TargetPrint can improve the reliability of hormone receptor and HER2 testing for those centers with a lower rate of concordance with the reference laboratory, with the limitation of a positive agreement of 75% for HER2. TargetPrint consequently has important implications for treatment decisions in clinical practice and is a reliable alternative to local assessment for ER.
Keywords: breast cancer, concordance, FISH, hormone receptor, IHC, TargetPrint
Abstract
Background
To investigate the correlation of TargetPrint with local and central immunohistochemistry/fluorescence in situ hybridization assessment of estrogen (ER), progesterone (PgR), and human epidermal growth factor receptor 2 (HER2) in the first 800 patients enrolled in the MINDACT trial.
Patients and methods
Data from local (N = 800) and central (N = 626) assessments of receptor status were collected and compared with TargetPrint results.
Results
For ER, the positive agreement (the percentage of central pathology positive assessments that were also TargetPrint/local laboratory positive) for TargetPrint in comparison to centralized assessment was 98% with a negative agreement (the percentage of central pathology negative assessments that were also TargetPrint/local laboratory negative) of 96%. For PgR, the positive agreement was 83% with a negative agreement of 92%. For HER2, the positive agreement was 75% with a negative agreement of 99%. Even though the local assessment showed higher positive agreement for PgR (89%) and higher positive agreement for HER2 (85%), the range of discordant local versus central assessments were as high as 6.7% for ER, 12.9% for PgR, and 4.3% for HER2.
Conclusion
TargetPrint and local assessment of ER, PgR, and HER2 show high concordance with central assessment in the first 800 MINDACT patients. However, there are concerns about the higher discordance rates for some local sites. TargetPrint can improve the reliability of hormone receptor and HER2 testing for those centers with a lower rate of concordance with the reference laboratory, with the limitation of a positive agreement of 75% for HER2. TargetPrint consequently has important implications for treatment decisions in clinical practice and is a reliable alternative to local assessment for ER.
Clinical Trials number
introduction
This study was undertaken to determine the correlation of mRNA readout of estrogen receptor (ER), progesterone receptor (PgR), and human epidermal growth factor receptor 2 (HER2) by TargetPrint® (commercially available microarray-based test) with immunohistochemistry (IHC)/fluorescence in situ hybridization (FISH) assessments determined locally and centrally in the first 800 patients enrolled in the MINDACT trial [1]. Measurement of ER, PgR, and HER2 status in early-stage breast cancer is critical for informing treatment recommendations [2]. Controversy remains about the optimal stratification of patients with indeterminate risk and considerable differences exist among physicians regarding the selection of patients for adjuvant chemotherapy. Accordingly, several guidelines have been issued, e.g. the St Gallen [3], ESMO [4], and NCCN [5].
MINDACT is an international, prospective, randomized, phase III trial investigating the clinical utility of MammaPrint® versus standard clinicopathological criteria (Adjuvant! Online) to select patients with breast cancer for adjuvant chemotherapy and has enrolled 6694 patients. The trial's hypothesis is that the molecular assay will outperform the established clinicopathological assessment, reducing the number of patients receiving adjuvant chemotherapy by 10%–15% without impairing long-term outcomes [6, 7]. A pilot phase comprising the first 800 enrolled patients was predefined to ensure the trial's feasibility [1].
We present the results of the preplanned translational research project on the patients from this pilot phase. The aim of this analysis was to investigate the correlation of microarray mRNA readouts with local and central IHC/FISH assessments of ER, PgR, and HER2 status. Interlaboratory variability in ER, PgR, and HER2 assessments remains a major concern worldwide [8] and a more reliable assessment is highly desirable.
patients and methods
Women aged ≥18 years with histologically proven operable invasive breast cancer, 0–3 positive lymph nodes, and a frozen tumor sample containing ≥30% tumor cells were eligible to enroll in MINDACT from February 2007 to July 2011 (closed to accrual). Further eligibility criteria included tumor stage T1–2, or operable T3 and treatment with breast conserving surgery or mastectomy combined with a sentinel node procedure or full axillary clearance. A WHO performance status of 0 or 1 and adequate bone marrow, liver, and renal function were required. Main exclusion criteria included previous or concurrent cancer, previous chemotherapy, endocrine therapy, or radiotherapy, and clinically significant cardiac disease. The protocol was approved by independent ethics committees and medical authorities. All patients provided written informed consent. The study was conducted in accordance with the Declaration of Helsinki and good clinical practice guidelines.
Data from local (N = 800) and central (N = 626; European Institute of Oncology, Milan, Italy) assessments of ER, PgR, and HER2 status were collected and compared with microarray readout (TargetPrint) provided by Agendia (Amsterdam, The Netherlands). Baseline characteristics are shown in Table 1.
