Abstract
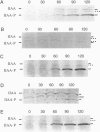
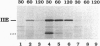
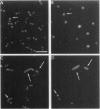
Cell-specific activation of the transcription factor sigma F during sporulation in Bacillus subtilis is controlled by a regulatory pathway involving the proteins SpoIIE, SpoIIAA, and SpoIIAB. SpoIIAB is an antagonist of sigma F, and SpoIIAA, which is capable of overcoming SpoIIAB-mediated inhibition of sigma F, is an antagonist of SpoIIAB. SpoIIAA is, in turn, negatively regulated by SpoIIAB, which phosphorylates SpoIIAA on serine 58. SpoIIAA is also positively regulated by SpoIIE, which dephosphorylates SpoIIAA-P, the phosphorylated form of SpoIIAA. Here, isoelectric focusing and Western blot analysis were used to examine the phosphorylation state of SpoIIAA in vivo. SpoIIAA was found to be largely in the phosphorylated state during sporulation in wild-type cells but a significant portion of the protein that was unphosphorylated could also be detected. Consistent with the idea that SpoIIE governs dephosphorylation of SpoIIAA-P, SpoIIAA was entirely in the phosphorylated state in spoIIE mutant cells. Conversely, overexpression of spoIIE led to an increase in the ratio of unphosphorylated SpoIIAA to SpoIIAA-P and caused inappropriate activation of sigma F in the predivisional sporangium. We also show that a mutant form of SpoIIAA (SpoIIAA-S58T) in which serine 58 was replaced with threonine was present exclusively as SpoIIAA-P, a finding that confirms previous biochemical evidence that the mutant protein is an effective substrate for the SpoIIAB kinase but that SpoIIAA-S58T-P cannot be dephosphorylated by SpoIIE. We conclude that SpoIIE plays a crucial role in controlling the phosphorylation state of SpoIIAA during sporulation and thus in governing the cell-specific activation of sigma F.
Full text
PDF

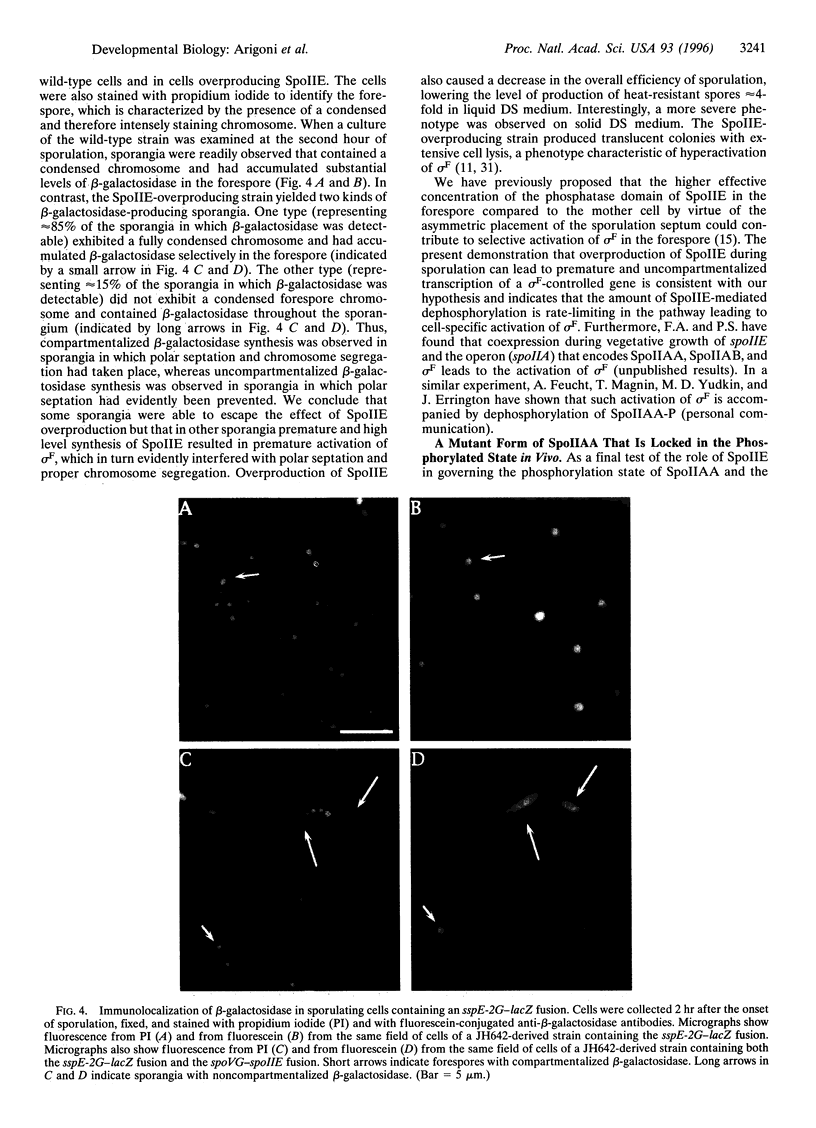

Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alper S., Duncan L., Losick R. An adenosine nucleotide switch controlling the activity of a cell type-specific transcription factor in B. subtilis. Cell. 1994 Apr 22;77(2):195–205. doi: 10.1016/0092-8674(94)90312-3. [DOI] [PubMed] [Google Scholar]
- Arigoni F., Pogliano K., Webb C. D., Stragier P., Losick R. Localization of protein implicated in establishment of cell type to sites of asymmetric division. Science. 1995 Oct 27;270(5236):637–640. doi: 10.1126/science.270.5236.637. [DOI] [PubMed] [Google Scholar]
- Benson A. K., Haldenwang W. G. Characterization of a regulatory network that controls sigma B expression in Bacillus subtilis. J Bacteriol. 1992 Feb;174(3):749–757. doi: 10.1128/jb.174.3.749-757.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boylan S. A., Rutherford A., Thomas S. M., Price C. W. Activation of Bacillus subtilis transcription factor sigma B by a regulatory pathway responsive to stationary-phase signals. J Bacteriol. 1992 Jun;174(11):3695–3706. doi: 10.1128/jb.174.11.3695-3706.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coppolecchia R., DeGrazia H., Moran C. P., Jr Deletion of spoIIAB blocks endospore formation in Bacillus subtilis at an early stage. J Bacteriol. 1991 Nov;173(21):6678–6685. doi: 10.1128/jb.173.21.6678-6685.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Diederich B., Wilkinson J. F., Magnin T., Najafi M., Erringston J., Yudkin M. D. Role of interactions between SpoIIAA and SpoIIAB in regulating cell-specific transcription factor sigma F of Bacillus subtilis. Genes Dev. 1994 Nov 1;8(21):2653–2663. doi: 10.1101/gad.8.21.2653. [DOI] [PubMed] [Google Scholar]
