Abstract
Cardiometabolic diseases can be present long before becoming clinically apparent. Accurate predictors of disease are of particular importance since the delay or prevention of morbidity is possible via pharmacological and behavioral interventions. Metabolomics is increasingly applied to biomarker discovery. Understanding how metabolites relate with established cardiometabolic risk factors is critical in evaluating their potential value as clinical biomarkers. Large epidemiological cohort studies can assess whether metabolite biomarkers improve upon existing disease markers. Furthermore, experimental work in model systems and integration with other functional genomic approaches will facilitate the discovery of causal links between select biomarkers and disease pathogenesis.
The need for new cardiometabolic biomarkers
Cardiometabolic diseases are difficult for physicians to manage because they can be present for years before becoming clinically apparent. For example, patients presenting with acute coronary syndromes due to focal plaque instability often have extensive subclinical atherosclerosis in other vascular beds. Similarly, by the time the diagnosis of type 2 diabetes mellitus (DM) is made based on hyperglycemia, significant β-cell insufficiency has often occurred. Although knowledge of conventional risk predictors can help in detection of future disease, prediction remains imperfect, mirroring our incomplete understanding of underlying mechanisms. Despite strong statistical associations with outcomes, new biomarkers often contribute less than expected to the accuracy of risk models because many report on the same pathways as pre-existing disease markers. As highlighted in this review, there is a mandate for “orthogonal” (i.e., uncorrelated) disease biomarkers that provide additional clinical information along new biological axes. A robust set of predictors for identifying at risk individuals is of particular importance because the delay or prevention of diseases such as atherosclerosis or DM is now possible via pharmacological interventions such as statins and metformin, as well as by behavioral approaches including weight loss and exercise interventions. New predictors may also identify patients likely to respond to specific interventions, towards a goal of more personalized medicine (Losalzo, 2012).
One avenue for identification of novel risk markers is being opened by the global analysis of the human metabolome. While decades of research in biochemistry, nutrition, and physiology have revealed specific metabolic pathways, systematic surveys of pathways altered in human disease states such as atherosclerosis or DM are now possible. An emerging set of metabolite profiling capabilities, based on techniques such as mass spectrometry and nuclear magnetic resonance (NMR) spectroscopy, enable the monitoring of hundreds of analytes from biological samples (“metabolomics” or “metabonomics”). These technologies promise to transform our ability to profile samples with the goal of illuminating biology and discovering valuable clinical biomarkers.
From a clinical perspective, biomarkers can serve a variety of functions, corresponding to different stages in disease evolution. Biomarkers can assist in the care of patients without apparent disease (screening biomarkers), with suspected disease (diagnostic biomarkers), or with progression or remission of overt disease (prognostic biomarkers). Biomarkers that participate in disease pathways are the most likely candidates for disease surrogates – substitutes for a clinical endpoint. A surrogate endpoint might predict clinical benefit (or harm) of a given therapeutic with an ultimate goal of facilitating the drug discovery process.
In the case of coronary heart disease, the majority of individuals have only one or none of the classic cardiovascular risk factor (Khott et al. 2003), which underscores in particular the need for screening biomarkers. The majority of current biomarkers for cardiovascular screening fall along pathways, such as inflammation and cholesterol biosynthesis, already known to be associated with cardiovascular disease. Similarly, many biomarkers for diabetes solely interrogate glucose homeostasis. Consequently, available biomarkers often provide information that is correlated with what is already known or being measured. Although correlated biomarkers can underscore the importance of a biological pathway, they may not provide a substantial increase in diagnostic or predictive value.
This point has been highlighted by recent investigations. Wang and colleagues explored factors that influence whether new biomarkers can add to diagnosis and risk prediction above and beyond established markers (Wang et al. 2006). They performed a simulation based on adding 1 to 100 hypothetical biomarkers to a traditional risk model for cardiovascular disease (Figure 1). When assessing biomarker sensitivity and specificity traditional risk models (including cholesterol concentrations, age, and hypertension) have an approximate area under the curve (AUC) of 0.75 across populations. For their simulation, each of the hypothetical biomarkers had a similar magnitude of association with cardiovascular events similar to established cardiovascular biomarkers such as C-reactive protein (CRP) or B-type natriuretic peptide (nBNP). They demonstrated that a key determinant of improvement in the c statistic is the degree of correlation between biomarkers. With a set of biomarkers that has a mean marker-marker correlation of r = 0.4 (moderately correlated), >50 biomarkers are needed before the c statistic is increased by 0.05. By contrast, when the average marker-marker correlation is r = 0.05 (weakly correlated), <10 biomarkers are needed to raise the c statistic by 0.05. Thus, new methodologies must be brought to bear to identify signals across different physiological axes to improve disease prediction.
Figure 1.
Increment in discrimination from adding hypothetical biomarkers, according to the degree of marker-marker correlation (r). The simulated hazards ratio for the outcome is 1.35 per SD increment in the biomarker. The y axis shows the C statistic from a model containing traditional risk factors plus a variable number of simulated biomarkers (x axis), each with a fixed association with the outcome.
