Abstract
We propose a technique to simultaneously estimate the local fiber orientations and perform multi-fiber tractography. Existing techniques estimate the local fiber orientation at each voxel independently so there is no running knowledge of confidence in the measured signal or estimated fiber orientation. Further, to overcome noise, many algorithms use a filter as a post-processing step to obtain a smooth trajectory. We formulate fiber tracking as causal estimation: at each step of tracing the fiber, the current estimate of the signal is guided by the previous. To do this, we model the signal as a discrete mixture of Watson directional functions and perform tractography within a filtering framework. Starting from a seed point, each fiber is traced to its termination using an unscented Kalman filter to simultaneously fit the signal and propagate in the most consistent direction. Despite the presence of noise and uncertainty, this provides an accurate estimate of the local structure at each point along the fiber. We choose the Watson function since it provides a compact representation of the signal parameterized by the principal diffusion direction and a scaling parameter describing anisotropy, and also allows analytic reconstruction of the oriented diffusion function from those parameters. Using a mixture of two and three components (corresponding to two-fiber and three-fiber models) we demonstrate in synthetic experiments that this approach reduces signal reconstruction error and significantly improves the angular resolution at crossings and branchings. In vivo experiments examine the corpus callosum and internal capsule and confirm the ability to trace through regions known to contain such crossing and branching while providing inherent path regularization.
Keywords: Diffusion-weighted MRI, tractography, Kalman filtering, Watson directional function
1. Introduction
The advent of diffusion weighted magnetic resonance imaging has provided the opportunity for non-invasive investigation of neural architecture. Using this imaging technique, clinicians and neuroscientists want to ask how neurons originating from one region connect to other regions, or how well-defined those connections may be. For such studies, the quality of the results relies heavily on the chosen fiber representation and the method of reconstructing pathways.
To begin studying the microstructure of fibers, we need a model to interpret the diffusion weighted signal. Such models fall broadly into two categories: parametric and nonparametric. One of the simplest parametric models is the diffusion tensor which describes a Gaussian estimate of the diffusion orientation and strength at each voxel [6, 10]. While robust, this model can be inadequate in cases of mixed fiber presence or more complex orientations [3, 18]. To handle more complex diffusion patterns, various parametric models have been introduced: weighted mixtures [1, 51, 33, 19, 41], higher order tensors [25, 8], directional functions [37, 32, 45], and diffusion oriented transforms [39].
Nonparametric models often provide more information about the diffusion pattern. Instead of estimating a discrete number of fibers as in parametric models, nonparametric techniques estimate an oriented distribution function (ODF) describing an arbitrary configuration of fibers. For this estimation, Tuch [50] introduced Q-ball imaging to numerically compute the ODF using the Funk-Radon transform. The use of spherical harmonics simplified the computation with an analytic form [4, 24, 16] and spherical ridgelets further reduced the coefficients required [38]. Recently, Poupon et al. [42] demonstrated online direct estimation of single-tensor and harmonic coefficients using a linear Kalman filter. Another approach to producing an ODF is to assume a model for the signal response of a single-fiber and use spherical deconvolution [29, 28, 49, 32, 34]. A good review of both parametric and nonparametric models can be found in [2, 17].
Based on these models, several techniques attempt to reconstruct pathways. Deterministic tractography involves directly following the diffusion pathways. In the single tensor model, this means simply following the principal diffusion direction [7], while multi-fiber models often include techniques for determining the number of fibers present or when pathways branch [23, 33, 22, 43]. Kalman and particle filters have been used with single tensor streamline tractography [21, 12, 54, 27], but these have been used for path regularization and not to estimate the full underlying fiber model. Another approach to regularizing single tensor tractography uses a moving least squares estimate weighted with the previous tensor [55]. While this present study focuses on deterministic techniques, probabilistic methods have been developed to form connections by diffusing out a connectivity map according to the ODF [40, 14, 26, 9].
While parametric methods directly describe the principal diffusion directions, interpreting the ODFs from model independent representations typically involves determining the number and orientation of principal diffusion directions present [53, 47, 30]. For example, Bloy et al. [13] and Ghosh et al. [20] find them as maxima on the spherical surface; Descoteaux et al. [17] deconvolve with a sharpening kernel before extracting maxima; and Schultz and Seidel [46] decompose a high-order tensor into a mixture of rank-1 tensors. Ramirez-Manzanares et al. [44] provide a quantitative comparison of several such techniques.
Finally, [42] proposed using a linear Kalman filter for online, direct estimation of either single-tensor or harmonic coefficients while successive diffusion image slices are acquired, while [15] revisited the technique to account for proper regularization and proposed a method to quickly determine optimal gradient set orderings.
1.1. Our contributions
Of the approaches listed above, almost all fit the model at each voxel independent of other voxels. In this paper, we describe a method to estimate the model parameters and perform tractography simultaneously within a causal filter. In this way, the estimation at each position builds upon the previous estimates along the fiber.
To begin estimating within a finite dimensional filter, we model the diffusion signal using a mixture of either two or three Watson directional functions [45]. This enables estimation directly from the raw signal data without separate preprocessing or regularization steps. Because the signal reconstruction is nonlinear, we use the unscented Kalman filter to perform model estimation and then propagate in the most consistent direction. Using causal estimation in this way yields inherent path regularization, low signal reconstruction error, and accurate fiber resolution at crossing angles not found with independent optimization. We further note that the approach presented here generalizes to arbitrary fiber model with finite dimensional parameter space, and since the estimation is inherently smooth, it does not require arbitrary termination criteria such as curvature.
2. Approach
The main idea of our approach is to trace the local fiber orientations using the estimation at previous positions to guide estimation at the current position. In a loop, the Kalman filter estimates the model at the current position, moves a step in that direction, and then begins estimation again. Iterative estimation in this manner greatly improves the accuracy of resolving individual orientations and yields inherently smooth tracts despite the presence of noise and uncertainty. Further, since each iteration begins with a near-optimal solution provided by the previous estimation, the convergence of model fitting is improved and many local minima are naturally avoided.
Section 2.1 provides the necessary background on modeling the measurement signal using Watson functions and defines the specific fiber model employed in this study. Then, Section 2.2 describes how this model may be estimated using an unscented Kalman filter, and finally Section 2.3 summarizes the entire algorithm.