Table 1.
Baseline characteristics of the first 800 patients enrolled in the MINDACT trial (adapted from Rutgers et al. [1], with permission, with ER, PgR, and HER2 status updated at the time of analysis)
| Characteristic | Number of patients (%) | ||
|---|---|---|---|
| Age, years | |||
| <50 | 264 (33.0) | ||
| ≥50 | 536 (67.0) | ||
| Tumor size, cm | |||
| ≤2 | 601 (75.1) | ||
| >2–5 | 195 (24.4) | ||
| >5 | 4 (0.5) | ||
| Lymph node status | |||
| Negative | 794 (99.3) | ||
| 1–3 node positive | 4 (0.5) | ||
| Positive before amendment | 1 (0.1) | ||
| Missing | 1 (0.1) | ||
| Histological grade | |||
| 1 | 169 (21.1) | ||
| 2 | 361 (45.1) | ||
| 3 | 266 (33.3) | ||
| Undefined | 4 (0.5) | ||
| ER status (local) | |||
| Positive | 674 (84.3) | ||
| Negative | 126 (15.8) | ||
| PgR status (local) | |||
| Positive | 570 (71.3) | ||
| Negative | 228 (28.5) | ||
| Unknown | 2 (0.3) | ||
| HER2 status (local) | |||
| Negative | 680 (85.0) | ||
| Positive | 92 (11.5) | ||
| Unknown | 28 (3.5) | ||
| ER + PgR status (local) | |||
| ER positive – PgR positive | 562 (70.3) | ||
| ER positive – PgR negative | 110 (13.8) | ||
| ER negative – PgR positive | 8 (1.0) | ||
| ER negative – PgR negative | 118 (14.8) | ||
| Unknown | 2 (0.3) | ||
| Data availability | Central IHC/FISH | Local IHC/FISH | TargetPrint |
| ER | N = 619 | N = 800 | N = 800 |
| PgR | N = 619 | N = 798 | N = 800 |
| HER2 | N = 608 | N = 772 | N = 800 |
ER, estrogen receptor; PgR, progesterone receptor; HER2, human epidermal growth factor receptor 2; IHC, immunohistochemistry; FISH, fluorescence in situ hybridization.
data availability
At the time of analysis, central pathology results were unavailable for 174 patients due to delayed submission of tumor samples. Among the 626 patients with central pathology results, 15 had incomplete data (7 for ER and PgR, 15 for HER2) and 3 equivocal HER2 IHC and FISH. For local pathological laboratory assessments, two patients had a missing PgR assessment and 28 had unknown HER2 status. TargetPrint readout was available for all 800 patients (Table 1).
immunohistochemistry and FISH
In the central laboratory, ER and PgR status were assessed on formalin-fixed paraffin-embedded tissue blocks by IHC using the ER/PgR PharmDX kit (Dako, Glostrup, Denmark). Tumors were classified as ER- or PgR-positive when ≥1% invasive tumor cells showed definite nuclear staining, irrespective of staining intensity [2]. HER2 expression was evaluated with the HercepTest kit (Dako) and scored as 0, 1+, 2+, or 3+, according to the FDA scoring system. Tumors scored as 2+ or 3+ were re-tested with FISH using the PathVysion HER2 DNA probe kit (Vysis-Abbott, Chicago, IL). Cases were considered HER2-positive if scored 3+ by IHC and/or amplified by FISH (ratio ≥2).
Details of the assays and protocols for ER, PgR, and HER2 status assessment used in local centers were not available, but according to the MINDACT protocol, ER/PgR-positive disease was defined as a tumor with ≥1% immunoreactive cells, an Allred score >2, or a biochemical protein concentration ≥10 fmol/mg; HER2 status was determined according to local policies.
microarray readout by targetprint
Gene-expression data for ER, PgR, and HER2 (blinded) were obtained by TargetPrint on frozen samples; tumors were considered as receptor-positive or -negative using previously determined and validated thresholds [9]. RNA isolation, labeling, and hybridization were carried out as described previously [10]. Fluorescence intensities on scanned images were quantified and normalized using Feature Extraction software (Agilent Technologies, Santa Clara, CA).
statistical analysis
Statistical calculations were conducted using SAS® 9.2 (SAS Institute, Inc., Cary, NC). Statistics summarizing the agreement between TargetPrint and local pathology versus central pathology included: positive agreement (percentage of central pathology positives that were TargetPrint/local laboratory positive) and negative agreement [11]; positive (PPV) and negative (NPV) predictive value (PPV: percentage of TargetPrint/local positives that are central pathology positive); percentage of concordance; and Cohen's κ coefficient [12]. To test for a difference between centers in the amount of discordance, a contingency table was constructed with the 51 centers in rows and the number of concordances and discordances in columns (data not shown). A two-sided Fisher's exact test was carried out (5% significance level). Patient data were omitted if one of the two assessments were missing in the comparison.
results
Table 2 cross tabulates local assessments and TargetPrint results versus central assessments and the TargetPrint results versus local assessments. (No statistical trend was seen for local lab-TargetPrint concordance versus percentage missing central pathology assessments, data not shown.)
