Abstract
We provide here a comparative genome analysis of 31 strains within the genus Paenibacillus including 11 new genomic sequences of N2-fixing strains. The heterogeneity of the 31 genomes (15 N2-fixing and 16 non-N2-fixing Paenibacillus strains) was reflected in the large size of the shell genome, which makes up approximately 65.2% of the genes in pan genome. Large numbers of transposable elements might be related to the heterogeneity. We discovered that a minimal and compact nif cluster comprising nine genes nifB, nifH, nifD, nifK, nifE, nifN, nifX, hesA and nifV encoding Mo-nitrogenase is conserved in the 15 N2-fixing strains. The nif cluster is under control of a σ70-depedent promoter and possesses a GlnR/TnrA-binding site in the promoter. Suf system encoding [Fe–S] cluster is highly conserved in N2-fixing and non-N2-fixing strains. Furthermore, we demonstrate that the nif cluster enabled Escherichia coli JM109 to fix nitrogen. Phylogeny of the concatenated NifHDK sequences indicates that Paenibacillus and Frankia are sister groups. Phylogeny of the concatenated 275 single-copy core genes suggests that the ancestral Paenibacillus did not fix nitrogen. The N2-fixing Paenibacillus strains were generated by acquiring the nif cluster via horizontal gene transfer (HGT) from a source related to Frankia. During the history of evolution, the nif cluster was lost, producing some non-N2-fixing strains, and vnf encoding V-nitrogenase or anf encoding Fe-nitrogenase was acquired, causing further diversification of some strains. In addition, some N2-fixing strains have additional nif and nif-like genes which may result from gene duplications. The evolution of nitrogen fixation in Paenibacillus involves a mix of gain, loss, HGT and duplication of nif/anf/vnf genes. This study not only reveals the organization and distribution of nitrogen fixation genes in Paenibacillus, but also provides insight into the complex evolutionary history of nitrogen fixation.
Author Summary
We sequenced the genomes of 11 N2-fixing Paenibacillus strains and demonstrated the genomic diversity of the genus Paenibacillus by comparing these strains to each other and to 20 other strains (4 N2-fixing and 16 non-N2-fixing strains) that were sequenced previously. Phylogenetic analysis of the concatenated 275 single-copy core genes suggests that ancestral Paenibacillus did not fix nitrogen and the N2-fixing strains fall into two sub-groups, which were likely originated from a N2-fixing common ancestor. A minimal and compact nif cluster comprising nine nif genes encoding Mo-nitrogenase is highly conserved in the 15 N2-fixing strains. Variations in the nif cluster and in the chromosomal regions surrounding the nif cluster between two sub-groups imply at least two independent acquisitions with insertion of distinct nif cluster variants in different genomic sites of Paenibacillus in early evolutionary history. Phylogeny of the concatenated NifHDK sequences suggests that Paenibacillus and Frankia are sister groups. The nif cluster, a functional unit for nitrogen fixation, was lost, producing some non-N2-fixing strains. There were recent events of acquisition of vnf and anf genes, causing further diversification of some strains. The evolution of nitrogen fixation in Paenibacillus involves a mix of gain, loss, HGT and duplication of nif/anf/vnf genes.
Introduction
Biological nitrogen fixation, the conversion of atmospheric N2 to NH3, plays an important role in the global nitrogen cycle and in world agriculture [1]. Nitrogen fixation is mainly catalyzed by the Mo-nitrogenase. The ability to fix nitrogen is widely, but sporadically distributed among Archaea and Bacteria which includes these families: Proteobacteria, Firmicutes, Cyanobacteria, Actinobacteria and Chlorobi [2]. Also, the contents and organization of nitrogen fixation (nif) genes vary significantly among the different N2-fixing organisms. For example, in Klebsiella pneumoniae, twenty nif genes are co-located within a ∼24 kb cluster [3], whereas in Azotobacter vinelandii the nif genes are more dispersed and distributed as two clusters in the genome [4]. The random distribution pattern and the difference in contents and organization of nif genes raise the question of origins and evolution of Mo-nitrogenase. Phylogenetic inference based on the sequences of nif genes is generally used to understand the evolution of nif genes [5]–[7]. Two conflicting hypotheses for origins of Mo-nitrogenase have been proposed on the basis of phylogenetic examination of Mo-nitrogenase protein sequences (NifHDK) [8]–[11]. One is the last common ancestor (LCA) hypothesis which implies that the Mo-nitrogenase had its origin in a common ancestor of the bacterial and archaeal domains. According to the LCA model gene loss has been extensive and accounts for the fact that nitrogenase is found neither in eukaryotes nor in many entire phyla of prokaryotes. The other is the methanogen origin hypothesis which implies that nitrogen fixation was originated in methanogenic archaea and subsequently was transferred into a primitive bacterium via horizontal gene transfer (HGT).
Remarkable progress in sequencing technology has advanced in understanding genetics and phylogenetic history of nitrogen fixation. For example, genome sequences of several diazotrophs, such as Pseudomonas stutzeri A1501 [12], Herbaspirillum seropedicae SmR1 [13] and Wolinella succinogenes [14], revealed that the Mo-nitrogenase genes constitute a nitrogen fixation cluster or island. The nif genes of P. stutzeri, including nifQ, nifA, nifL, nifH, nifD, nifK, nifT, nifY, nifE, nifN, nifX, nifS, nifU, nifW, nifZ, nifM and nifF are distributed in a 49-kb region. The nif genes of H. seropedicae, including nifA, nifB, nifZ, nifZ1, nifH, nifD, nifK, nifE, nifN, nifX, nifQ, nifW, nifV, nifU and nifS are in a region spanning 37 kb interspersed with fix, mod, hes, fdx, hsc and other genes. Variation of G+C content between the nif cluster and the genome average in P. stutzeri A1501 and existence of transposase near the nif cluster in H. seropedicae SmR1 are indicative of HGT of nif gene clusters [13]. However, since nitrogen fixation is an ancient complex process and is widely, but sporadically distributed among prokaryote families, further extensive genome sequences are needed to completely resolve the evolutionary history of nitrogenase.
Mo-nitrogenase is composed of two proteins, dinitrogenase (MoFe protein) and dinitrogenase reductase (Fe protein). The MoFe protein is an α2β2 heterotetramer (encoded by nifDK) that contains the iron–molybdenum cofactors (FeMo-co) and P clusters. The FeMo-co is a [Mo-7Fe-9S-homocitrate] cluster which serves as the active site of substrate binding and reduction. The P-cluster is a [8Fe-7S] cluster which shuttles electrons to the FeMo-co. The Fe protein is a γ2 homodimer (encoded by nifH) bridged by an intersubunit [4Fe-4S] cluster that serves as the obligate electron donor to the MoFe protein. In addition to the structural genes nifHDK, other genes nifE nifN, nifX nifB, nifQ, nifV, nifY, nifU, nifS, nifZ and nifM contribute to the synthesis of FeMo-co and maturation of nitrogenase [15]–[17]. Although the majority of present-day biological N2 reduction is catalyzed by the Mo-nitrogenase, two homologous alternative nitrogenases: V- and Fe-nitrogenase are important biological sources of fixed nitrogen in environments where Mo is limiting [18]. V- and Fe-nitrogenase are encoded by the vnf and anf genes. The Mo-, V- and Fe-nitrogenases are not equally distributed in nature. Most of diazotrophs, such as K. pneumoniae, possesses only the Mo-nitrogenase [19]. While some organisms, like A. vinelandii, possess all three types of nitrogenases [20] and other organisms, like Rhodobacter capsulatus and Rhodospirillum rubrum, carry the Mo- and Fe-nitrogenases [21], [22].
Paenibacillus is a large genus of Gram-positive, facultative anaerobic, endospore-forming bacteria. Members of this genus are biochemically and morphologically diverse and are found in various environments, such as soil, rhizosphere, insect larvae, and clinical samples [23]–[26]. Originally Paenibacillus was included within the genus Bacillus, however in 1993 it was reclassified as a separate genus [27]. At that time, the genus Paenibacillus encompassed 11 species including the three N2-fixing species Paenibacillus polymyxa, Paenibacillus macerans and Paenibacillus azotofixans [27]. The genus Paenibacillus currently comprises more than 120 named species, more than 20 of which have nitrogen fixation ability, including the following 8 novel species described by our laboratory: Paenibacillus sabinae, Paenibacillus zanthoxyli, Paenibacillus forsythiae, Paenibacillus sonchi, Paenibacillus sophorae, Paenibacillus jilunlii, P. taohuashanense and P. beijingensis [28]–[35]. Although diazotrophic Paenibacillus strains have potential uses as a bacterial fertilizer in agriculture, genomic information to date is limited and the genetics and evolution of nitrogen fixation of these diazotrophs are unknown.
Here we sequenced 11 N2-fixing Paenibacillus strains and compared these strains to each other and to 20 other strains (4 N2-fixing and 16 non-N2-fixing strains) that were sequenced previously. These strains were obtained from plant rhizhospheres, hot spring and human body and from Brazil, China, Korea, Israel, France, Belgium, United States of America, etc. (Table 1). Our study revealed that a nif gene cluster comprising nifB, nifH, nifD, nifK, nifE, nifN, nifX, hesA and nifV encoding Mo-nitrogenase is highly conserved in the 15 N2-fixing strains. Also, two homologous alternative nitrogenases: V- and Fe-nitrogenase encoded by the vnf and anf genes, respectively, are found in some Paenibacillus species. HGT, gene loss and gene duplication of nif, vnf and anf genes have contributed to evolution of nitrogen fixation in Paenibacillus. This study not only reveals the organization and distribution of nif/anf/vnf genes and the evolutionary patterns of nitrogen fixation in Paenibacillus, but also provides support for the methanogen origin hypothesis for nif evolution [10], [11].
Table 1. Paenibacillus strains used in study.
| Strains | Source | Nitrogen fixer | Genome sequence |
| Paenibacillus sp. JDR2 | Sweetgum stem wood, Florida, USA | − | [36] |
| Paenibacillus sp. Y412MC10 | Obsidian hot spring, Montana, USA | − | [37] |
| P. mucilaginosus KNP414 | Soil of Tianmu Mountain, Zhejiang, China | − | Unpublished |
| P. mucilaginosus K02 | Soil of maize-farming fields, Guizhou, China | − | Unpublished |
| P. mucilaginosus 3016 | Rhizosphere soil, Shandong, China | − | [38] |
| P. polymyxa E681 | Rhizosphere of winter barley, Chonnam, South Korea | − | [39] |
| P. polymyxa SC2 | Rhizosphere of pepper, Guizhou, China | − | [40] |
| P. curdlanolyticus YK9 | Soil, Kobe city, Japan | − | Unpublished |
| Paenibacillus sp. HGF5 | Human intestinal microflora, USA | − | Unpublished |
| Paenibacillus sp. HGF7 | Human intestinal microflora, USA | − | Unpublished |
| P. dendritiformis C454 | Soil, Tel Aviv, Israel | − | [41] |
| P. elgii B69 | Soil samples, Hangzhou, China | − | [42] |
| P. lactis 154 | Milk, Belgium | − | Unpublished |
| P. peoriae KCTC 3763 | Soil, Republic of Korea | − | [43] |
| Paenibacillus sp. oral taxon786D14 | Oral swab from female patient, USA | − | Unpublished |
| P. vortex V453 | Rhizosphere, Tel Aviv, Israel | − | [44] |
| P. polymyxa WLY78 | Bamboo rhizosphere, Beijing, China | + | Unpublished |
| P. polymyxa TD94 | Scutellaria rhizosphere, Liaoning, China | + | This study |
| P. polymyxa 1–43 | Corn rhizosphere, Shanxi, China | + | This study |
| P.beijingensis 1–18 | Wheat rhizosphere, Beijing, China | + | This study |
| Paenibacillus sp. 1–49 | Corn rhizosphere, Shanxi, China | + | This study |
| Paenibacillus sp. Aloe-11 | Root of Aloe chinensis, Chongqing, China | + | [45] |
| P. terrae HPL-003 | Soil of forest residue, Daejeon, Republic of Korea | + | [46] |
| P. azotofixans ATCC35681 | Wheat roots, Parana state, Brazil | + | This study |
| P. graminis RSA19 | Maize rhizosphere soil, Ramonville, France | + | This study |
| P. sonchi X19-5 | Rhizosphere of Ku Caihua, Xinjiang, China | + | This study |
| P. sophorae S27 | Rhizosphere of Sophora japonica, Beijing, China | + | This study |
| P. massiliensis T7 | Willow rhizosphere, Beijing, China | + | This study |
| P. zanthoxyli JH29 | Pepper rhizosphere, Hubei, China | + | This study |
| P. forsythia T98 | Forsythia rhizosphere, Beijing, China | + | This study |
| P. sabinae T27 | Rhizosphere of Sabina squamata, Beijing, China | + | Unpublished |
Results
Genomic features
A summary of the features of each of the 11 newly-sequenced genomes of N2-fixing Paenibacillus strains and 20 previously-sequenced genomes of Paenibacillus strains (4 N2-fixers and 16 non-N2-fixers) is shown in Table 2. The characteristics (size, GC content, predicted number of coding sequences, and number of tRNA genes) of the 11 newly-sequenced genomes are within the range of previously-sequenced genomes of Paenibacillus strains (Table 2, Table S1). The 31 genomes vary in size by approximately three megabases (ranging from 4.90–8.77 Mb) with the number of CDSs ranging from 4460–9087, indicating substantial strain-to-strain variation. The G+C contents of the 31 genomes range from 44.2–58.4. The genome of Paenibacillus sophorae S27 has a larger size than those of the newly-sequenced strains.
