Abstract
Rationale: Mutations in genes encoding proteins important in the function and metabolism of pulmonary surfactant are recognized causes of lung disease. Clinical genetic testing is available for these disorders, but children with phenotypes consistent with surfactant dysfunction and no identifiable mutations in the known causative genes have been reported.
Objectives: To identify the mechanism(s) for lung disease in two children with the phenotype of surfactant dysfunction who had negative testing in clinical laboratories for gene mutations causing surfactant dysfunction.
Methods: Amplicons spanning multiple exons of candidate genes were generated by polymerase chain reaction and sequenced.
Measurements and Main Results: A 4,335-base deletion that included all of exon 12 of the gene encoding member A3 of the adenosine triphosphate–binding cassette transporter was identified in a full-term infant with respiratory failure. A 333-base deletion involving part of exon 4 and the adjacent intron of the gene encoding surfactant protein C was identified in a child with interstitial lung disease.
Conclusions: Large deletions are a cause of surfactant dysfunction disorders and may need to be sought for specifically in children whose phenotypes suggest these syndromes but in whom clinical genetic testing is unrevealing.
Keywords: pulmonary surfactant, respiratory distress syndrome, interstitial lung disease, genetic basis of disease
Mutations in genes important for surfactant function and metabolism result in the phenotypes of neonatal respiratory failure and children’s interstitial lung disease (1, 2). These include biallelic loss-of-function mutations in the gene encoding member A3 of the adenosine triphosphate (ATP) family of transporters (ABCA3; adenosine triphosphate–binding cassette family member A3) and heterozygous mutations that alter the coding sequence of the surfactant protein C gene (SFTPC). To date, only single-base mutations in the coding regions, altered splicing, or small deletions or insertions have been reported in these genes as causes of lung disease. We investigated the mechanism for lung disease in two children who presented with phenotypes consistent with surfactant dysfunction, including typical lung histopathology findings on lung biopsy, but whose testing did not identify mutations in genes known to cause surfactant dysfunction (SFTPC, ABCA3, SFTPB [gene encoding surfactant protein B], and NKX2-1 [gene encoding thyroid transcription factor-1]), and who were found to have novel deletions in ABCA3 and SFTPC.
Methods
Case Reports
Subject 1 was born after an uncomplicated pregnancy at 39 weeks of gestation and developed respiratory distress within hours of birth, necessitating intubation and mechanical ventilation. Chest radiographs showed diffuse, coarse granular infiltrates and high-resolution chest computed tomography demonstrated diffuse parenchymal disease with cysts and septal thickening (Figure 1). Because of persistent clinical and radiographic diffuse lung disease, genetic testing was done for surfactant dysfunction disorders. Normal results were obtained for SFTPB, SFTPC, and NKX2-1. No deviations from the reference ABCA3 sequence were noted, but exon 12 could not be amplified. A lung biopsy was performed, which showed changes typical for surfactant dysfunction with alveolar type II cell (AEC2) hyperplasia, focal aggregates of macrophages embedded within proteinaceous material in the airspaces, and interstitial thickening (Figure 2A). Electron microscopy demonstrated small, electron-dense lamellar bodies. He continued to require mechanical ventilation and high levels of supplemental oxygen and was referred for evaluation for lung transplantation because of the lack of clinical improvement and a diagnosis of surfactant dysfunction based on the lung biopsy findings.
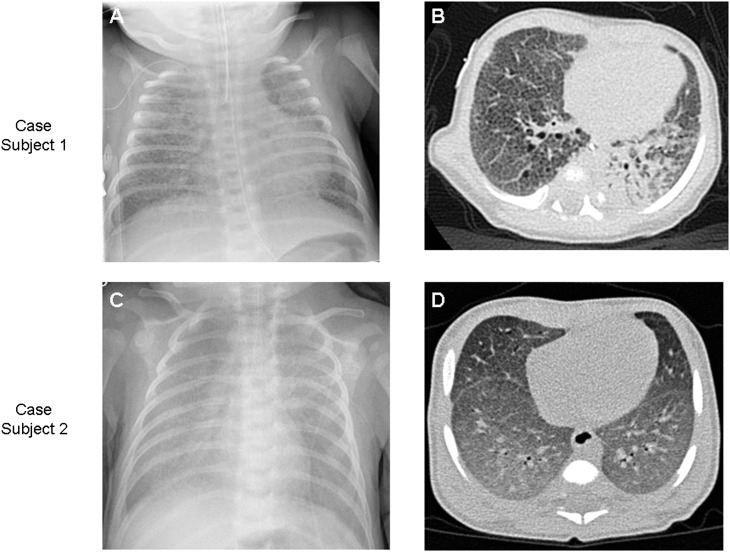
Figure 1.
