Abstract
Nitrate is a primary nutrient for plant growth, but its levels in soil can fluctuate by several orders of magnitude. Previous studies have identified Arabidopsis NRT1.1 as a dual-affinity nitrate transporter, which can take up nitrate over a wide range of concentrations. The mode of action of NRT1.1 is controlled by phosphorylation of a key residue, Thr101. Yet how this posttranslational modification switches the transporter between two affinity states remains unclear. Here we report the crystal structure of unphosphorylated NRT1.1, which reveals an unexpected homodimer in the inward-facing conformation. In this low-affinity state, the Thr101 phosphorylation site is embedded in a pocket immediately adjacent to the dimer interface, linking the phosphorylation status of the transporter to its oligomeric state. Using a cell-based fluorescence resonance energy transfer assay, we show that functional NRT1.1 indeed dimerizes in the cell membrane and the phosphomimetic mutation of Thr101 converts the protein into a monophasic high affinity transporter by structurally decoupling the dimer. Together with analyses of the substrate transport tunnel, our results establish a phosphorylation-controlled dimerization switch that allows NRT1.1 to uptake nitrate with two distinct affinity modes.
Introduction
Active nitrate (NO3−) uptake by roots represents the critical first step of nitrogen acquisition in plants, which render the essential element to animals in organic forms. The abundance of nitrate in soil is affected by many environmental factors. As a result, the soil concentrations of nitrate can undergo rapid changes, varying from low μM to high mM. In adaptation to the fluctuating nitrate levels, plants have evolved two complementary nitrate transporter systems with distinct kinetic properties1–3. The high affinity transport system (HATS), which consists of members of the NRT2/NNP family, drives nitrate uptake with KM values in the μM range, while the low affinity transport system (LATS), constituted by the NRT1/PTR family, transports nitrate at mM concentrations. Facing changes in nitrogen availability and demands, the activity of select components of these two systems can be further fine-tuned by transcriptional regulation and post-translational modifications. Transporters of both families belong to the major facilitator superfamily (MFS) of secondary active transporters and are dependent on protons for nitrate transport4,5.
The Arabidopsis NRT1.1/CHL1/NPF6.3 protein is the first nitrate transporter identified in higher plants and belongs to the NRT1 family6,7. Distinct from most of the NRT1 and NRT2 family members, NRT1.1 functions as a dual-affinity transporter and contributes to both HATS and LATS8–10. In comparison to the wild type plant, nrt1.1 mutants showed marked nitrate uptake defects in both high and low affinity ranges. In the heterologous Xenopus oocyte expression system, the transporter activity of NRT1.1 exhibited a characteristic biphasic kinetics with two different KM values of ~50μM and ~4mM. Importantly, recent studies have shown that phosphorylation of a single residue, Thr101, is responsible for switching NRT1.1 from the low-affinity to the high-affinity mode11. Mutations of Thr101 preventing or mimicking phosphorylation can effectively convert the dual-affinity transporter into a monophasic low-affinity or high-affinity transporter, respectively. Nitrate is not only a nutrient but also a signaling molecule, which modulates many aspects of plant physiology and optimal nitrate acquisition12–15. Independent of its transporter function, NRT1.1 also acts as a nitrate sensor, regulating differential expression of primary nitrate response genes at different nitrate levels16–18. Remarkably, the sensor function of NRT1.1 also shows a biphasic pattern and is affected by the phosphorylation status of Thr101.
The MFS permeases transport a wide spectrum of substrates and represent one of the largest families of secondary carriers in all species of life. Despite the growing number of available crystal structures of MFS transporters19–29, little is known about the structural mechanisms regulating their activities. Here we report the crystal structure of the full-length Arabidopsis thaliana NRT1.1, which reveals an unexpected phosphorylation-controlled dimerization switch that enables the transporter to operate with a dual-affinity mode.
RESULTS
Overall structure of NRT1.1
The Arabidopsis thaliana NRT1.1 gene encodes a 590-amino acid protein, which is highly conserved among plant NRT1.1 orthologs, but not Arabidopsis NRT1 family members (Extended Data Fig. 1 & 2). The recombinant NRT1.1 protein was overexpressed, solubilized and purified from insect cells with dodecyl maltoside (DDM) and crystallized in the presence of 10 mM NaNO3. With combined phases from Rosetta-improved molecular replacement30 and single wavelength anomalous diffraction, we determined and refined the NRT1.1 structure at 3.25 Å resolution (Extended Data Fig. 3 & Table 1).
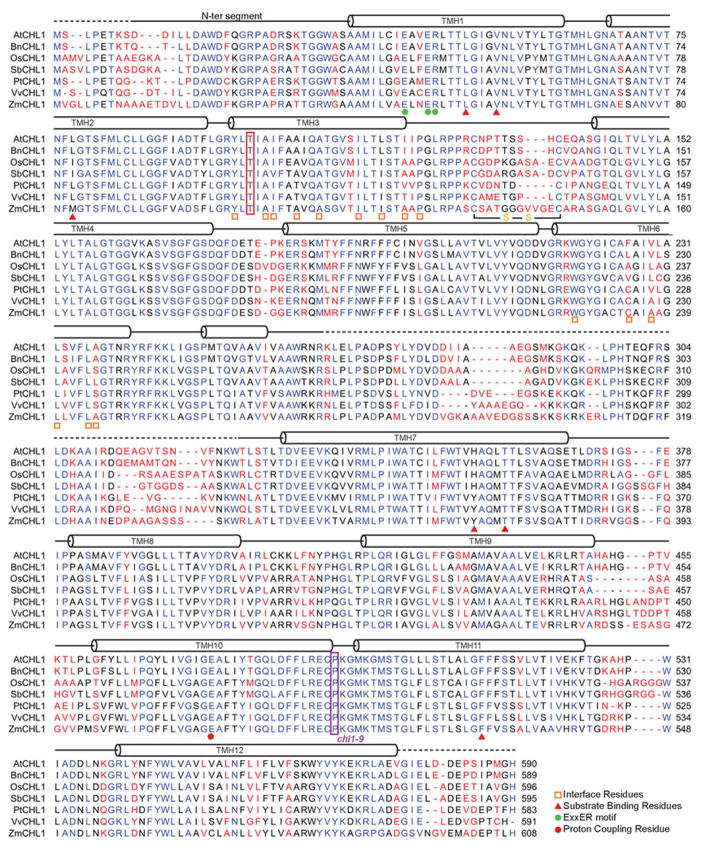
Extended Data Figure 1. Sequence alignment of plant NRT1.1 orthologs.
Alignment and secondary structure assignments of NRT1.1 orthologs from Arabidopsis thaliana (At), Brassica napus (Bn), Oryza sativa (Os), Sorghum bicolor (Sb), Populus trichocarpa (Pt), Vitis vinifera (Vv), and Zea mays (Zm). Strictly conserved residues are colored in blue. Green dots indicate the ExxER motif. Orange empty squares indicate dimer interface residues. Red triangles indicate residues in the substrate-binding pocket. Red dot indicates the energy-coupling residue. Dashed lines represent the disordered region in the crystal structure.
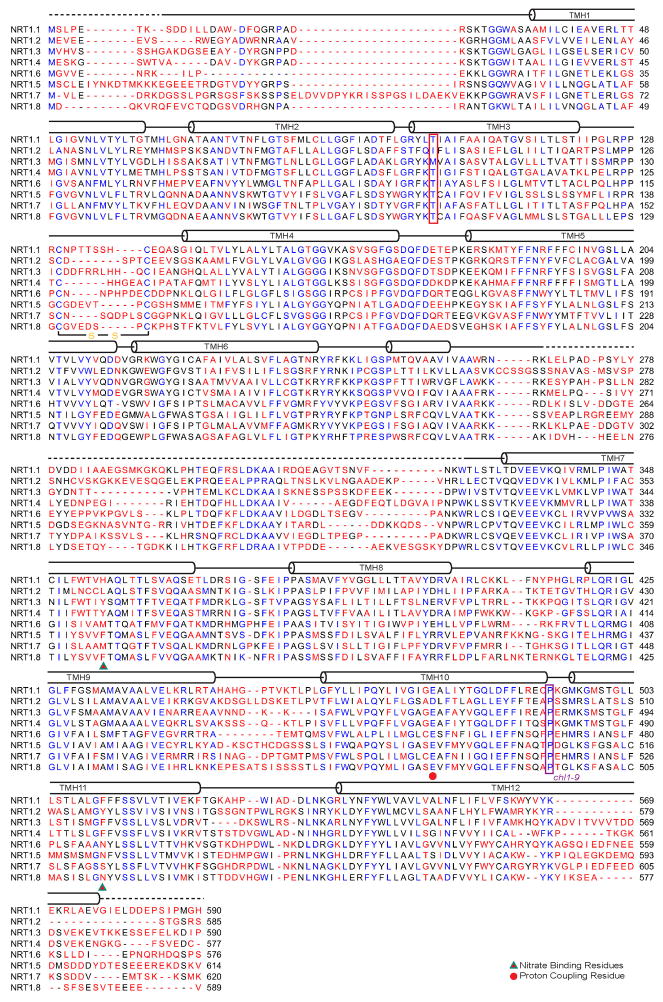
Extended Data Figure 2. Sequence alignment of Arabidopsis NRT1 family members.