2.1. Modeling local fiber orientations
In diffusion weighted imaging, image contrast is related to the strength of water diffusion, and our goal is to accurately relate these signals to an underlying model of fiber orientation. At each image voxel, diffusion is measured along a set of distinct gradients, u1, …, un ∈𝕊3 (on the unit sphere), producing the corresponding signal, s = [s1, …, sn ]T ∈ ℝn. For voxels containing a mixed diffusion pattern, a general weighted formulation may be written as,
| (1) |
where s0 is a baseline signal intensity, b is an acquisition-specific constant, wj are convex weights, and Dj is a tensor describing a diffusion pattern [51, 33].
Considering a single tensor, we now follow the formulation of Rathi et al. [45] to define the Watson directional function which approximates the apparent diffusion pattern. We begin by noting that any diffusion tensor D can be decomposed as D = UΛ UT , where U is a rotation matrix and Λ is a diagonal matrix with eigenvalues {λ1, λ2, λ3}. These eigenvalues determine the shape of the tensor: ellipsoidal, planar, and spherical. For example, if λ1 > λ2 > λ3, then the shape is ellipsoidal with the major axis of the ellipsoid pointing to the eigenvector corresponding to λ1. Intuitively, it represents strong diffusion along that particular direction. When λ1 = λ2 > λ3, the shape is planar indicating diffusion along orthogonal directions, and finally, when λ1 = λ2 = λ3, the diffusion is spherical (isotropic).
The most common of these configurations is ellipsoidal with principal diffusion direction m and eigenvalue λ1, and hence the first step to introducing directional functions is to approximate the tensor by its first eigenvector expansion: D ≈ λ1mmT . Using this, each exponent in Eq. 1 may then be rewritten,
| (2) |
| (3) |
| (4) |
where the scalar k concentration parameter determines the degree of anisotropy. Finally, the general model may be restated:
| (5) |
where A is a normalization constant such that ‖s‖ = 1. For purposes of comparison, this normalization will also be done to signals obtained from scanner. Note that, while the diffusion tensor requires six parameters, these Watson functions require four parameters: three for the orientation vector m and one concentration parameter k. Employing spherical coordinates further reduces the unit vector m to two parameters. Fig. 1a demonstrates how adjusting the k-value produces different diffusion patterns, and Fig. 1b illustrates two multi-fiber configurations.
Figure 1.
Watson directional functions are capable of representing various diffusion patterns and fiber orientations.
From this general mixture model, we choose to start with a restricted form involving two equally-weighted Watson functions. This choice is guided by several previous studies. Behrens et al. [9] showed that at a b-value of 1000 the maximum number of detectable fibers is two. Several other studies have also found two-fiber models to be sufficient [51, 33, 53, 41]. Using this as a practical guideline, we started with a mixture of two Watson functions as our local fiber model. Further, following the study of [53], we assume an equal combination (50%-50%) of the two Watson functions. While the effect of this second choice appears to have little to no effect on experiments, we have yet to quantify any potential loss in accuracy. These assumptions leave us with the following two-fiber model used in this study:
| (6) |
where k1 and m1 parameterize the first Watson function, and k2 and m2 parameterize the second, and A is again a normalization constant such that ‖s‖ = 1. Thus the equally-weighted two-fiber model is fully described by the following parameters: k1, m1, k2, m2. Extending off the two-Watson model, we can directly formulate the equally-weighted three-Watson model:
| (7) |
with the additional parameters k3 and m3.
Finally, from such parameters, Rathi et al. [45] describe how one may compute the ODF analytically by applying the Funk-Radon transform directly to Eq. 5. The ODF can be reconstructed directly from the same parameters describing the signal without a separate estimation process. For the two-Watson model (Eq. 6) the ODF is approximated by,
| (8) |
and for the three-Watson model (Eq. 7) this becomes,
| (9) |
where B is a normalization factor such that Σi fi = 1.
2.2. Estimating the fiber model
Given the measured signal at a particular voxel, we want to estimate the underlying model parameters that explain this signal. As in streamline tractography, we treat the fiber as the trajectory of a particle which we trace out. At each step, we examine the measured signal at that position, estimate the underlying model parameters, and propagate forward in the most consistent direction. Fig. 2 illustrates this filtering process.
Figure 2.
System overview illustrating relation between the neural fibers, the scanner signals, and the unscented Kalman filter as it is used to estimate the local model. At each step, the filter uses its current model state (xt) to predict the observed scanner signal (ȳt+1|t) and then compares that against the actual measured signal (yt) from the scanner in order to update its internal model state (xt+1).
To use a state-space filter for estimating the model parameters, we need the application-specific definition of four filter components:
The system state x: the model parameters
The state transition f[·]: how the model changes as we trace the fiber
The observation h[·]: how the signal appears given a particular state
The measurement y: the actual signal obtained from the scanner
For our state, we directly use the model parameters, thus the two-fiber model in Eq. 6 has the following state vector:
| (10) |
Similarly, the three-Watson model adds additional state variables for the third component:
| (11) |
While each m could be represented in a reduced spherical form, the antipodes of the spherical parameterization would then introduce nonlinearities which complicate estimation. For the state transition we assume identity dynamics; the local fiber configuration does not undergo drastic change from one position to the next. Our observation is the signal reconstruction, y = s = [s1, …, sn]T using si from Eq. 6, and our measurement is the actual signal interpolated at the current position. Theoretically, the measurement noise model is Rician and not additive Gaussian as the Kalman filter assumes. A specific formulation of the Kalman filter for Rician noise might provide better performance under severe noise, but such a formulation is beyond the focus of this paper.
Since the signal reconstruction is a nonlinear processes, we employ an unscented Kalman filter to perform nonlinear estimation. Similar to classical linear Kalman filtering, the unscented version seeks to reconcile the predicted state of the system with the measured state and addresses the fact that those two processes (prediction and measurement) may be nonlinear or unknown. It does this in two phases: first it uses the system transition model to predict the next state and observation, and then it uses the new measurement to correct that state estimate. In what follows, we present the algorithmic application of the filter. For more thorough treatments, see [31, 52].
It is important to note two alternative techniques for nonlinear estimation. First, particle filters are commonly used to provide a multi-modal estimate of unknown systems. With respect to an n-dimensional state space, particle filters require the number of particles to be exponential to properly explore the state space. In contrast, the unscented filter requires only 2n + 1 particles (sigma points) for a Gaussian estimate that space. Further, for many slowly varying systems, the multimodal estimate is unnecessary: from one voxel to the next, fibers tend not to change direction drastically. Second, the extended Kalman filter may also be used to provide a Gaussian estimate after linearizing the system; however, the unscented Kalman filter provides a more accurate estimate with equivalent computational cost and altogether avoids the attempt at linearization [31, 52, 35].