Imaging studies. (A) Chest radiograph at age 2 weeks from subject 1, demonstrating diffuse ground-glass opacification in all lung fields. (B) High-resolution chest computed tomography (CT) from subject 1 obtained at age 3 weeks, demonstrating diffuse areas of ground-glass opacification with intraparenchymal cysts, thickening of the interlobular septae, and dependent atelectasis. (C) Chest radiograph obtained at time of presentation from subject 2 shows diffuse, relatively homogeneous alveolar infiltrates. (D) High-resolution chest CT obtained at age 3 weeks shows diffuse ground-glass densities, more prominent in the dependent lobes, and thickening of the interlobular septae.
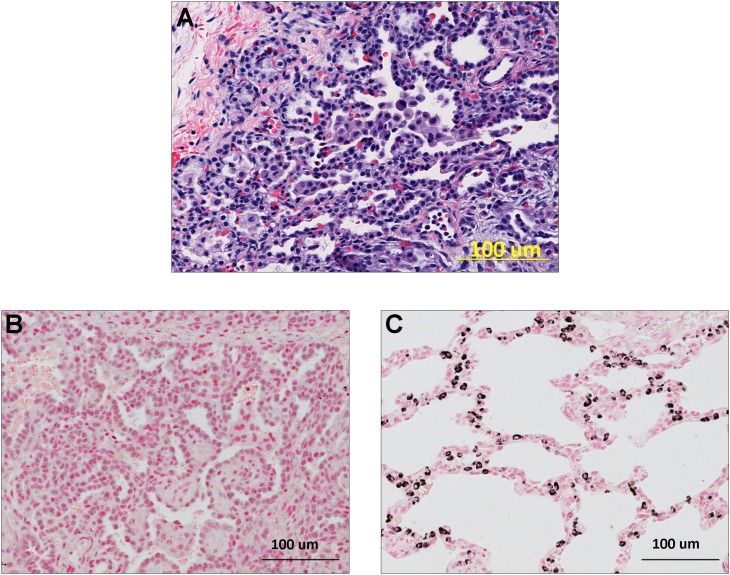
Figure 2.
Histology and immunohistochemical staining for ABCA3 (adenosine triphosphate–binding cassette family member A3). (A) Hematoxylin and eosin stain of lung biopsy from case subject 1. The alveolar septa are thickened, alveolar type II epithelial cell hyperplasia is present, and there are focal accumulations of macrophages in the alveolar lumen. (B) No staining for ABCA3 is detected in tissue from the subject. (C) Positive staining for ABCA3 in alveolar type II epithelial cells is detected in tissue from a control subject. The alveolar architecture is normal and the airspaces are free of cells or debris.
Subject 2 was born at term after an uneventful pregnancy and delivery. At 7 days of age he presented to an emergency room in respiratory distress and required intubation and admission to a pediatric intensive care unit. His chest radiograph had a bilateral ground-glass appearance and high-resolution chest computed tomography showed diffuse ground-glass changes and thickening of the interlobular septa (Figure 1). He remained dependent on continuous positive airway pressure at 5 weeks of age. Clinical genetic testing revealed no mutations in SFTPB, SFTPC, and ABCA3 other than known common polymorphisms. Lung biopsy was performed at 8 weeks of age and the findings included proliferation of AEC2s, focal intraalveolar accumulations of foamy macrophages, and marked thickening of the interstitium with simplification of alveolar architecture, suggestive of surfactant dysfunction. Electron microscopy showed numerous lamellar bodies within AEC2s. He was treated with oral and intravenous steroids and hydroxychloroquine with some improvement in his respiratory status, but continued to require supplemental oxygen and a gastrostomy feeding tube was placed because of poor weight gain. He remained hospitalized until discharged home at age 14 months. Although developing normally, he still has a continuous supplemental oxygen requirement and persistent respiratory distress at 3 years of age.