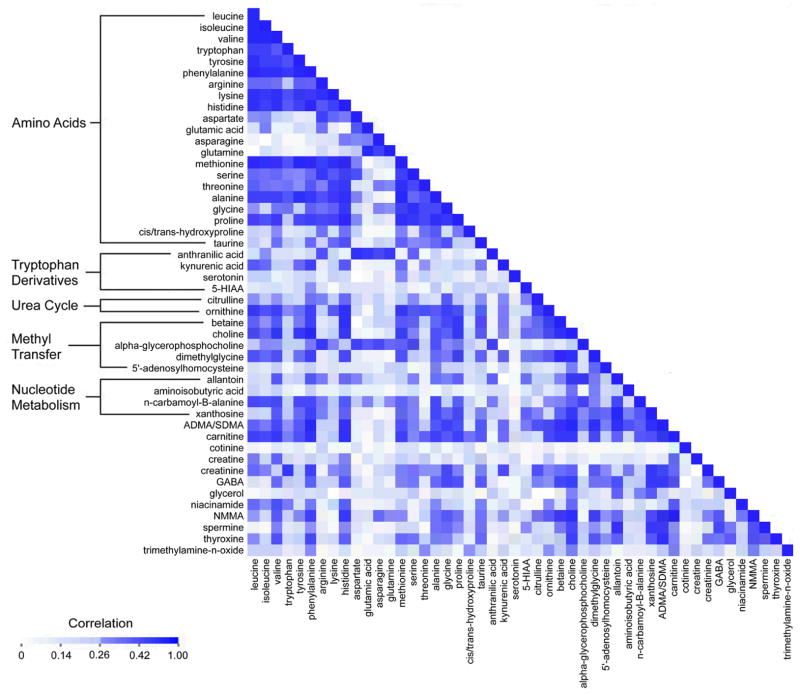
To illustrate the potential contribution of metabolite profiling to clinical biomarker discovery, Figure 2 depicts the correlations for 48 plasma metabolite concentrations in approximately 200 fasting individuals in the population based Framingham Heart Study. Mean correlations within groups of molecules were highest for urea cycle metabolites (r = 0.49), metabolites involved in nucleotide metabolism (r = 0.38), amino acids (r = 0.34) and methyl transfer metabolites (r = 0.34). Correlations across other metabolites or groups of metabolites were far weaker. As more metabolites are measured in large cohorts, similar correlation clusters, both expected and unexpected, are likely to emerge. Understanding how metabolites relate with each other and with established risk markers will be important in assessing their value as potential biomarkers and may also lead to previously unappreciated connections between metabolic pathways.
Figure 2.
Intercorrrelation of circulating metabolites in 200 normal individuals from the Framingham Heart Study.
Incorporation of Metabolomics Platforms for Biomarker Discovery
The limitations of currently available biomarkers for screening thus underscores the importance of identifying “uncorrelated” or “orthogonal” biomarkers associated with novel disease pathways. Emerging technologies are beginning to permit the systematic, unbiased characterization of variation in genes, RNA, proteins, and metabolites associated with disease conditions. Interrogation of metabolites as disease markers is particularly attractive since small molecules are most proximal to any disease phenotype.
During the 1970’s, Arthur Robinson and Linus Pauling postulated that the quantitative and qualitative pattern of metabolites in biological fluids reflected the functional status of the complex biological system from which they were derived (Pauling et al., 1971). Concomitantly, the term metabolic profiling was introduced to describe data obtained from gas chromatography analysis of a patient sample (Horning and Horning, 1971). Seminal studies of inborn errors of metabolism in infants can also be used to inform the application of metabolite profiling to human diagnostics. Millington and colleagues pioneered the use of mass spectrometry-based methods for monitoring fatty acid oxidation, as well as organic and selected amino acids (Roe et al. 1986). In many instances, rapid identification of inborn errors allows for dietary modulation and amelioration of symptoms or disease onset. It is anticipated that a global metabolomic analysis of aberrant pathways in more common diseases such as diabetes or atherosclerosis might similarly spotlight new biomarkers or pathways for dietary or drug modulation.
The metabolomics approach, built upon decades of fundamental research in biochemistry, aims to measure the metabolites present within a cell, biofluid, tissue or organism at baseline or in the context of a perturbed state, such as a genetic alteration or a physiological or environmental stimulus (Oliver et al., 1998; Nicholson et al., 1998; Fiehn, 2001). Metabolomics was initially employed to identify seemingly “silent phenotypes” in model systems such as yeast. That is, biochemical profiling techniques were used to identify the downstream functions of genes that when perturbed have no apparent influence on physical characteristics or behavior on initial routine evaluation (Griffin et al., 2002). The reason for this powerful analytical advantage becomes apparent when we consider that the metabolome is the terminal downstream product of the genome and consists of the total complement of all the low molecular weight molecules (metabolites) in a cell, biofluid, tissue or organism required for growth, maintenance or normal function in a specific physiological state (Goodacre, 2003; Goodacre et al., 2004). It could be envisioned that metabolic perturbations developing prior to the onset of disease symptoms (a silent phenotype) could be used as biomarkers for screening at risk populations, or that a particular metabolic profile corresponds to specific disease states or progression.
The metabolome itself represents an incredible variety of chemical classes and physical properties ranging from the small polar hydrophilic volatiles to large complex hydrophobic lipids. The dynamic range of metabolite concentrations exists across orders of magnitude, varying from the picomolar to the millimolar. This variability in chemical property and dynamic range provides a considerable challenge for metabolomics, especially viewed in light of other “omic” techniques such as genomics and transcriptomics where the chemical variations in the analytes (DNA or mRNA respectively) are minimal in comparison. To optimize coverage of the metabolome the use of an extensive array of analytical techniques within the field of metabolomics is necessary.
Metabolomic Analytical Techniques
The principle profiling technologies utilized in metabolomic analysis are NMR spectroscopy and mass spectrometry, which is often coupled to a prior chromatographic technique, most commonly liquid or gas chromatography (LC or GC). However, with an estimated 5000 currently detectable human serum metabolites (a number likely to increase with advancement in technology) (Wishart et al, 2013), a comprehensive metabolomic map remains an unattained goal. Using a combined analytical approach one can build a more inclusive picture of the metabolome by overcoming the limitations of individual techniques (Table 1).
Table 1.