- Dufour A., Haldenwang W. G. Interactions between a Bacillus subtilis anti-sigma factor (RsbW) and its antagonist (RsbV). J Bacteriol. 1994 Apr;176(7):1813–1820. doi: 10.1128/jb.176.7.1813-1820.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duncan L., Alper S., Arigoni F., Losick R., Stragier P. Activation of cell-specific transcription by a serine phosphatase at the site of asymmetric division. Science. 1995 Oct 27;270(5236):641–644. doi: 10.1126/science.270.5236.641. [DOI] [PubMed] [Google Scholar]
- Duncan L., Losick R. SpoIIAB is an anti-sigma factor that binds to and inhibits transcription by regulatory protein sigma F from Bacillus subtilis. Proc Natl Acad Sci U S A. 1993 Mar 15;90(6):2325–2329. doi: 10.1073/pnas.90.6.2325. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Errington J. Bacillus subtilis sporulation: regulation of gene expression and control of morphogenesis. Microbiol Rev. 1993 Mar;57(1):1–33. doi: 10.1128/mr.57.1.1-33.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Errington J., Mandelstam J. Use of a lacZ gene fusion to determine the dependence pattern of sporulation operon spoIIA in spo mutants of Bacillus subtilis. J Gen Microbiol. 1986 Nov;132(11):2967–2976. doi: 10.1099/00221287-132-11-2967. [DOI] [PubMed] [Google Scholar]
- Gholamhoseinian A., Piggot P. J. Timing of spoII gene expression relative to septum formation during sporulation of Bacillus subtilis. J Bacteriol. 1989 Oct;171(10):5747–5749. doi: 10.1128/jb.171.10.5747-5749.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harry E. J., Pogliano K., Losick R. Use of immunofluorescence to visualize cell-specific gene expression during sporulation in Bacillus subtilis. J Bacteriol. 1995 Jun;177(12):3386–3393. doi: 10.1128/jb.177.12.3386-3393.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karmazyn-Campelli C., Bonamy C., Savelli B., Stragier P. Tandem genes encoding sigma-factors for consecutive steps of development in Bacillus subtilis. Genes Dev. 1989 Feb;3(2):150–157. doi: 10.1101/gad.3.2.150. [DOI] [PubMed] [Google Scholar]
- Losick R., Stragier P. Crisscross regulation of cell-type-specific gene expression during development in B. subtilis. Nature. 1992 Feb 13;355(6361):601–604. doi: 10.1038/355601a0. [DOI] [PubMed] [Google Scholar]
- Losick R., Youngman P., Piggot P. J. Genetics of endospore formation in Bacillus subtilis. Annu Rev Genet. 1986;20:625–669. doi: 10.1146/annurev.ge.20.120186.003205. [DOI] [PubMed] [Google Scholar]
- Magnin T., Lord M., Errington J., Yudkin M. D. Establishing differential gene expression in sporulating Bacillus subtilis: phosphorylation of SpoIIAA (anti-anti-sigmaF) alters its conformation and prevents formation of a SpoIIAA/SpoIIAB/ADP complex. Mol Microbiol. 1996 Feb;19(4):901–907. doi: 10.1046/j.1365-2958.1996.434964.x. [DOI] [PubMed] [Google Scholar]