Research-based genetic studies on both children were subsequently conducted under a protocol approved by the Johns Hopkins Medicine Institutional Review Board (IRB) (Baltimore, MD) after written informed consent was obtained from the parents of each child. Immunostaining for ABCA3 was performed on paraffin-embedded tissue obtained from the lung biopsy of subject 1 under a protocol approved by the Cincinnati Children’s Hospital IRB (Cincinnati, OH) and using a mouse monoclonal antibody directed against a recombinant peptide corresponding to the first loop (amino acids 43–233) of the human ABCA3 protein (WMAB-17G524; Seven Hills Bioreagents, Cincinnati, OH) as previously described (3, 4).
Analysis of the portion of the ABCA3 gene spanning exon 12 in subject 1 was performed with primers located in exons 11 and 13 to generate an amplicon spanning intron 11, exon 12, and intron 12 with a predicted size of 8,417 bases based on the reference sequence (NG_011790.1) using the Expand long template PCR system (Roche Diagnostics, Indianapolis, IN) according to the manufacturer’s directions. For SFTPC analysis, multiple overlapping amplicons spanning two or more SFTPC exons were generated by PCR, using modifications of previously published methods (5). Primer sequences used to generate ABCA3 and SFTPC amplicons and long-range PCR conditions are listed in the online supplement. Amplicons were isolated after gel electrophoresis, purified using spin columns (Qiagen, Valencia, CA), and sequenced through the Johns Hopkins University DNA Analysis Core Facility as previously described (5, 6).
Results
No immunostaining for ABCA3 was detected in the lung tissue of subject 1, whereas positive staining was readily detected in control tissue (Figure 2). Amplification of a region of ABCA3 spanning exons 11 to 13 from subject 1 yielded an approximately 4,000-base product from the subject’s DNA, in contrast to the 8,417-base predicted fragment observed in control subjects with known normal ABCA3 sequence (Figure 3). Sequence analysis of the amplicon from subject 1 indicated that the region beginning 652 bases after the end of exon 11 and ending 3,317 bases before the start of exon 13 was deleted, a 4,335-base span encompassing all of exon 12 (c.1285+652_c.1468–3317del4335). Notably, the 100 bases directly preceding the deletion shared 91% homology with the 100 bases at the 3′ end of the deletion, including complete homology for the last 26 bases. Analysis of parental DNA demonstrated that the mother carried the same deletion whereas only the amplicon of the predicted size was amplified from the father’s DNA (Figure 3). A single-nucleotide polymorphism (SNP) array performed for clinical indications identified an almost 8-Mb region of homozygosity on the short arm of chromosome 16 that included the ABCA3 locus. Given that the subject’s mother, but not the father, carried the ABCA3 deletion, these findings are consistent with maternal uniparental disomy of chromosome 16, where the child inherited both copies of chromosome 16 from the mother, rather than one from each parent, with each copy from the mother containing the partial ABCA3 deletion, resulting in the child being homozygous for the deletion.
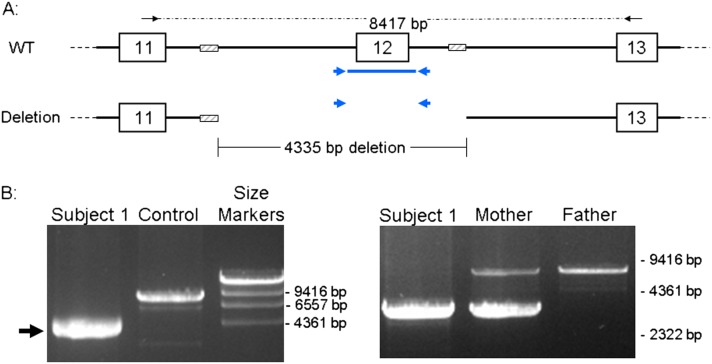
Figure 3.
Polymerase chain reaction (PCR) analysis of ABCA3 (gene encoding adenosine triphosphate–binding cassette family member A3). (A) Schematic of exons 11, 12 and 13 of ABCA3 along with the locations of the primers (arrows) used for PCR. The sizes of the normal (wild-type; WT) and deleted amplicons are shown, with the hatched boxes indicating a 100-base region where the sequences are 91% homologous. The blue arrows indicate the approximate locations of PCR primers used to amplify exon 12 in the clinical laboratory. A product (thick blue line) will be amplified from the normal allele, but no product would be amplified from the allele containing the deletion. (B) The PCR product from the subject is considerably smaller than that observed from the control subject, which was the predicted size (8,417 bases). On the right, only the smaller PCR product is seen in the lane for subject 1. The lane containing the product amplified from the father’s DNA contains only the normal-sized product whereas the lane from the mother has both the normal and smaller-sized products, indicating that she is heterozygous for the allele containing the deletion.