Metabolomics technologies.
| Chromatography | None | GC | • Ideal for volatile nonpolar analytes |
| LC | • Ideal for ionizable analytes in solution | ||
| Other | • Direct infusion (no chromatography) | ||
| • Capillary electrophoresis | |||
| • Thin-layer chromatography | |||
| ↓ | ↓ |
| MS | ||||
|---|---|---|---|---|
| Analytical method | NMR | TOF | Triple quadrupole | Ion trap |
| Metabolite identification | Chemical shift | m/z (Flight time) | m/z (Filters for m/z) | m/z (Trapping frequency) |
| Advantages | • Robust | • Mass accuracy | • Sensitivity | • Sensitivity |
| • No sample destruction | • Wide mass range | • Can perform tandem MS | • Can perform tandem MS | |
| • No chromatography or analyte ionization | • Dynamic range | |||
| • Unambiguous identification of abundant analytes | ||||
| Disadvantages | • Limited sensitivity | • Limited dynamic range | • Poor mass accuracy | • Limited dynamic range |
| • Isotopic standards required for absolute quantitation | • Isotopic standards required for absolute quantitation | • Isotopic standards required for absolute quantitation | ||
1H-Nuclear Magnetic Resonance Spectroscopy in Metabolomics
The use of 1H-NMR spectroscopy to simultaneously detect an array of metabolites in biological fluids was pioneered during the early 1980’s (Nicholson et al., 1983; Nicholson et al., 1984; Nicholson et al., 1984b; Bales et al., 1984). NMR utilizes the magnetic properties of select atomic nuclei (e.g. 1H, 13C, or 31P) to determine the structure and abundance of metabolites in a biological specimen. In a strong magnetic field, a given NMR-active nucleus absorbs electromagnetic radiation at a characteristic frequency. Since this NMR signal is influenced in an identifiable way by the nature of neighboring atoms, chemical shifts in its resonance frequency can be used to assign local molecular structure. By measuring all frequencies, metabolite identity can be determined, although this is not feasible for all compounds in a complex mixture. The popularity of NMR spectroscopy arises from the technique’s spectroscopic, quantitative, robust and reproducible nature and its low cost per sample analysis (Bothwell et al. 2011). It is non-destructive, permitting multiple analyses of the same sample. NMR is capable of absolute quantification without the use of isotope labeled standards. Although the majority of NMR-based metabolomics experiments are performed on samples in the solution state, NMR has a further advantage; through the use of high resolution magic angle spinning NMR, metabolites can be measured within intact tissue samples (Moestue et al., 2011). However, a major criticism of NMR spectroscopy in metabolomic analyses is that, compared to many other techniques, it suffers from relatively poor sensitivity in the analysis of low abundance metabolites. The relative lack of sensitivity in NMR analysis has limitations when it comes to biomarker discovery where alterations in low abundance metabolites may be missed or obscured by signals from species at a higher abundance. The analysis of blood plasma metabolite extracts using 1H-NMR typically detects 20–50 metabolites (Rooney et al., 2003; Griffin et al., 2001). However, when applied to urine the number of metabolites observed may be double that number (Griffin et al., 2004). High-resolution NMR using stronger magnetic fields or two-dimensional NMR permits higher information content and increased analytical sensitivity especially when combined with the use of cryoprobes, but at the expense of increased cost and throughput.
Mass Spectrometry in Metabolomics
The field of metabolomics has been strongly influenced by recent developments in mass spectrometry. Mass spectrometry provides effective and versatile analyte separation, and is highly sensitive especially when compared to other metabolomic techniques such as NMR, facilitating the analysis of hundreds, and potentially thousands, of metabolite species (Schiller et al., 2000). As discussed below, mass spectrometric analysis integrates several distinct events, including chromatographic separation, analyte ionization, and mass dependent ion separation.
Chromatography
It is possible to infuse samples directly into the mass spectrometer, a technique which is sometimes described as a “shotgun” approach (Han and Gross, 2005). However, most metabolomics analyses employ an upfront chromatographic time-dependent separation of the molecular species within each biological sample to facilitate metabolite identification and quantification. These chromatographic processes involve a mobile solvent phase into which the analyte mixture is dissolved. The mobile phase passes through a column containing surfaces coated in specific interaction chemistries, the stationary phase. The individual chemical species dissolved within the mobile phase are retained in the column, or pass through at varying speeds, depending on their interaction with the chemistry of the stationary phase. By gradually altering the composition of the mobile phase the interactions between stationary phase chemistry and analyte species can be disrupted and the analytes eluted from the column. The chemical properties of the analyte species results in differential retention on the stationary phase, and therefore separation of compounds.
The primary techniques employed are gas chromatography (GC) and liquid chromatography (LC). GC vaporizes the analytes and separates them in the gas phase, therefore it requires the analyte species to be non-polar volatiles. A relatively time consuming and laborious pre-chromatography chemical derivatization step is included to form volatile derivatives of the species of interest. For example, fatty acids can be derivatized to fatty acid methyl esters. By contrast, LC separates the analytes species in the liquid phase and is therefore suited for the analysis of thermally labile, polar and high molecular mass compounds in solution. A variety of LC column chemistries are available facilitating the analysis of a range of metabolites with differing chemical properties. Non polar columns with carbon chain chemistries are especially well suited for the analysis of lipid species, whereas polar chemistries as found in hydrophilic interaction chromatography columns are appropriate for the separation of small charged metabolite species such as amino acids, nucleotides and organic acids. The duration of chromatography methods tends to be under an hour, with most analyses lasting between 15 – 30 minutes per sample for each column depending on the class of metabolite measured.
Ionization
The foremost methods of analyte ionization in metabolomic mass spectrometry are electrospray ionization (ESI) and electron impact (EI) ionization. EI ionization is used almost exclusively in combination with GC as it requires the analyte to be volatile. In EI, high energy electrons bombard the volatile analyte causing it to fragment. The fragmentation generates a reproducible pattern of signals, which can be matched to spectral databases, such as the National Institute of Standards and Technology Mass Spectral Library, for the purpose of identification.
ESI is the most commonly applied ionization technique in metabolomic mass spectrometry since it facilitates ionization of non-volatiles in solution, making it optimal for coupling an analytical LC to a mass spectrometer for upfront molecular separation of complex biological fluids (Wong et al., 1988). Briefly, in ESI the analyte is eluted through a highly charged needle tip. The resultant charged analyte spray droplets are heated, evaporating the solvent and resulting in ionization, where they can then be introduced to the mass separation unit (Kebarle and Ho, 1997). The capabilities of different ionization modes vary in their ability to measure a range of metabolite species. In negative mode there is a preference for anionic species to ionize (eg, organic acids), while positive mode detects neutral and cationic metabolites (eg., protonated amino acids and amines).