Table 2. Genomic features of Paenibacillus strains.
| Species | Status | GenBank accession number | Genome size (Mb) | G+C content | tRNA genes | Protein-coding sequences (CDSs) |
| Paenibacillus sp. JDR 2 | Complete | CP001656. | 7.18 | 50.3 | 87 | 6213 |
| Paenibacillus sp. Y412MC10 | Complete | CP001793 | 7.12 | 51.2 | 73 | 6238 |
| P.mucilaginosus KNP414 | Complete | CP002869 | 8.66 | 58.4 | 108 | 7811 |
| P. mucilaginosus K02 | Complete | CP003422 | 8.77 | 58.2 | 189 | 7252 |
| P. mucilaginosus 3016 | Complete | CP003235 | 8.74 | 58.3 | 170 | 7057 |
| P. polymyxa E681 | Complete | CP000154 | 5.39 | 45.8 | 91 | 4805 |
| P. polymyxa SC2 | Complete | CP002213 | 6.24 | 44.6 | 91 | 6032 |
| P. curdlanolyticus YK9 | Complete | AEDD00000000 | 5.45 | 51.9 | 101 | 4824 |
| Paenibacillus sp. HGF5 | Draft | AEXS00000000 | 6.95 | 51.0 | 71 | 6496 |
| Paenibacillus sp. HGF7 | Draft | AFDH00000000 | 6.28 | 52.8 | 72 | 5992 |
| P. dendritiformis C454 | Draft | AHKH00000000 | 6.38 | 54.0 | 31 | 5660 |
| P. elgii B69 | Draft | AFHW00000000 | 7.96 | 52.4 | 51 | 7777 |
| P. lactis 154 | Draft | AGIP00000000 | 6.81 | 51.8 | 74 | 6149 |
| P. peoriae KCTC 3763 | Draft | AGFX00000000 | 5.77 | 46.4 | 81 | 5073 |
| Paenibacillus sp. oral taxon786 str. D14 | Draft | ACIH00000000 | 4.90 | 51.8 | 69 | 4460 |
| P. vortex V453 | Draft | ADHJ00000000 | 6.39 | 48.8 | 57 | 5928 |
| P. polymyxa WLY78 | Draft | ALJV00000000 | 5.92 | 45.1 | 54 | 5729 |
| P. polymyxa TD94 | Draft | ASSA00000000 | 6.10 | 45.0 | 50 | 5697 |
| P. polymyxa 1–43 | Draft | ASRZ00000000 | 6.00 | 44.2 | 69 | 5731 |
| P. beijingensis 1–18 | Draft | ASSB00000000 | 5.44 | 46.0 | 59 | 5599 |
| Paenibacillus sp. 1–49 | Draft | ASRY00000000 | 5.65 | 46.4 | 56 | 5628 |
| Paenibacillus sp. Aloe-11 | Draft | AGFI00000000 | 5.79 | 46.6 | 73 | 5275 |
| P. terrae HPL-003 | Complete | CP003107 | 6.08 | 46.8 | 89 | 5525 |
| P. massiliensis T7 | Draft | ASSE00000000 | 6.32 | 48.4 | 63 | 5722 |
| P. graminis RSA19 | Draft | ASSG00000000 | 7.08 | 50.4 | 61 | 7081 |
| P. sonchi X19-5 | Draft | AJTY00000000 | 7.61 | 50.4 | 46 | 7705 |
| P.azotofixans ATCC35681 | Draft | ASQQ00000000 | 5.44 | 50.8 | 37 | 5924 |
| P. sophorae S27 | Draft | ASSF00000000 | 8.52 | 47.9 | 83 | 9087 |
| P. zanthoxyli JH29 | Draft | ASSD00000000 | 5.12 | 50.9 | 50 | 5622 |
| P. forsythia T98 | Draft | ASSC00000000 | 5.19 | 53.0 | 37 | 5552 |
| P. sabinae T27 | Complete | CP004078 | 5.27 | 52.6 | 82 | 5250 |
Core and pan-genome analysis
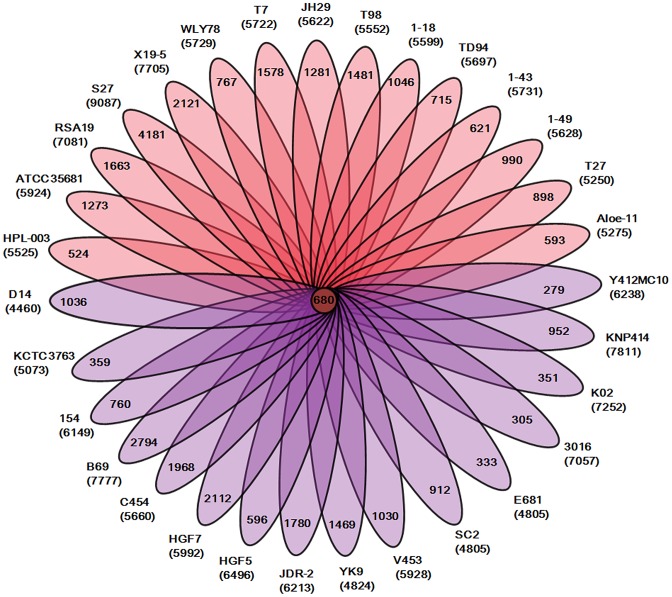
Our analysis of the total 31 genomes reveals that a pan genome contains 55504 putative protein-coding genes in the genus Paenibacillus. Of the 55504 putative protein-coding genes, 37105, which made up 66.9% of the genes in the pan genome, were represented in only one genome of Paenibacillus spp., suggesting a high frequency of horizontal gene acquisition from other taxa. In contrast to the pan-genome, the genus Paenibacillus had the core genome of 680 putative protein-coding genes, which represents only 9% to 15% of the repertoire of protein coding genes of each strain, illustrating a large degree of genomic diversity in this group of bacteria (Figure 1). The genomic data are consistent with the fact that Paenibacillus strains are morphologically and physiologically diverse.
Figure 1. Genomic diversity of strains in the genus Paenibacillus.
Each strain is represented by an oval that is colored: N2-fixing strains (red), non- N2-fixing strains (purple). The number of orthologous coding sequences (CDSs) shared by all strains (i.e., the core genome) is in the center. Overlapping regions show the number of CDSs conserved only within the specified genomes. Numbers in non-overlapping portions of each oval show the number of CDSs unique to each strain. The total number of protein coding genes within each genome is listed below the strain name.
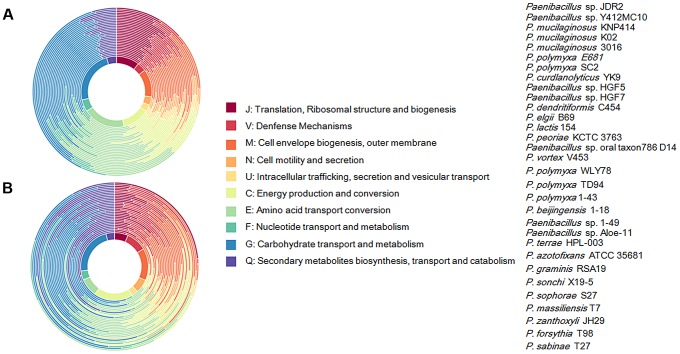
We further comparatively analyze the core genome of 15 N2-fixing and 16 non-N2-fixing Paenibacillus strains. We found that non-N2-fixing strains had the core genome of 908 putative protein-coding genes, which made up 12–20% of protein-coding genes in each strain. N2-fixing strains had the core genome of 1264 putative protein-coding genes, which code 14–24% of the protein pool in each genome. Further, we use Cluster of Orthologous Groups (COG) assignments to determine whether there were differences in the proportion of the core genome attributable to a particular cellular process (Figure 2 and Table S2). Interestingly, core genome of N2-fixing strains was found to be disproportionally enriched in cell motility and chemotaxis genes (Fisher's exact test; P value<0.01). Since these N2-fixing strains were isolated from plant rhizospheres, cell motility and chemotaxis are of importance for bacterial adaptation to the ever-changing rhizosphere environment [47].
Figure 2. Functional classification of gene content of the 31 Paenibacillus strains.
(A) Profiles of Cluster of Orthologous Groups (COG) showing percentage of genes in each category out of total annotated genes. Taxa from inside of circle to outside of circle are from Paenibacillus sp. JDR 2 (top in the strain list) to P. sabinae T27 (down in the strain list). (B) Profiles of COG showing function categories for genes in core genomes. Taxa from inside of circle to outside of circle are from Paenibacillus sp. JDR 2 (top in the strain list) to P. sabinae T27 (down in the strain list).
Transposable elements
In this study, transposons were identified using the ISfinder database (http://www-is.biotoul.fr/) and only expectation values of 10−5 and below were considered as significant matches during searches. Each Paenibacillus genome in this study contains a unique set of transposons (Table S3). The number of transposon copies pergenome ranges from 3 (P. polymyxa SC2) to 118 (P. sophorae S27). Members of the IS3, IS4, IS5, IS1182 and IS200/IS605 families are most common. However, there is not a large difference in numbers of transposable elements between other N2-fixing and non-N2-fixing strains.
Prophage
Here prophages were identified using PHAST. Each genome of the 31 strains contains 1–10 prophages and/or prophage remnants, ranging in size from 14.4 to 59.1 kb. Collectively, the 31 genomes have 16 intact prophages and 69 prophage remnants. The newly-sequenced genomes have 38 prophages, most of which have a set of cargo genes that encode putative bacteriocins, DNA replication protein DnaD, ABC transporter ATP-binding protein, Non-ribosomal peptide synthase module containing protein adenine- and cytosine-specific DNA methyltransferases, and DNA/RNA helicase (Table S4). However, there is not a large difference in numbers of prophages between other N2-fixing and non-N2-fixing strains.
The nif gene cluster is highly conserved in Paenibacillus
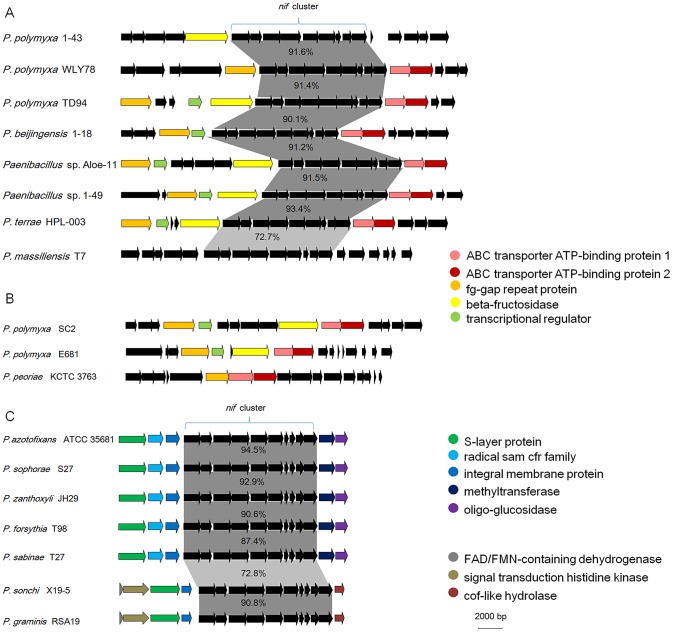
Comparison of COG assignments between non-N2-fixing and N2-fixing Paenibacillus strains (Table S2) revealed that 9 core genes in the N2-fixing strains: nifB, nifH, nifD, nifK, nifE, nifN, nifX, hesA and nifV, which are organized as a nif gene cluster arranged within an 10.5–12 kb genomic region, are conserved in all of the 15 N2-fixing strains (Figure 3, Table S5). The nifH, nifD and nifK are structural genes for Mo-nitrogenase, and the nifB, nifE, nifN, nifX and nifV are involved in synthesis of FeMo-cofactor. The gene hesA, which is located between nifX and nifV, is also found in the nif clusters of Frankia [48] and cyanobacteria [49]. HesA (also being called NAD/FAD-binding protein) is a member of the ThiF-MoeB-HesA family, which is involved in molybdopterin and thiamine biosynthesis. Our recent studies demonstrated that HesA is required for efficient nitrogen fixation in Paenibacillus [50]. As shown in Figure S1, the numbers of nif genes and size of the nif cluster of Paenibacillus are much smaller than those of Frankia, cyanobacteria, Chlorobia (green sulfur) and Proteobacteria.