Table 2.
Assessment of ER, PgR, and HER2 status by local versus central pathology (immunohistochemistry/fluorescence in situ hybridization) and TargetPrint versus central and local pathology
| ER (central pathology) | |||||
| Positive (N = 537), n (%) | Negative (N = 82), n (%) | Missing (N = 181), n (%) | Total (N = 800), n (%) | ||
| ER (local site) | |||||
| Positive | 524 (97.6) | 2 (2.4) | 148 (81.8) | 674 (84.3) | |
| Negative | 13 (2.4) | 80 (97.6) | 33 (18.2) | 126 (15.8) | |
| PgR (central pathology) | |||||
| Positive (N = 490), n (%) | Negative (N = 129), n (%) | Missing (N = 181), n (%) | Total (N = 800), n (%) | ||
| PgR (local site) | |||||
| Positive | 438 (89.4) | 12 (9.3) | 120 (66.3) | 570 (71.3) | |
| Negative | 52 (10.6) | 116 (89.9) | 60 (33.1) | 228 (28.5) | |
| Missing | 0 (0.0) | 1 (0.8) | 1 (0.6) | 2 (0.3) | |
| HER2 (central pathology) | |||||
| Negative (N = 537), n (%) | Positive (N = 71), n (%) | IHC2 + FISH equivocal (N = 3), n (%) | Missing (N = 189), n (%) | Total (N = 800), n (%) | |
| HER2 (local site) | |||||
| Negative | 506 (94.2) | 10 (14.1) | 3 (100.0) | 161 (85.2) | 680 (85.0) |
| Positive | 12 (2.2) | 57 (80.3) | 0 (0.0) | 23 (12.2) | 92 (11.5) |
| Missing | 19 (3.5) | 4 (5.6) | 0 | 5 (2.6) | 28 (3.5) |
| ER (central pathology) | |||||
| Positive (N = 537), n (%) | Negative (N = 82), n (%) | Missing (N = 181), n (%) | Total (N = 800), n (%) | ||
| ER (TargetPrint) | |||||
| Positive | 525 (97.8) | 3 (3.7) | 144 (79.6) | 672 (84.0) | |
| Negative | 12 (2.2) | 79 (96.3) | 37 (20.4) | 128 (16.0) | |
| PgR (central pathology) | |||||
| Positive (N = 490), n (%) | Negative (N = 129), n (%) | Missing (N = 181), n (%) | Total (N = 800), n (%) | ||
| PgR (TargetPrint) | |||||
| Positive | 408 (83.3) | 11 (8.5) | 104 (57.5) | 523 (65.4) | |
| Negative | 82 (16.7) | 118 (91.5) | 77 (42.5) | 277 (34.6) | |
| HER2 (central pathology) | |||||
| Negative (N = 537), n (%) | Positive (N = 71), n (%) | IHC2 + FISH equivocal (N = 3), n (%) | Missing (N = 189), n (%) | Total (N = 800), n (%) | |
| HER2 (TargetPrint) | |||||
| Negative | 532 (99.1) | 18 (25.4) | 3 (100.0) | 168 (88.9) | 721 (90.1) |
| Positive | 5 (0.9) | 53 (74.6) | 0 (0.0) | 21 (11.1) | 79 (9.9) |
| ER (local site) | |||||
| Positive (N = 674), n (%) | Negative (N = 126), n (%) | Total (N = 800), n (%) | |||
| ER (TargetPrint) | |||||
| Positive | 661 (98.1) | 11 (8.7) | 672 (84.0) | ||
| Negative | 13 (1.9) | 115 (91.3) | 128 (16.0) | ||
| PgR (local site) | |||||
| Positive (N = 570), n (%) | Negative (N = 228), n (%) | Missing (N = 2), n (%) | Total (N = 800), n (%) | ||
| PgR (TargetPrint) | |||||
| Positive | 490 (86.0) | 33 (14.5) | 0 (0.0) | 523 (65.4) | |
| Negative | 80 (14.0) | 195 (85.5) | 2 (100.0) | 277 (34.6) | |
| HER2 (local site) | |||||
| Negative (N = 680), n (%) | Positive (N = 92), n (%) | Missing (N = 28), n (%) | Total (N = 800), n (%) | ||
| HER2 (TargetPrint) | |||||
| Negative | 677 (99.6) | 20 (21.7) | 24 (85.7) | 721 (90.1) | |
| Positive | 3 (0.4) | 72 (78.3) | 4 (14.3) | 79 (9.9) | |
ER, estrogen receptor; PgR, progesterone receptor; HER2, human epidermal growth factor receptor.