Alignment and secondary structure assignments of the NRT1 family members from Arabidopsis thaliana. Strictly conserved residues are colored in blue. The two residues potentially important for substrate binding are indicated by red triangle with green stroke. Energy coupling residue is indicated by red dot. Dashed lines represent the disordered region in the crystal structure.
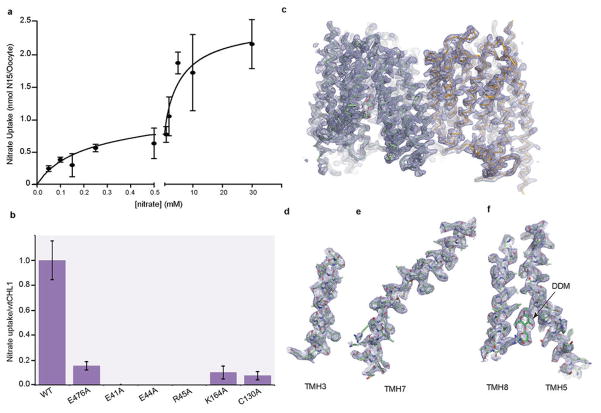
Extended Data Figure 3. Nitrate uptake and electron density map of NRT1.1.
a, Measurement of nitrate uptake by NRT1.1 was carried out in Xenopus oocytes in the presence of increasing concentrations of nitrate. Q-test was used to identify statistical outliers in data. All data points are mean ± s.d. of one experiment in quintuplicates or sextuplicates. The data were fit with a two site nonlinear binding curve using Prism. b, Relative nitrate uptake activities of NRT1.1 mutants to the wild type protein measured in the Xenopus oocyte-based assay in the presence of 10mM nitrate. All results are the mean ± s.d. of one experiment in quintuplicates or sextuplicates. c, The overall 2Fo-Fc map of the NRT1.1 dimer contoured at 1.5σ. d–e, Two representative helixes and their 2Fo-Fc maps contoured at 1.5σ. f, An island of density from the 2Fo-Fc map contoured at 1.5σ is assigned to the head group of DDM bound between TMH5 and TMH8..
Extended Data Table 1.
Data collection, phasing and refinement statistics (SAD).
| Native | Hg Derivative | |
|---|---|---|
| Data collection | ||
| Space group | C2221 | C2221 |
| Cell dimensions | ||
| a, b, c (Å) | 84.8, 188.5, 262.8 | 84.7, 189.9, 262.8 |
| α, β, γ (°) | 90, 90, 90 | 90, 90, 90 |
| Resolution (Å) | 50.0-3.2 (3.26-3.20)* | 50-3.50 (3.56-3.50)* |
| Rsym or Rmerge | 0.07 (0.91) | 0.08 (0.65) |
| I/σ/ | 17.92 (2.05) | 26.96 (1.84) |
| Completeness (%) | 99.1 (94.6) | 98.2 (96.5) |
| Redundancy | 5.9 (4.9) | 7.0 (5.5) |
| Refinement | ||
| Resolution (Å) | 3.25 | |
| No. reflections | 30437 | |
| Rwork/ Rfree | 0.236/0.305 | |
| No. atoms | 8060 | |
| Protein | 8004 | |
| Ligand/ion | 56 | |
| Water | ||
| B-factors | 94.50 | |
| Protein | 94.50 | |
| Ligand/ion | 93.40 | |
| Water | ||
| R.m.s deviations | ||
| Bond lengths (Å) | 0.010 | |
| Bond angles (°) | 1.40 | |
Highest resolution shell is shown in parenthesis.
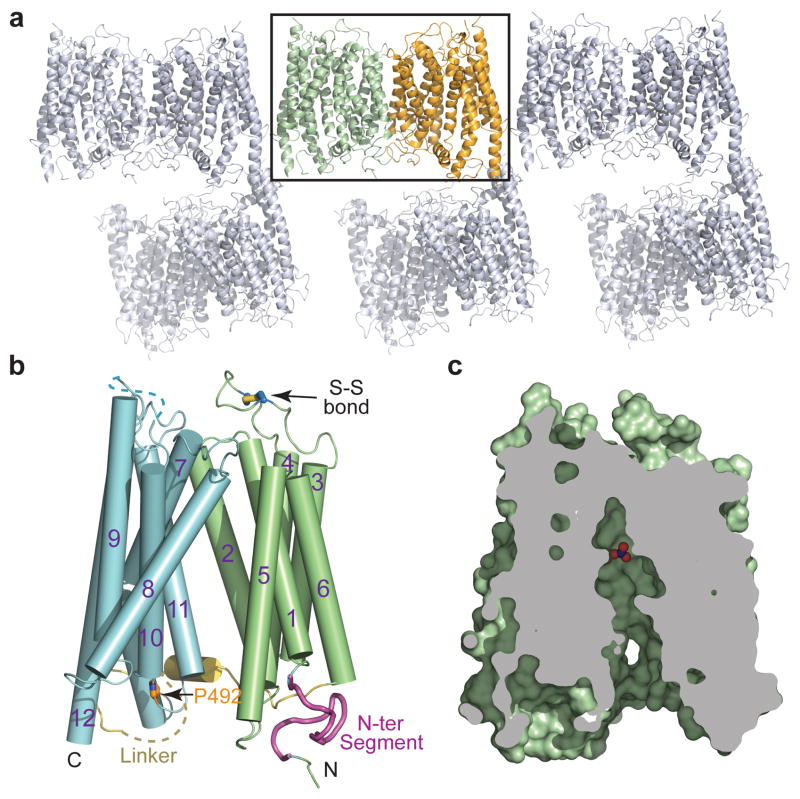
NRT1.1 crystallized with two molecules in the asymmetric unit. The two copies of NRT1.1 can be superimposed with a root mean square deviation of 0.9 Å over 504 Cα atoms, indicating a common overall structure (Fig. 1a). As predicted, the transporter adopts a typical MFS fold, which is characterized by 12 TMHs with a pseudo 2-fold axis relating the N-terminal (TMH1–6) and C-terminal (TMH7–12) domains (Fig. 1b). NRT1.1 is captured in an inward-facing conformation as previously observed in the LacY19, GlpT20, and PepTSt25 structures (Fig. 1c, Extended Data Fig. 4).
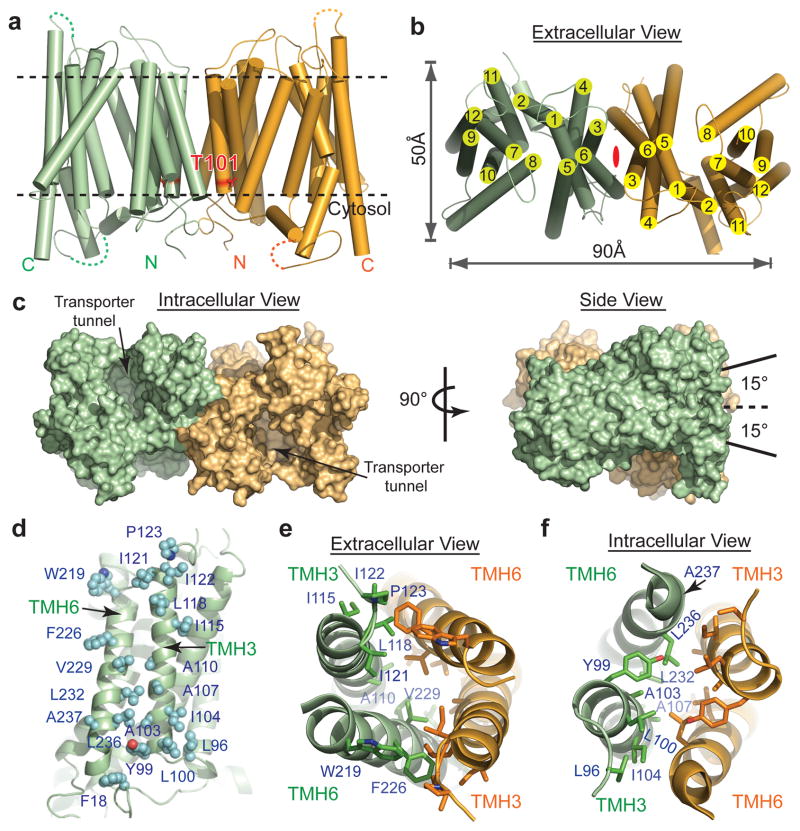
Figure 1. Crystal packing and overall structure of NRT1.1.
a, Crystal packing of NRT1.1 in C2221 space group with two molecules in each asymmetric unit. b, Overall structure of NRT1.1. The N-terminal and C-terminal domains, the N-terminal conserved segment, the inter-domain linker and Pro492 are colored in pale green, cyan, magenta, yellow and orange, respectively. A functional important extracellular disulfide bond is indicated. c, Cutaway view showing that NRT1.1 is captured in an inward conformation with nitrate displayed in spheres.
