcription-factor-binding loci) occupancies at non-promoter regions. d, Boxplot expression (reads per kilobase per million reads (RPKM) as y axis) of the genes with versus without interactions (x axis) in ESCs.
Extended Data Figure 3. Promoter-mediated interactions and associated gene expression levels in NSCs and NPCs.
a, Distribution of defined interaction between promoters, inter- and intragenic regions in NSCs (top) and NPCs (bottom). b, Boxplots of the expression level (RPKM, y axes) between genes tethered by RNAPII and genes without tethered interactions (x axes) in NSCs (top) and NPCs (bottom).
Extended Data Figure 4.
Enhancers for Nanog (top left), Phc1 (top right), Lefty1 (bottom right) and Oct4 (bottom left) uncovered through 3C analysis in mESC V6.5 (middle black track) and ChIA-PET analysis in mESC E14 (bottom red track).
Extended Data Figure 5.
Phylogenetic conservation represented by PhastCon scores30 of the putative enhancer regions in comparison with other types of genomic regions.
To directly confirm the in vivo enhancer activities of these non-promoter distal interaction loci, we carried out enhancer assays in transgenic zebrafish with green fluorescent protein (GFP) reporter genes17. Eleven out of 21 selected putative DRE loci drove reproducible and specific GFP expression patterns, indicating their spatiotemporal enhancer activity in zebrafish. (Fig. 2, Supplementary Table 3 and Supplementary Information section 6). Among them, a poised enhancer of tcf12, a neuronal developmental gene, shows no activity at early developmental stages but displays forebrain-specific activity at later developmental stages in zebrafish (Fig. 2d). Likewise in mouse, this locus is associated with the bivalent promoter, is bound by Suz12 and exhibits repressive trimethylated H3K27 (H3K27me3) modification in ESCs. This locus subsequently acquires p300 binding in NSCs and in correlation with high Tcf12 expression (reads per kilobase per million reads (RPKM) 7, 119 and 75 in ESCs, NSCs and NPCs, respectively). Taken together, these distal interacting loci presumably not only function as developmental regulators but also retain their targeting specificities across different vertebrates.
Figure 2. Distal interaction loci display enhancer activities in zebrafish.
a, The PET-5 cluster showing an intergenic region, MTL12, on chromosome 15 that targets in trans the npm1 promoter on chromosome 11 in ESCs. Embryos from 8, 12 and 24 hours post fertilization (hpf) are shown with the endogenous npm1a expression pattern as detected by whole-mount in situ hybridization (adapted from ref. 29 with permission). b, A cis-acting interaction region, MTL22, on chromosome 12, 9 kb upstream to the dact1 target gene in ESCs. Zebrafish enhancer assays from 8, 12 and 24 hpf and endogenous dact1 in situ hybridization are shown (adapted from ZFIN (http://zfin.org/) with permission). c, A zebrafish-validated enhancer, mE12, located in the 13th intron of the slc13a4 gene interacts with cell-specific target genes: 1810058I24Rik (cis in NSCs) and sema5b (trans in ESCs). At 24 and 48 hpf, this enhancer drove GFP expression in the somitic muscles and olfactory vesicle. d, Interactions mediated from the ‘poised’ enhancer in ESCs (top), NSCs (middle) and NPCs (bottom). The interactions are schematically depicted (left) and shown in the browser (right). Suz12, Zfx and p300 (ref. 12) binding are illustrated. Zebrafish embryo with representative forebrain expression at 48 hpf is shown at the bottom.