positive and negative agreement, PPV and NPV
Of the 537 central ER-positive cases, 525 (98%) were TargetPrint ER-positive, well above the set target of 90%. For PgR, the positive agreement was 83%, which is below the desirable 90% and also below the positive agreement for local assessment (89%). For the negative agreement, the target of 95% was attained for ER (96%); while for PgR, this was slightly lower (92%). For PgR, the negative agreement was also <95% for local assessment (91%). For HER2, the positive agreement for TargetPrint (75%) was quite low versus that for local assessment (85%). For PPV and NPV, there were strong similarities in performance for TargetPrint and local assessment of ER compared with central pathology. For PgR, local IHC assessment was more in line with central assessment compared with the TargetPrint results. The PPV for HER2 status by TargetPrint was higher than that for local assessment (Table 3).
Table 3.
Positive and negative agreement and predictive value for local and TargetPrint assessment of ER, PgR, and HER2 versus central pathology
| Positive and negative agreement, % (n/N) | ||
| Local versus central pathology | ||
| Positive agreement | Negative agreement | |
| ER | 97.6 (524/537) | 97.6 (80/82) |
| PgR | 89.4 (438/490) | 90.6 (116/128) |
| HER2 | 85.1 (57/67) | 97.7 (506/518) |
| TargetPrint versus central pathology | ||
| Positive agreement | Negative agreement | |
| ER | 97.8 (525/537) | 96.3 (79/82) |
| PgR | 83.3 (408/490) | 91.5 (118/129) |
| HER2 | 74.7 (53/71) | 99.1 (532/537) |
| Positive and negative predictive value | ||
| Local versus central pathology | ||
| PPV | NPV | |
| ER | 99.6 (524/526) | 86.0 (80/93) |
| PgR | 97.3 (438/450) | 69.0 (116/168) |
| HER2 | 82.6 (57/69) | 98.1 (506/516) |
| TargetPrint versus central pathology | ||
| PPV | NPV | |
| ER | 99.4 (525/528) | 86.8 (79/91) |
| PgR | 97.4 (408/419) | 59.0 (118/200) |
| HER2 | 91.4 (53/58) | 96.7 (532/550) |
ER, estrogen receptor; PgR, progesterone receptor; HER2, human epidermal growth factor receptor; NPV, negative predictive value; PPV, positive predictive value.
Data for ER and PgR using 10% invasive tumor cells as cutoff are provided in supplementary Table S1, available at Annals of Oncology online.
overall agreement
The percentage of concordant assessments and κ coefficients were used to summarize the overall agreement. Comparison of local with central assessments (Table 4) indicated highly similar results for ER (98% concordance; κ = 0.90), good concordance for HER2 (96%; κ = 0.82), and somewhat lower concordance for PgR (90%; κ = 0.72). Comparison of central assessment with TargetPrint (Table 4) indicated a highly similar overall performance, with a concordance of 98% (κ = 0.90) for ER, 96% for HER2 (κ = 0.80), and lower concordance for PgR (85%; κ = 0.62).
Table 4.
Inter-rater agreement statistics for ER, PgR, and HER2 assessments: local versus central pathology and TargetPrint versus central and local pathology
| % Concordance | 95% CI | Cohen's κ coefficient | 95% CI | N | |
|---|---|---|---|---|---|
| Local versus central pathology | |||||
| ER | 97.6 | 96.4–98.8 | 0.900 | 0.851–0.950 | 619 |
| PgR | 89.6 | 87.2–92.1 | 0.717 | 0.653–0.781 | 618 |
| HER2 | 96.2 | 94.7–97.8 | 0.817 | 0.743–0.891 | 585 |
| TargetPrint versus central pathology | |||||
| ER | 97.6 | 96.4–98.8 | 0.899 | 0.849–0.949 | 619 |
| PgR | 85.0 | 82.2–87.8 | 0.621 | 0.554–0.689 | 619 |
| HER2 | 96.2 | 94.7–97.7 | 0.801 | 0.722–0.879 | 608 |
| TargetPrint versus local pathology | |||||
| ER | 97.0 | 95.6–98.1 | 0.888 | 0.844–0.932 | 800 |
| PgR | 85.8 | 83.2–88.2 | 0.673 | 0.618–0.728 | 798 |
| HER2 | 97.0 | 95.6–98.1 | 0.846 | 0.784–0.907 | 772 |
CI, confidence interval; ER, estrogen receptor; PgR, progesterone receptor; HER2, human epidermal growth factor receptor.