Extended Data Figure 4. Structural comparison of NRT1.1 and other MSF transporters.
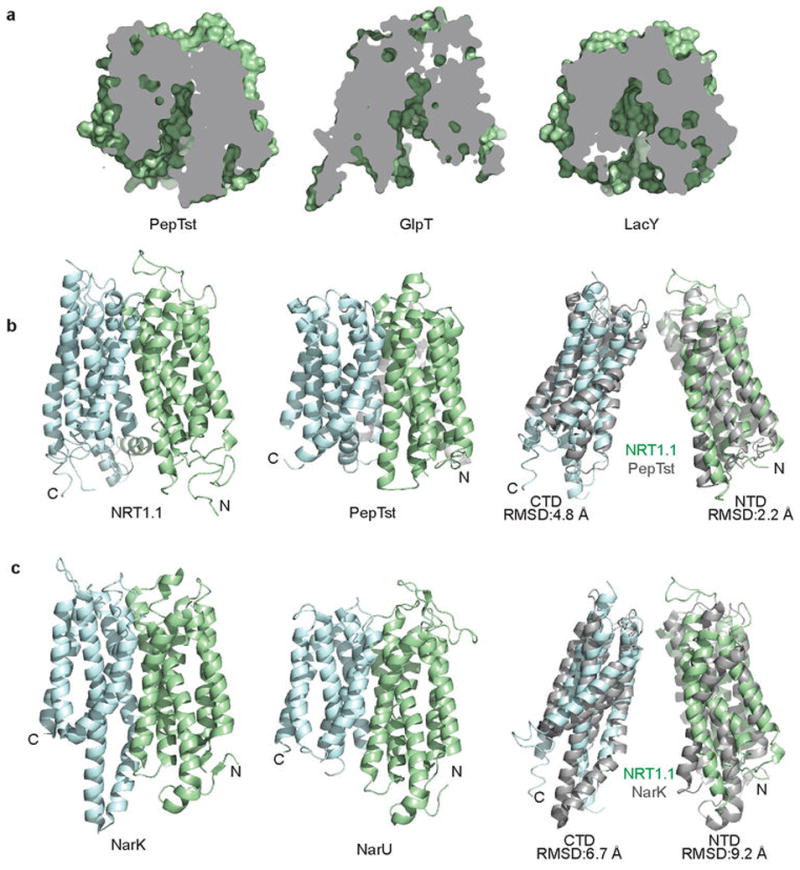
a, Cutaway views of PepTst, GlpT and LacY showing their shared inward conformation as observed for NRT1.1. b, Overall structural comparison of NRT1.1 and PepTst with their N-terminal and C-terminal domains (NTD & CTD) colored in pale green and cyan, respectively. Superposition of their NTDs and CTDs are shown separately. c, Comparisons between NRT1.1 and two bacterial NRT2 family nitrate transporters, NarK and NarU.
By comparing the structures of NRT1.1 and a bacterial peptide transporter, PepTSt, we confirm that eukaryotic and prokaryotic members of the NRT1/PTR family of MFS transporters share a similar overall architecture (Extended Data Fig. 4). The plant nitrate transporter, nonetheless, has three unique and conserved structural elements, including a well-structured N-terminal cytoplasmic segment, a disulfide bond-stabilized extracellular loop, and a partially ordered central linker sequence (Fig. 1c). Consistent with the sequence divergence between the NRT1/PTR and NRT2/NNP nitrate transporter families, the structures of NRT1.1 and two bacterial NRT2/NNP family members, NarK24 and NarU26, share few common features except the MFS fold (Extended Data Fig. 4). The residue mutated (P492L) in the chl1-9 mutant, which lost the transporter but not the sensor function of NRT1.117, is located at the short TMH10–TMH11 loop (Fig. 1c). Its mutation likely affects the structural coordination of the two helices.
NRT1.1 dimer in the crystal
To date, crystal structures of more than ten MFS transporters have been determined in the monomeric form19–29. The DDM-solubilized NRT1.1 protein was also isolated in a monomeric state as determined by size exclusion chromatography-coupled multi-angle light scattering measurements31 (Extended Data Fig. 5a). A closer examination of the two NRT1.1 molecules in the asymmetric unit, however, reveals a possible biological dimer.
Extended Data Figure 5. Assessment of the oligomerization state of detergent-solubilized NRT1.1.
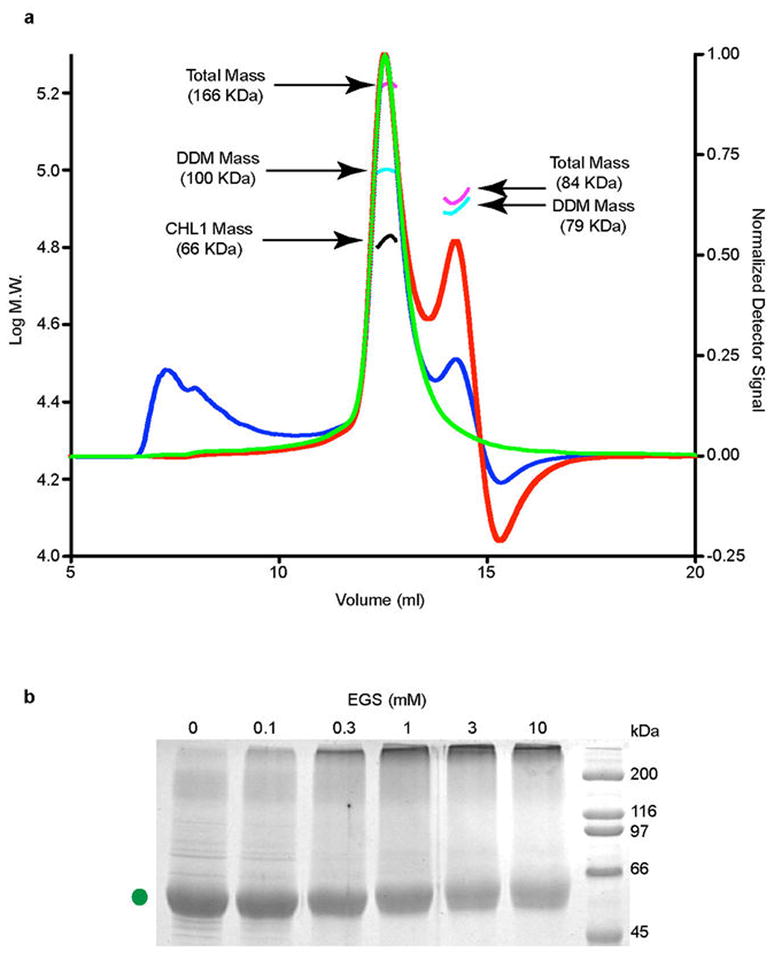
a, SEC-LS-RI-UV analysis of DDM-solubilized NRT1.1. Normalized light scattering signal from the 90° detector, UV absorption signal, and the refractive index signals are plotted in blue, green, and red lines, respectively. The highest peak contains NRT1.1 bound to DDM. The calculated masses of the protein-detergent micelle complex (magenta), DDM micelle (cyan), and the NRT1.1 protein (black) are shown. The complex contains about 196 detergent molecules (MW 510.62) and 1 NRT1.1 molecule (67 kDa). The second peak belongs to the detergent micelle with a mass of 79 kDa or 155 detergent molecules. b, Crosslinking of the NRT1.1 T101D mutant protein with increasing concentrations of EGS. The protein was purified in the presence of 0.1% digitonin.