While ESI is now the backbone of MS analysis, one disadvantage that occurs when analyzing complex biological molecular mixtures is ion suppression (Petkovic et al., 2001). Ion suppression occurs when analytes compete for charge during the ionization process, in which an individual metabolite’s ionization efficiency is based on its chemical and physical characteristics (Knochenmuss, 2003). Therefore, the observed relative abundance for a particular ion can change, depending on the other analytes or contaminants co-ionized in the matrix, even when the interfering compound is not detected in the mass spectrum (Wenk et al., 2003). Thus, metabolomic mass spectrometric analysis must assume that mixtures are roughly the same between the groups of samples analyzed. Inferences with regards to absolute concentrations of metabolites across tissue matrices must be considered preliminary. In the clinical setting, the interfering compound could take the form of a drug present in the circulation in a subset of the study population but not present in the total population. The upfront chromatographic separation in LC and GC prior to mass spectrometric analysis functions to reduce ion suppression effects, but may not eliminate the problem entirely (Peterson and Cummings, 2006).
Mass Analysis
Ionic separation in metabolomic mass spectrometry can be achieved by a variety of techniques including time-of-flight (TOF), quadrupole, and ion trap mass analyzers (Downard, 2004). The specifications of the analyzers vary across dynamic range, resolution, capability for tandem-MS (MS/MS) and mass accuracy. One limitation is the direct correlation between the resolution of an instrument and its cost. To an extent, lower resolution and mass accuracy can be compensated for by MS/MS capability. MS/MS is the use of two or more stages of mass analysis to selectively examine the fragmentation of an ion within a mixture of ions (de Hoffman 1996). MS/MS thus isolates individual mass-to-charge ratio (m/z) species for fragmentation. In a process known as collision induced dissociation (CID), the molecular ions are accelerated by an electrical potential increasing their kinetic energy. The ions then collide with neutral gas molecules such as helium or nitrogen within a collision cell in the mass spectrometer. As the molecules collide, the kinetic energy is converted into internal energy, resulting in the breakage of chemical bonds and the fragmentation of the molecular ion. The fragments are then separated by m/z in a second mass analyzer. The m/z values of the fragments aid the identification of the isolated ions. For example, phospholipid headgroups give characteristic fragment patterns in the form of a neutral loss or a specific fragment ion (Han and Gross, 2005). The speed, relative robustness and potential for automation of this technique have been extensively exploited. As the range of mass spectrometers available has increased so too has their application to metabolomics research.
Time of flight (TOF) instruments offer a high mass accuracy and wide m/z range. Ions are accelerated by the application of a fixed electric field through a flight tube of know length. The time it takes for an ion to reach a detector at the end of the flight tube is measured. Since the electric field is fixed, all ions with the same charge gain the same kinetic energy. However, since heavier particles travel at a lower velocity, and conversely lighter particles at a higher velocity, the m/z ratio can be determined from the time-of-flight. In metabolomics experiments it is more common to see instruments where TOF is combined with another upfront mass analyzer, typically a quadrupole (QTOF) or an ion trap instrument as discussed below. The upfront analyzer offers the ability to actively select ions of interest and to fragment them for the purposes of identification while the TOF allows for high resolution and mass accuracy.
Quadrupole mass spectrometers apply oscillating electrical fields between four parallel rods to selectively stabilize the flight path of ions of a particular m/z, thus serving as a mass-selective filter. The potentials across the rods can be adjusted to sweep across a range of m/z values, or to settle only on select ion masses of interest. Although mass accuracy and m/z range are generally inferior compared to TOF instruments, the linear dynamic range for relative quantification is generally better with quadrupole instruments. Further, in triple quadrupole mass spectrometers, the first quadrupole can act as a mass filter for select ‘precursor’ ions, the second quadrupole can be used as a collision cell to allow collision-induced dissociation (CID) of these precursor ions, and the third quadrupole can either allow all fragments to pass to the detector or serve as a mass filter for a single ‘product’ fragment. Operated in the latter mode, triple quadrupole instruments permit MS/MS spectrometry, with subsequent improvements in both the analytical sensitivity and specificity for preselected analytes of interest, which can include as many as 300 metabolites
Ion trap mass spectrometers, which can have different structural configurations, apply an electrical field to accumulate and hold ions of a specific m/z. Trapped metabolites can then be ejected from the ion trap, or fragmented in the ion trap for tandem MS analysis; m/z is determined on the basis of the radio frequency potential required to retain ions in the trap. By accumulating molecules of interest, ion traps maximize analytical sensitivity, but provide less robust quantification across a narrower dynamic range.
Orbitrap mass analyzers were first described in 2000 (Makarov, 2000). The mass analyzers radially trap ions around a central electrode. An outer electrode positioned coaxially with the inner electrode allows m/z values to be determined from the frequency of harmonic ion oscillations of the ions trapped in orbitals. The axial frequency of the ions depends on their energy and dispersion in space. Ion frequencies are measured and a mass spectrum obtained using fast Fourier transforms. Orbitraps achieve high mass resolution, high mass accuracy and a robust dynamic range.
Instruments that combine mass analyzers are available, e.g. quadrupole-TOF and ion trap-TOF instruments, in which the upstream module can be used to select and fragment ions of interest, and the downstream TOF component provides high resolution mass spectra. Further, alternative methodologies for m/z discrimination are available, including Fourier transform ion cyclotron resonance mass spectrometers. To date, these instruments have been applied less frequently for human biomarker discovery.