The unscented Kalman filter considers a system of interest at time t given a Gaussian estimate of its current state with mean (xt ∈ ℝn) and covariance (Pt ∈ ℝn × n). Prediction begins with the formation of a set Xt = {χi} ⊂ ℝn of (2n + 1) sigma point states with associated convex weights, wi ∈ ℝ, each a perturbed version of the current state. We use the covariance, Pt, to distribute this set:
| (12) |
where [A]i denotes the ith column of matrix A and κ is an adjustable scaling parameter (κ = 0.01 in all experiments). The sigma points are spread in this manner so as to retain the mean and covariance, yet alternative set arrangements are sometimes used to control additional characteristics of the distribution [31].
This set is then propagated through the state transition function, χ̂ = f[χ] ∈ ℝn, to obtain a new predicted sigma point set: Xt+1|t = {f[χi]} = {χ̂i}. Since in this study we assume the fiber configuration does not change drastically as we follow it from one voxel to the next, we may write this identity transition as, xt+1|t = f[xt] = xt. These are then used to calculate the predicted system mean state and covariance,
| (13) |
where Q is the injected process noise bias. This procedure comprises the unscented transform used to estimate the behavior of a nonlinear function: spread sigma points based on your current uncertainty, propagate those using your transform function, and measure their spread.
To obtain the predicted observation, we again apply the unscented transform this time using the predicted states, Xt+1|t, to estimate what we expect observe from the measurement of each state: γ = h[χ̂] ∈ ℝm. Keep in mind that, for this study, our observation is the signal reconstruction from Eq. 6, and the measurement itself is the diffusion-weighted signal, s, interpolated at the current position. From these, we obtain the predicted set of observations, Yt+1|t = {h[χ̂i]} = {γi}, and may calculate its mean and covariance,
| (14) |
where R is the injected measurement noise bias. The cross correlation between the estimated state and measurement may also be calculated:
| (15) |
As is done in the classic linear Kalman filter, the final step is to use the Kalman gain, , to correct our prediction and provide us with the final estimated system mean and covariance,
| (16) |
| (17) |
where yt ∈ ℝm is the actual signal measurement taken at this time.
2.3. The algorithm
To summarize the proposed technique, we are using the unscented Kalman filter to estimate the local model parameters as we trace out each fiber. For each fiber, we maintain the position at which we are currently tracing it and the current estimate of its model parameters (mean and covariance).
At each step of tracing a fiber, we predict its new orientation, which in this case is simply identity: xt+1|t = xt. Our actual measurement yt in Eq. 16 is the diffusion-weighted signal s recorded by the scanner at this position. At subvoxel positions we use trilinear interpolation directly on the diffusion-weighted images. Further, we normalize this measurement to place it on the same scale as Eq. 6 and Eq. 7. With these, we step through the equations above to find the new estimated model parameters, xt+1.
Last, this filtering procedure is used to drive second-order Runge-Kutta path integration through the volume: at each step we use the filter to estimate and then report the most consistent of the principal diffusion directions for integration. Algorithm 1 outlines these steps.
3. Experiments
We first use experiments with synthetic data to validate our technique against ground truth. We confirm that our approach accurately estimates the true underlying signal and reliably recognizes crossing fibers over a broad range of angles. Comparing against two alternative multi-fiber optimization techniques, we find the filtered approach gives consistently superior results in both respects (Section 3.1). We follow this with the estimation of three-fiber crossings (Section 3.2). Next, we perform tractography through crossing fiber fields and examine the underlying orientations and branchings (Section 3.3). Lastly, we examine a real dataset to demonstrate how causal estimation is able to pick up fibers and branchings known to exist in vivo yet absent using other techniques (Section 3.4).
Following the experimental method of generating synthetic data found in [49, 17, 46], we pull from our real data set the 300 voxels with highest fractional anisotropy (FA) and compute the average eigenvalues among these voxels: {1200, 100, 100} µm2/msec (FA=0.91). We generated synthetic MR signals according to Eq. 1 using these eigenvalues to form an anisotropic tensor at both b = 1000 and b = 3000, using 81 gradient directions uniformly spread on the hemisphere, and assume s0 = 1. We generate two separate data sets, each with a different level of Rician noise: low-noise (σ = 0.1) and high-noise (σ = 0.2). To get an idea of this level of noise, Fig. 3 visualizes a sample voxel with two fibers at a 60° angle.
Figure 3.
Synthetic two-fiber voxel signals at a 60° angle (black wires indicate principal diffusion directions). Each column shows the same surface from two view-points. (a) shows the ground truth signal and corresponding true ODF (left to right). (b) shows the corrupted versions of the ground truth signal (left to right).
Throughout the experiments, we draw comparison to three alternative techniques. First, we use the same two-Watson model from Section 2.1 with a variant of matching pursuit for brute force, dictionary-based optimization [36]. In our implementation, we construct a finite dictionary of two-6 Watson signals at a range of various k-values, essentially discretizing the search space across orientations and k-values. Given a new measured signal, the signal from the dictionary with highest inner product provides an estimate of orientation and concentration. While our signal is produced at 81 gradient directions, we use 341 directions to construct the dictionary, thus any error is due to the method’s sensitivity to noise and discretization. Note that by using 341 orientation directions there is roughly an 8° angular difference between offset orientations, hence we see that the angular error is often at most 8°. This approach highlights the effect of using the same model but changing the optimization technique to one that treats each voxel independently. Second, we use spherical harmonics for modeling [49] and fiber-ODF sharpening for peak detection as described in [17] (order l = 8, regularization L = 0.006). This provides a comparison with an independently estimated, model-free representation. Note that this technique is very similar to spherical deconvolution. Last, when performing tractography on real data, we use single-tensor streamline tractography as a baseline2.
Algorithm 1.
Main loop repeated for each fiber
| repeat |
| Form the new weighted sigma points around the current mean xt and covariance Pt |
| Predict the observed signals from these sigma points (Eq. 6) |
| Update estimate using Kalman gain K and scanner measurement yt |
| Proceed to new position along most consistent mj |
| until estimated model appears isotropic |
The unscented Kalman filter conveniently requires few parameters. Specifically, of importance are the matrices for injecting model noise Q and injecting measurement noise R (see Eq. 13 and Eq. 14). Fortunately, the relative magnitude of each can be determined off-line from the data itself. We found that values on the order of qm = 0.001 (roughly 2°), qλ = 10, and rs = 0.02 were quite robust for the appropriate diagonal entries of Q and R. Off-diagonal entries were left at zero.