In the crystal, the two adjacent noncrystallographically related NRT1.1 molecules are juxtaposed in a side-to-side fashion with their N-terminal halves facing and interacting with each other (Fig. 1a & 2a, b). The intermolecular packing is predominantly mediated by TMH3 and TMH6, which are located at a peripheral edge of the canonical MFS fold. Although crystal contacts may not always reflect biological interactions, two prominent features of the crystallographic dimer arrangement support its physiological relevance. First, the overall topology of the putative NRT1.1 dimer is perfectly compatible with its transporter function at the membrane (Fig. 2a, b). Second, the interface between the two NRT1.1 molecules is extensive and complementary with a total surface area of ~2160 Å2 ((Fig. 2d–f, Extended Data Fig. 6). Overall, the two inward-facing NRT1.1 molecules give rise to a putative “in-phase” dimer assembly, which is about 90 Å wide and 50 Å thick (Fig. 2b). When viewed from the side, the two substrate-transporting tunnels are not in parallel with the central two-fold axis but slant at an ~15° angle in opposite directions (Fig. 2c).
Figure 2. NRT1.1 dimer interface.
a, Cylinder representation of the NRT1.1 dimer with Thr101 shown in red sticks. b, Extracellular view of the NRT1.1 dimer with a central 2-fold axis indicated in red. c, Two orthogonal views of the NRT1.1 dimer in surface representation. The dashed line represents the central 2-fold axis. d, NRT1.1 dimer interface. The side chains of all interface residues are shown in cyan spheres. e–f, Extracellular and intercellular views of the NRT1.1 dimer interface with TMH3 and TMH6 shown in ribbon and the side chains of interacting residues shown in sticks. The interface residues of one chain are labeled.
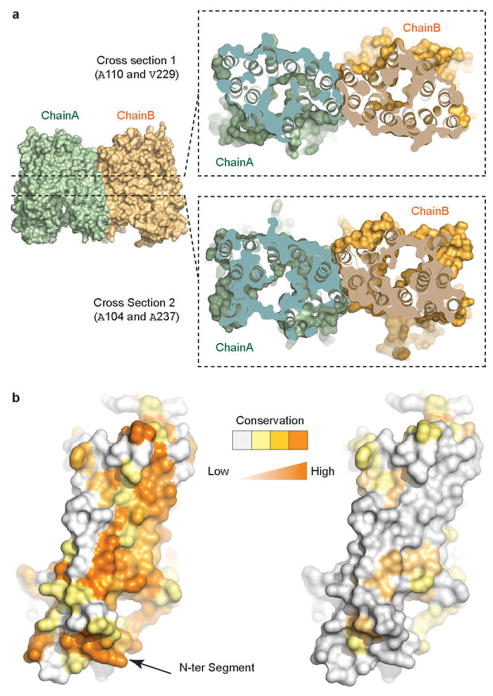
Extended Data Figure 6. Shape complementarity and conservation of the NRT1.1 dimer interface.
a, Two representative cross section views of the NRT1.1 dimer interface that are parallel to the membrane. The top cross section goes through the plane defined by Ala110 and Val229 in the two NRT1.1 protomers. The bottom cross section goes through Ala104 and Ala237. b, Conservation surface mapping of NRT1.1 residues at the dimer interface among NRT1.1 orthologs (left) and among Arabidopsis NRT1 family members (right). A color ramp (white, pale yellow, bright orange, to deep orange) is used to indicate the degree of conservation of surface residues. The arrow indicates the N-terminal segment.
Functional dimerization of NRT1.1
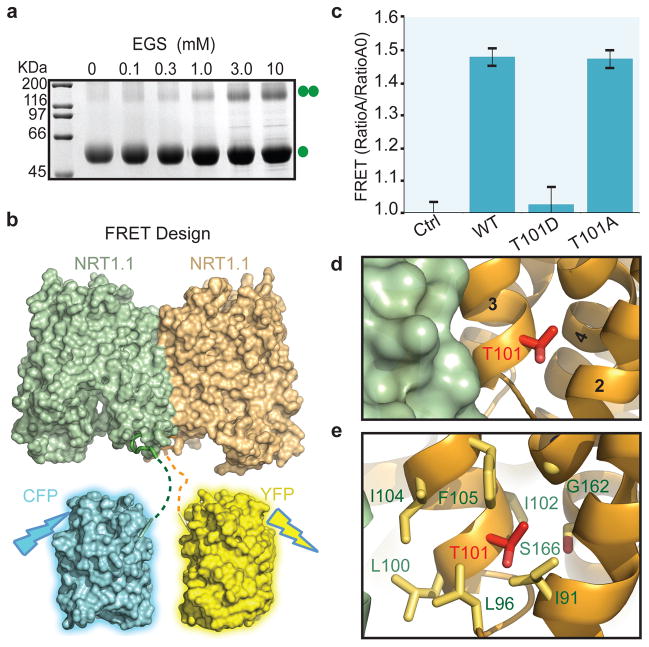
To dissect the biological relevance of the NRT1.1 dimer observed in the crystal, we first used a crosslinking experiment to assess the potential of detergent solubilized NRT1.1 to dimerize in solution. Despite its low efficiency, an amine reactive crosslinker was able to crosslink NRT1.1 in a concentration dependent manner (Fig. 3a). The crosslinked products migrated on SDS-PAGE with a size corresponding to a NRT1.1 dimer, indicating that DDM-solubilized NRT1.1 is capable of forming a transient dimer in a membrane-free environment.
Figure 3. NRT1.1 dimerization controlled by T101 phosphorylation.
a, Crosslinking of NRT1.1 with increasing concentrations of EGS. b, The design of FRET assay. Dashed lines indicate the eleven residue long linkers between the fluorescence proteins and the structurally resolved NRT1.1 N-terminus. c, FRET measurements of wild type and mutant NRT1.1. The mCFP-HCN-mYFP-NRT1.1 pair was used as negative control. Consistent with the lose of FRET signal, the T101D mutant failed to be crosslinked in solution (Extended Data Fig. 5b). d, A close-up view of Thr101 at the NRT1.1 dimer interface. e, Thr101-interacting residues with their side chains shown in sticks.
Because solubilization by DDM might interfere with NRT1.1 dimer formation, we next performed Förster resonance energy transfer (FRET) spectroscopy experiments with the nitrate transporter expressed in the membrane of Xenopus oocytes32, which allowed us to examine the oligomeric state of NRT1.1 in the same lipid environment where its dual-affinity transporter activity has been measured. We separately fused the N-terminus of NRT1.1 with either the mCerulean variant of cyan fluorescent protein (mCFP) or the mCitrine variant of yellow fluorescent protein (mYFP), which constitute a FRET pair with an R0 of ~ 50 Å for 50% energy transfer efficiency33 (Fig. 3b). In the structure of the putative NRT1.1 dimer, the N-terminal ends of the two NRT1.1 molecules are about 40 Å away from each other (Extended Data Fig. 7a). Therefore, FRET is expected to occur if NRT1.1 dimerizes in the membrane in the same fashion as seen in the crystal structure. As shown in Fig. 3c, a strong FRET signal measured by a spectrum-based approach was indeed detected between the co-expressed mCFP-NRT1.1 and mYFP-NRT1.1 fusion proteins, but not in the negative control (Extended Data Fig. 7b). This result strongly suggests that the plant dual-affinity nitrate transporter can form a homo-dimer not only in the crystal but also in the cellular membrane.
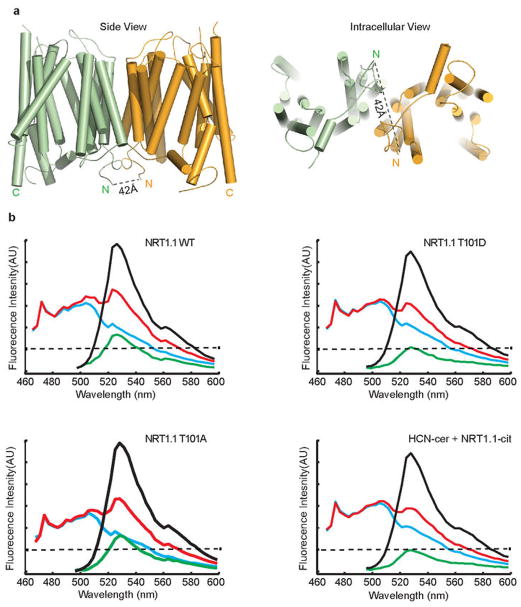
Extended Data Figure 7. Spatial relationship between the N-terminus of the two NRT1.1 protomers.
a, The N-termini of the two NRT1.1 protomers in the crystal structure are about 42 Å apart and are shown in two orthogonal orientations. b, Spectral quantification of the FRET. Emission spectra measured from an oocyte expressing WT NRT1.1 (top left), NRT1.1-T101D (top right), NRT1.1-T101A (bottom left), and WT mCitrine-NRT1.1 and mCerulean-HCN2 (bottom right). The spectra are color coded as follows: cyan, 458 nm excitation of oocytes expressing mCerulean constructs alone; black, 488 nm excitation of oocytes expressing both mCitrine and mCerulean constructs; red, 458 nm excitation of oocytes expressing both mCitrine and mCerulean constructs; green, subtracted spectrum (red minus cyan). The dashed line is the position of the peak of the fluorescence signal after excitation at 458nm of the mCitrine-NRT1.1 only expressing oocytes (the position of the Azero or no FRET peak).