Untargeted vs. Targeted Metabolic Profiling
Both NMR and MS can be used to characterize metabolite data either in a targeted manner, or in a non-targeted, pattern recognition manner. The approaches have their own inherent advantages and disadvantages, but when used in combination can be highly complementary. Untargeted metabolomics is the attempt to measure all the analytes in a sample, including chemical unknowns. Due to its comprehensive nature, analysis of untargeted metabolomic data must be conducted with advanced chemometric techniques such as multivariate analysis, designed to reduce the extensive datasets generated into smaller, more manageable signals. These unidentified signals then require annotation using either in silico libraries or experimental identification using analytical chemistry with standards. From a perspective of biomarker discovery, untargeted analysis offers the tantalizing opportunity for novel species discovery, and therefore the identification of unique, specific and accurate biomarkers.
In untargeted analysis, coverage of the metabolome is unbiased and only restricted by the methodologies of sample preparation and the inherent sensitivity and specificity of the analytical techniques employed. However, the untargeted approach does suffer from a number of limitations; the protocols and time constraints required to process the extensive raw data sets generated, the analytical chemistry challenges of identifying and characterizing unknown metabolites, the time demands of identifying unknown metabolites that can run to weeks and months, the over-reliance on the innate analytical coverage of the profiling technique utilized, and a strong bias towards detection of highly abundant small molecule species can all have detrimental effects.
Targeted metabolomics, on the other hand, is the measurement of distinct sets of characterized and chemically annotated small molecules within a biological sample. Absolute quantification of metabolite levels using mass spectrometry can be performed via the standard addition method. Internal standards, typically isotopically labeled versions of the endogenous metabolite, are spiked into the matrix across a range of concentrations to generate a standard curve. The labeled standards, which often contain 13C or 2H isotopes, can be easily differentiated from the endogenous metabolite of interest due to the isotopic difference in mass. It is also imperative that the isotope-labeled standard is pure to avoid interference with, and modulation of, the endogenous metabolite signal. The targeted approach exploits the extensive a priori knowledge of a vast array of metabolites and the metabolic pathways to which they belong in order to select specific species for analysis. By analyzing specific key metabolites across all characterized metabolic pathways, these metabolites can function as sentinels to identify perturbations in the pathways, highlighting them for further analysis. When performing targeted metabolomics on a mass spectrometer the instrument is more sensitive since it is measuring fewer compounds. Furthermore, the sample preparation step can be modified to reduce the over representation of abundant molecules in the analyses. From a biomarker perspective, this can facilitate the association of low abundance metabolites with disease states. While their unambiguous identification is known a priori, their association with a given phenotype may be novel. If a particular pathway appears to be highlighted in the analyses, additional metabolites in the pathway can be added to the mass spectrometric metabolomic assay for a more detailed interrogation. In addition, since all analyzed species are clearly defined, analytical artifacts are not carried through to downstream analyses, and laborious chemical classification steps are not required.
Applications of Metabolomics to Biomarker Discovery
In an initial proof of principle study, Newgard and colleagues profiled obese versus lean humans to gain a broad understanding of the metabolic and physiologic differences in these two disparate groups. Their studies identified a branched-chain amino acid signature that was highly correlated with metrics of insulin resistance (Newgard et al., 2009). In complementary studies in two large population based cohorts, branched chain and aromatic amino acid concentrations were demonstrated to have a significant association with future type 2 diabetes, up to 12 years before the onset of overt disease (Wang et al., 2011). These findings were essentially unchanged after adjustment for established clinical risk factors. The initial discovery experiment utilized a nested, case-control design in the Framingham Heart Study (FHS) with approximately 200 individuals who ultimately developed DM and a similar number of control individuals who were matched for age, body mass index, and fasting glucose who did not develop DM. A validation experiment in the Malmo Diet and Cancer study employed the same matching scheme. Since the study design enriched the control population for high-risk features such as obesity and increased fasting glucose, analyses were also performed on randomly selected controls as well. In a distinct study, similar findings were obtained in a much larger population of Finnish men (Stancakova et al., 2012). Although the association between metabolites and incident diabetes was significant across each experiment, the strength of association was attenuated in the case versus random cohort analysis. Whereas amino acid increases led to large improvements in model fit and discrimination (c statistics) between cases and matched controls, changes in these parameters were less significant between cases and random controls. These findings suggest that amino acid profiling might have greater value in high-risk individuals, and underscores how study design modulates study outcome and interpretation; although metabolomics technologies are new, old lessons in epidemiology still apply.
Limitations to Human Biomarker Studies
Several limitations to metabolomic biomarker discovery efforts in humans warrant consideration. While serum or plasma samples from peripheral blood are easily obtained or already available in many cohorts, the circulation integrates the inputs of multiple organ systems making the source of a given signal difficult to ascertain. It is notable, however, that metabolomics profiling can measure metabolites in plasma typically thought to be confined to intracellular compartments based on their phosphorylation state or site of bioactivity. For example, while intracellular TCA cycle and adenine nucleotide changes in muscle during exercise are well characterized from animal tissues or even human tissue punch biopsies, high sensitivity LC-MS platform have also been used to examine whether these changes occur in plasma in a fitness-dependent way (Lewis et al., 2010). In certain unique human conditions, selective catheterization across tissues beds can be performed, for example by simultaneously acquiring coronary sinus or pulmonary artery and peripheral arterial blood samples. Of course, some metabolic abnormalities in a given tissue bed may remain “silent” in the peripheral blood until severe organ dysfunction has occurred.
Although capable of generating metabolic ‘snapshots’, metabolite profiling of clinical plasma samples does not provide information on pathway flux – for example, it is unable to determine whether a metabolite is increased because it is being produced in excess or because of a downstream inhibition. To unambiguously define the rate of flux of a metabolic pathway stable isotope labeled substrates can be employed (Roberts et al., 2011). In flux analysis the incorporation of the label, for example 13C, into metabolites downstream of the labeled substrate in a metabolic pathway can be measured using mass spectrometry or NMR to determine the metabolic rate.
Further, the development of clinically useful biomarkers will require a more comprehensive understanding of how exogenous factors impact the metabolome, e.g. how do diet and medications impact metabolite markers of interest, and what is the normal intra- and inter-individual variability for these metabolites? Finally, although adequately powered clinical studies enable strategies to minimize confounding, e.g. multivariable adjustment or population stratification by known risk factors, they are unable to establish a causal link between disease biomarkers and disease pathogenesis.