For subject 2, those amplicons that spanned SFTPC exons 1 to 2, 1 to 3 (data not shown), and 2 through part of exon 4 yielded products of the expected size that were comparable to those observed in control subjects (Figure 4). However, amplicons spanning exons 3 to 5, exons 4 to 6 (data not shown), and exons 4 to 5 contained two bands (Figure 4), one migrating at the expected size comparable to that observed in control subjects, and one ∼350 bases shorter in length. Sequence analysis revealed a 333-base deletion involving the last 57 bases of exon 4 through 60 bases upstream of exon 5 (c.379_c.436–61del333), eliminating the last 19 codons of and normal donor splice sequence for exon 4. The deletion was inherited from the mother, who had a history of respiratory symptoms until the age of 6 years that were believed to be due to asthma and had received multiple courses of corticosteroids in childhood. She is clinically well at the time of this report and a detailed clinical and radiological assessment showed no evidence of lung disease. Sequence analysis of NKX2-1 did not identify any deviations from the reference sequence in this subject.
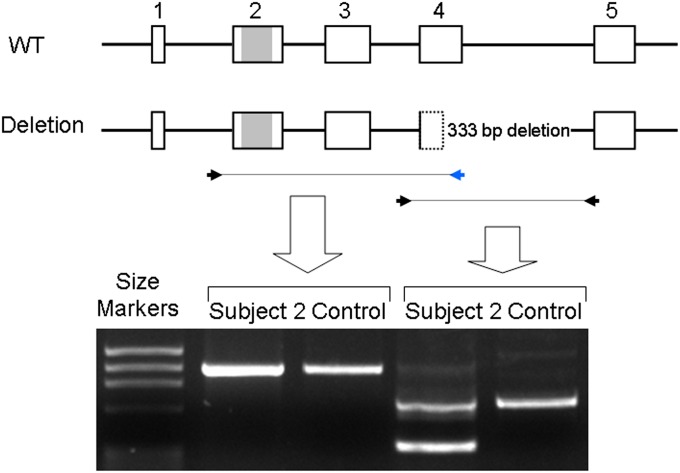
Figure 4.
Polymerase chain reaction (PCR) analysis of surfactant protein C gene (SFTPC). The SP-C gene and its five coding exons are depicted at the top, with both the normal allele (wild type; WT) and allele with the deletion represented. The shaded portion in exon 2 corresponds to the sequence encoding mature SP-C. The lanes containing amplicons spanning exons 2 to 4 contain a single product in both the subject and the control subject, as the reverse primer past exon 4 (blue arrow) will not amplify from the deleted allele. The lane containing the amplicon spanning exons 4 and 5 from the subject contains two products, the smaller representing the allele with the deletion, whereas the lane containing the amplicons from the control subject contains a single product.
Discussion
Genetic surfactant dysfunction syndromes result in acute neonatal respiratory failure or chronic interstitial or diffuse lung disease in older children and adults. Mutations in the genes encoding SP-B, SP-C, ABCA3, and thyroid transcription factor-1 (TTF-1) have been recognized as causes of surfactant dysfunction (1, 2, 7–9). The identification of genetic causes provides a means for noninvasive diagnosis by analyses of these loci. However, such testing is not 100% sensitive (10). In general, only the regions including the exons and surrounding intron–exon boundaries are sequenced and thus functionally significant variants in untranslated regions will be missed. In addition, as PCR is used to amplify the portions of the gene to be sequenced, a deletion encompassing the location of one of the primers or a sequence variant in the primer-binding site may result in the region of interest not being amplified. Should the region that fails to amplify be present on both alleles, then no product will be generated, providing a clue to the deletion, as was the case for subject 1. However, should a deletion be present on only one allele, then the primers will amplify the other allele, which when sequenced will yield either a normal result or any sequence variants will appear to be homozygous.