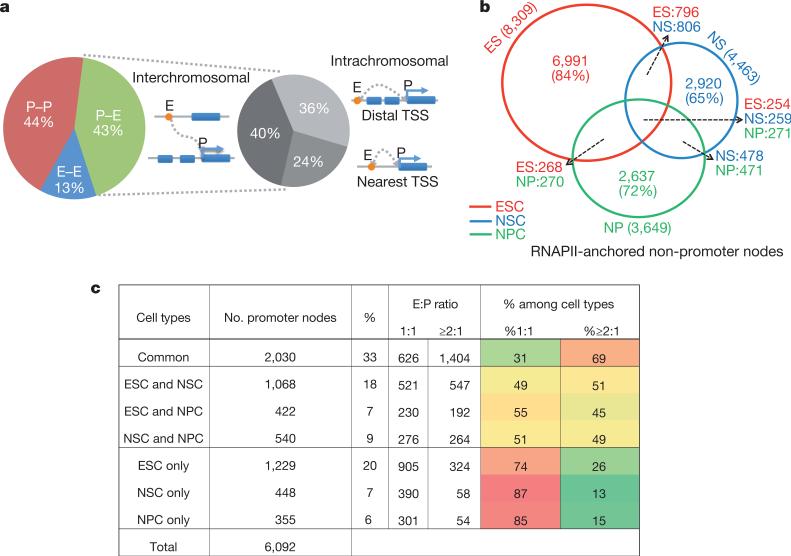
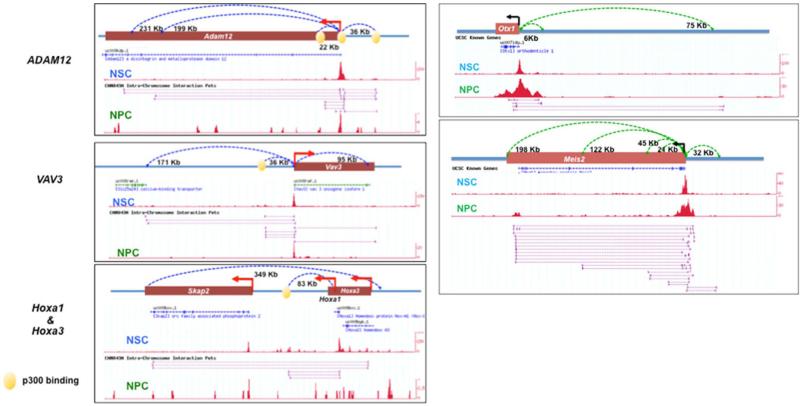
These putative E–P interactions defined by ChIA-PET were used to decipher the nature of regulatory elements and their targeted gene associations in mouse stem cells. Being different from many previous studies that assumed proximity as the governing rule for E–P associations11, our data suggest that a substantial fraction of these putative enhancers do not select their nearest promoters as their targets. In ESCs, 76% of the putative enhancer nodes interact beyond their closest active genes. Among them, 40% associated with interchromosomal interactions and 36% were involved in intrachromosomal interactions (Fig. 3a). Similar results were found in NSCs (77%) and NPCs (54%). Distal targeting has been reported in selected cases10,18. Our data suggest that this could be a pervasive phenomenon throughout the genome. Besides long-range targeting, putative E–P interactions also exhibit high specificity for targeted genes and cell types. In all three cell types examined, 60–70% of the total RNAPII-tethered promoters are associated with only one distinct putative enhancer (E:P ratio = 1) and over 90% of the potential enhancer loci are associated with only one targeted gene (P:E ratio = 1) (Supplementary Table 4). Supplementary Tables 5 and 6 list the numbers of enhancers and promoters associated with each of their corresponding partner nodes in each cell type. Among the cell-specific interactions, the putative E–P interactions are the most prevalent type, suggesting that cell-specific genes targeted by their corresponding enhancers are the most distinctive features of the RNAPII-mediated chromatin interactions (Supplementary Table 7). For example, Otx1 and Meis2, key forebrain-expressed homeobox genes, are connected with their cell-specific enhancers in NPCs, but not in NSCs. By contrast, NSCs contain specific E–P interactions involving genes expressed in the more posterior neuraxis, early development and in response to FGF2/EGF-like Adam12, VAV3 and Hoxa (Extended Data Fig. 6) are found. When all the interacting promoter and distal non-promoter nodes are compared separately, only a low percentage of the potential enhancers but a high percentage of the promoter nodes overlap among the three cell types (Fig. 3b, c), suggesting that the putative enhancers are mostly cell-type-specific but their targeted genes are largely common. When the cell-specific nature of these promoters was evaluated together with their interaction specificity, cell-specific promoters were found to be largely (74–87%) monogamous (E–P ratio of 1:1), whereas the universally expressed promoters are mostly (69%) promiscuous (E–P ratio of ≥ 2:1) (Fig. 3c). Supplementary Table 8 lists the 6,092 interacting promoter nodes found in all three cell types, their cell specificity and enhancer connectivity, and examples are shown in Extended Data Fig. 7. Therefore, we speculate that distinct regulatory elements are used to target ubiquitously expressed genes in different cell types and such widespread differential enhancer usage not only functions to upregulate target genes but can also serve as a distinct mode to organize cell-specific transcription regulatory networks.
Figure 3. Characterization of putative enhancer–promoter targeting.
a, Distributions of P–P, E–E and E–P interactions in ESCs (left) and interchromosomal, intrachromosomal with nearest promoters and intrachromosomal with distal targeting promoters (right). The dotted lines indicate the relative portions of different targeting patterns derived from only the P–E interactions. b, Venn diagram of non-promoter nodes defined among ESCs, NSCs and NPCs. c, Putative enhancer–promoter pairing relations versus their cell specificities.