While a growing number of studies have used mass spectrometry as a tool for biomarker discovery, many of the studies have been cross-sectional, providing limited information regarding the relation of metabolomic (or proteomic) biomarkers to the future development of disease. Long periods of observation are also important features of a good study design, because it enables investigators to identify changes in metabolites that occur well before any secondary alteration from the overt disease process.
From Association to Causation: integrating human and animal studies
To investigate causal relationships, metabolomic findings derived in human studies can be further interrogated in model systems. Studies in rodents have extended the clinical observations of elevated branched chain amino acids in prediabetes. To test the impact of BCAA on metabolism, Newgard and colleagues placed rats on diets consisting of standard chow, high-fat, or high fat supplemented with BCAA (Newgard et al. 2009). Despite having food intake and weights equivalent to the standard chow group, high-fat and BCAA fed mice were as insulin resistant as the high-fat fed animals. The underlying cellular mechanisms may include activation of the mTOR, JUN and IRS1 signaling pathways in skeletal muscle. While not all studies of branched chain amino acid supplementation in murine systems have led to identical results, future exploration of this pathway is clearly warranted.
In a translational study utilizing non-targeted LC-MS based metabolite profiling, Wang et al. first profiled plasma from 75 individuals from a hospital based cohort who experienced a myocardial infarction, stroke, or death in the ensuing three years and 75 age- and gender-matched controls that did not (Wang, Z. et al. 2011). Of 18 analytes that were significantly different between cases and controls, three demonstrated significant correlations among one another, suggesting a potential common biochemical pathway. Using complementary analytical methods, these metabolites were identified as betaine, choline, and trimethylamine-N-oxide, all metabolites of dietary phosphatidylcholine. Dietary supplementation of choline was sufficient to promote atherosclerosis in mice, and suppression of intestinal bacteria responsible for the conversion of phosphatidylcholine to choline inhibited this atherogenesis. In addition to reinforcing the interaction between diet, gut bacteria, and the metabolome, this study demonstrates how metabolomic biomarker discovery can elucidate novel pathways to disease.
From Association to Causation: integrating genetic findings
Integrating genomic and metabolomic data in humans represents an alternative strategy to establish a causal link between metabolite biomarkers and disease. For a biomarker that has a causal role, the expected random distribution in a population of a polymorphism that determines high or low biomarker concentrations would be skewed in individuals depending on their disease status. The association between a disease or a metabolite and a genetic polymorphism that mimics the biological link between a proposed exposure and disease is not confounded by the reverse causation affecting many epidemiological observational studies. Data from so-called “Mendelian randomization” studies are accumulating for several biomarkers. Analyses demonstrating that single nucleotide polymorphisms that modulate plasma LDL cholesterol concentrations are independently associated with incident cardiovascular disease is consistent with the known causal role LDL cholesterol plays in atherogenesis (Kathiresan et al., 2008). By contrast, genetic loci associated with plasma C-reactive protein concentrations (including in the CRP locus) had no association with coronary heart disease, arguing against a causal association between C-reactive protein and cardiovascular disease (Elliott et al., 2009).
Recent work has begun to explore the genetic determinants of plasma metabolite levels in large human cohorts. One of the first publications to successfully combine Genome-Wide Association Studies (GWAS) with metabolomics analyzed 363 metabolites in fasting serum from 284 males enrolled in the KORA study using LC-MS (Gieger et al., 2008). GWAS analysis of these individuals demonstrated that associations between SNPs and metabolite concentrations accounted for 12% of the observed variance in metabolism overall. Illig et al, in a larger scale study of 1809 individuals also from the KORA study, combined GWAS with an LC-MS based metabolomics analysis of 169 serum metabolites to demonstrate that SNPs located in or near a range of genes accounted for 5.6 – 36.3% of variance in metabolite concentrations (Illig et al 2010). The data was validated in a population of 422 individuals from the Twins UK study. This approach has also been applied to the urine metabolome. Suhre et al published a study analyzing GWAS in relation to 59 metabolite concentrations measured by NMR in urine samples from two large cohorts (Suhre et al., 2011). Many of the findings in these studies highlight locus-metabolite associations with strong biological plausibility. For example, the locus of interest includes a gene encoding a protein directly responsible for the metabolism or transport of the given metabolite. However, many other locus-metabolite associations have been identified in these studies without a clear biochemical relationship with the given metabolite, providing strong motivation for future investigation, particularly when the metabolite of interest is associated with a disease phenotype. An alternative approach that can be taken is to begin with disease associated SNPs, and then integrate the NMR or MS data to begin to understand the metabolic consequences of the human variant of interest. Despite these advances, however, convincing links between SNPs, metabolite levels and disease states remain to be validated. As ongoing studies further delineate the genetic determinants of plasma metabolite profiles, efforts to “triangulate” gene-metabolite-disease associations will provide insight into if and how metabolite markers contribute to disease pathogenesis – studies which can be further buttressed by pathway modulation in animal models.
Conclusions
Emerging technologies now permit higher resolution phenotyping of biological specimens. Whereas robust technologies capable of genomic and transcriptomic profiling are now well established, no single analytical method provides comprehensive coverage of the human metabolome. Thus, investigators have employed various technologies, including NMR and MS (coupled to either upfront LC or GC), and utilized alternative analytical strategies, i.e. targeted versus pattern recognition. Metabolite profiling has demonstrated feasibility in large epidemiologic human cohorts. However, further efforts to understand the diversity of inputs to the metabolome will be an important resource for future metabolomic biomarker studies. In parallel, investigation in model systems and integration with other functional genomic approaches in humans will provide insight into the pathophysiologic interactions between metabolite markers and disease.
Acknowledgments
We would like to acknowledge our research support from the NIH R01-DK-HL081572, R01 HL098280. NIH-1U01HL107440, a Leducq Foundation Transatlantic Network Grant, and the American Heart Association. Dr. Roberts is supported by a Leducq Foundation Career Development Award.