3.1. Signal reconstruction and angular resolution
While the independent optimization techniques can be run on individually generated voxels, care must be taken in constructing reasonable scenarios to test the causal filter. For this purpose, we constructed an actual 2D field through which to navigate (see Fig. 7 and Fig. 8a). In the middle is one long fiber pathway where the filter begins estimating a single component but then runs into a field of voxels with two crossed fibers at a fixed angle. In this crossing region we calculated error statistics. Similarly, we computed the angular error over this region using both sharpened spherical harmonics and matching pursuit. We generated several similar fields, each at a different fixed angle. By varying the size of the crossing region or the number of fibers run, we ensured that each technique performed estimation on at least 500 voxels.
Figure 7.
Fiber passing through 40°, 50°, and 60° synthetic crossings (b=3000, noisy). Blue dashes represent the orientation of the second fiber when detected.
Figure 8.
One fiber passing through an example synthetic field (b = 1000, noisy) and the estimated ODFs using spherical harmonics and the filter as it passes through the crossing region (blue box). The filter provides consistent angular resolution along the fiber while independent spherical harmonic modeling at those same locations misses the second fiber in two voxels.
In the first experiment, we look at signal reconstruction error. We calculate the mean squared error of the reconstructed signal, s, against the ground truth signal, ŝ (pure, no noise): ‖s − ŝ‖2/‖ŝ‖2. In essence, this is exactly what the filter is trying to minimize: the error between the reconstructed signal and the measured signal. Fig. 4 shows the results of using the proposed filter, matching pursuit, and spherical harmonics. Over each technique’s series of estimations, the trend-lines indicate the mean error while the bars indicate one standard deviation. Spherical harmonics (red) appear to produce a smooth fit to the given noisy data, while matching pursuit (blue) shows the effect of discretization and sensitivity to noise. The two raised areas are a result of the dictionary being constructed with an 8° minimum separation between any pair of orientations. This experiment demonstrates that the proposed filter (black) accurately and reliably estimates the true underlying signal.
Figure 4.
Mean squared error (MSE) between reconstructed signal and ground truth signal at various crossing angles (low-noise on left, high-noise on right). Notice how the increased noise has little effect on the filter (black) compared to using matching pursuit (blue) or sharpened spherical harmonics (red).
In the second experiment, we looked at the error in angular resolution by comparing the filtered approach to matching pursuit and sharpened spherical harmonics. Fig. 5a and Fig. 5b show the sensitivity of matching pursuit. Consistent with the results reported in [17, 16], spherical harmonics are generally unable to detect and resolve angles below 50° for b = 1000 or below 40° for b = 3000. Fig. 5c and Fig. 5d confirm this, respectively. This experiment demonstrates that for b=1000, the filtered approach consistently resolves angles down to 20–30° with 5° error compared to independent optimization which fails to reliably resolve below 60° with as much as 15° error. For b=3000, the filtered approach consistently resolves down to 20–30° with 2–3° error compared to independent optimization which cannot resolve below 50° with 5° error.
Figure 5.
Average angle error at various crossing angles comparing all three techniques: matching pursuit (blue), sharpened spherical harmonics (red), and the proposed filter (black). The filter provides stable and consistent estimation compared to either alternative technique. Each subfigure shows both the low-noise and high-noise experiments (left, right).
3.2. Three-fiber crossings
Resolving three-fiber crossings has proven difficult for many techniques, especially at the lower b-values typically used in in vivo studies. For ex-ample, Tuch et al. [51] found the general multi-tensor model to be unstable for three or more components using data at b=1077 with 126 gradients. Bergmann et al. [11] only reported results for up to two-tensors (b = 700, 30 gradients). Behrens et al. [9] found that b-values at upwards of 3000–4000 were required for detecting more than two fibers in simulations and none were found in vivo (b = 1000, 60 gradients). Further, many studies specifically use at most two orientations [1, 33, 41]. However, detection of three-fiber crossings has been reported using spherical harmonics. Tournier et al. [49] reported such crossings using spherical deconvolution (b=2971, 60 gradients) and Descoteaux et al. [17] also found three-fiber voxels using spherical harmonics with fiber sharpening (b = 1000, 60 gradients). Most recently, Schultz and Seidel [46] demonstrated tensor decomposition as a promising technique for resolving such configurations.
Following the experimental setup of Schultz and Seidel [46], we constructed an additional set of synthetic fields this time with three equally-weighted Gaussian components. As in the synthetic fields shown in Fig. 7 and Fig. 8a, one fiber is angled up and is the intended orientation to track through the region. The other two orientations were set so that the endpoints of the three principal axes formed an equilateral triangle with any two separated by the specified angle. With this setup, Fig. 6 shows that the filtered approach provides an accurate estimate that reaches roughly 60° compared to 70° for spherical harmonics [17]. A significant bias is apparent at more acute angles using either technique.
Figure 6.
The filtered approach (black) is able to resolve three-fiber crossings with improved accuracy and at sharper angles compared to using sharpened spherical harmonics (red). Both the low-noise and high-noise experiments are shown at b=3000 (left, right).
3.3. Synthetic tractography
Having verified the technique’s accuracy, we now turn to the resulting tractography. Fig. 7 provides examples of synthetic crossing fiber fields each at different fixed angles: 40°, 50°, 60° (b = 3000, noisy). In our experiments, we start fibers from the bottom and propagate upward where they encounter the crossing region. Here we show one such fiber and use blue glyphs to indicate the second component detected as the it passes through the crossing region. In general, we found that in regions with only one true fiber present (those outside the crossing), the second component either aligned with the first or adjusted its concentration parameter to fill out the isotropic component of the signal. Further, we found the filtering strategy to be robust with respect to initial configuration and choice of injected noise matrices Q and R (Eq. 13 and Eq. 14).
In Fig. 8a we show another 60° field (b = 1000, noisy) but take a closer look at several points along a single fiber as it passes through the crossing region. We also examine the corresponding ODFs reconstructed using sharpened spherical harmonics and the proposed filter. As expected, the sharpened spherical harmonics often do not detect the crossing but result in a single angle as seen in the middle two samples in Fig. 8b. A close examination of the reported axes shows the bias toward a single averaged axis as reported in [53, 48, 46]. In contrast, the filtered results are consistent and accurate.