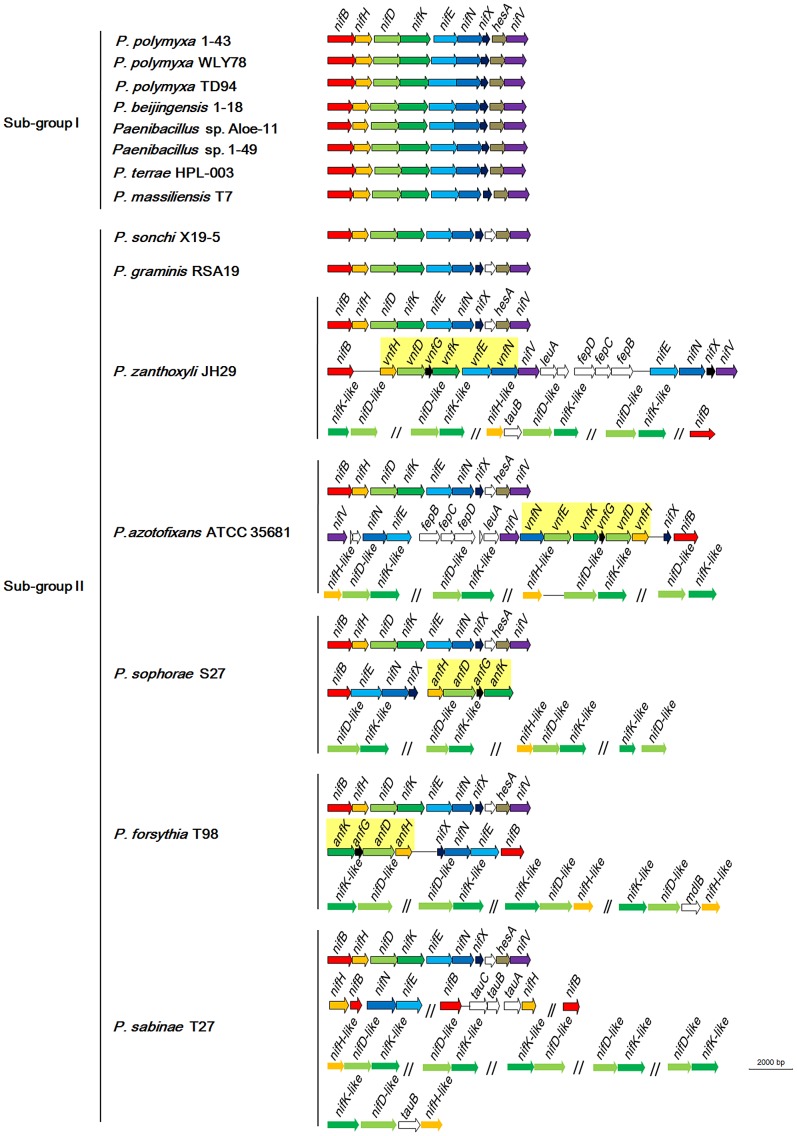
Figure 3. Organization of nif, vnf, anf and nif-like genes in N2-fixing Paenibacillus strains.
nif, vnf, anf and nif-like genes are marked with different colors. The 9 nif genes nifBHDKENXhesAnifV are contiguous within Sub-group I and there is a orf between nifX and hesA within Sub-group II.
Although the nif gene cluster composed of nifB, nifH, nifD, nifK, nifE, nifN, nifX, hesA and nifV is highly conserved among the 15 N2-fixing Paenibacillus strains, there are some variations in DNA sequences of the nif clusters, which can be divided to two sub-groups: Sub-group I and Sub-group II. The 9 genes nifBHDKENXhesAnifV of the nif gene cluster within Sub-group I are contiguous, while there is an orf of 261–561 bp, whose predicted product is unknown, between nifX and hesA within Sub-group II. Except those of P. massiliensis T7 within Sub-group I, and P. sonchi X19-5 and P. graminis RSA19 within Sub-group II, the nif gene clusters generally exhibited more than 90% identity among each Sub-group and about 80% identity between two Sub-groups,
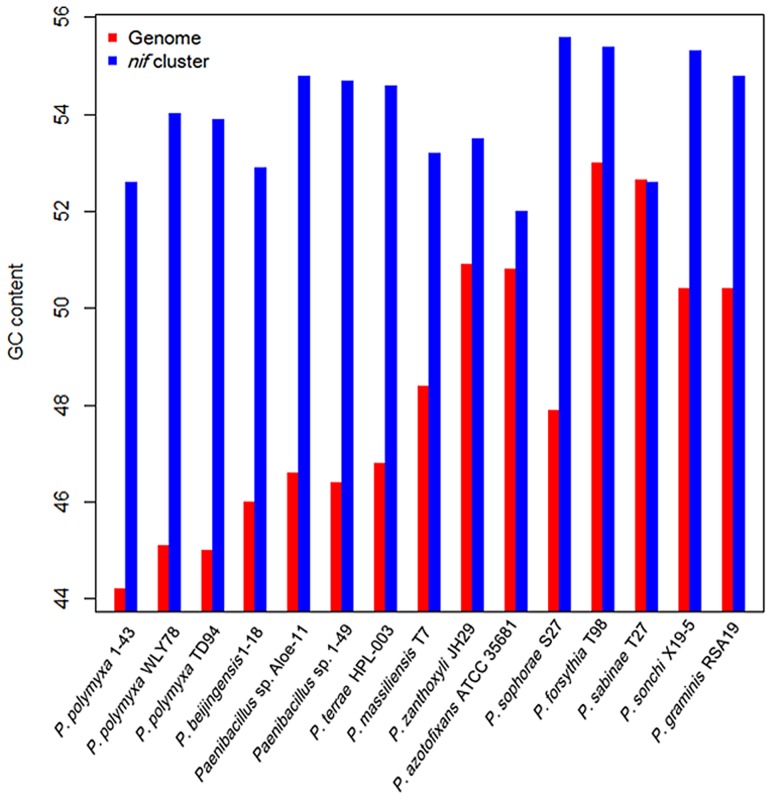
The G+C contents of the nif clusters are higher than those of the average of the entire genomes in other 14 N2-fixing Paenibacillus strains (52–55 vs. 44–54) except that the nif cluster of P. sabinae T27 has the same G+C with the genome (Figure 4). There is a transposase gene, an indicative of HGT, near the nif clusters of Paenibacillus sp. Aloe-11 and P. sabinae T27 (Figure S2). These data suggest that the nif clusters were acquired in Paenibacillus strains by HGT.
Figure 4. Comparison of G+C contents of the nif clusters with those of the average of the chromosomal genomes.
Evolution of the nif gene cluster in Paenibacillus
To elucidate the evolution of the nif gene cluster in Paenibacillus strains, we further compared the chromosomal regions flanking the nif gene clusters to each other among the 15 N2-fixing Paenibacillus strains and to the corresponding chromosomal regions of the non-N2-fixing Paenibacillus strains. We found that ABC transporter ATP-binding protein gene and beta-fructosidase gene/fg-gap repeat protein gene were conserved in the downstream and upstream, respectively, of the nif clusters in the 7 N2-fixing Paenibacillus strains (P. polymyxa 1–43, P. polymyxa WLY78, P. polymyxa TD-94, P. beijingensis 1–18, Paenibacillus. sp. Aloe-11, Paenibacillus sp. 1–49 and P. terrae HPL-003) within Sub-group I (Figure 5A). Unlike in Sub-group I, integral membrane protein gene and FAD/FMN-containing dehydrogenase gene/methyltranferase gene were conserved in the downstream and upstream, respectively, of the nif clusters in all of the 7 N2-fixing Paenibacillus species (P. sonchi X19-5, P. graminis RSA19, P. azotofixans ATCC 35681, P. sophorae S27, P. zanthoxyli JH29, P. forsythia T98 and P. sabinae T27) within Sub-group II (Figure 5C). Combination of the findings that nif clusters fall into two sub-groups according to their identities, these data imply at least two independent acquisitions with insertion of distinct nif variants in different genomic sites of Paenibacillus.
Figure 5. Synteny of the chromosomal regions flanking the nif gene cluster among each sub-group.
(A) nif clusters of Sub-group I. (B) The chromosomal regions of non-N2-fixing strains corresponding to those flanking the nif gene cluster of Sub-group I. (C) nif clusters of Sub-group II.
Notably, the chromosomal regions flanking the nif gene clusters within Sub-group I are homologous to the corresponding regions of the non-N2-fixing P. polmyxa SC2, P. polmyxa E681 and P. peoriae KCTC 3763, suggesting that the nif cluster was lost in these strains (Figure 5B). Our results are consistent with the report that nif gene cluster was lost in cyanobacteria [49].
Sporadic occurrence of alternative nitrogenase
As shown in Figure 3, in addition to the nif cluster encoding Mo-nitrogenase, 2 strains have vnfHDGKEN encoding V-nitrogenase and 2 strains have anfHDGK encoding Fe-nitrogenase. In P. sophorae S27 and P. forsythia T98, anfHDGK are linked with nifBENX, forming a 9.1–9.7 kb cluster. In P. zanthoxyli JH29 and P. azotofixans ATCC 35681, vnfHDGKEN are linked with nifBENXV, fepBCD (encoding iron-enterobactin transporter subunits), leuA and other unknown genes, forming a 20.4–20.9 kb cluster. These anf/vnf clusters are flanked by genes coding for hypothetical proteins. Each alternative nitrogenase cluster contains, as a minimum, vnf/anfH, D, G, and K. The organizations of vnf or anf are largely consistent, but distinct with those of A. vinelandii and Methanococcus maripaludis [4], [51]. It is most likely that anf or vnf gene cluster was recently horizontally transferred to N2-fixing strains which have already had a nif cluster, producing the P. sophorae S27, P. forsythia T98, P. zanthoxyli JH29 and P. azotofixans.