References
- Bales JR, Higham DP, Howe I, Nicholson JK, Sadler PJ. Use of high-resolution proton nuclear magnetic resonance spectroscopy for rapid multi-component analysis of urine. Clin Chem. 1984;30:426– 32. [PubMed] [Google Scholar]
- Bothwell JHF, Griffin JL. An introduction to biological nuclear magnetic resonance spectroscopy. 2011;86:493–510. doi: 10.1111/j.1469-185X.2010.00157.x. [DOI] [PubMed] [Google Scholar]
- Downard K. Mass Spectrometry - A Foundation Course. Cambridge UK: Royal Society of Chemistry; 2004. [Google Scholar]
- Elliott P, Chambers JC, Zhang W, Clarke R, Hopewell JC, Peden JF, Erdmann J, Braund P, Engert JC, Bennett D. Genetic Loci associated with C-reactive protein levels and risk of coronary heart disease. JAMA. 2009;302:37– 48. doi: 10.1001/jama.2009.954. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fiehn O. Combining genomics, metabolome analysis, and biochemical modelling to understand metabolic networks. Comp Funct Genomics. 2001;2:155–68. doi: 10.1002/cfg.82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gieger C, Geistlinger L, Altmaier E, Hrabe de Angelis M, Kronenberg F, Meitinger T, Mewes HW, Wichmann HE, Weinberger KM, Adamski J. Genetics meets metabolomics: a genome-wide association study of metabolite profiles in human serum. PLoS Genet. 2008;4:e1000282. doi: 10.1371/journal.pgen.1000282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goodacre R. Metabolic Profiling: Its role in Biomarker Discovery and Gene Function Analysis. Kluwer Academic Publishers; London: 2003. [Google Scholar]
- Goodacre R, Vaidyanathan S, Dunn WB, Harrigan GG, Kell DB. Metabolomics by numbers: acquiring and understanding global metabolite data. Trends Biotechnol. 2004;22:245–252. doi: 10.1016/j.tibtech.2004.03.007. [DOI] [PubMed] [Google Scholar]
- Griffin JL, Nicholls AW, Keun HC, Mortishire-Smith RJ, Nicholson JK, Kuehn T. Metabolic profiling of rodent biological fluids via 1H NMR spectroscopy using a 1 mm microlitre probe. Analyst. 2002;127:582–584. doi: 10.1039/b201207c. [DOI] [PubMed] [Google Scholar]
- Griffin JL, Shockcor JP. Metabolic profiles of cancer cells. Nat Rev Cancer. 2004;4:551–561. doi: 10.1038/nrc1390. [DOI] [PubMed] [Google Scholar]
- Griffin JL, Williams HJ, Sang E, Nicholson JK. Magn. Reson Med. 2001;46:249. doi: 10.1002/mrm.1185. [DOI] [PubMed] [Google Scholar]
- Han X, Gross RW. Shotgun lipidomics: multi-dimensional mass spectrometric analysis of cellular lipidomes. Gross Expert Rev Proteomics. 2005;2:253–264. doi: 10.1586/14789450.2.2.253. [DOI] [PubMed] [Google Scholar]
- de Hoffman E. Tandem Mass Spectrometry: A Primer. J Mass Spectrom. 1996;31:129– 137. [Google Scholar]
- Kathiresan S, Melander O, Anevski D, Guiducci C, Burtt NP, Roos C, Hirschhorn JN, Berglund G, Hedblad B, Groop L, et al. Polymorphisms associated with cholesterol and risk of cardiovascular events. N Engl J Med. 2008;358:1240–9. doi: 10.1056/NEJMoa0706728. [DOI] [PubMed] [Google Scholar]
- Kebarle P, Ho Y. In: Electrospray Ionization Mass Spectrometry; Fundamentals Instrumentation & Applications. Cole RB, editor. New York: John Wiley & Sons, Inc; 1997. [Google Scholar]
- Khot UN, Khot MB, Bajzer CT, Sapp SK, Ohman EM, Brener SJ, Ellis SG, Lincoff AM, Topol EJ. Prevalence of conventional risk factors in patients with coronary heart disease. JAMA. 2003;290:898– 904. doi: 10.1001/jama.290.7.898. [DOI] [PubMed] [Google Scholar]
- Knochenmuss R. A quantitative model of ultraviolet matrix-assisted laser desorption/ionization including analyte ion generation. Anal Chem. 2003;75:2199–207. doi: 10.1021/ac034032r. [DOI] [PubMed] [Google Scholar]
- Lewis GD, Farrell L, Wood MJ, Martinovic M, Arany Z, Rowe GC, Souza A, Cheng S, McCabe EL, Yang E, et al. Metabolic signatures of exercise in human plasma. Sci Transl Med. 2010;2:33ra7. doi: 10.1126/scitranslmed.3001006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Losalzo J. Personalized cardiovascular medicine and drug development: time for a new paradigm. Circulation. 2012;125:638–45. doi: 10.1161/CIRCULATIONAHA.111.089243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Makarov A. Electrostatic axially harmonic orbital trapping: a high-performance technique of mass analysis. 2000;72:1156–62. doi: 10.1021/ac991131p. [DOI] [PubMed] [Google Scholar]
- Moestue S, Sitter B, Bathen TF, Tessem MB, Gribbestad IS. HR MAS MR spectroscopy in metabolic characterization of cancer. Curr Top Med Chem. 2011;11:2– 26. doi: 10.2174/156802611793611869. [DOI] [PubMed] [Google Scholar]
- Newgard CB, An J, Bain JR, Muehlbauer MJ, Stevens RD, Lien LF, Haqq AM, Shah SH, Arlotto M, Slentz CA, Rochon J, et al. A branched-chain amino acid-related metabolic signature that differentiates obese and lean humans and contributes to insulin resistance. Cell Metab. 2009;9:311–26. doi: 10.1016/j.cmet.2009.02.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nicholson JK, Buckingham MJ, Sadler PJ. High resolution 1H n. m.r studies of vertebrate blood and plasma. Biochem J. 1983;211:605–15. doi: 10.1042/bj2110605. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nicholson JK, Sadler PJ, Bales JR, Juul SM, MacLeod AF, Sonksen PH. Monitoring metabolic disease by proton NMR of urine. Lancet. 1984;2:751–2. doi: 10.1016/s0140-6736(84)92656-4. [DOI] [PubMed] [Google Scholar]