3.4. In vivo tractography
We tested our approach on a real human brain scan acquired on a 3-Tesla GE system using an echo planar imaging (EPI) diffusion weighted image sequence. A double echo option was used to reduce eddy-current related distortions. To reduce impact of EPI spatial distortion, an eight channel coil was used to perform parallel imaging using Array Spatial Sensitivity Encoding Techniques (GE) with a SENSE-factor (speed-up) of 2. Acquisitions have 51 gradient directions with b=900 and eight baseline scans with b=0. The original GE sequence was modified to increase spatial resolution, and to further minimize image artifacts. The following scan parameters were used: TR 17000 ms, TE 78 ms, FOV 24 cm, 144×144 encoding steps, 1.7 mm slice thickness. All scans had 85 axial slices parallel to the AC-PC line covering the whole brain. In addition, b=0 field inhomogeneity maps were collected and calculated.
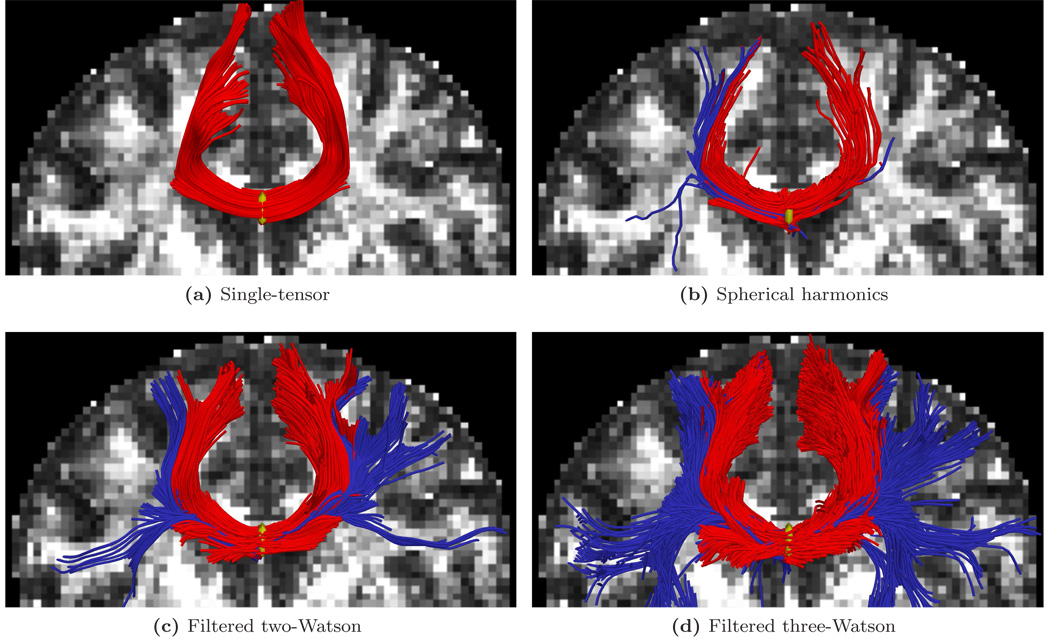
We first focused on fibers originating in the corpus callosum. Specifically, we sought to trace out the lateral transcallosal fibers that run through the corpus callosum out to the lateral gyri. It is known that single-tensor streamline tractography only traces out the dominant pathways forming the U-shaped callosal radiation (Fig. 9a and Fig. 15a). Several studies document this phenomena, among them the works of Descoteaux et al. [17] and Schultz and Seidel [46] have side-by-side comparisons. These fibers have been reported in using diffusion spectrum imaging [23], probabilistic tractography [32, 5, 17], and more recently with tensor decomposition [46].
Figure 9.
Filtered tractography picks up many fiber paths consistent with the underlying structures. Both single-tensor streamline and sharpened spherical harmonics are unable to find the majority of these pathways. Fibers existing ±22 mm around the mid-sagittal plane are indicated in blue. Seed region indicated in yellow.
Figure 15.
Tracing fibers originating from the center of the entire corpus callosum with views from above (top rows) and front-to-back (bottom rows). The proposed filtered tractography is able to find many of the lateral projections (blue) while single-tensor is unable to find any and few are found with sharpened spherical harmonics. Seed region indicated in yellow.
We start with two basic experiments: first examining the tracts surrounding a single coronal slice and second looking at all tracts passing through the corpus callosum. We seed each algorithm multiple times in voxels at the intersection of the mid-sagital plane and the corpus callosum. To explore branchings found using the proposed technique, we considered a component to be branching if it was separated from the primary component by less than 40° with k ≥ 0.6. Similarly, with sharpened spherical harmonics, we considered it a branch if we found additional maxima over the same range. We terminated fibers when the general fractional anisotropy of the estimated signal (std/rms) fell below 0.1. While such heuristics are somewhat arbitrary, we found little qualitative difference in adjusting these values.
To demonstrate the flexibility of the proposed filtering strategy with respect to model choice, we use both the two-Watson fiber model (Eq. 6) and the three-Watson fiber model (Eq. 7). While this introduced differences in the quantity of branchings detected, we found that using either model resulted in generally finding the same pathways.
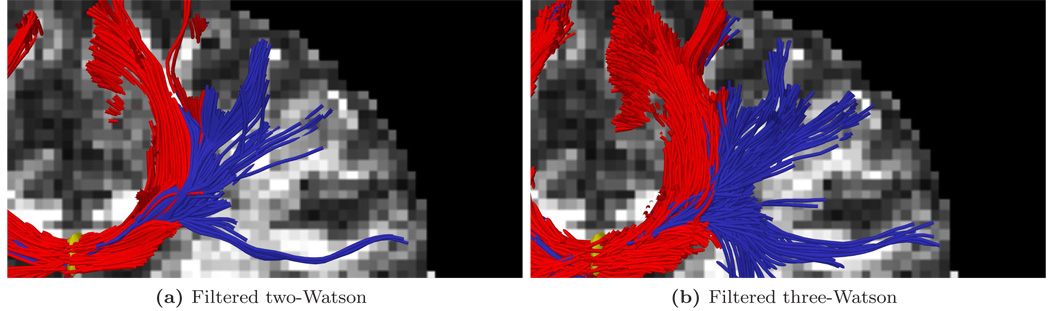
For the first experiment, Fig. 9 shows tracts originating from within a few voxels intersecting a particular coronal slice. For a reference backdrop, we use a coronal slice showing the intensity of fractional anisotropy (FA) placed a few voxels behind the seeded coronal position. Keeping in mind that these fibers are intersecting or are in front of the image plane, this roughly shows how the fibers navigate the areas of high anisotropy (bright regions). Similar to the results in [17, 46], Fig. 9b shows that sharpened spherical harmonics only pick up a few fibers intersecting the U-shaped callosal radiata. In contrast, our proposed method traces out many pathways consistent with the apparent anatomy using either the two-fiber or three-fiber model. To emphasize transcallosal tracts, we color as blue those fibers exiting a corridor of ±22 mm around the mid-sagittal plane. Fig. 10 provides a closer inspection of Fig. 9c and Fig. 9d where, to emphasize the underlying anatomy influencing the fibers, we use as a backdrop the actual coronal slice passing through the voxels used to seed this run. Such results are obtained in minutes in our current Matlab implementation. At each step, the cost of reconstructing the signal for few sigma points approaches the cost of a few iterations of weighted least-squares estimation of a single tensor.