Extended Data Figure 6.
NSC- and NPC-specific interactions detected from promoters of early developmental genes (left) Adam12 (top; chromosome 7:141165832—141495831), Vav3 (middle; chromosome 3:108932769—109282768) and Hoxa (bottom; chromosome 6:51730841—52230840) as well as key telencephalic homeobox transcription factors (right) Otx1 (top, chromosome 11:21878211—21998210) and Meis2 (bottom, chromosome 2:115603679—116003678). Dotted connecting lines depict the defined interactions with the distances labelled. The RNAPII binding peaks are shown in the middle track, followed by PET mapping in NSCs and NPCs, respectively.
Extended Data Figure 7. Examples for interacting promoter nodes with their cell specificity and enhancer connectivity found in all three cell types.
a, ESC-specific promoter nodes (Fzd7) with E:P (2:1). b, NSC- and NPC-specific promoter nodes (Fabp7) with E:P (1:1 and M:1 M: multiple, ≥2). c, Promoter nodes (Sox2) found in three different cell lines with dynamic E:P interactions.
To evaluate whether transcription regulatory circuitry is reflected by the cell-specific chromatin organization, we explored the spatial chromatin connectivity of core reprogramming genes19 (Pou5f1, Nanog, Lin28a, Sox2, Klf4 and Myc) in ESCs through the RNAPII interaction maps. The expression of these key reprogramming genes is known to govern pluripotency in ESCs through coordinated autoregulatory loops20. Using connectivity analysis, three Klf genes (Klf1, Klf2 and Klf4) were found to directly link to Sox2 (Extended Data Fig. 8). When the network analysis was extended from one to two hops of connectivity, all of the reprogramming genes were found to be connected within one major hub, except for Myc and Lin28a (Fig. 4a), suggesting that they could be colocalized in the nucleus within the same ‘transcriptional factory’. Among them, Nanog and Pou5f1 have limited connected edges whereas Sox2 has extensive connectivity.
Extended Data Figure 8. Connectivity constructed from the one-hop interactions mediated from reprogramming factor genes in ESCs.
Different colours represent different categories of cell specificity; the different sizes of the nodes represent non-promoter, promoter and iPS (induced pluripotent stem cell) factor nodes.
Figure 4. Connectivity networks converged by key transcription regulator genes.
a, Key re-programming gene network in ESCs. The connectivity was built through two hops of all interactions (light grey lines) mediated from 14 genes. Size of the circles represents the features of the nodes. Colours represent their cell specificities. b, Sox2-centric interaction map in ESCs. All of the Sox2-directly-interacting genes are labelled. Thick grey lines highlight the connectivity between Sox2, Klf1, Klf2 and Klf4. c, d, Olig1–Olig2 interaction networks in NSCs (c) and NPCs (d). Thick black lines highlight their common interactions. Genes involved in neuronal developments are in bold.
Sox2 can reprogram somatic cells to ESCs or NSCs21. In ESCs, the Sox2 promoter connects to clusters of ESC-specific enhancers to other pluripotency related genes like Sall1, Asf1b, Dusp6 and Jund (Fig. 4b). In NSCs, a very different Sox2 connectivity profile is observed (Extended Data Fig. 9). Such cell-specific connectivity is mediated through differential enhancer usage. Similarly, distinct connectivity maps constructed by oligodendrocyte transcription factors Olig1 and Olig2, genes important for neural cell fate determination22,23, are found in NSCs (Fig. 4c) and NPCs (Fig. 4d), respectively. As shown, Olig1 and Olig2 are directly connected to many genes critical for neuronal development in NSCs, including neuropilin (Neto2), Fabp7 and Bsg.
Extended Data Figure 9. Sox2-centric interaction map in NSCs.
All the interaction nodes directly connecting Sox2 are highlighted and gene names are labelled. The connectivities between Sox2, Myc and Pou3f2 are highlighted by thick grey lines.
RNAPII–chromatin interacting complexes captured here result from pre-initiation events and not all of them will proceed to active transcription or elongation24. Therefore, the rich repertoires of in vivo chromatin interactions presumably comprise a mixture of promoters with various transcriptional activities. Using the wealth of other genome-wide transcription and epigenetic data sets, one can further discriminate the transcription states of the identified interacting promoters and derive the significance of different ‘transcription factories’.