Thr101 phosphorylation as a dimerization switch
The phosphorylation site residue, Thr101, is strictly conserved among plant NRT1.1 orthologs and represents one of the hallmarks of the dual-affinity nitrate transporter11 (Extended Data Fig. 1). In the NRT1.1 structure, Thr101 is located at the N-terminal end of TMH3 and is entirely buried in a hydrophobic pocket formed among TMH2, TMH3, and THM4 (Fig. 3d, e). Strikingly, this pocket is directly adjacent to the dimer interface with one of its walls demarcated by three hydrophobic interface residues, Leu96, Leu100, and Ile104 (Fig. 2d, 3e). While the side chain of Thr101 is unmodified in the crystal, its phosphorylation is expected to induce major electrostatic and conformational changes in its vicinity and have a direct impact on the dimer interface. This structural clue prompted us to postulate that the formation of the NRT1.1 dimer might be determined by the phosphorylation status of Thr101 and the two distinct affinity states of the transporter might be enabled by the difference in its oligomerization state.
To test this hypothesis, we compared the wild type NRT1.1 with the phosphomimetic mutant, T101D, and the phosphorylation defective mutant, T101A, in the oocyte-based FRET experiments (Fig. 3c, Extended Data Fig. 7b). Consistent with the prediction, the phosphomimetic mutant, T101D, which has been previously shown to bear a monophasic high affinity nitrate transporter activity11, completely lost the FRET signal, indicating a spatial separation of the two NRT1.1 N-terminal domains, if not a complete disruption of the NRT1.1 dimer. By contrast, the phosphorylation defective mutant, T101A, which is known to transport nitrate in the low affinity state11, generated a robust FRET signal similar to the wild type protein. The matching levels of FRET between wild type NRT1.1 and the T101A mutant indicates that the wild type protein is mostly in the unmodified form under the nitrate-free condition of the FRET measurement. This is in agreement with previous studies showing that phosphorylated NRT1.1 only started to accumulate when plants were exposed to nitrate. Together, these results not only confirm the functional relevance of the NRT1.1 dimer, but also suggest a dimerization-based switching mechanism for the dual affinity nitrate transporter — unmodified NRT1.1 forms a structurally coupled homo-dimer and functions as a low-affinity transporter, whereas phosphorylated NRT1.1 undergoes dimer decoupling and adopts a high-affinity state.
Substrate Binding Site and Proton Coupling
In both protomers of the refined NRT1.1 dimer structure, an island of strong electron density is present in the middle of the transport tunnel between the N-terminal and C-terminal domains (Fig. 4a, Extended Data Fig. 8). The overall location of the density coincides with the substrate binding sites of other MFS structures, suggesting that it belongs to the substrate molecule, nitrate. Indeed, when nitrate was omitted from the cryo-protection buffer, this electron density completely disappeared from both NRT1.1 molecules (Extended Data Fig. 8). The position of the nitrate density is slightly shifted between the two NRT1.1 molecules, which might reflect the mode of substrate release (Extended Data Fig. 8).
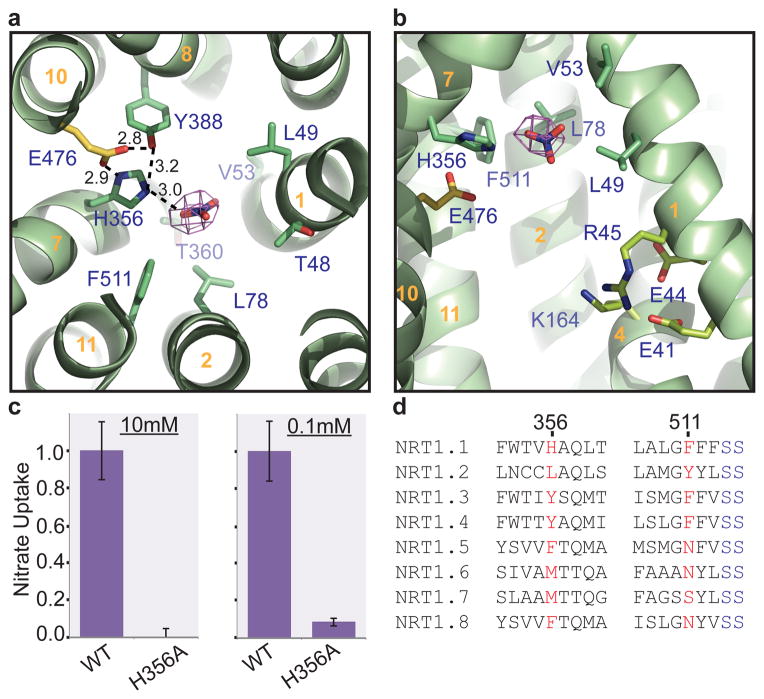
Figure 4. Substrate binding and energy coupling in NRT1.1.
a, Intracellular view of the nitrate binding pocket. Nitrate is shown in sticks together with electron density contoured at 4σ from a Fo-Fc map calculated before the nitrate was modeled in. THMs are numbered (orange). E476 is a His356-interacting residue. whose mutation abolishes the transporter function of NRT1.1 (Extended Data Fig. 3b). b, Side view of the putative nitrate binding site and the transporter tunnel with the clustered ExxER motif and K164. c, Nitrate uptake activities of the H356A mutant relative to wild type NRT1.1 in the presence of 10mM or 0.1mM nitrate. All results are the mean ± s.d. of one experiment in quintuplicates or sextuplicates. d, Sequence alignment of eight NRT1 family members from Arabidopsis thaliana in regions surrounding H356 and F511 of NRT1.1.
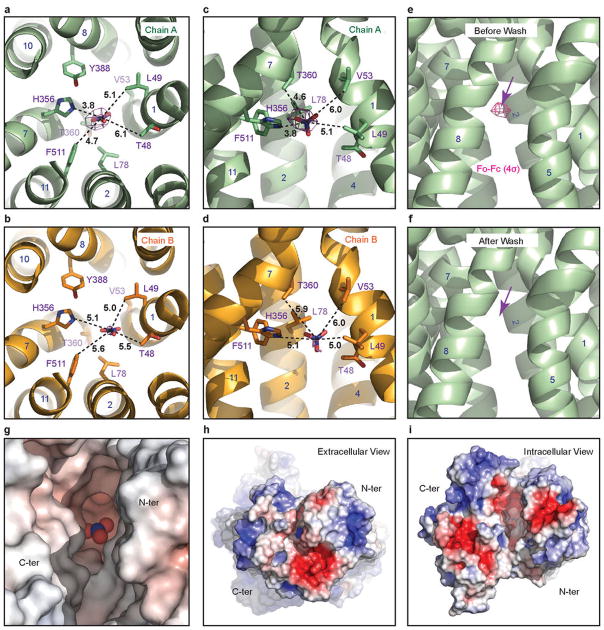
Extended Data Figure 8. Putative nitrate-binding site in the two NRT1.1 protomers.
a–b, Intracellular view of the substrate binding site in the two copies of NRT1.1 within the dimer. To compare the relative position of the substrate to its surrounding residues, distances between the nitrogen atom of the modeled nitrate and select amino acid atoms in its vicinity are shown in dashed lines and indicated. Nitrate is shown in sticks with electron density countered at 4σ from a Fo-Fc map calculated without the substrate. TMHs are numbered. c–d, Side view of the substrate binding site. e–f, A comparison of the putative substrate density between the NRT1.1 structures determined with a cryo-protectant solution containing 10 mM or 0 mM nitrate. g–i, Electrostatic potential surface of the NRT1.1 substrate pocket. The surface colors are clamped between red (−20 kT/e) and blue (+20 kT/e). Nitrate is shown as spheres. Two global views of the electrostatic potential surface of NRT1.1 are shown for comparison.