Figure 10.
Closeup of upper right in Fig. 9c and Fig. 9d.
For the second experiment, Fig. 15 shows a view of the whole brain to see the overall difference between the different methods. Here again we emphasize with blue the transcallosal fibers found using the proposed filter. Comparing Fig. 15c and Fig. 15d we see several regions that appear to have different fiber density using the two models. This suggests that incorporating model selection into filtered approaches may have a significant effect. To show the various pathways infiltrating the gyri, Fig. 11 provides a closeup of the frontal lobe from above (without blue emphasis).
Figure 11.
Closeup of frontal fibers in Fig. 15c and Fig. 15d viewed from above.
Next we examined fibers passing through the internal capsule to trace out the pathways reaching up into the primary motor cortex at the top of the brain as well as down into the hippocampal regions near the brain stem. Fig. 12 shows frontal views for each technique with seeding near the cerebral peduncles (blue). Fig. 14 shows this same result from a side view where we can see that the filtered approach picks up the corticospinal pathways. Notice that the two-Watson model picks up the temporo-pontine and parietopontine tracts and the three-Watson model further reveals the occipitopontine pathways, another indication that the chosen fiber model often affects the results. As reported elsewhere [9, 43], single-tensor tractography follows the dominant corticospinal tract to the primary motor cortex. The same pathways were also found with spherical harmonics. Fig. 13 shows a view from above where we use a transverse FA image slice near the top of the brain as a backdrop so we can focus on the fiber endpoints. From this we can see how each method infiltrates the sulci grooves, and specifically we see that the filtered method is able to infiltrate sulci more lateral compared to single-tensor tractography.
Figure 12.
Frontal view with seeding in the internal capsule (blue). While both single-tensor and spherical harmonics tend to follow the dominant corticospinal tract to the primary motor cortex, the filtered approach follows many more pathways. Seed region indicated in yellow.
Figure 14.
Side view with seeding in the internal capsule (yellow). Filtered tractography finds many insertions into cortical regions of the parietal and occipital lobes. Seed region indicated in yellow.
Figure 13.
View from above showing cortical insertion points for each method. FA backdrop is taken near the top of the brain. The filtered approach shows more lateral insertions compared to single-tensor and spherical harmonic tracts.
Note that in the region of intersection between the transcallosal fibers, the corticospinal, and the superior longitudinal fasciculus, the partial voluming of each of these pathways leads the filter to report several end-to-end connections that are not necessarily present, e.g. fibers originating in the left internal capsule do not pass through this region, through the corpus callosum and then insert into the right motor cortex. Many of the lateral extensions are callosal fibers that are picked up while passing through this juncture. It is our hope that such connections may be avoided with the introduction of weighted mixtures, alternative filter formulations, or different heuristic choices in the algorithm.
4. Conclusion
Studies involving deterministic tractography rely on the underlying model estimated at each voxel as well as the reconstructed pathways. In this work, we demonstrated that using a causal filter provides robust estimates of much higher accuracy than independent estimation techniques. While the model we employed has been introduced previously [45], we primarily focused on the optimization technique used to estimate that model. Framing that estimation within a causal filter allowed us to apply a standard technique for nonlinear estimation. The proposed approach gives significantly lower angular error (5–10°) in regions with fiber crossings compared to using sharpened spherical harmonics (15–20°), and it is able to reliably resolve crossings down to 20–30° compared to spherical harmonics which reaches only down to 50–60°.
We believe that exploring both alternative models and filtering techniques will provide more accurate and comprehensive information about neural pathways and ultimately enhance non-invasive diagnosis and treatment of brain disease.
Acknowledgements
This work was supported in part by a Department of Veteran Affairs Merit Award (Shenton, R McCarley), the VA Schizophrenia Center Grant (McCarley, Shenton) and NIH grants: P41 RR13218 (Shenton), K05 MH 070047 (Shenton), 1P50MH080272-01 (Shenton), R01 MH 50740 (Shenton), R01 MH 082918 (Bouix), P50 MH080272-01 (McCarley), R01 MH50740-15 (Shenton), R03 TW008134-01 (Shenton), and the Center for Integration of Medicine and Innovative Technology Soldier in Medicine Award (Bouix).
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
Using the freely available Slicer 2.7 (http://www.slicer.org).