- Nicholson JK, O’Flynn MP, Sadler PJ, Macleod AF, Juul SM, Sonksen PH. Proton-nuclear-magnetic-resonance studies of serum, plasma and urine from fasting normal and diabetic subjects. Biochem J. 1984b;217:365–75. doi: 10.1042/bj2170365. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nicholson JK, Lindon JC, Holmes E. ‘Metabonomics’: understanding the metabolic responses of living systems to pathophysiological stimuli via multivariate statistical analysis of biological NMR spectroscopic data. Xenobiotica. 1998;29:1181–1189. doi: 10.1080/004982599238047. [DOI] [PubMed] [Google Scholar]
- NIH Definitions Working Group. Biomarkers and Surrogate Endpoints. Amsterdam: Elsevier; 2000. pp. 1–9. Biomarkers and surrogate endpoints in clinical research: definitions and conceptual model. [Google Scholar]
- Oliver SG, Winson MK, Kell DB, Baganz F. Systematic functional analysis of the yeast genome. Trends Biotechnol. 1998;16:373–378. doi: 10.1016/s0167-7799(98)01214-1. [DOI] [PubMed] [Google Scholar]
- Peterson BL, Cummings BS. A review of chromatographic methods for the assessment of phospholipids in biological samples. Biomed Chromatogr. 2006;20:227–43. doi: 10.1002/bmc.563. [DOI] [PubMed] [Google Scholar]
- Petkovic M, Schiller J, Muller M, Benard S, Reichl S, Arnold K, Arnhold J. Detection of individual phospholipids in lipid mixtures by matrix-assisted laser desorption/ionization time of-flight mass spectrometry: phosphatidylcholine prevents the detection of further species. Anal Biochem. 2001;289:202–216. doi: 10.1006/abio.2000.4926. [DOI] [PubMed] [Google Scholar]
- Roberts LD, Murray AJ, Menassa D, Ashmore T, Nicholls AW, Griffin JL. The contrasting roles of PPARδ and PPARγ in regulating the metabolic switch between oxidation and storage of fats in white adipose tissue. Genome Biol. 2011;12:R75. doi: 10.1186/gb-2011-12-8-r75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roe CR, Millington DS, Maltby DA. Identification of 3-methylglutarylcarnitine. A new diagnostic metabolite of 3-hydroxy-3-methylglutaryl-coenzyme A lyase deficiency. J Clin Invest. 1986;77:1391–4. doi: 10.1172/JCI112446. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rooney OM, Troke J, Nicholson JK, Griffin JL. High-resolution diffusion and relaxation-edited magic angle spinning 1H NMR spectroscopy of intact liver tissue. Magn Reson Med. 2003;50:925–30. doi: 10.1002/mrm.10620. [DOI] [PubMed] [Google Scholar]
- Schiller J, Arnold K. Mass Spectrometry in Structural Biology. In: Meyers RA, editor. Encyclopedia of Analytical Chemistry. Chichester: Willey and Sons; 2000. [Google Scholar]
- Stancáková A, Civelek M, Saleem NK, Soininen P, Kangas AJ, Cederberg H, Paananen J, Pihlajamäki J, Bonnycastle LL, Morken MA, et al. Hyperglycemia and a common variant of GCKR are associated with the levels of eight amino acids in 9,369 Finnish men. Diabetes. 2012;61:1895–1902. doi: 10.2337/db11-1378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suhre K, Shin SY, Petersen AK, Mohney RP, Meredith D, Wägele B, Altmaier E, Deloukas P, Erdmann J, Grundberg E, et al. Human metabolic individuality in biomedical and pharmaceutical research. Nature. 2011;477:54–60. doi: 10.1038/nature10354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang TJ, Gona P, Larson MG, Tofler GH, Levy D, Newton-Cheh C, Jacques PF, Rifai N, Selhub J, Robins SJ, Benjamin EJ, D’Agostino RB, Vasan RS. Multiple biomarkers for the prediction of first major cardiovascular events and death. N Engl J Med. 2006;355:2631–9. doi: 10.1056/NEJMoa055373. [DOI] [PubMed] [Google Scholar]
- Wang TJ, Larson MG, Vasan RS, Cheng S, Rhee EP, McCabe E, Lewis GD, Fox CS, Jacques PF, Fernandez C, et al. Metabolite profiles and the risk of developing diabetes. Nat Med. 2011;17:448–53. doi: 10.1038/nm.2307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Z, Klipfell E, Bennett BJ, Koeth R, Levison BS, Dugar B, Feldstein AE, Britt EB, Fu X, Chung YM, et al. Gut flora metabolism of phosphatidylcholine promotes cardiovascular disease. Nature. 2011;472:57–63. doi: 10.1038/nature09922. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wenk MR, Lucast L, Di Paolo G, Romanelli AJ, Suchy SF, Nussbaum RL, Cline GW, Shulman GI, McMurray W, De Camilli P. Phosphoinositide profiling in complex lipid mixtures using electrospray ionization mass spectrometry. Nat Biotech. 2003;21:813–817. doi: 10.1038/nbt837. [DOI] [PubMed] [Google Scholar]
- Wishart DS, Jewison T, Guo AC, Wilson M, Knox C. HMDB 3.0—The Human Metabolome Database in 2013. Nucleic Acids Res. 2013;41:D801–7. doi: 10.1093/nar/gks1065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wong SF, Meng CK, Fenn JB. Multiple Charging in Electrospray Ionization of Poly(ethylene glycols) J Phys Chem. 1988;92:546–550. [Google Scholar]