Distinct from the nitrate/nitrite binding sites of NarK and NarU, which coordinate the substrate(s) with two opposing conserved Arg residues24,26, the nitrate-binding pocket in NRT1.1 is predominantly formed by hydrophobic residues, including Leu49, Val53, Leu78, and Phe511 (Fig. 4a, b). His356 on TMH7 is the only polar residue that is in close contact with nitrate, whose precise binding mode cannot be resolved due to the resolution limit of the structure. Based on its close proximity to the nitrate density and the crystallization condition (pH=4.5), His356 likely stabilizes the substrate in the pocket through a charge-charge interaction. Its side chain conformation, meanwhile, is supported by two nearby residues, Tyr388 and Glu476. Although Tyr388 and two other polar residues, Thr360 and Thr48, are also around the substrate, their hydroxyl groups do not appear to be at the optimal hydrogen bond distance (Fig. 4a).
To validate the substrate-binding site, we mutated His356 and compared the nitrate uptake activities of the wild type and mutant transporters11. In support of a critical role of His356 in substrate transport, its mutation to alanine completely abolished the transport activity of NRT1.1 at both high and low nitrate concentration (Fig. 4c). Interestingly, His356 is not conserved among plant NRT1.1 orthologs and Arabidopsis NRT1 family members, which harbor either a tyrosine or a hydrophobic amino acid (Leu/Met/Phe) at the equivalent position (Fig. 4d). This key residue, nevertheless, has closely co-evolved with the adjacent residue, Phe511. Among all NRT1.1 orthologs and paralogs, a combination of a polar and a hydrophobic side chain has been generally maintained between the two residues, suggesting that one of them is responsible for specific nitrate binding (Fig. 4d, Extended Data Fig. 1, 2). NRT1.1 is unique among all Arabidopsis NRT1 family members by featuring a histidine at the nitrate-binding pocket. This charged residue provides a plausible explanation for the high affinity nitrate uptake activity acquired by NRT1.1, which is otherwise a member of LATS. On the other hand, the replacement of the histidine residue by tyrosine in some of the plant NRT1.1 orthologs raises a question about their dual affinity transporter function (Extended Data Fig. 1).
In the PepTSt structure, a conserved motif, ExxERFxYY, on TMH1 has been identified to play an important role in proton coupling. Part of this motif, ExxER, is also found in all plant NRT1.1 orthologs (Extended Data Fig. 1). Together with the conserved residue, Lys164, this motif presents a cluster of interacting residues under the nitrate-binding pocket and facing toward the transport tunnel (Fig. 4b). Consistent with a key function in the symport cycle, alanine mutation of each of the four residues abrogated the transporter activity of NRT1.1 in the oocyte-based nitrate uptake assay (Extended Data Fig. 3b). Surprisingly, these four residues have been simultaneously evolved into non-charged residues in two Arabidopsis NRT1 family members, AtNRT1.5 and AtNRT1.8 (Extended Data Fig. 2). Their documented pH-dependent nitrate transporter activities necessitate an alternative proton coupling mechanism.
Cytoplasmic Structural Elements
NRT1.1 has an ~30 amino acids long N-terminal cytoplasmic segment, which is highly conserved among its plant orthologs (Extended Data Fig. 1 & 2). In the crystal, this sequence adopts a well-ordered loop structure and forms a pronounced cleft between the NRT1.1 dimer (Extended Data Fig. 9). With several strictly conserved residues exposed to the solvent, this cleft presents a putative two-fold symmetric protein-protein interaction site with a potential role in recruiting kinases and phosphatases. Although the central linker sequence is mostly disordered in the crystal, its N-terminal region forms a stable amphipathic α-helix (Fig. 1c, 2a), providing yet another potential protein docking site.
Extended Data Figure 9. The conserved cleft formed by the N-terminal segment of NRT1.1.
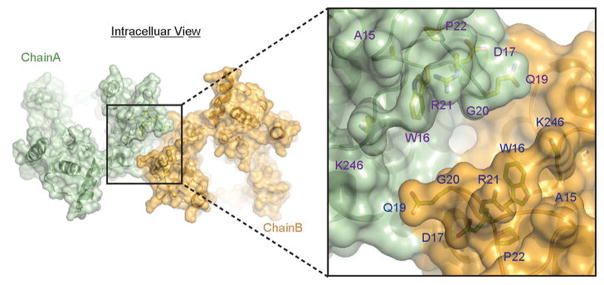
Overall and close-up views of the N-terminal segments of NRT1.1 within the dimer are shown in surface representation. The cleft forming residues, which are strictly conserved in the NRT1.1 orthologs, are labeled and shown in sticks.
Discussion
The crystal structure of NRT1.1 reveals a biologically relevant dimer, whose dynamic coupling and decoupling is controlled by the phosphorylation of a single residue near the dimer interface. Since the same post-translational modification switches the mode of action of the dual-affinity transporter, we propose that dimer assembly and disassembly enables NRT1.1 to toggle between the low-affinity and high-affinity states with an overlapping, if not the same, substrate-binding site. In this model (Fig. 5), structural engagement of two protomers at the interface allosterically regulates the affinity of substrate binding at the central transport tunnel. Whether dimer decoupling itself is sufficient to shift NRT1.1 into the high-affinity mode awaits future analysis.
Figure 5. A dimerization switch model.
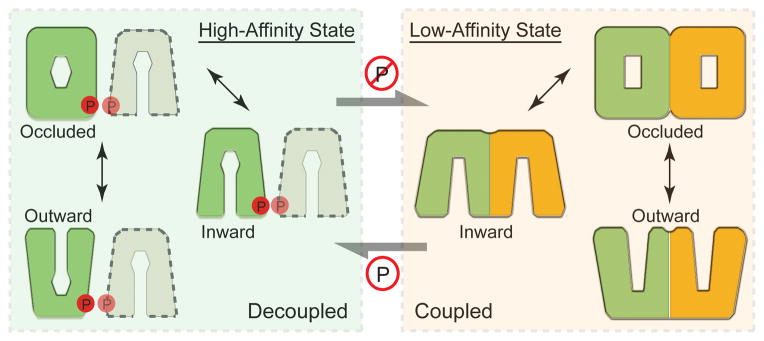
The non-phosphorylated and structurally coupled NRT1.1 dimer functions as an “in-phase” homodimeric low-affinity nitrate transporter (right). Once phosphorylated, the NRT1.1 dimer is decoupled, and each molecule functions as an independent high-affinity nitrate transporter (left). Different shapes of the putative substrate-binding site at the central transport tunnel reflect its differential nitrate binding properties.
Previous structural studies of several MFS members have established a “rocker-switch” mechanism of substrate transport, in which the transporters cycle through outward-facing, occluded, and inward-facing conformations21,25. The structure of the dimeric unmodified NRT1.1 reveals a buried Thr101 phosphorylation site in the inward-facing conformation. Thr101 phosphorylation, therefore, likely occurs when the dimeric transporter adopts either the outward-facing or occluded conformation. It is equally possible that the unmodified dimeric transporter is in equilibrium with the monomeric form, which is susceptible to phosphorylation.
Despite the current structure, questions remain as to how NRT1.1 senses nitrate and transduces the signal. Our structure reveals only one nitrate-binding site within the substrate transport tunnel. However, there might be an additional nitrate binding site responsible for signaling, which is excluded from the inward-facing conformation. Previous studies have suggested that dephosphorylation of Thr101 is required for the low-affinity sensor function of NRT1.117. It is possible that dimerization mediates this signaling function of the transporter in the same manner as found in many common cell surface receptors. The high-affinity sensor function of NRT1.1, on the other hand, might involve an entirely different mechanism.
Transporter oligomerization and phosphorylation have been implicated in the proper functions of a number of MFS members, such as LacS34, GLUT135, TetL36, and hRFC37. The crystal structure of the NRT1.1 dimer not only establishes a structural framework for understanding its dual-affinity nitrate transporter and receptor activities, but also reveals for the first time how protein oligomerization and post-translational modification can synergistically expand the functional capacity of an MFS transporter.
ONLINE METHODS
Protein expression and purification
The full-length Arabidopsis thaliana NRT1.1 was cloned into pFastBac vector with a C-terminal 8xHis tag. Recombinant baculovirus was generated using Bac-to-Bac system (Invitrogen). Monolayer High Five insect cells were infected for protein expression. Cells were harvested 72 hours after infection, washed and lysed with a buffer containing 20 mM Tris-HCl, pH 8.0, 200 mM NaCl, 10 mM NaNO3 (Buffer A). NRT1.1 was subsequently solubilized by DDM (Anatrace) at a final concentration of 1.5% with 1–2 hr incubation at 4°C. Solubilized NRT1.1 was separated from the insoluble fraction by high-speed ultra-centrifugation and applied to a Ni-NTA gravity column (GE Healthcare). The bound NRT1.1 protein was then washed and eluted in the presence of 0.02% DDM with Buffer A supplemented with 20 mM and 200 mM imidazole, respectively. The 8xHis tag was cleaved by thrombin overnight and the NRT1.1 protein was further purified by size exclusion chromatography with a Superdex 200 column (GE Healthcare) in Buffer A supplemented with 0.02% DDM. The peak fractions containing NRT1.1 were pooled and the protein was concentrated to 10 mg/ml using a 50 kDa MWCO centrifugal device (Ambion).