Our current study illustrates the complexity and dynamics of the underlying chromatin structures in the nucleus; however, we recognize that not all the interactions are functional. With additional ChIA-PET analyses from various forms of RNAPII, deeper sequencing and increased coverage, we expect to capture more promoter-mediated interaction pairs and diverse types of regulatory elements. Before concluding new insights and principles, any functional model made solely from colocalization association has to be examined together with other integrated (epi)genomic information and requires extensive functional validations, like the zebrafish transgenic assays or genome editing tools like CRISPR/Cas25. Finally, we expect that the recapitulation of such structure-based framework on a greater diversity of cell types and its integrated analyses shall further elucidate the mechanisms driving genome re-configuration, and to what extent it contributes to transcriptional regulation and cell specification.
METHODS SUMMARY
The ChIA-PET assay was performed using an RNAPII monoclonal antibody (8WG16, Covance) with E14 mouse ESC, NSC NS5 and NPC chromatin extracts. The sequence data were analysed as described26. The interactome networks were constructed using non-overlapping genomic regions. Nodes were annotated using the UCSC Genes annotations as ‘Promoter’ if they were within 2.5 kb of any transcription start site. Interactomes from various gene sets were visualized in Gephi27. ‘Parallel Force Atlas’ layout was used.
Supplementary Material
Acknowledgements
The authors thank J. Mariani for the preparation of RNA from NPC; K. Murphy and A. Ku for their assistance with zebrafish enhancer assays; and A. Visel and A. Nord for discussion and their comments on the manuscript. S.N. and R.F. were supported by grants from ASTIL Regione Lombardia (SAL-19 ref. no. 16874), Telethon (GGP12152), Cariplo (Rif. 2010-0673) and AIRC (IG-5801). N.A. is supported by NINDS grant number R01NS079231, NICHD grant number R01HD059862, NHGRI grant numbers R01HG005058 and R01HG006768, NIDDK award number R01DK090382, NIGMS award number GM61390 and Simons Foundation SFARI no. 256769. R.Y.B. is supported by NINDS grant number R01NS079231 and the UCSF Program for Biomedical Breakthrough Research (PBBR). This work was supported by Agency for Science, Technology and Research (A*STAR), Singapore, the Office of Science of the U.S. Department of Energy under contract no. DE-AC02-05CH11231 and National Institutes of Health ENCODE grants (R01 HG004456-01, R01HG003521-01 and 1U54HG004557-01) to Y.R. and C.-L.W.
Footnotes
Author Contributions Y.Z. constructed ChIA-PET experiments and data evaluation. R.Y.B. and N.A. designed and performed zebrafish enhancer assays. C.-H.W. carried out data analysis. C.-Y.N. and Y.Z. performed the RNA-seq experiments. J.L. performed DNA-FISH. H.M.P., E.T., R.F. and E.W. prepared the cells and ChIP material. G.L., F.H.M., W.-K.S. and Y.R. designed the data processing pipeline. C.-L.W. and S.N. designed the experiments. Y.Z. and C.-L.W. wrote the paper. All authors provided intellectual input and approved the final manuscript.
The authors declare no competing financial interests.
Online Content Any additional Methods, Extended Data display items and Source Data are available in the online version of the paper; references unique to these sections appear only in the online paper.
Supplementary Information is available in the online version of the paper.
References
- 1.Neph S, et al. Circuitry and dynamics of human transcription factor regulatory networks. Cell. 2012;150:1274–1286. doi: 10.1016/j.cell.2012.04.040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Visel A, et al. ChIP-seq accurately predicts tissue-specific activity of enhancers. Nature. 2009;457:854–858. doi: 10.1038/nature07730. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Li G, et al. Extensive promoter-centered chromatin interactions provide a topological basis for transcription regulation. Cell. 2012;148:84–98. doi: 10.1016/j.cell.2011.12.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Handoko L, et al. CTCF-mediated functional chromatin interactome in pluripotent cells. Nature Genet. 2011;43:630–638. doi: 10.1038/ng.857. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Fullwood MJ, et al. An oestrogen-receptor-a-bound human chromatin interactome. Nature. 2009;462:58–64. doi: 10.1038/nature08497. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Phatnani HP, Greenleaf AL. Phosphorylation and functions of the RNA polymerase II CTD. Genes Dev. 2006;20:2922–2936. doi: 10.1101/gad.1477006. [DOI] [PubMed] [Google Scholar]
- 7.Evans MJ, Kaufman MH. Establishment in culture of pluripotential cells from mouse embryos. Nature. 1981;292:154–156. doi: 10.1038/292154a0. [DOI] [PubMed] [Google Scholar]