- Margolis P., Driks A., Losick R. Establishment of cell type by compartmentalized activation of a transcription factor. Science. 1991 Oct 25;254(5031):562–565. doi: 10.1126/science.1948031. [DOI] [PubMed] [Google Scholar]
- Min K. T., Hilditch C. M., Diederich B., Errington J., Yudkin M. D. Sigma F, the first compartment-specific transcription factor of B. subtilis, is regulated by an anti-sigma factor that is also a protein kinase. Cell. 1993 Aug 27;74(4):735–742. doi: 10.1016/0092-8674(93)90520-z. [DOI] [PubMed] [Google Scholar]
- Najafi S. M., Willis A. C., Yudkin M. D. Site of phosphorylation of SpoIIAA, the anti-anti-sigma factor for sporulation-specific sigma F of Bacillus subtilis. J Bacteriol. 1995 May;177(10):2912–2913. doi: 10.1128/jb.177.10.2912-2913.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Partridge S. R., Errington J. The importance of morphological events and intercellular interactions in the regulation of prespore-specific gene expression during sporulation in Bacillus subtilis. Mol Microbiol. 1993 May;8(5):945–955. doi: 10.1111/j.1365-2958.1993.tb01639.x. [DOI] [PubMed] [Google Scholar]
- Salamitou S., Lemaire M., Fujino T., Ohayon H., Gounon P., Béguin P., Aubert J. P. Subcellular localization of Clostridium thermocellum ORF3p, a protein carrying a receptor for the docking sequence borne by the catalytic components of the cellulosome. J Bacteriol. 1994 May;176(10):2828–2834. doi: 10.1128/jb.176.10.2828-2834.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Savva D., Mandelstam J. Synthesis of spoIIA and spoVA mRNA in Bacillus subtilis. J Gen Microbiol. 1986 Nov;132(11):3005–3011. doi: 10.1099/00221287-132-11-3005. [DOI] [PubMed] [Google Scholar]
- Schaeffer P., Millet J., Aubert J. P. Catabolic repression of bacterial sporulation. Proc Natl Acad Sci U S A. 1965 Sep;54(3):704–711. doi: 10.1073/pnas.54.3.704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmidt R., Margolis P., Duncan L., Coppolecchia R., Moran C. P., Jr, Losick R. Control of developmental transcription factor sigma F by sporulation regulatory proteins SpoIIAA and SpoIIAB in Bacillus subtilis. Proc Natl Acad Sci U S A. 1990 Dec;87(23):9221–9225. doi: 10.1073/pnas.87.23.9221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stragier P., Bonamy C., Karmazyn-Campelli C. Processing of a sporulation sigma factor in Bacillus subtilis: how morphological structure could control gene expression. Cell. 1988 Mar 11;52(5):697–704. doi: 10.1016/0092-8674(88)90407-2. [DOI] [PubMed] [Google Scholar]
- Sun D., Fajardo-Cavazos P., Sussman M. D., Tovar-Rojo F., Cabrera-Martinez R. M., Setlow P. Effect of chromosome location of Bacillus subtilis forespore genes on their spo gene dependence and transcription by E sigma F: identification of features of good E sigma F-dependent promoters. J Bacteriol. 1991 Dec;173(24):7867–7874. doi: 10.1128/jb.173.24.7867-7874.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zuber P., Losick R. Use of a lacZ fusion to study the role of the spoO genes of Bacillus subtilis in developmental regulation. Cell. 1983 Nov;35(1):275–283. doi: 10.1016/0092-8674(83)90230-1. [DOI] [PubMed] [Google Scholar]