References
- 1.Alexander A, Hasan K, Tsuruda J, Parker D. Analysis of partial volume effects in diffusion-tensor MRI. Magnetic Resonance in Medicine. 2001;45:770–780. doi: 10.1002/mrm.1105. [DOI] [PubMed] [Google Scholar]
- 2.Alexander DC. Multiple-fiber reconstruction algorithms for diffusion MRI. Annals of the New York Academy of Sciences. 2005;1046(1):113–133. doi: 10.1196/annals.1340.018. [DOI] [PubMed] [Google Scholar]
- 3.Alexander DC, Barker G, Arridge S. Detection and modeling of non-Gaussian apparent diffusion coefficient profiles in human brain data. Magnetic Resonance in Medicine. 2002;48:331–340. doi: 10.1002/mrm.10209. [DOI] [PubMed] [Google Scholar]
- 4.Anderson A. Measurement of fiber orientation distributions using high angular resolution diffusion imaging. Magnetic Resonance in Medicine. 2005;54(5):1194–1206. doi: 10.1002/mrm.20667. [DOI] [PubMed] [Google Scholar]
- 5.Anwander A, Descoteaux M, Deriche R. Probabilistic Q-Ball tractography solves crossings of the callosal fibers. Human Brain Mapping. 2007:342. [Google Scholar]
- 6.Basser P, Jones D. Diffusion-tensor MRI: theory, experimental design and data analysis - A technical review. NMR in Biomedicine. 2002;15:456–467. doi: 10.1002/nbm.783. [DOI] [PubMed] [Google Scholar]
- 7.Basser P, Pajevic S, Pierpaoli C, Duda J, Aldroubi A. In vivo fiber tractography using DT-MRI data. Magnetic Resonance in Medicine. 2000;44:625–632. doi: 10.1002/1522-2594(200010)44:4<625::aid-mrm17>3.0.co;2-o. [DOI] [PubMed] [Google Scholar]
- 8.Basser PJ, Pajevic S. Spectral decomposition of a 4th-order covariance tensor: Applications to diffusion tensor MRI. Signal Processing. 2007;87:220–236. [Google Scholar]
- 9.Behrens T, Johansen-Berg H, Jbabdi S, Rush-worth M, Woolrich M. Probabilistic diffusion tractography with multiple fibre orientations: What can we gain? NeuroImage. 2007;34:144–155. doi: 10.1016/j.neuroimage.2006.09.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Behrens T, Woolrich M, Jenkinson M, Johansen-Berg H, Nunes R, Clare S, Matthews P, Brady J, Smith S. Characterization and propagation of uncertainty in diffusion-weighted MR imaging. Magnetic Resonance in Medicine. 2003;50:1077–1088. doi: 10.1002/mrm.10609. [DOI] [PubMed] [Google Scholar]
- 11.Bergmann O, Kindlmann G, Lundervold A, Westin C-F. Diffusion k-tensor estimation from Q-Ball imaging using discretized principal axes. Medical Image Computing and Computer Assisted Intervention (MICCAI) 2006:268–275. doi: 10.1007/11866763_33. [DOI] [PubMed] [Google Scholar]
- 12.Björnemo M, Brun A, Kikinis R, Westin C-F. Regularized stochastic white matter tractography using diffusion tensor MRI. Medical Image Computing and Computer Assisted Intervention (MICCAI) 2002:435–442. [Google Scholar]
- 13.Bloy L, Verma R. On computing the underlying fiber directions from the diffusion orientation distribution function. Medical Image Computing and Computer Assisted Intervention (MICCAI) 2008:1–8. doi: 10.1007/978-3-540-85988-8_1. [DOI] [PubMed] [Google Scholar]
- 14.Campbell JSW, Siddiqi K, Rymar VV, Sadikot AF, Pike GB. Flow-based fiber tracking with diffusion tensor and Q-ball data: Validation and comparison to principal diffusion direction techniques. NeuroImage. 2005;27(4):725–736. doi: 10.1016/j.neuroimage.2005.05.014. [DOI] [PubMed] [Google Scholar]
- 15.Deriche R, Calder J, Descoteaux M. Optimal real-time Q-Ball imaging using regularized Kalman filtering with incremental orientation sets. Medical Image Analysis. 2009;13(4):564–579. doi: 10.1016/j.media.2009.05.008. [DOI] [PubMed] [Google Scholar]
- 16.Descoteaux M, Angelino E, Fitzgibbons S, Deriche R. Regularized, fast, and robust analytical Q-ball imaging. Magnetic Resonance in Medicine. 2007;58:497–510. doi: 10.1002/mrm.21277. [DOI] [PubMed] [Google Scholar]
- 17.Descoteaux M, Deriche R, Knoesche T, Anwander A. Deterministic and probabilistic tractography based on complex fiber orientation distributions. Trans. on Med. Imag. 2009;28(2):269–286. doi: 10.1109/TMI.2008.2004424. [DOI] [PubMed] [Google Scholar]
- 18.Frank L. Characterization of anisotropy in high angular resolution diffusion-weighted MRI. Magnetic Resonance in Medicine. 2002;47:1083–1099. doi: 10.1002/mrm.10156. [DOI] [PubMed] [Google Scholar]
- 19.Friman O, Farnebäck G, Westin C-F. A Bayesian approach for stochastic white matter tractography. Trans. on Med. Imag. 2006;25(8):965–978. doi: 10.1109/tmi.2006.877093. [DOI] [PubMed] [Google Scholar]
- 20.Ghosh A, Tsigaridas E, Descoteaux M, Comon P, Mourrain B, Deriche R. A polynomial based approach to extract the maxima of an antipodally symmetric spherical function and its application to extract fiber directions from the Orientation Distribution Function in diffusion MRI. Workshop on Computational Diffusion MRI (in MICCAI) 2008:237–248. [Google Scholar]
- 21.Gössl C, Fahrmeir L, P utz B, Auer L, Auer D. Fiber tracking from DTI using linear state space models: Detectability of the pyramidal tract. NeuroImage. 2002;16:378–388. doi: 10.1006/nimg.2002.1055. [DOI] [PubMed] [Google Scholar]
- 22.Guo W, Zeng Q, Chen Y, Liu Y. Using multiple tensor deflection to reconstruct white matter fiber traces with branching; Int. Symp. on Biomedical Imaging; 2006. pp. 69–72. [Google Scholar]
- 23.Hagmann P, Reese T, Tseng W-Y, Meuli R, Thiran J-P, Wedeen VJ. Diffusion spectrum imaging tractography in complex cerebral white matter: An investigation of the centrum semiovale; Int. Symp. on Magnetic Resonance in Medicine (ISMRM); 2004. p. 623. [Google Scholar]
- 24.Hess C, Mukherjee P, Han E, Xu D, Vigneron D. Q-ball reconstruction of multimodal fiber orientations using the spherical harmonic basis. Magnetic Resonance in Medicine. 2006;56:104–117. doi: 10.1002/mrm.20931. [DOI] [PubMed] [Google Scholar]
- 25.Hlawitschka M, Scheuermann G. HOT-lines: Tracking lines in higher order tensor fields. Visualization. 2005:27–34. [Google Scholar]
- 26.Hosey T, Williams G, Ansorge R. Inference of multiple fiber orientations in high angular resolution diffusion imaging. Magnetic Resonance in Medicine. 2005;54:1480–1489. doi: 10.1002/mrm.20723. [DOI] [PubMed] [Google Scholar]
- 27.Imperati D, Frosio I, Tittgemeyer M, Borghese NA. Prediction correction tractography through statistical tracking; IEEE Nuclear Science Symp. Conf. Record; 2008. pp. 4140–4146. [Google Scholar]