We identified deletions in the genes encoding ABCA3 and SP-C that resulted in lung disease. In subject 1, all of ABCA3 exon 12 along with ∼3,500 bases upstream and ∼500 bases downstream were deleted. PCR primers designed to amplify exon 12 located relatively close to the intron–exon boundaries but falling within the deleted region would be expected to fail amplification, and thus account for the lack of a product corresponding to this exon. The inability to amplify this exon provided a clue to the possible presence of a deletion, which was confirmed by long-range PCR and direct sequence analysis. The complete loss of exon 12 would result in the corresponding sequence not being present in the ABCA3 transcript. As exon 12 contains 182 bases, this would result in a frameshift and the introduction of a premature codon for the termination of translation, which would likely be subject to nonsense-mediated decay, thus resulting in a null allele. We cannot exclude the possibility of alternative splicing resulting in the introduction of a cryptic exon, but the complete absence of ABCA3 protein as assessed by immunohistochemistry supports the finding that a null allele was responsible for disease in this child. The highly homologous regions upstream of and at the 3′ end of the deletion could pair in opposite directions, resulting in a looping out of the intervening sequence and providing a potential mechanism for the deletion, which could thus also occur de novo in other subjects. The child’s ABCA3 deletion was inherited from the mother and a large contiguous region of homozygosity was detected on the short arm of chromosome 16 by SNP array. These findings are consistent with maternal uniparental disomy as the mechanism for the child bearing two defective copies of the ABCA3 gene. This most likely resulted from an error in maternal meiosis that generated an egg with two chromosome 16s, each containing the defective ABCA3 gene (see Figure E1 in the online supplement). Uniparental disomy has been reported as a mechanism for ABCA3 deficiency, although previously reported cases were paternally inherited (11).
Subject 2 was found to be heterozygous for a deletion in SFTPC that encompassed part of exon 4 and the first 266 bases of intron 4. A primer designed to amplify SFTPC exon 4 located in the region of the deletion would thus fail to amplify from this allele, but a product would be generated from the opposite allele. As the corresponding sequence from the opposite allele did not contain any DNA sequence variations, this would explain the apparently normal result obtained by the clinical laboratory, which used an approach of sequencing amplicons corresponding to each individual exon, with the reverse primer for exon 4 located within the deleted region (E. Ashton [Great Ormond Street Hospital, London, UK], personal communication). Heterozygous SFTPC mutations are sufficient to cause disease through the production of an abnormal proprotein, which may exert a toxic gain of function (12). Although the functional effects of this deletion could not be determined as suitable tissue was not available for RNA or protein blotting studies, this deletion almost certainly accounted for the child’s lung disease. The normal donor splice site for exon 4 was eliminated, making it likely that the mutation resulted in the skipping of exon 4 in the SP-C transcript, a recognized cause of SP-C–related disease (13). Alternatively, the elimination of 19 amino acids due to the 57 deleted bases in exon 4 would also be highly likely to be deleterious to the structure of the SP-C proprotein. The mutation was inherited from the mother, who was initially not known to have a history of lung disease. After the results of the genetic studies were known a history of recurrent respiratory illness in infancy for which she was repeatedly treated with corticosteroids was obtained. Whether the mother’s symptoms were related to the SFTPC deletion is unknown as she currently has no clinical or radiographic evidence of lung disease. Marked phenotypic variability and improvement after childhood lung disease have been recognized in other subjects with SFTPC mutations (5, 14, 15).
Moderate to large-sized deletions in other genes involving one or more exons have been recognized as the cause for disease in multiple genes, including NKX2-1 (brain–thyroid–lung syndrome), SFTPB (surfactant dysfunction), and BMPR2 (pulmonary hypertension) and thus these results are not surprising (7, 16, 17). However, specific methods are needed to detect such deletions. SNP arrays or array comparative hybridization may detect large deletions, but a sufficient density of probes must be located in the deleted region to sensitively detect small or moderate-sized deletions. Multiplex ligation-dependent probe amplification assays may be needed for the detection of such deletions (16, 18). These are the first reported deletions in the genes encoding ABCA3 and SP-C but unlikely to be the only ones, and clinicians and laboratories offering testing for these disorders need to incorporate this information into their interpretations of the sensitivity of mutation detection.