The origin of nif/vnf/anf in Paenibacillus
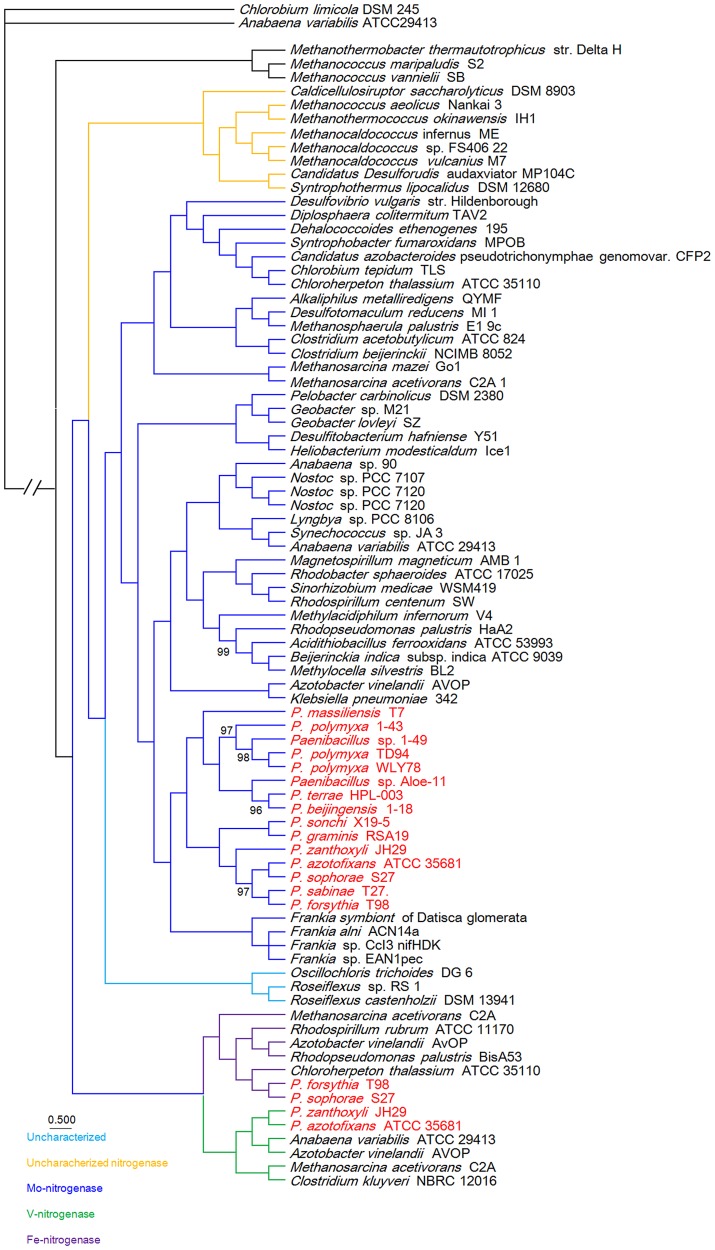
To gain insights into the origin of nif/vnf/anf genes in Paenibacillus, a Bayesian inferred phylogenetic tree was constructed based on the concatenated Nif/Vnf/AnfHDK proteins. Results shown in Figure 6 indicate that Nif/Vnf/AnfHDK proteins of Paenibacillus strains fall into three distinct lineages. This phylogeny exhibits that NifHDK protein homologs formed two distinct clades, one of which was comprised of proteins from hydrogenotrophic methanogens and the other was comprised of proteins from both bacterial and methanogen genomes, in agreement with methanogen origin hypothesis of nitrogen fixation proposed by Boyd et al [10]. Our phylogenetic analysis of the concatenated NifHDK derived from the nifHDK of the nif clusters reveals that all of the 15 N2-fixing Paenibacillus strains form a coherent cluster consisting of two sub-groups, in agreement with the two sub-groups of nif clusters (Figure 7). Notably, the phylogeny reveals that Paenibacillus and Frankia are sister groups to the exclusion of the Firmicute Clostridium, implying that Paenibacillus and Frankia have a common nif gene ancestor. Phylogenies derived from each of the individual NifB, H, D, K, E, N, X and V are congruent with the phylogeny of the concatenated NifHDK (Figure S3, S4, S5, S6, S7, S8, S9, S10).
Figure 6. Bayesian inferred phylogenetic tree of concatenated NifHDK homologs.
The interior node values of the tree are clade credibility values, values lower than 100% are indicated. Branches are colored blue (Mo-nitrogenase, Nif), green (V-nitrogenase, Vnf), purple (Fe-nitrogenase, Anf), light blue (uncharacterized homolog), dark yellow (uncharacterized nitrogenase). The text colored red was Paenibacillus.
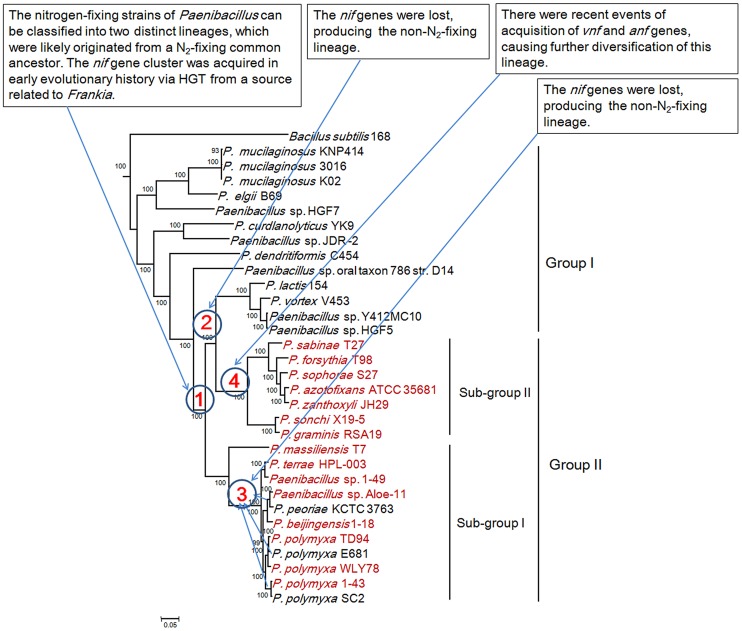
Figure 7. Maximum-likelihood phylogenetic tree of Paenibacillus strains and the 4 possible evolutionary pathways of nitrogen fixation in Paenibacillus.
The tree was constructed based on 275 single-copy core proteins shared by the 31 Paenibacillus genomes and the rooting strain B. subtilis 168. Four likely pathways are marked with number 1–4.
This phylogeny shows that Vnf/Anf proteins of Paenibacillus strains fall into the corresponding homologous lineages. Phylogeny derived from each of the individual VnfH/AnfH, D, G, K, E, N and X is congruent with the phylogeny of the concatenated Vnf/AnfHDK (Figure S3, S4, S5, S6, S7, S8, S9, S10). anf and vnf of Paenibacillus are nested with those of archaeon M. acetivorans, supporting that the ancestor of anf and vnf may originate from archaea.
Phylogenetic analysis
We reconstructed the phylogeny of the 31 genomes based on the concatenation of the 275 core genes that are present in single copy in a genome. The 18 strains including 15 N2-fixing strains and 3 non-N2-fixing strains form a large group including two sub-groups and the other 13 non-N2-fixing strains fall into a large group (Figure 7). The clustering resulting from phylogenetic analysis corresponds well with the species assignments based on average nucleotide identity (ANI) using MUMmer (ANIm) (Table S6) [52]. For examples, P. mucilaginosus K02, P. mucilaginosus 3016 and P. mucilaginosus KNP414 have higher ANIm (98%). N2-fixing strains P. polymyxa 1–43, P. polymyxa WLY78 and P. polymyxa TD94 isolated from China, and non-N2-fixing strains P. polymyxa SC2 and P. polymyxa E681 isolated from China and South Korea, respectively, have higher ANIm (>95%). It is noteworthy that the other 2 unnamed strains Aloe-11 (ANIm≤87%) and 1–49 (ANIm<93%) may represent a novel species, respectively.
This phylogeny suggests that the Paenibacillus ancestor was probably non-fixing and the N2-fixing Paenibacillus strains appeared to occur much later than non-N2-fixing strains. Combination of the data that the nif cluster is conversed in the 15 N2-fixing Paenibacillus strains and the G+C contents of the nif clusters are higher than those of the average of the entire genomes, we proposed that N2-fixing Paenibacillus strains were generated by acquiring the nif cluster via HGT.
The N2-fixing strains of Paenibacillus fall into a large group composed of 2 distinct sub-groups (Sub-group I and Sub-group II), which were likely originated from a N2-fixing common ancestor. This species phylogeny is congruent with the phylogeny of nif genes. The phylogeny suggests that the 8 N2-fixing strains and the 3 non-N2-fixing strains within Sub-group I are most closely related. Nitrogen fixation may have been present in the ancestor of the 8 N2-fixing strains (P. polymyxa 1–43, P. polymyxa WLY78, P. polymyxa TD-94, P. beijingensis1–18, Paenibacillus. sp. Aloe-11, Paenibacillus sp. 1–49, P. terrae HPL-003 and P. massiliensis T7) and the 3 non-N2-fixing strains (P. polymyxa SC2, P. polymyxa E681 and P. peoriae KCTC 3763), and was later lost in the 3 non-N2-fixing strains. This phylogeny also shows that the 7 N2-fixing strains within Sub-group II (P. sonchi X19-5, P. graminis RSA19, P. azotofixans ATCC 35681, P. sophorae S27, P. zanthoxyli JH29, P. forsythia T98 and P. sabinae T27) are sister group with the 4 non-N2-fixing strains P. lactis 154, P. vortex V453, Paenibacillus sp. Y412MC10 and Paenibacillus sp. HGF5. Nitrogen fixation may have been present in the ancestor of the 7 N2-fixing and 4 non-N2-fixing strains and the nif genes were lost, producing the non-N2-fixing P. lactis 154 lineage.
Taken together, the Paenibacillus ancestor was probably non-fixing and the N2-fixing strains of Paenibacillus can be classified into 2 distinct sub-groups, which were likely originated from a N2-fixing common ancestor with minor variation in nif sequences. N2-fixing Paenibacillus strains were generated by acquiring the nif cluster in early evolutionary history via HGT from a source related to Frankia. After these initial acquisitions of the nif gene clusters, the strains that have them now have inherited them by vertical transmission. However, during the process of evolution, the nif cluster was lost, producing the 3 non-N2-fixing strains P. polmyxa SC2, P. polmyxa E681 and P. peoriae KCTC 3763 and the non-N2-fixing lineage P. lactis 154. There were recent events of acquisition of vnf and anf genes, causing further diversification of strains within Sub-group II. The most likely pathways of nitrogen fixation evolution are summarized in Figure 7.
The nif gene cluster is a functional unit for nitrogen fixation
To investigate that the nif gene cluster is a functional unit for nitrogen fixation, the contiguous nine genes nifBHDKENXhesAnifV of the nif cluster and the nifB promoter from P. beijinesis 1–18, a representative of N2-fixing Paenibacillus strains, was PCR amplified and then constructed to vector pHY300PLK and further transferred to E. coli JM109. This yielded the recombinant E. coli strain 1–18. Nitrogenase activity was determined using the acetylene reduction assay (expressed as nmol C2H4/hr/mg protein) [53] and a 15N2 enrichment assay (expressed as δ15N) [54]. As shown in Figure S11, the nine genes nifBHDKENXhesAnifV within the nif cluster enabled E. coli to fix nitrogen, in agreement with our recent report obtained in P. polmyxa WLY78 [50]. The results indicate that the nif cluster is a functional unit for nitrogen fixation, and also a unit of HGT.
The nif gene cluster possesses a σ70-dependent promoter and a GlnR/TnrA-binding site
We recently determined that the nine genes nifB, nifH, nifD, nifK, nifE, nifN, nifX, hesA and nifV within the nif gene cluster in P. polmyxa WLY78 were organized as an operon and that the nifB promoter of the nif cluster is a σ70-dependent promoter −35 (TTGACT) and −10 (TAAGAT) [50]. Here we revealed using bioinformatics analysis that the nif genes within the nif gene clusters among the other 14 N2-fixing Paenibacillus strains are organized as an operon and each of the nif clusters has a σ70-dependent promoter (Figure S12). The σ70-dependent promoter is very distinct from the typical σ54-dependent −24/−12 promoters found upstream of nif genes in Gram-negative N2-fixing bacteria, such as K. pneumoniae and A. vinelandii, whose nif gene expression requires the activation of the transcriptional activator NifA according to the concentration of ammonium and oxygen [55]. Although the σ70-dependent promoter is highly conserved among the 15 N2-fixing Paenibacillus strains, there are some variations in length of interval sequence between the putative transcriptional start site (TSS) and translation start codon (ATG) of nifB (Figure S12).
Unlike in Gram-negative diazotrophs, there is neither nifA gene encoding transcriptional activator NifA, nor NifA-binding site in the promoter region of the nif gene cluster. However, the genomes of the 15 N2-fixing Paenibacillus strains have glnR gene. In the Gram-positive model organism Bacillus subtilis, two transcriptional factors, TnrA and GlnR, control gene expression in response to nitrogen availability [56], [57]. TnrA activates and represses gene transcription when nitrogen is limiting for growth, while GlnR represses gene expression during growth with excess nitrogen. The two proteins bind to DNA sequences (GlnR/TnrA-sites) with a common consensus sequence (TGTNAN7TNACA) [56], [57]. Here we found that the GlnR/TnrA-binding sites exist in the nif promoter regions of the 15 N2-fixing Paenibacillus genomes (Figure S12). The GlnR/TnrA-binding sites are located upstream of the σ70-dependent promoter (−35 and −10) region in Sub-group I strains and some Sub-group II strains, while they are located downstream of the −35 and −10 regions in some Sub-group II strains. The existence of GlnR/TnrA-sites in nif promoter region suggests that regulation mechanisms of nitrogen fixation in Paenibacillus may be different from those of Gram-negative N2-fixing organisms.