Crystallization, data collection and structure determination
The NRT1.1 crystals were grown at 4°C by the hanging-drop vapour diffusion method, using 2 μl protein sample mixed with 1 μl reservoir solution containing 100 mM sodium acetate, pH 4.5, 30% PEG300 and 3% MPD. Crystals of maximal sizes were obtained after 1 month. 7.5% ethylene glycol was added during crystal harvest and data collection as cryo-protectant. The heavy atom derivative crystals were obtained by soaking in the presence of 1 mM ethyl mercury thiosalicylate (EMTS) for 2 hours. All data sets were collected at the BL8.2.1 and BL8.2.2 beamlines at the Advanced Light Source. The single anomalous dispersion (SAD) data set was collected near the mercury absorption edge (λ = 1.008 Å). X-ray diffraction data were integrated and scaled with HKL2000 package38. Improved molecular replacement by Rosetta30 combined with SAD was used to determine the initial phase using PHENIX39 with a 3.5 Å mercury derivative data set. Initial structural models were built, refined and rebuilt using COOT40 and PHENIX. The final model was built and refined with a native data set of 3.25 Å resolution. All the structure model figures in the paper were prepared using PyMol.
Transporter assay
The transporter assays in oocytes were carried out as previously described11. Briefly, genes were constructed into pGEMHE vector and the plasmids were then linearized by NheI (NEB). cRNAs were prepared using mMessage mMachine T7 Ultra Kit (Ambion) and diluted to a final concentration of 2 μg/ml. 50 nl cRNA was manually injected into each oocyte. Oocytes were then incubated at 16°C in regular Barth’s Solution supplemented with gentamicin. After 3 days, transporter assays were carried out by incubating oocytes in the following solution: 230 mM Mannitol, 0.3 mM CaCl2, 10 mM MES-TRIS, pH 5.5 for 2 hours with N15 labeled K15NO3 (Sigma) supplemented when required. The oocytes were rinsed 5 times with ND96 buffer after incubation and then dried individually at 80 °C for 24 hours. The retained 15N was analyzed on a continuous flow isotope ratio mass spectrometry coupled with a nitrogen elemental analyzer (UC Davis Stable Isotope Facility). To prepare for structural studies, we verified that the affinity-tagged NRT1.1 construct still retains its characteristic dual-affinity nitrate transporter activity in the Xenopus oocyte system (Extended Data Fig. 3a). For NRT1.1 mutants, the background signal from the oocytes injected with water was subtracted from the final uptake data, which were subsequently normalized to the wild type protein. Every data point in the experiments was measured and averaged on more than 5 oocytes.
FRET assay
The mCerulean variant of cyan fluorescent protein and mCitrine variant of yellow fluorescent protein were individually fused to the N-terminus of NRT1.1 with a three-alanine linker. Then cRNAs were prepared using the same vector and kit as in the transporter assay and injected into oocytes. The FRET signal from the surface membrane was measured after incubation for 3 days using the Zeiss LS-710 confocal microscope as previously described32. Briefly, emission spectra of mCerulean and mCitrine were collected using laser excitation at 458 and 488 nm, respectively, and an emission window of 3.2 nm. FRET was calculated using a spectrum-based approach to remove contaminations caused by donor emission and direct excitation of the acceptor fluorophore. mCerulean spectra were collected from oocytes expressing mCerulean-tagged NRT1.1 transporters alone and then subtracted from spectra taken from oocytes expressing both mCerulean and mCitrine tagged transporters. The resulting extracted mCitrine emission spectrum, F458, contains two components: one caused by direct excitation, Fdirect458, and the other by FRET, FFRET458. F458 was normalized by total mCitrine emission excited directly with 488nm light (F488). The resulting ratio, termed RatioA, can be expressed as:
The direct excitation component, , termed RatioA0, was experimentally determined with oocytes expressing mCitrine-tagged transporters only. The ratio between RatioA and RatioA0 (FR), directly proportional to FRET efficiency, was calculated as follows:
Images were analyzed in ImageJ (National Institute of Health) by drawing regions of interest around the fluorescent membranes and using the measure stack feature. The data were then analyzed using programs written in MATLAB (MathWorks, Natick, NJ)
Crosslinking experiment
Purified NRT1.1 protein was dialyzed in PBS buffer, pH 7.4 supplemented with 0.01% DDM for 24 hours with one buffer exchange. The protein was then concentrated to 5 mg/ml, and 2 μl ethylene glycol bis-succinimidylsuccinate (EGS) (Pierce) was added into 18 μl protein sample at increasing concentrations (0, 0.1 mM, 0.3 mM, 1 mM, 3 mM, and 10 mM). The reaction was carried out at room temperature for 30 minutes and stopped by a 50 mM Tris-HCl buffer, pH 8.0. 1/5 of the reaction solution was analyzed by SDS-PAGE.
SEC-LS-RI-UV measurement
The SEC-MAL system consisted of a P900 HPLC pump (GE), a UV-2077 detector (Jasco), a Tri Star Mini Dawn light scattering instrument (Wyatt), and an Opti Lab T-Rex refractive index instrument (Wyatt). 20 μL of purified and DDM-solubilized NRT1.1 (5 mg/ml) was injected into a Superdex 200 (10/300GL) gel filtration column and eluted isocratically at 0.5 ml/min in a buffer containing 20 mM Tris, 200 mM NaCl, 5mM NaNO3, 0.02% DDM, pH 8.0. The extinction coefficient of NRT1.1 at 280 nm was calculated from the amino acid sequence (E=1.268 ml•mg−1•cm−1). DDM has no absorbance at 280 nm. The specific refractive index of NRT1.1 and DDM was assumed to be 0.186 g/ml and 0.147ml/g, respectively. Data collection and analysis was performed with Astra 6 software (Wyatt). Total molecular mass and individual masses of the protein and the detergent were determined with Astra6 software using protein conjugate analysis. Both peak over lap and peak broadening were corrected with Astra 6 software. The SEC-MAL system was pre-calibrated with BSA.
Acknowledgments
We thank the beamline staff of the Advanced Light Source at the University of California at Berkeley. We also thank members of the Zheng laboratory, Zagotta laboratory, Xu laboratory and Hongjin Zheng for discussion and help. This work is supported by the Howard Hughes Medical Institute (N. Z.), National Institutes of Health (R01EY10329 to W.N.Z., NS074545 to J.R.B.) and the National Science Foundation (N.Z.).
Footnotes
Author Contributions. J.S. and N.Z. conceived, and J.S. conducted the protein purification and crystallization experiments. J.P. provided experimental suggestions. J.S. and N.Z. determined and analyzed the structures. J.S., J.R.B., W.N.Z., and N.Z. conceived, and J.S. and J.R.B. conducted FRET experiments. J.S., and N.Z. conceived and J.S. conducted mutational and transporter assays. J.S. and N.Z wrote the manuscript with inputs from all authors.
Structural coordinates and structural factors will be deposited in the Protein Data Bank under accession numbers 4OH3.
Authors declare no financial interest.