These cases illustrate the need for a comprehensive and flexible approach to the diagnoses of these rare disorders which incorporates clinical presentation, imaging, genetic testing, lung biopsy including ultrastructural studies, and research approaches. A high index of suspicion based on the unusual clinical presentations and imaging findings prompted genetic testing for surfactant dysfunction disorders in the hopes of making definitive diagnoses noninvasively. When the results from genetic testing were not conclusive, lung biopsies were performed to establish diagnoses to guide treatment decisions, due to the severity of each child’s illness. A diagnosis of a surfactant dysfunction disorder was made in each subject on the basis of characteristic histopathology findings. Those findings do not indicate which gene is involved, although the electron microscopy findings suggested that ABCA3 was responsible in the first subject and argued against ABCA3 in the second. Identification of the responsible gene has important implications for prognosis, as ABCA3 deficiency often has a more severe course than SP-C dysfunction, and for recurrence risk, and the modes of inheritance differ (1). The findings from the clinical laboratory in subject 1 provided an important clue to the cause of lung disease and led us to focus on the possibility of a deletion in ABCA3. This was confirmed, but found to result from a rare mechanism wherein the child inherited two copies of a disease-causing ABCA3 allele from one parent with a recurrence risk of much less than the usual 25% for an autosomal recessive condition. In contrast, for subject 2, the finding that one parent carried the same mutation has implications for that individual’s health and also predicts a recurrence risk as high as 50% for future pregnancies. Thus further clinical (SNP array) and research-based studies (immunohistochemistry, genetic) were able to establish definitively the mechanism for lung disease in each child and provided additional important information for counseling the families.
Acknowledgments
Acknowledgment
The authors thank Dr. Emma Ashton (North East Thames Regional Genetics Laboratory, Great Ormond Street Hospital, London, UK) for sharing information on the details of their assay, and Sarah Hirsch for technical assistance.
Footnotes
Supported by a grant from the U.S. National Institutes of Health (HL-54703) and the Eudowood Foundation (L.M.N.). A.B. was supported by the NIHR Respiratory Disease Biomedical Research Unit at the Royal Brompton and Harefield NHS Foundation Trust and Imperial College London.
Author Contributions: L.B.H. and L.M.N. were involved in the conception and design of the study; all authors participated in the acquisition of the data and its interpretation; L.M.N. and L.B.H. wrote the initial draft and along with K.M., S.W., J.C., and A.B. were substantially involved in its revision before submission.
This article has an online supplement, which is available from this issue’s table of contents at www.atsjournals.org
Author disclosures are available with the text of this article at www.atsjournals.org.
References
- 1.Gower WA, Nogee LM. Surfactant dysfunction. Paediatr Respir Rev. 2011;12:223–229. doi: 10.1016/j.prrv.2011.01.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Wert SE, Whitsett JA, Nogee LM. Genetic disorders of surfactant dysfunction. Pediatr Dev Pathol. 2009;12:253–274. doi: 10.2350/09-01-0586.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Stahlman MT, Besnard V, Wert SE, Weaver TE, Dingle S, Xu Y, von Zychlin K, Olson SJ, Whitsett JA. Expression of ABCA3 in developing lung and other tissues. J Histochem Cytochem. 2007;55:71–83. doi: 10.1369/jhc.6A6962.2006. [DOI] [PubMed] [Google Scholar]
- 4.Uchida DA, Wert SE, Nogee LM, Carroll TR, Chatfield BA. Pulmonary nodules in a newborn with ATP-binding cassette transporter A3 (ABCA3) mutations. Pediatrics. 2011;127:e1347–e1351. doi: 10.1542/peds.2010-1477. [DOI] [PubMed] [Google Scholar]