Suf system encoding [Fe–S] cluster is highly conserved in N2-fixing and non-N2-fixing Paenibacillus strains
Mo-nitrogenase is a complex [Fe-S] enzyme and the [Fe-S] clusters of nitrogenase play a critical function in electron transfer and in the reduction of substrates driven by the free energy liberated from Mg-ATP hydrolysis [19]. NifU and NifS are generally thought to be specialized for the nitrogenase [Fe-S] cluster assembly of nitrogen-fixing bacteria [58]. However, the genomes of the 15 N2-fixing Paenibacillus strains involved in this study do not possess homologues of nifU and nifS. Here we discovered that a Suf system (sufCDSUB operon) responsible for the formation of [Fe-S] clusters is highly conserved in N2-fixing and non-N2-fixing Paenibacillus strains. Suf system has been reported in E. coli (sufABCDSE) and some other organisms [59]. We deduce that sufCDSUB operon in N2-fixing Paenibacillus strains are involved in synthesis of the [Fe-S] clusters of nitrogenase and other FeS proteins. Perhaps it is because there is a sufCDSUB operon in non-N2-fixing Paenibacillus strain, a single event of HGT of the nif gene cluster will transfer a non-N2-fixing Paenibacillus strain to a N2-fixing Paenibacillus strain.
Multiple nif genes in Paenibacillus
In addition to nifBHDKENXhesAnifV within the nif gene cluster, there is a set of additional nifBEN which are linked together with vnf or anf in the 4 species: P. zanthoxyli JH29 and P. azotofixans ATCC 35681, P. sophorae S27 and P. forsythia T98. Since the additional nifBEN form a cluster with vnf or anf, it is likely that they were horizontally transferred to the 4 species with vnf or anf. There are a cluster of nifHBEN, 2 nifB and 1 nifH located at different sites outside of the nif gene cluster in P. sabinae T27. The phylogenetic trees based on each of the individual NifB, NifH, NifE and NifN protein sequences (Figure S3, S4, S5, S6, S7, S8, S9, S10) show that each of them is clustered with its homolog derived from the nif gene clusters of Paenibacillus, suggesting that these genes derived from gene duplication. Transposases near the nifBHEN and nifB in P. sabinae T27 suggest that these genes may originate from gene duplication (Figure S2). Our previous results demonstrated that the 3 nifH genes from P. sabinae T27 could complement the K. pneumoniae nifH− mutant [60], suggesting that these nifH genes are functional in nitrogen fixation. However, we are not sure that the multiple nifHBEN are positively related to high nitrogenase activity.
Multiple nitrogenase-like genes in Paenibacillus
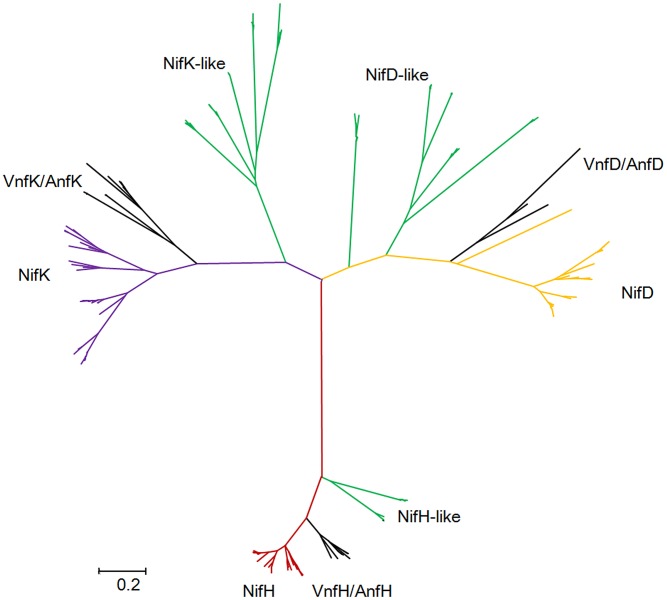
Our studies revealed that there are nitrogenase-like genes including 1–2 nifH-like and 4–6 pairs of nifDK-like genes in the 5 species within Sub-group II: P. azotofixans ATCC 35681, P. sophorae S27, P. zanthoxyli JH29, P. forsythia T98 and P. sabinae T27 (Figure 3). Alignments of NifH-like sequences with NifH sequences show that 4Fe-4S iron sulfur cluster ligating cysteines (Cys97 and Cys132), ADP-ribose binding arginine (Arg101) and the P-loop/MgATP binding motif are invariant, suggesting that NifH-like proteins may function analogously to NifH (γ subunit of nitrogenase) (Figure S13). Conversely, NifD/NifK-like sequences are highly diverged from both α and β subunits of nitrogenase. For example, FeMoco-ligating residues at αCys275 and αHis442, and P-cluster-ligating residues at Cys62, Cys88 and Cys154 of NifD, are not conserved in NifD-like sequences (Figure S14). The residues ligating P-cluster at Cys70, Cys95 and Cys153 of NifK are not conserved in NifK-like sequences (Figure S15). Our results are in agreement with previous reports obtained in studies with Archaea and Firmicutes Clostridium [4], [8]. Further, phylogenetic analysis reveals that the NifH/NifD/NifK-like sequences form distinct groups which are clearly divergent from conventional nitrogenase (Figure 8).
Figure 8. Maximum-likelihood phylogenetic tree of Mo-, Fe- and V-nitrogenases and nitrogenase-like sequences.
Nif/Vnf/AnfH, Nif/Vnf/AnfD, Nif/Vnf/AnfK, Nif/Vnf/AnfH-like, Nif/Vnf/AnfD-like and Nif/Vnf/AnfK sequences were derived from the 15 N2-fixing Paenibacillus strains and other representative species.
Discussion
In this study, we sequenced the genomes of 11 N2-fixing Paenibacillus strains and made a comparative genomic analysis with 20 other strains (4 N2-fixing and 16 non-N2-fixing strains) that were sequenced previously. Our analysis of the total 31 genomes revealed that of the 55504 putative protein-coding genes, 37105, which made up 66.9% of the genes in the pan genome, were represented in only one genome of Paenibacillus spp., suggesting a remarkable degree of HGT in shaping the genomes of each of the genus. It is generally accepted that abundance of mobile genetic elements correlates positively with the frequency of HGT. We discovered that each genome of all of the 31 strains contains 1–10 prophages and/or prophage remnants and 3–118 IS elements, supporting that these strains are rich in mobile genetic elements. The existence of transposable elements and prophage near the nif gene and nif gene cluster suggest that they may be involved in HGT and loss of nif genes. Our demonstration that the nif cluster from P. beijinesis 1–18 enabled E. coli to have nitrogen fixation ability supports that the nif cluster is a functional unit for nitrogen fixation and also a unit of HGT.
Genomic islands are known to have contributed to the evolution of microbial genomes by HGT in many bacteria, influencing traits such as antibiotic resistance, symbiosis and fitness, and adaptation in general [61]. The evolutionary advantage of genomic islands is that a large number of genes (e.g. operon, gene clusters encoding related functions) may be horizontally transferred and incorporated en bloc into the recipient genome in a single step [62]. Genome sequence analysis here revealed that nine genes nifB, nifH, nifD, nifK, nifE, nifN, nifX, hesA and nifV which are organized as a cluster arranged within 10.5–12 kb region are highly conserved in the 15 N2-fixing Paenibacillus strains. The sizes of nif clusters of Paenibacillus fall into the range of 10–200 kb genome islands in length. Also, the G+C contents of the nif clusters are higher than those of the average of the genomes in 14 N2-fixing strains except P. sabinae T27, in agreement with genome islands whose G+C content often differs from that of the rest of the genome. This favored the hypothesis that the nif region in Paenibacillus constitutes a nitrogen fixation island, as discovered in other nitrogen fixers [14], [63]. For example, nif genes are part of an island in Wolinella succinogenes [14] and in Rhizobium leguminosarum [63]. nif genes organized as clusters are also found in many other N2-fixing organisms. For examples, 20 nif genes are organized in 8 operons (nifJC, nifHDKTY, nifEN, nifUSVW, nifZM, nifF, nifLA, nifBQ) within ca. 24 kb of DNA in the chromosome of K. pneumoniae [3]. A total of 17–20 ORFs including 9–11 nif genes were organized as a cluster arranged within 17.3–18.5 kb regions among 4 Frankia strain: Frankia sp. EuIK1, Frankia sp. EAN1pec, Frankia sp. ACN14a and Frankia sp. HFPCcI3 [48]. In the Cyanothece 51142 genome, a representative of nonheterocystous cyanobacteria, the majority of genes involved in nitrogen fixation are located in a contiguous 28 kb cluster of 34 genes [49]. The different gene content and organization of nif genes indicate that complex evolutionary history of nif genes, and also suggest differences in protein requirements for nitrogenase synthesis and regulation of nitrogen fixation.
Phylogeny of the concatenated NifHDK proteins revealed that Paenibacillus and Frankia are sister groups to the exclusion of the Firmicute Clostridium, implying that Paenibacillus and Frankia have a common nif gene ancestor. Our results are consistent with the previous reports that Frankia and cyanobacterium Anabaena were sister groups to the exclusion of the Firmicute Clostridium [7]. Some common features found in the nif clusters support that Paenibacillus and Frankia are closely related. The first common feature is hesA, which is conserved in the nif clusters of Paenibacillus, Frankia and cyanobacteria, but not in N2-fixing Gram-negative and other Gram-positive bacteria, such as Clostridium. The second common feature is the compact organized nifHDKENX which is found in the nif clusters of Paenibacillus and Frankia, but not in Clostridium spp. In contrast, gene content and organization varied greatly between the nif clusters of Paenibacillus and Clostridium, although both genera Paenibacillus and Clostridium belong to the low G+C and Gram-positive Firmicutes. For example, nifN-B fusion gene was found in the nif gene clusters of the three species of Clostridia: C. acetobutylicum, C. beijerinckii, and C. pasteurianum [59], [64]. Also, the nif gene clusters of C. acetobutylicum and C. beijerinckii have nifI1 and nifI2 (homologs of glnB), which are involved in post-translational regulation of nitrogenase activity in response to fixed nitrogen [65]. These data suggest that the gene content and organization of the nif cluster of anaerobic Clostridium spp. are similar with those of M. acetovorans and M. maripaudis whose nif clusters also contain nifI1 and nifI2 located between nifH and nifDK [51], [65].
Phylogeny of the concatenated 275 single-copy core genes (Figure 7) suggests that the ancestral Paenibacillus did not fix nitrogen. Genome sequencing revealed that the nif cluster is highly conserved in all of the 15 N2-fixing strains and the G+C contents of the nif clusters are higher than those of the average of the genomes in 14 N2-fixing strains except P. sabinae T27. Also, phylogeny of the concatenated NifHDK proteins (Figure 6) revealed that Paenibacillus and Frankia are sister groups. All of these facts and evidences indicate that N2-fixing Paenibacillus strains may be generated by acquiring the nif cluster via HGT from a source related to Frankia in early evolutionary history. Strain phylogeny (Figure 7) also shows that the 15 N2-fixing strains of Paenibacillus fall into 2 distinct sub-groups, consistent with phylogeny of nif genes (Figure 6). The nif clusters show some variation between two sub-groups, and the genes surrounding the nif clusters from two Sub-groups are conserved and distinct. These data imply at least two independent acquisitions with insertion of distinct nif variants in different genomic sites of Paenibacillus.
Furthermore, strain phylogeny suggests that nitrogen fixation may have been present in the ancestor of the 8 N2-fixing strains (P. polymyxa 1–43, P. polymyxa WLY78, P. polymyxa TD94, P. beijingensis1–18, Paenibacillus. sp. Aloe-11, Paenibacillus sp. 1–49, P. terrae HPL-003 and P. massiliensis T7) and the 3 non-N2-fixing strains (P. polmyxa SC2, P. polmyxa E681 and P. peoriaeKCTC 3763) within Sub-group I, and was later lost in the 3 non-N2-fixing strains (P. polmyxa SC2, P. polmyxa E681 and P. peoriae KCTC 3763). Notably, the model P. polymyxa is a N2-fixing species, and now this species includes both N2-fixing and non-N2-fixing strains. These closely related strains of this group were isolated from plant rhizospheres and from different geological locations of China, South Korea and Republic of Korea. Likewise, it is likely that nitrogen fixation may have been present and was later lost in the non-N2-fixing lineage P. lactis 154. The members of this lineage were isolated from complex locations. For examples, P. lactis 154 was isolated from milk, Paenibacillus sp. HGF5 from human intestinal microflora and Paenibacillus sp. Y412MC10 from hot spring, and P. vortex V453 is known to develop complex colonies with intricate architectures.