References
- 1.Wang YY, Hsu PK, Tsay YF. Uptake, allocation and signaling of nitrate. Trends in plant science. 2012;17:458–467. doi: 10.1016/j.tplants.2012.04.006. [DOI] [PubMed] [Google Scholar]
- 2.Tsay YF, Chiu CC, Tsai CB, Ho CH, Hsu PK. Nitrate transporters and peptide transporters. FEBS letters. 2007;581:2290–2300. doi: 10.1016/j.febslet.2007.04.047. [DOI] [PubMed] [Google Scholar]
- 3.Nacry PB, Gojon EA. Nitrogen acquisition by roots: physiological and developmental mechanisms ensuring plant adaptation to a fluctuating resource. Plant and Soil. 2013;370:29. doi: 10.1007/s11104-013-1645-9. [DOI] [Google Scholar]
- 4.Pao SS, Paulsen IT, Saier MH., Jr Major facilitator superfamily. Microbiology and molecular biology reviews: MMBR. 1998;62:1–34. doi: 10.1128/mmbr.62.1.1-34.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Law CJ, Maloney PC, Wang DN. Ins and outs of major facilitator superfamily antiporters. Annual review of microbiology. 2008;62:289–305. doi: 10.1146/annurev.micro.61.080706.093329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Leran S, et al. A unified nomenclature of NITRATE TRANSPORTER 1/PEPTIDE TRANSPORTER family members in plants. Trends in plant science. 2013 doi: 10.1016/j.tplants.2013.08.008. [DOI] [PubMed] [Google Scholar]
- 7.Tsay YF, Schroeder JI, Feldmann KA, Crawford NM. The herbicide sensitivity gene CHL1 of Arabidopsis encodes a nitrate-inducible nitrate transporter. Cell. 1993;72:705–713. doi: 10.1016/0092-8674(93)90399-b. [DOI] [PubMed] [Google Scholar]
- 8.Huang NC, Chiang CS, Crawford NM, Tsay YF. CHL1 encodes a component of the low-affinity nitrate uptake system in Arabidopsis and shows cell type-specific expression in roots. The Plant cell. 1996;8:2183–2191. doi: 10.1105/tpc.8.12.2183. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Wang R, Liu D, Crawford NM. The Arabidopsis CHL1 protein plays a major role in high-affinity nitrate uptake. Proceedings of the National Academy of Sciences of the United States of America. 1998;95:15134–15139. doi: 10.1073/pnas.95.25.15134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Liu KH, Huang CY, Tsay YF. CHL1 is a dual-affinity nitrate transporter of Arabidopsis involved in multiple phases of nitrate uptake. The Plant cell. 1999;11:865–874. doi: 10.1105/tpc.11.5.865. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Liu KH, Tsay YF. Switching between the two action modes of the dual-affinity nitrate transporter CHL1 by phosphorylation. The EMBO journal. 2003;22:1005–1013. doi: 10.1093/emboj/cdg118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Guo FQ, Young J, Crawford NM. The nitrate transporter AtNRT1.1 (CHL1) functions in stomatal opening and contributes to drought susceptibility in Arabidopsis. The Plant cell. 2003;15:107–117. doi: 10.1105/tpc.006312. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Wang R, Okamoto M, Xing X, Crawford NM. Microarray analysis of the nitrate response in Arabidopsis roots and shoots reveals over 1,000 rapidly responding genes and new linkages to glucose, trehalose-6-phosphate, iron, and sulfate metabolism. Plant physiology. 2003;132:556–567. doi: 10.1104/pp.103.021253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Krouk G, et al. Nitrate-regulated auxin transport by NRT1.1 defines a mechanism for nutrient sensing in plants. Developmental cell. 2010;18:927–937. doi: 10.1016/j.devcel.2010.05.008. [DOI] [PubMed] [Google Scholar]
- 15.Walch-Liu P, et al. Nitrogen regulation of root branching. Annals of botany. 2006;97:875–881. doi: 10.1093/aob/mcj601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Munos S, et al. Transcript profiling in the chl1-5 mutant of Arabidopsis reveals a role of the nitrate transporter NRT1.1 in the regulation of another nitrate transporter, NRT2.1. The Plant cell. 2004;16:2433–2447. doi: 10.1105/tpc.104.024380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Ho CH, Lin SH, Hu HC, Tsay YF. CHL1 functions as a nitrate sensor in plants. Cell. 2009;138:1184–1194. doi: 10.1016/j.cell.2009.07.004. [DOI] [PubMed] [Google Scholar]
- 18.Bouguyon E, Gojon A, Nacry P. Nitrate sensing and signaling in plants. Seminars in cell & developmental biology. 2012;23:648–654. doi: 10.1016/j.semcdb.2012.01.004. [DOI] [PubMed] [Google Scholar]
- 19.Abramson J, et al. Structure and mechanism of the lactose permease of Escherichia coli. Science. 2003;301:610–615. doi: 10.1126/science.1088196. [DOI] [PubMed] [Google Scholar]
- 20.Huang Y, Lemieux MJ, Song J, Auer M, Wang DN. Structure and mechanism of the glycerol-3-phosphate transporter from Escherichia coli. Science. 2003;301:616–620. doi: 10.1126/science.1087619. [DOI] [PubMed] [Google Scholar]
- 21.Dang S, et al. Structure of a fucose transporter in an outward-open conformation. Nature. 2010;467:734–738. doi: 10.1038/nature09406. [DOI] [PubMed] [Google Scholar]
- 22.Newstead S, et al. Crystal structure of a prokaryotic homologue of the mammalian oligopeptide-proton symporters, PepT1 and PepT2. The EMBO journal. 2011;30:417–426. doi: 10.1038/emboj.2010.309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Pedersen BP, et al. Crystal structure of a eukaryotic phosphate transporter. Nature. 2013;496:533–536. doi: 10.1038/nature12042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Zheng H, Wisedchaisri G, Gonen T. Crystal structure of a nitrate/nitrite exchanger. Nature. 2013;497:647–651. doi: 10.1038/nature12139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Solcan N, et al. Alternating access mechanism in the POT family of oligopeptide transporters. The EMBO journal. 2012;31:3411–3421. doi: 10.1038/emboj.2012.157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Yan H, et al. Structure and mechanism of a nitrate transporter. Cell reports. 2013;3:716–723. doi: 10.1016/j.celrep.2013.03.007. [DOI] [PubMed] [Google Scholar]
- 27.Sun L, et al. Crystal structure of a bacterial homologue of glucose transporters GLUT1-4. Nature. 2012;490:361–366. doi: 10.1038/nature11524. [DOI] [PubMed] [Google Scholar]
- 28.Yin Y, He X, Szewczyk P, Nguyen T, Chang G. Structure of the multidrug transporter EmrD from Escherichia coli. Science. 2006;312:741–744. doi: 10.1126/science.1125629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Doki S, et al. Structural basis for dynamic mechanism of proton-coupled symport by the peptide transporter POT. Proceedings of the National Academy of Sciences of the United States of America. 2013;110:11343–11348. doi: 10.1073/pnas.1301079110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.DiMaio F, et al. Improved molecular replacement by density- and energy-guided protein structure optimization. Nature. 2011;473:540–543. doi: 10.1038/nature09964. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Slotboom DJ, Duurkens RH, Olieman K, Erkens GB. Static light scattering to characterize membrane proteins in detergent solution. Methods. 2008;46:73–82. doi: 10.1016/j.ymeth.2008.06.012. [DOI] [PubMed] [Google Scholar]
- 32.Zheng J, Trudeau MC, Zagotta WN. Rod cyclic nucleotide-gated channels have a stoichiometry of three CNGA1 subunits and one CNGB1 subunit. Neuron. 2002;36:891–896. doi: 10.1016/s0896-6273(02)01099-1. [DOI] [PubMed] [Google Scholar]
- 33.Taraska JW, Zagotta WN. Fluorescence applications in molecular neurobiology. Neuron. 2010;66:170–189. doi: 10.1016/j.neuron.2010.02.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Veenhoff LM, Heuberger EH, Poolman B. The lactose transport protein is a cooperative dimer with two sugar translocation pathways. The EMBO journal. 2001;20:3056–3062. doi: 10.1093/emboj/20.12.3056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Pessino A, et al. Evidence that functional erythrocyte-type glucose transporters are oligomers. The Journal of biological chemistry. 1991;266:20213–20217. [PubMed] [Google Scholar]
- 36.Safferling M, et al. TetL tetracycline efflux protein from Bacillus subtilis is a dimer in the membrane and in detergent solution. Biochemistry. 2003;42:13969–13976. doi: 10.1021/bi035173q. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Hou Z, Cherian C, Drews J, Wu J, Matherly LH. Identification of the minimal functional unit of the homo-oligomeric human reduced folate carrier. The Journal of biological chemistry. 2010;285:4732–4740. doi: 10.1074/jbc.M109.086033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Otwinowski Z, Minor W. Processing of X-ray diffraction data collected in oscillation mode. Method Enzymol. 1997;276:307–326. doi: 10.1016/S0076-6879(97)76066-X. [DOI] [PubMed] [Google Scholar]
- 39.Adams PD, et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta crystallographica. Section D, Biological crystallography. 2010;66:213–221. doi: 10.1107/S0907444909052925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Emsley P, Lohkamp B, Scott WG, Cowtan K. Features and development of Coot. Acta crystallographica. Section D, Biological crystallography. 2010;66:486–501. doi: 10.1107/S0907444910007493. [DOI] [PMC free article] [PubMed] [Google Scholar]