- 5.Cameron HS, Somaschini M, Carrera P, Hamvas A, Whitsett JA, Wert SE, Deutsch G, Nogee LM. A common mutation in the surfactant protein C gene associated with lung disease. J Pediatr. 2005;146:370–375. doi: 10.1016/j.jpeds.2004.10.028. [DOI] [PubMed] [Google Scholar]
- 6.Bullard JE, Wert SE, Whitsett JA, Dean M, Nogee LM. ABCA3 mutations associated with pediatric interstitial lung disease. Am J Respir Crit Care Med. 2005;172:1026–1031. doi: 10.1164/rccm.200503-504OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Hamvas A, Deterding RR, Wert SE, White FV, Dishop MK, Alfano DN, Halbower AC, Planer B, Stephan MJ, Uchida DA, et al. Heterogeneous pulmonary phenotypes associated with mutations in the thyroid transcription factor gene NKX2-1. Chest. 2013;144:794–804. doi: 10.1378/chest.12-2502. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Somaschini M, Nogee LM, Sassi I, Danhaive O, Presi S, Boldrini R, Montrasio C, Ferrari M, Wert SE, Carrera P. Unexplained neonatal respiratory distress due to congenital surfactant deficiency. J Pediatr. 2007;150:649–653, 653.e641. doi: 10.1016/j.jpeds.2007.03.008. [DOI] [PubMed] [Google Scholar]
- 9.Turcu S, Ashton E, Jenkins L, Gupta A, Mok Q. Genetic testing in children with surfactant dysfunction. Arch Dis Child. 2013;98:490–495. doi: 10.1136/archdischild-2012-303166. [DOI] [PubMed] [Google Scholar]
- 10.Gower WA, Wert SE, Ginsberg JS, Golan A, Whitsett JA, Nogee LM. Fatal familial lung disease caused by ABCA3 deficiency without identified ABCA3 mutations. J Pediatr. 2010;157:62–68. doi: 10.1016/j.jpeds.2010.01.010. [DOI] [PubMed] [Google Scholar]
- 11.Hamvas A, Nogee LM, Wegner DJ, Depass K, Christodoulou J, Bennetts B, McQuade LR, Gray PH, Deterding RR, Carroll TR, et al. Inherited surfactant deficiency caused by uniparental disomy of rare mutations in the surfactant protein-B and ATP binding cassette, subfamily A, member 3 genes. J Pediatr. 2009;155:854–859, e851. doi: 10.1016/j.jpeds.2009.06.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Beers MF, Mulugeta S. Surfactant protein C biosynthesis and its emerging role in conformational lung disease. Annu Rev Physiol. 2005;67:663–696. doi: 10.1146/annurev.physiol.67.040403.101937. [DOI] [PubMed] [Google Scholar]
- 13.Nogee LM, Dunbar AE, III, Wert SE, Askin F, Hamvas A, Whitsett JA. A mutation in the surfactant protein C gene associated with familial interstitial lung disease. N Engl J Med. 2001;344:573–579. doi: 10.1056/NEJM200102223440805. [DOI] [PubMed] [Google Scholar]
- 14.Guillot L, Epaud R, Thouvenin G, Jonard L, Mohsni A, Couderc R, Counil F, de Blic J, Taam RA, Le Bourgeois M, et al. New surfactant protein C gene mutations associated with diffuse lung disease. J Med Genet. 2009;46:490–494. doi: 10.1136/jmg.2009.066829. [DOI] [PubMed] [Google Scholar]
- 15.Thomas AQ, Lane K, Phillips J, III, Prince M, Markin C, Speer M, Schwartz DA, Gaddipati R, Marney A, Johnson J, et al. Heterozygosity for a surfactant protein C gene mutation associated with usual interstitial pneumonitis and cellular nonspecific interstitial pneumonitis in one kindred. Am J Respir Crit Care Med. 2002;165:1322–1328. doi: 10.1164/rccm.200112-123OC. [DOI] [PubMed] [Google Scholar]
- 16.Cogan JD, Pauciulo MW, Batchman AP, Prince MA, Robbins IM, Hedges LK, Stanton KC, Wheeler LA, Phillips JA, III, Loyd JE, et al. High frequency of BMPR2 exonic deletions/duplications in familial pulmonary arterial hypertension. Am J Respir Crit Care Med. 2006;174:590–598. doi: 10.1164/rccm.200602-165OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Wegner DJ, Hertzberg T, Heins HB, Elmberger G, MacCoss MJ, Carlson CS, Nogee LM, Cole FS, Hamvas A. A major deletion in the surfactant protein-B gene causing lethal respiratory distress. Acta Paediatr. 2007;96:516–520. doi: 10.1111/j.1651-2227.2006.00188.x. [DOI] [PubMed] [Google Scholar]
- 18.Teissier R, Guillot L, Carre A, Morandini M, Stuckens C, Ythier H, Munnich A, Szinnai G, de Blic J, Clement A, et al. Multiplex ligation-dependent probe amplification improves the detection rate of NKX2.1 mutations in patients affected by brain–lung–thyroid syndrome. Hormone Res Paediatr. 2012;77:146–151. doi: 10.1159/000337214. [DOI] [PubMed] [Google Scholar]