The newly sequenced genomes revealed that the 4 Paenibacillus species P. sophorae S27, P. forsythia T98, P. zanthoxyli JH29 and P. azotofixans have the second nif cluster which carrying vnf or anf, in addition to the nif cluster. anfHDGK are clustered with nifBENX in a 9.1–9.7 kb region in P. sophorae S27 and P. forsythia T98, vnfHDGKEN are clustered with nifBENXV, fepBCD, leuA and other unknown genes in a 20.4–20.9 kb region in P. zanthoxyli JH29 and P. azotofixans ATCC 35681. Phylogeny of the concatenated Nif/Anf/VnfHDK proteins indicates that anfHDGK and vnfHDGKEN of Paenibacillus originate differently from nifHDK, and may be not duplicated from their nifHDK. It is most likely that the nif cluster carrying anf/vnf genes was recently horizontally transferred to N2-fixing strains which have already had the nif cluster, producing P. sophorae S27, P. forsythia T98, P. zanthoxyli JH29 and P. azotofixans. These species were isolated from plant rhizosphere from China and Brazil. Our results are consistent with the recent reports that both Nif and Anf evolved in the methanogenic archaea, and anf or vnf derived from duplication of nif [8]. As described above, phylogenies of the concatenated Anf/VnfHDK and each of individual Anf/VnfH, D and K show that Paenibacillus strains fall into Anf and Vnf clusters, respectively. However, we found that the conserved residues in the P-loop binding motif of AnfH do not exist in P. sophorae S27, and the residues ligating P-cluster at Cys70 and Cys95 of VnfK do not exist in P. zanthoxyli JH29. Perhaps the residues ligating P-cluster or in P-loop binding motif are located on the other sites in VnfK and AnfH, respectively.
This study reveals that HGT of nif/anf/vnf gene cluster contributed to evolution of nitrogen fixation in Paenibacillus. Usually, a vehicle is needed to transfer genes efficiently between different species. It is thought that foreign DNAs are mainly transferred by means of plasmids or bacteriophages, as well as direct uptake by the host itself [58], [66], [67]. The best studied example of HGT of nif genes is symbiosis island of Mesorhizobium loti. The symbiosis island, a 502-kb chromosomally integrated element containing nif genes, was integrated into a phenylalanine tRNA gene mediated by a P4-type integrase encoded at the left end of the symbiosis island [68]–[70]. However, a phenylalanine tRNA gene near the nif cluster is not found, suggesting that it may be not transferred by P4-type integrase. But we found that there is a transposase gene, an indicative of HGT, near the nif clusters of Paenibacillus sp. Aloe-11 and P. sabinae T27 and near the anf cluster of P. sophorae S27. Also, a transcriptional regulator gene of araC type, which is known to be involved primarily in regulating pathogenicity islands in some bacteria but is also present in nonpathogenic organisms [62], neighbors the nif clusters of P. polmyxa TD94 and Paenibacillus sp. 1–11.
The deviant G+C content is one of the indicative used to detect HGT [67]. The G+C contents of the nif clusters are higher than those of the average of the entire genomes (52–55 vs. 44–53) in the 14 N2-fixing Paenibacillus strains except P. sabinae T27, supporting that the nif gene clusters in these strains are acquired by HGT. The similar G+C contents and high identities of nif genes among the 15 nif clusters suggest that these nif clusters originated from a common ancestor with minor variation. The G+C contents of the anf cluster is higher than the average of the genome in P. sophorae S27 (51% vs. 40%), and is lower than the average of the genome in P. forsythia T98 (51% vs. 53%). The G+C contents of the vnf cluster is the same (51% vs. 51%) as the average of the chromosomal genome in P. azotofixans ATCC 35681 and P. zanthoxyli JH29. A higher G+C contents of the nif cluster were found in some N2-fixing bacteria, such as P. stutzeri A1501 (66.8% vs. 63.8%) [12]. In rhizobia, the nif genes are located on either plasmids or genomic islands, which are prone to transfer between related bacteria [71]. However, the G+C contents of these plasmids and genomic islands are generally lower than the average of the chromosomal genome [72]–[74]. However, the G+C contents of the nif clusters are similar with those of the average of the entire genomes in the sequenced Frankia strains (69% vs. 70% in Frankia sp. HFPCcI3, 70% vs. 71% in Frankia sp. EAN1pec and 71% vs. 72% in Frankia alni ACN14a). It is generally accepted that although the deviant G+C content can be used to detect HGT, detection of HGT depends on a combination of several methods. This is because it is hard to detect HGT via deviant G+C content, if HGT occurred between the organisms with the same G+C contents [67].
Our genome sequencing revealed that there are nitrogenase-like genes including 1–2 nifH-like and 4–6 pairs of nifDK-like genes in the 5 species within Sub-group II: P. azotofixans ATCC 35681, P. sophorae S27, P. zanthoxyli JH29, P. forsythia T98 and P. sabinae T27 (Figure 3 and Table S5). Alignment of conserved residues ligating 4Fe-4S in NifH and ligating P-cluster and FeMoco and phylogenetic analysis in in NifD/K revealed that the nif-like and nifDK-like genes are clustered with those of archaea and Firmicutes such as Clostridia [4]. The data that NifH/NifD/NifK-like sequences fall into distinct groups by phylogenetic analysis suggest that multiple nifH-like and nifDK-like genes may result from gene duplication. The existence of transposases near the nifDK-like genes also suggested that multiple nifDK-like genes may result from gene duplication. It was proposed that Nif emerged from a nitrogenase-like ancestor approximately 1.5–2.2 Ga [10]. We wonder why there are so many nifDK-like genes in these Paenibacillus species. The determination of the function of nitrogenase-like genes will clarify their relation with nitrogen fixation.
Materials and Methods
Genome sequencing, assembly, and annotation
The draft sequences of 11 test Paenibacillus strains were produced by using Illumina paired-end sequencing technology at the BGI–Shenzhen (Table 2). Assembly was conducted by using SOAPdenovo v. 1.04 assembler [75]. Gene prediction was made using Glimmer v3.0 [76]. Annotation of protein coding sequence was performed by using the Basic Local Alignment Search Tool (BLAST) against the COG, Kyoto Encyclopedia of Genes and Genomes (KEGG) databases and NCBI nr protein database. The draft genomes of the 11 test Paenibacillus strains have been deposited in GenBank and the project accession numbers are listed in Table 2. Prophage was identified using PHAST [77].
Comparative genomics
Pan Genome Analysis Pipeline of PGAP [78] was used to identify all of the orthologous pairs between test Paenibacillus genomes. The common dataset of shared genes among test strains was defined as their core genome. The total set of genes within test genomes was defined as the pan genome. The set of genes in each strain not shared with other strains was defined as unique genes. The average nucleotide identity (ANI) between strains of the 31 sequenced genomes were calculated using MUMmer [52]. Multiple alignment of conserved genomic sequence was using Mauve [79]. The genomes sequenced in this study are listed in Table S1.
Phylogenetic analysis
Single gene alignments were aligned with molecular evolutionary genetics analysis (MEGA) [80]. The neighbor-joining trees were constructed by using the same software, and 1,000 bootstraps were done. Bayesian inferred phylogenetic tree of concatenated HDK homologs was generated using the MrBayes package [81]. A maximum-likelihood phylogenetic tree of Paenibacillus species was constructed based on 275 single-copy core proteins shared by 31 Paenibacillus genomes and the genome of Bacillus subtilis 168 according to the following methods: (i) multiple alignment of amino acid sequences were carried out by ClustalW (version 2.1) [82] (ii) conserved blocks from multiple alignment of test protein were selected by using Gblocks [83] (iii) ML tree were constructed using PhyML (version 3.0) [84] software (iv) CONSEL program [85] was used to select the best model of the trees.
Construction of the recombinant plasmid and E. coli strain
Genomic DNA of diazotrophic P. beijingensis 1–18 was used as a template for cloning nif genes. A 10.7 kb Xba I -BamH I DNA fragment containing the nif cluster (a 300 bp promoter region and the contiguous nine genes nifBHDKENXhesAnifV and 184 bp downstream of the stop codon TAA of nifV) was PCR amplified with primers nif cluster-up (5′-TGCTCTAGAGGGAATATAACGTGGAGAGG-3′) and nif cluster-down (5′-CGCGGATCCCATTATACAGCACTATATTG-3′) and then ligated to Xba I and BamH I sites of pHY300PLK, yielding plasmid pHY300-18 (Pnif+nif cluster). The plasmid was then transferred to E. coli JM109, yielding the recombinant E. coli 1–18.
Acetylene reduction assays
For acetylene reduction assays, P. beijingensis 1–18 and the recombinant E. coli strain 18 were grown overnight in LD medium, then diluted into nitrogen-deficient medium and grown for 15–18 h. Following this stage, the cultures were collected and resuspended in an N-free medium to an OD600 of 0.2–0.4 in a serum bottle for nitrogenase derepression. The serum bottle was vacuumed and charged with argon gas. After 5–6 h, C2H2 (10% of the headspace volume) was injected into the serum bottle. After 30 min to 1 h, C2H4 was analyzed by Gas Chromatography [53].
15N2 incorporation assay
Paenibacillussp.1–18 and the recombinant E. coli strain 1–18 were grown overnight in LD medium. The cultures were collected and resuspended in 70 ml N-free medium to an OD600 of 0.4 in the 120 ml serum bottle. The serum bottles were filled with N2 gas, and then 8-ml gas was removed and 5 ml 15N2 (99%+, Shanghai Engineering Research Center for Stable Isotope) gas was injected. After 72 hours of incubation at 30°C, the cultures were collected, freeze dried, ground, weighed and sealed into tin capsules. Isotope ratios are expressed as δ15N whose values are a linear transform of the isotope ratios 15N/14N, representing the per mille difference between the isotope ratios in a sample and in the atmospheric N2 [54].
Data access
The genome sequences used in this study were submitted to the GenBank, the accession number was shown in Table 2.
Supporting Information
Comparison of the nif gene cluster of Paenibacillus with those of the representative N2-fixing bacteria and archaea. (A) Paenibacillus polymyxa 1–43, (B) Azotobacter vinelandii, (C) Klebsiella oxytoca M5al, (D) Nostoc punctiforme PCC 73102, (E) Frankia sp. EAN1pec, (F) Clostridium acetobutylicum, (G) Methanococus maripaludis.
(TIF)
IS elements or prophages linked with the nif gene, nif cluster and nif-like genes.
(TIF)
Neighbor joining phylogenetic tree of the NifB sequences derived from Paenibacillus and other representative species. A total of 1,000 bootstrap replicates were made, and bootstrap values are indicated at each node.
(TIF)
Neighbor joining phylogenetic tree of the NifH, VnfH, AnfH and NifH-like protein sequences derived from Paenibacillus and other representative species. A total of 1,000 bootstrap replicates were made, and bootstrap values are indicated at each node.
(TIF)
Neighbor joining phylogenetic tree of the NifD, VnfD, AnfD and NifD-like protein sequences derived from Paenibacillus and other representative species. A total of 1,000 bootstrap replicates were made, and bootstrap values are indicated at each node.
(TIF)
Neighbor joining phylogenetic tree of the NifK, VnfK, AnfK and NifK-like protein sequences derived from Paenibacillus and other representative species. A total of 1,000 bootstrap replicates were made, and bootstrap values are indicated at each node.
(TIF)
Neighbor joining phylogenetic tree of the NifE, VnfE and NifE-like protein sequences derived from Paenibacillus and other representative species. A total of 1,000 bootstrap replicates were made, and bootstrap values are indicated at each node.
(TIF)
Neighbor joining phylogenetic tree of the NifN, VnfN and NifN-like protein sequences derived from Paenibacillus and other representative species. A total of 1,000 bootstrap replicates were made, and bootstrap values are indicated at each node.
(TIF)
Neighbor joining phylogenetic tree of the NifX protein sequences derived from Paenibacillus and other representative species. A total of 1,000 bootstrap replicates were made, and bootstrap values are indicated at each node.
(TIF)
Neighbor joining phylogenetic tree of the NifV protein sequences derived from Paenibacillus and other representative species. A total of 1,000 bootstrap replicates were made, and bootstrap values are indicated at each node.
(TIF)
Nitrogen fixation abilities of P. beijingensis 1–18 (WT) and recombinant E. coli 1–18 strain. (A) Nitrogenase activities determined by using acetylene reduction assay. (B) Nitrogen fixation ability determined by using for 15N2 incorporation. Error bars indicate the standard deviation observed from at least two independent experiments.