- 28.Jansons K, Alexander DC. Persistent angular structure: New insights from diffusion MRI data. Inverse Problems. 2003;19:1031–1046. [Google Scholar]
- 29.Jian B, Vemuri B. A unified computational framework for deconvolution to reconstruct multiple fibers from diffusion weighted MRI. Trans. on Med. Imag. 2007;26(11):1464–1471. doi: 10.1109/TMI.2007.907552. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Jian B, Vemuri B, Özarslan E, Carney PR, Mareci TH. A novel tensor distribution model for the diffusion-weighted MR signal. NeuroImage. 2007;37(1):164–176. doi: 10.1016/j.neuroimage.2007.03.074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Julier S, Uhlmann J. Unscented filtering and nonlinear estimation. IEEE. 2004;92(3):401–422. [Google Scholar]
- 32.Kaden E, Knøsche T, Anwander A. Parametric spherical deconvolution: Inferring anatomical connectivity using diffusion MR imaging. NeuroImage. 2007;37:474–488. doi: 10.1016/j.neuroimage.2007.05.012. [DOI] [PubMed] [Google Scholar]
- 33.Kreher B, Schneider J, Mader I, Martin E, Hennig J, Il’yasov K. Multitensor approach for analysis and tracking of complex fiber configurations. Magnetic Resonance in Medicine. 2005;54:1216–1225. doi: 10.1002/mrm.20670. [DOI] [PubMed] [Google Scholar]
- 34.Kumar R, Barmpoutis A, Vemuri BC, Carney PR, Mareci TH. Multi-fiber reconstruction from DW-MRI using a continuous mixture of von Mises-Fisher distributions. Mathematical Methods in Biomedical Image Analysis (MMBIA) 2008:1–8. [Google Scholar]
- 35.Lefebvre T, Bruyninckx H, Schutter JD. Kalman filters for non-linear systems: a comparison of performance. International Journal of Control. 2004;77(7):639–653. [Google Scholar]
- 36.Mallat S, Zhang Z. Matching pursuits with time-frequency dictionaries. Trans. on Signal Processing. 1993;41:3397–2415. [Google Scholar]
- 37.McGraw T, Vemuri B, Yezierski B, Mareci T. Von Mises-Fisher mixture model of the diffusion ODF; Int. Symp. on Biomedical Imaging; 2006. pp. 65–68. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Michailovich O, Rathi Y. On approximation of orientation distributions by means of spherical ridgelets; Int. Symp. on Biomedical Imaging; 2008. pp. 939–942. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Özarslan E, Shepherd T, Vemuri B, Blackband S, Mareci T. Resolution of complex tissue microarchitecture using the diffusion orientation transform. NeuroImage. 2006;31(3):1086–1103. doi: 10.1016/j.neuroimage.2006.01.024. [DOI] [PubMed] [Google Scholar]
- 40.Parker G, Alexander DC. Probabilistic Monte Carlo based mapping of cerebral connections utilizing whole-brain crossing fiber information. Information Processing in Medical Imaging (IPMI) 2003:684–696. doi: 10.1007/978-3-540-45087-0_57. [DOI] [PubMed] [Google Scholar]
- 41.Peled S, Friman O, Jolesz F, Westin C-F. Geometrically constrained two-tensor model for crossing tracts in DWI. Magnetic Resonance in Medicine. 2006;24(9):1263–1270. doi: 10.1016/j.mri.2006.07.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Poupon C, Roche A, Dubois J, Mangin J-F, Poupon F. Real-time MR diffusion tensor and Q-ball imaging using Kalman filtering. Medical Image Analysis. 2008;12:5527–5534. doi: 10.1016/j.media.2008.06.004. [DOI] [PubMed] [Google Scholar]
- 43.Qazi A, Radmanesh A, O’Donnell L, Kindlmann G, Peled S, Whalen S, Westin C-F, Golby AJ. Resolving crossings in the corticospinal tract by two-tensor streamline tractography: Method and clinical assessment using fMRI. NeuroImage. doi: 10.1016/j.neuroimage.2008.06.034. In press. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Ramirez-Manzanares A, Cook P, Gee J. A comparison of methods for recovering intra-voxel white matter fiber architecture from clinical diffusion imaging scans. Medical Image Computing and Computer Assisted Intervention (MICCAI) 2008:305–312. doi: 10.1007/978-3-540-85988-8_37. [DOI] [PubMed] [Google Scholar]
- 45.Rathi Y, Michailovich O, Shenton ME, Bouix S. Directional functions for orientation distribution estimation. Medical Image Analysis. 2009;13:432–444. doi: 10.1016/j.media.2009.01.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Schultz T, Seidel H. Estimating crossing fibers: A tensor decomposition approach. Trans. on Visualization and Computer Graphics. 2008;14(6):1635–1642. doi: 10.1109/TVCG.2008.128. [DOI] [PubMed] [Google Scholar]
- 47.Seunarine K, Cook P, Hall M, Embleton K, Parker G, Alexander DC. Exploiting peak anisotropy for tracking through complex structures. Mathematical Methods in Biomedical Image Analysis (MMBIA) 2007:1–8. [Google Scholar]
- 48.Tournier J-D, Calamante F, Connelly A. Robust determination of the fibre orientation distribution in diffusion MRI: Non-negativity constrained super-resolved spherical deconvolution. NeuroImage. 2007;35:1459–1472. doi: 10.1016/j.neuroimage.2007.02.016. [DOI] [PubMed] [Google Scholar]
- 49.Tournier J-D, Calamante F, Gadian D, Connelly A. Direct estimation of the fiber orientation density function from diffusion-weighted MRI data using spherical deconvolution. NeuroImage. 2004;23:1176–1185. doi: 10.1016/j.neuroimage.2004.07.037. [DOI] [PubMed] [Google Scholar]
- 50.Tuch D. Q-ball imaging. Magnetic Resonance in Medicine. 2004;52:1358–1372. doi: 10.1002/mrm.20279. [DOI] [PubMed] [Google Scholar]
- 51.Tuch D, Reese T, Wiegell M, Makris N, Belliveau J, Wedeen V. High angular resolution diffusion imaging reveals intravoxel white matter fiber heterogeneity. Magnetic Resonance in Medicine. 2002;48:577–582. doi: 10.1002/mrm.10268. [DOI] [PubMed] [Google Scholar]
- 52.van der Merwe R, Wan E. Sigma-point Kalman filters for probabilistic inference in dynamic state-space models. Workshop on Advances in Machine Learning. 2003 [Google Scholar]
- 53.Zhan W, Yang Y. How accurately can the diffusion profiles indicate multiple fiber orientations? A study on general fiber crossings in diffusion MRI. J. of Magnetic Resonance. 2006;183:193–202. doi: 10.1016/j.jmr.2006.08.005. [DOI] [PubMed] [Google Scholar]
- 54.Zhang F, Hancock E, Goodlett C, Gerig G. Probabilistic white matter fiber tracking using particle filtering and von Mises-Fisher sampling. Medical Image Analysis. 2009;13:5–18. doi: 10.1016/j.media.2008.05.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Zhukov L, Barr A. Oriented tensor reconstruction: Tracing neural pathways from diffusion tensor MRI. Visualization. 2002:387–394. [Google Scholar]