- 8.Conti L, et al. Niche-independent symmetrical self-renewal of a mammalian tissue stem cell. PLoS Biol. 2005;3:e283. doi: 10.1371/journal.pbio.0030283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Zappone MV, et al. Sox2 regulatory sequences direct expression of a b-geo transgene to telencephalic neural stem cells and precursors of the mouse embryo, revealing regionalization of gene expression in CNS stem cells. Development. 2000;127:2367–2382. doi: 10.1242/dev.127.11.2367. [DOI] [PubMed] [Google Scholar]
- 10.Spilianakis CG, Lalioti MD, Town T, Lee GR, Flavell RA. Interchromosomal associations between alternatively expressed loci. Nature. 2005;435:637–645. doi: 10.1038/nature03574. [DOI] [PubMed] [Google Scholar]
- 11.Shen Y, et al. A map of the cis-regulatory sequences in the mouse genome. Nature. 2012;488:116–120. doi: 10.1038/nature11243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Chen X, et al. Integration of external signaling pathways with the core transcriptional network in embryonic stem cells. Cell. 2008;133:1106–1117. doi: 10.1016/j.cell.2008.04.043. [DOI] [PubMed] [Google Scholar]
- 13.Kagey MH, et al. Mediator and cohesin connect gene expression and chromatin architecture. Nature. 2010;467:430–435. doi: 10.1038/nature09380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Hardison RC, Taylor J. Genomic approaches towards finding cis-regulatory modules in animals. Nature Rev. Genet. 2012;13:469–483. doi: 10.1038/nrg3242. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Bernstein BE, et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell. 2006;125:315–326. doi: 10.1016/j.cell.2006.02.041. [DOI] [PubMed] [Google Scholar]
- 16.Rada-Iglesias A, et al. A uniquechromatin signature uncovers early developmental enhancers in humans. Nature. 2011;470:279–283. doi: 10.1038/nature09692. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Li Q, et al. A systematic approach to identify functional motifs within vertebrate developmental enhancers. Dev. Biol. 2010;337:484–495. doi: 10.1016/j.ydbio.2009.10.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Nolis IK, et al. Transcription factors mediate long-range enhancer–promoter interactions. Proc. Natl Acad. Sci. USA. 2009;106:20222–20227. doi: 10.1073/pnas.0902454106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Yu J, et al. Induced pluripotent stem cell lines derived from human somatic cells. Science. 2007;318:1917–1920. doi: 10.1126/science.1151526. [DOI] [PubMed] [Google Scholar]
- 20.Jaenisch R, Young R. Stem cells, the molecular circuitry of pluripotency and nuclear reprogramming. Cell. 2008;132:567–582. doi: 10.1016/j.cell.2008.01.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Ring KL, et al. Direct reprogramming of mouse and human fibroblasts into multipotent neural stem cells with a single factor. Cell Stem Cell. 2012;11:100–109. doi: 10.1016/j.stem.2012.05.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Pollard SM, Wallbank R, Tomlinson S, Grotewold L, Smith A. Fibroblast growth factor induces a neural stem cell phenotype in foetal forebrain progenitors and during embryonic stem cell differentiation. Mol. Cell. Neurosci. 2008;38:393–403. doi: 10.1016/j.mcn.2008.03.012. [DOI] [PubMed] [Google Scholar]
- 23.Ligon KL, et al. Olig2-regulated lineage-restricted pathway controls replication competence in neural stem cells and malignant glioma. Neuron. 2007;53:503–517. doi: 10.1016/j.neuron.2007.01.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Levine M. Paused RNA polymerase II as a developmental checkpoint. Cell. 2011;145:502–511. doi: 10.1016/j.cell.2011.04.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Jinek M, et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. 2012;337:816–821. doi: 10.1126/science.1225829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Li G, et al. ChIA-PET tool for comprehensive chromatin interaction analysis with paired-end tag sequencing. Genome Biol. 2010;11:R22. doi: 10.1186/gb-2010-11-2-r22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Bastian M, Heymann S, Jacomy M. Gephi: an open source software for exploring and manipulating networks. International AAAI Conference on Weblogs and Social Media. 2009 http://www.aaai.org/ocs/index.php/ICWSM/09/paper/view/154 and https://gephi.org/users/publications/
- 28.Meissner A, et al. Genome-scale DNA methylation maps of pluripotent and differentiated cells. Nature. 2008;454:766–770. doi: 10.1038/nature07107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Bolli N, et al. Expression of the cytoplasmic NPM1 mutant (NPMc1) causes the expansion of hematopoietic cells in zebrafish. Blood. 2010;115:3329–3340. doi: 10.1182/blood-2009-02-207225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Siepel A, et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res. 2005;15:1034–1050. doi: 10.1101/gr.3715005. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.