(TIF)
The σ70-depedent promoters of the nif clusters and the GlnR/TnrA-binding sites in the nif promoter regions in Paenibacillus strains.
(TIF)
Alignments of crucial residues surrounding the P-loop/MgATP binding motif, cysteine ligating 4Fe-4S and arginine ligating ADP-ribose in NifH and NifH-like protein sequences from Paenibacillus and other organisms.
(TIF)
Alignments of crucial residues ligating FeMo-co or P-cluster in NifD and NifD-like protein sequences from Paenibacillus and other organisms.
(TIF)
Alignments of crucial residues ligating FeMo-co or P-cluster in in NifK and NifK-like protein sequences from Paenibacillus and other organisms.
(TIF)
The genomes sequenced in this study.
(DOCX)
Comparison of COG assignments between non-N2-fixing and N2-fixing Paenibacillus strains.
(DOCX)
Transposons present in the genomes of 31 Paenibacillus strains. The following information is provided for each putative transposon in genomes: transposon family, transposases, and numbers of copies of intact or remnant transposons in each genome.
(DOCX)
Prophages present in the genomes of 31 Paenibacillus strains. The following information is provided for each prophage: insertion site, size, locus tags, and selected cargo genes.
(DOCX)
The nitrogen fixation genes and nitrogenase-like genes in the nitrogen-fixing Paenibacillus strains.
(DOCX)
Average Nucleotide Identity (%) based on whole genome alignments.
(XLSX)
Acknowledgments
We thank Dr. Peter Young for valuable comments on the phylogenetic analysis, Dr. Ray Dixon for thoughtful comments on the nitrogenase-like genes and Dr. Xu-Xian Zhang for reading the manuscript.
Funding Statement
This work was supported by funds from the National “973” Project (Grant No. 2010CB126504) and the National Nature Science Foundation of China (Grant No. 31270129). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1. Falkowski PG (1997) Evolution of the nitrogen cycle and its influence on the biological sequestration of CO2 in the ocean. Nature 387: 272–275. [Google Scholar]
- 2. Dos Santos PC, Fang Z, Mason SW, Setubal JC, Dixon R (2012) Distribution of nitrogen fixation and nitrogenase-like sequences amongst microbial genomes. BMC Genomics 13: 162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Arnold W, Rump A, Klipp W, Priefer UB, Pühler A (1988) Nucleotide sequence of a 24,206-base-pair DNA fragment carrying the entire nitrogen fixation gene cluster of Klebsiella pneumoniae . J Mol Biol 203: 715–738. [DOI] [PubMed] [Google Scholar]
- 4. Setubal JC, dos Santos P, Goldman BS, Ertesvåg H, Espin G, et al. (2009) Genome sequence of Azotobacter vinelandii, an obligate aerobe specialized to support diverse anaerobic metabolic processes. J Bacteriol 191: 4534–4545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Normand P, Bouquet J (1989) Phylogeny of nitrogenase sequences in Frankia and other nitrogen-fixing microorganisms. J Mol Evol 29: 436–447. [DOI] [PubMed] [Google Scholar]
- 6. Normand P, Gouy M, Cournoyer B, Simonet P (1992) Nucleotide sequence of nifD from Frankia alni strain ArI3: phylogenetic inferences. Mol Biol Evol 9: 495–506. [DOI] [PubMed] [Google Scholar]
- 7. Hartmann LS, Barnum SR (2010) Inferring the evolutionary history of Mo-dependent nitrogen fixation from phylogenetic studies of nifK and nifDK . J Mol Evol 71: 70–85. [DOI] [PubMed] [Google Scholar]
- 8. Raymond J, Siefert JL, Staples CR, Blankenship RE (2004) The natural history of nitrogen fixation. Mol Biol Evol 21: 541–554. [DOI] [PubMed] [Google Scholar]
- 9. Leigh JA (2000) Nitrogen fixation in methanogens: the archaeal perspective. Crit Rev Microbiol 2: 125–131. [PubMed] [Google Scholar]
- 10. Boyd E, Hamilton T, Peters J (2011) An alternative path for the evolution of biological nitrogen fixation. Front Microbiol 2: 205 doi: 10.3389/fmicb.2011.00205 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Boyd E, Anbar A, Miller S, Hamilton T, Lavin M, et al. (2011) A late methanogen origin for molybdenum-dependent nitrogenase. Geobiology 9: 221–232. [DOI] [PubMed] [Google Scholar]
- 12. Yan Y, Yang J, Dou Y, Chen M, Ping S, et al. (2008) Nitrogen fixation island and rhizosphere competence traits in the genome of root-associated Pseudomonas stutzeri A1501. Proc Natl Acad Sci U S A 105: 7564–7569. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Pedrosa FO, Monteiro RA, Wassem R, Cruz LM, Ayub RA, et al. (2011) Genome of Herbaspirillum seropedicae strain SmR1, a specialized diazotrophic endophyte of tropical grasses. PLoS Genet 7: e1002064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Baar C, Eppinger M, Raddatz G, Simon J, Lanz C, et al. (2003) Complete genome sequence and analysis of Wolinella succinogenes . Proc Natl Acad Sci U S A 100: 11690–11695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Hu Y, Fay AW, Lee CC, Yoshizawa J, Ribbe MW (2008) Assembly of nitrogenase MoFe protein. Biochemistry 47: 3973–3981. [DOI] [PubMed] [Google Scholar]
- 16. Rubio LM, Ludden PW (2008) Biosynthesis of the iron-molybdenum cofactor of nitrogenase. Annu Rev Microbiol 62: 93–111. [DOI] [PubMed] [Google Scholar]
- 17. Kaiser JT, Hu Y, Wiig JA, Rees DC, Ribbe MW (2011) Structure of precursor-bound NifEN: a nitrogenase FeMo cofactor maturase/insertase. Science 331: 91–94. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Joerger RD, Bishop PE, Evans HJ (1988) Bacterial alternative nitrogen fixation systems. Crit Rev Microbiol 16: 1–14. [DOI] [PubMed] [Google Scholar]
- 19. Rubio LM, Ludden PW (2005) Maturation of nitrogenase: a biochemical puzzle. J Bacteriol 187: 405–414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Chisnell J, Premakumar R, Bishop P (1988) Purification of a second alternative nitrogenase from a nifHDK deletion strain of Azotobacter vinelandii . J Bacteriol 170: 27–33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Davis R, Lehman L, Petrovich R, Shah VK, Roberts GP, et al. (1996) Purification and characterization of the alternative nitrogenase from the photosynthetic bacterium Rhodospirillum rubrum . J Bacteriol 178: 1445–1450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Schneider K, Muller A, Schramm U, Klipp W (1991) Demonstration of a molybdenum- and vanadium-dependent nitrogenase in a nifHDK-deletion mutant of Rhodobacter capsulatus . Eur J Biochem 195: 653–661. [DOI] [PubMed] [Google Scholar]
- 23. Lal S, Tabacchioni S (2009) Ecology and biotechnological potential of Paenibacillus polymyxa: a minireview. Indian J Microbiol 49: 2–10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. McSpadden Gardener BB (2004) Ecology of Bacillus and Paenibacillus spp. in agricultural systems. Phytopathology 94: 1252–1258. [DOI] [PubMed] [Google Scholar]
- 25. Montes MJ, Mercadé E, Bozal N, Guinea J (2004) Paenibacillus antarcticus sp. nov., a novel psychrotolerant organism from the Antarctic environment. Int J Syst Evol Microbiol 54: 1521–1526. [DOI] [PubMed] [Google Scholar]
- 26. Ouyang J, Pei Z, Lutwick L, Dalal S, Yang L, et al. (2008) Paenibacillus thiaminolyticus: a new cause of human infection, inducing bacteremia in a patient on hemodialysis. Ann Clin Lab Sci 38: 393–400. [PMC free article] [PubMed] [Google Scholar]
- 27. Ash C, Priest FG, Collins MD (1993) Molecular identification of rRNA group 3 bacilli (Ash, Farrow, Wallbanks and Collins) using a PCR probe test. Antonie van Leeuwenhoek 64: 253–260. [DOI] [PubMed] [Google Scholar]
- 28. Ma Y, Xia Z, Liu X, Chen S (2007) Paenibacillus sabinae sp. nov., a nitrogen-fixing species isolated from the rhizosphere soils of shrubs. Int J Syst Evol Microbiol 57: 6–11. [DOI] [PubMed] [Google Scholar]
- 29. Ma Y, Zhang J, Chen S (2007) Paenibacillus zanthoxyli sp. nov., a novel nitrogen-fixing species isolated from the rhizosphere of Zanthoxylum simulans . Int J Syst Evol Microbiol 57: 873–877. [DOI] [PubMed] [Google Scholar]
- 30. Ma Y, Chen S (2008) Paenibacillus forsythiae sp. nov., a nitrogen-fixing species isolated from rhizosphere soil of Forsythia mira . Int J Syst Evol Microbiol 58: 319–323. [DOI] [PubMed] [Google Scholar]
- 31. Hong Y, Ma Y, Zhou Y, Gao F, Liu H, et al. (2009) Paenibacillus sonchi sp. nov., a nitrogen-fixing species isolated from the rhizosphere of Sonchus oleraceus . Int J Syst Evol Microbiol 59: 2656–2661. [DOI] [PubMed] [Google Scholar]
- 32. Jin H, Lv J, Chen S (2011) Paenibacillus sophorae sp. nov., a nitrogen-fixing species isolated from the rhizosphere of Sophora japonica . Int J Syst Evol Microbiol 61: 767–771. [DOI] [PubMed] [Google Scholar]
- 33. Jin H, Zhou Y, Liu H, Chen S (2011) Paenibacillus jilunlii sp. nov., a nitrogen-fixing species isolated from the rhizosphere of Begonia semperflorens . Int J Syst Evol Microbiol 61: 1350–1355. [DOI] [PubMed] [Google Scholar]
- 34. Xie J, Zhang L, Zhou Y, Liu H, Chen S (2012) Paenibacillus taohuashanense sp. nov., a nitrogen-fixing species isolated from rhizosphere soil of the root of Caragana kansuensis Pojark . Antonie van Leeuwenhoek 102: 735–741. [DOI] [PubMed] [Google Scholar]
- 35. Wang L, Li J, Li QX, Chen S (2013) Paenibacillus beijingensis sp. nov., a nitrogen-fixing species isolated from wheat rhizosphere soil. Antonie van Leeuwenhoek 104: 675–683. [DOI] [PubMed] [Google Scholar]
- 36. Chow V, Nong G, John FJS, Rice JD, Dickstein E, et al. (2012) Complete genome sequence of Paenibacillus sp. strain JDR-2. Stand Genomic Sci 6: 1–10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Mead DA, Lucas S, Copeland A, Lapidus A, Cheng J, et al. (2012) Complete genome sequence of Paenibacillus strain Y412MC10, a novel Paenibacillus lautus strain isolated from Obsidian hot spring in Yellowstone national park. Stand Genomic Sci 6: 381–400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Ma M, Wang Z, Li L, Jiang X, Guan D, et al. (2012) Complete genome sequence of Paenibacillus mucilaginosus 3016, a bacterium functional as microbial fertilizer. J Bacteriol 194: 2777–2778. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Kim JF, Jeong H, Park S, Kim S, Park YK, et al. (2010) Genome sequence of the polymyxin-producing plant-probiotic rhizobacterium Paenibacillus polymyxa E681. J Bacteriol 192: 6103–6104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Ma M, Wang C, Ding Y, Li L, Shen D, et al. (2011) Complete genome sequence of Paenibacillus polymyxa SC2, a strain of plant growth-promoting rhizobacterium with broad-spectrum antimicrobial activity. J Bacteriol 193: 311–312. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Sirota-Madi A, Olender T, Helman Y, Brainis I, Finkelshtein A, et al. (2012) Genome sequence of the pattern-forming social bacterium Paenibacillus dendritiformis C454 chiral morphotype. J Bacteriol 194: 2127–2128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Ding R, Li Y, Qian C, Wu X (2011) Draft Genome sequence of Paenibacillus elgii B69, a strain with broad antimicrobial activity. J Bacteriol 193: 4537–4537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Jeong H, Choi S, Park S, Kim S, Park S (2012) Draft genome sequence of Paenibacillus peoriae strain KCTC 3763T . J Bacteriol 194: 1237–1238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Sirota-Madi A, Olender T, Helman Y, Ingham C, Brainis I, et al. (2010) Genome sequence of the pattern forming Paenibacillus vortex bacterium reveals potential for thriving in complex environments. BMC Genomics 11: 710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Li N, Xia T, Xu Y, Qiu R, Xiang H, et al. (2012) Genome sequence of Paenibacillus sp. strain Aloe-11, an endophytic bacterium with broad antimicrobial activity and intestinal colonization ability. J Bacteriol 194: 2117–2118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Shin SH, Kim S, Kim JY, Song HY, Cho SJ, et al. (2012) Genome sequence of Paenibacillus terrae HPL-003, a xylanase-producing bacterium isolated from soil found in forest residue. J Bacteriol 194: 1266–1266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Merritt PM, Danhorn T, Fuqua C (2007) Motility and chemotaxis in Agrobacterium tumefaciens surface attachment and biofilm formation. J Bacteriol 189: 8005–8014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Oh CJ, Kim HB, Kim J, Kim WJ, Lee H, et al. (2012) Organization of nif gene cluster in Frankia sp. EuIK1 strain, a symbiont of Elaeagnus umbellata . Arch Microbiol 194: 29–34. [DOI] [PubMed] [Google Scholar]
- 49. Welsh EA, Liberton M, Stöckel J, Loh T, Elvitigala T, et al. (2008) The genome of Cyanothece 51142, a unicellular diazotrophic cyanobacterium important in the marine nitrogen cycle. Proc Natl Acad Sci U S A 105: 15094–15099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Wang L, Zhang L, Liu Z, Zhao D, Liu X, et al. (2013) A minimal nitrogen fixation gene cluster from Paenibacillus sp. WLY78 enables expression of active nitrogenase in Escherichia coli . PLoS Genet 9: e1003865. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Leigh J (2005) Genomics of diazotrophic archaea. Genomes and genomics of nitrogen-fixing organisms: Springer. pp. 7–12. [Google Scholar]
- 52. Richter M, Rosselló-Móra R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci U S A 106: 19126–19131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Xie JB, Bai LQ, Wang LY, Chen SF (2012) Phylogeny of 16S rRNA and nifH genes and regulation of nitrogenase activity by oxygen and ammonium in the genus Paenibacillus . Mikrobiologiia 81: 760–767. [PubMed] [Google Scholar]
- 54. Montoya JP, Voss M, Kahler P, Capone DG (1996) A simple, high-precision, high-sensitivity tracer assay for N (inf2) fixation. Appl Environ Microbiol 62: 986–993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Dixon R, Kahn D (2004) Genetic regulation of biological nitrogen fixation. Nat Rev Microbiol 2: 621–631. [DOI] [PubMed] [Google Scholar]
- 56. Kormelink TG, Koenders E, Hagemeijer Y, Overmars L, Siezen RJ, et al. (2012) Comparative genome analysis of central nitrogen metabolism and its control by GlnR in the class Bacilli . BMC Genomics 13: 191–206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Doroshchuk N, Gelfand M, Rodionov D (2006) Regulation of nitrogen metabolism in gram-positive bacteria. Mol Biol 40: 829–836. [PubMed] [Google Scholar]
- 58. Zhao D, Curatti L, Rubio LM (2007) Evidence for nifU and nifS participation in the biosynthesis of the iron-molybdenum cofactor of nitrogenase. J Biol Chem 282: 37016–37025. [DOI] [PubMed] [Google Scholar]
- 59. Johnson DC, Dean DR, Smith AD, Johnson MK (2005) Structure, function, and formation of biological iron-sulfur clusters. Annu Rev Biochem 74: 247–281. [DOI] [PubMed] [Google Scholar]
- 60. Hong Y, Ma Y, Wu L, Maki M, Qin W, et al. (2012) Characterization and analysis of nifH genes from Paenibacillus sabinae T27. Microbiol Res 167: 596–601. [DOI] [PubMed] [Google Scholar]
- 61. Hacker J, Carniel E (2001) Ecological fitness, genomic islands and bacterial pathogenicity. EMBO Rep 2: 376–381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62. Hacker J, Kaper JB (2000) Pathogenicity islands and the evolution of microbes. Annu Rev Microbiol 54: 641–679. [DOI] [PubMed] [Google Scholar]
- 63. Young JPW, Crossman LC, Johnston AW, Thomson NR, Ghazoui ZF, et al. (2006) The genome of Rhizobium leguminosarum has recognizable core and accessory components. Genome Biol 7: R34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Chen JS (2005) Genomic aspects of nitrogen fixation in the Clostridia. Genomes and genomics of nitrogen-fixing organisms: Springer. pp. 13–26. [Google Scholar]
- 65. Dodsworth JA, Leigh JA (2006) Regulation of nitrogenase by 2-oxoglutarate-reversible, direct binding of a PII-like nitrogen sensor protein to dinitrogenase. Proc Natl Acad Sci U S A 103: 9779–9784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66. Dobrindt U, Hochhut B, Hentschel U, Hacker J (2004) Genomic islands in pathogenic and environmental microorganisms. Nat Rev Microbiol 2: 414–424. [DOI] [PubMed] [Google Scholar]
- 67. Hirsch AM, McKhann HI, Reddy A, Liao J, Fang Y, et al. (1995) Assessing horizontal transfer of nifHDK genes in eubacteria: nucleotide sequence of nifK from Frankia strain HFPCcI3. Mol Biol Evol 12: 16–27. [DOI] [PubMed] [Google Scholar]
- 68. Nakamura Y, Itoh T, Matsuda H, Gojobori T (2004) Biased biological functions of horizontally transferred genes in prokaryotic genomes. Nat Genet 36: 760–766. [DOI] [PubMed] [Google Scholar]
- 69. Finan TM (2002) Evolving insights: symbiosis islands and horizontal gene transfer. J Bacteriol 184: 2855–2856. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70. Sullivan JT, Ronson CW (1998) Evolution of rhizobia by acquisition of a 500-kb symbiosis island that integrates into a phe-tRNA gene. Proc Natl Acad Sci U S A 95: 5145–5149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Young J (2005) The phylogeny and evolution of nitrogenases. Genomes and genomics of nitrogen-fixing organisms: Springer. pp. 221–241. [Google Scholar]
- 72. Galibert F, Finan TM, Long SR, Pühler A, Abola P, et al. (2001) The composite genome of the legume symbiont Sinorhizobium meliloti . Science 293: 668–672. [DOI] [PubMed] [Google Scholar]
- 73. Kaneko T, Nakamura Y, Sato S, Asamizu E, Kato T, et al. (2000) Complete genome structure of the nitrogen-fixing symbiotic bacterium Mesorhizobium loti . DNA research 7: 331–338. [DOI] [PubMed] [Google Scholar]
- 74. Kaneko T, Nakamura Y, Sato S, Minamisawa K, Uchiumi T, et al. (2002) Complete genomic sequence of nitrogen-fixing symbiotic bacterium Bradyrhizobium japonicum USDA110. DNA research 9: 189–197. [DOI] [PubMed] [Google Scholar]
- 75. Li R, Li Y, Kristiansen K, Wang J (2008) SOAP: short oligonucleotide alignment program. Bioinformatics 24: 713–714. [DOI] [PubMed] [Google Scholar]
- 76. Delcher AL, Bratke KA, Powers EC, Salzberg SL (2007) Identifying bacterial genes and endosymbiont DNA with Glimmer. Bioinformatics 23: 673–679. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77. Zhou Y, Liang Y, Lynch KH, Dennis JJ, Wishart DS (2011) PHAST: a fast phage search tool. Nucleic Acids Res 39: W347–W352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78. Zhao Y, Wu J, Yang J, Sun S, Xiao J, et al. (2012) PGAP: pan-genomes analysis pipeline. Bioinformatics 28: 416–418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79. Darling AC, Mau B, Blattner FR, Perna NT (2004) Mauve: multiple alignment of conserved genomic sequence with rearrangements. Genome Res 14: 1394–1403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80. Tamura K, Peterson D, Peterson N, Stecher G, Nei M, et al. (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28: 2731–2739. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81. Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, et al. (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61: 539–542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82. Thompson JD, Gibson T, Higgins DG (2002) Multiple sequence alignment using ClustalW and ClustalX. Curr Protoc Bioinformatics Chapter 2 Unit 2.3. doi: 10.1002/0471250953.bi0203s00 [DOI] [PubMed] [Google Scholar]
- 83. Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17: 540–552. [DOI] [PubMed] [Google Scholar]
- 84. Guindon S, Dufayard J, Lefort V, Anisimova M, Hordijk W, et al. (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59: 307–321. [DOI] [PubMed] [Google Scholar]
- 85. Shimodaira H, Hasegawa M (2001) CONSEL: for assessing the confidence of phylogenetic tree selection. Bioinformatics 17: 1246–1247. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Comparison of the nif gene cluster of Paenibacillus with those of the representative N2-fixing bacteria and archaea. (A) Paenibacillus polymyxa 1–43, (B) Azotobacter vinelandii, (C) Klebsiella oxytoca M5al, (D) Nostoc punctiforme PCC 73102, (E) Frankia sp. EAN1pec, (F) Clostridium acetobutylicum, (G) Methanococus maripaludis.
(TIF)
IS elements or prophages linked with the nif gene, nif cluster and nif-like genes.
(TIF)
Neighbor joining phylogenetic tree of the NifB sequences derived from Paenibacillus and other representative species. A total of 1,000 bootstrap replicates were made, and bootstrap values are indicated at each node.
(TIF)
Neighbor joining phylogenetic tree of the NifH, VnfH, AnfH and NifH-like protein sequences derived from Paenibacillus and other representative species. A total of 1,000 bootstrap replicates were made, and bootstrap values are indicated at each node.
(TIF)
Neighbor joining phylogenetic tree of the NifD, VnfD, AnfD and NifD-like protein sequences derived from Paenibacillus and other representative species. A total of 1,000 bootstrap replicates were made, and bootstrap values are indicated at each node.
(TIF)
Neighbor joining phylogenetic tree of the NifK, VnfK, AnfK and NifK-like protein sequences derived from Paenibacillus and other representative species. A total of 1,000 bootstrap replicates were made, and bootstrap values are indicated at each node.
(TIF)
Neighbor joining phylogenetic tree of the NifE, VnfE and NifE-like protein sequences derived from Paenibacillus and other representative species. A total of 1,000 bootstrap replicates were made, and bootstrap values are indicated at each node.
(TIF)
Neighbor joining phylogenetic tree of the NifN, VnfN and NifN-like protein sequences derived from Paenibacillus and other representative species. A total of 1,000 bootstrap replicates were made, and bootstrap values are indicated at each node.
(TIF)
Neighbor joining phylogenetic tree of the NifX protein sequences derived from Paenibacillus and other representative species. A total of 1,000 bootstrap replicates were made, and bootstrap values are indicated at each node.
(TIF)
Neighbor joining phylogenetic tree of the NifV protein sequences derived from Paenibacillus and other representative species. A total of 1,000 bootstrap replicates were made, and bootstrap values are indicated at each node.
(TIF)
Nitrogen fixation abilities of P. beijingensis 1–18 (WT) and recombinant E. coli 1–18 strain. (A) Nitrogenase activities determined by using acetylene reduction assay. (B) Nitrogen fixation ability determined by using for 15N2 incorporation. Error bars indicate the standard deviation observed from at least two independent experiments.
(TIF)
The σ70-depedent promoters of the nif clusters and the GlnR/TnrA-binding sites in the nif promoter regions in Paenibacillus strains.
(TIF)
Alignments of crucial residues surrounding the P-loop/MgATP binding motif, cysteine ligating 4Fe-4S and arginine ligating ADP-ribose in NifH and NifH-like protein sequences from Paenibacillus and other organisms.
(TIF)
Alignments of crucial residues ligating FeMo-co or P-cluster in NifD and NifD-like protein sequences from Paenibacillus and other organisms.
(TIF)
Alignments of crucial residues ligating FeMo-co or P-cluster in in NifK and NifK-like protein sequences from Paenibacillus and other organisms.
(TIF)
The genomes sequenced in this study.
(DOCX)
Comparison of COG assignments between non-N2-fixing and N2-fixing Paenibacillus strains.
(DOCX)
Transposons present in the genomes of 31 Paenibacillus strains. The following information is provided for each putative transposon in genomes: transposon family, transposases, and numbers of copies of intact or remnant transposons in each genome.
(DOCX)
Prophages present in the genomes of 31 Paenibacillus strains. The following information is provided for each prophage: insertion site, size, locus tags, and selected cargo genes.
(DOCX)
The nitrogen fixation genes and nitrogenase-like genes in the nitrogen-fixing Paenibacillus strains.
(DOCX)
Average Nucleotide Identity (%) based on whole genome alignments.
(XLSX)