range of discordance for local assessment
The percentage of discordant assessments was calculated for each site separately. Only those centers with ≥30 assessments for that receptor were taken into account. The range of discordance was 1.6%–6.7% for ER (eight centers), 5.7%–12.9% for PgR (eight centers), and 0%–4.3% for HER2 (six centers). Fisher's exact test (on all centers) was carried out to see whether there was a significant difference in the level of discordance with central pathology between centers (ER, P = 0.4; PgR, P = 0.03; HER2, P = 0.36). This was the case for PgR. A Cochran–Armitage trend test indicated no trend in discordance versus sample size for ER and HER2, only for PgR (data not shown). However, local centers can work with external and several laboratories. The latter can result in additional heterogeneity within the same local center.
three-way comparison
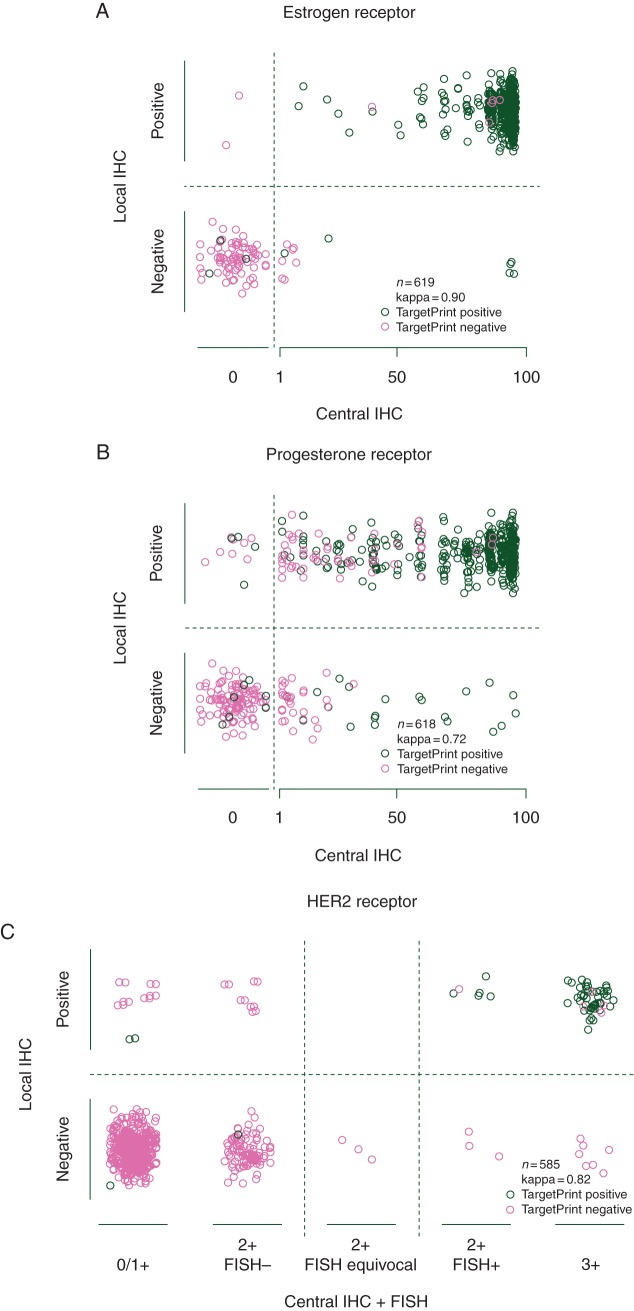
The scatterplots in Figures 1 and 2 show a three-way comparison of the different assessments. Central IHC assessments for ER and PgR were available as integer percentages and for HER2 as five categories. For local IHC, there is only a positive or negative assessment and the TargetPrint results are gene-expression scores on a continuous scale. To improve the visibility of individual observations, random trimmed noise was added to the pathology laboratory assessments.
Figure 1.
Three-way comparison of local pathology assessment (positive/negative) versus central pathology assessment as integer percentage for ER (A) and PgR (B) and as five categories for HER2 (C). The corresponding TargetPrint assessment is indicated by the color of the dots.
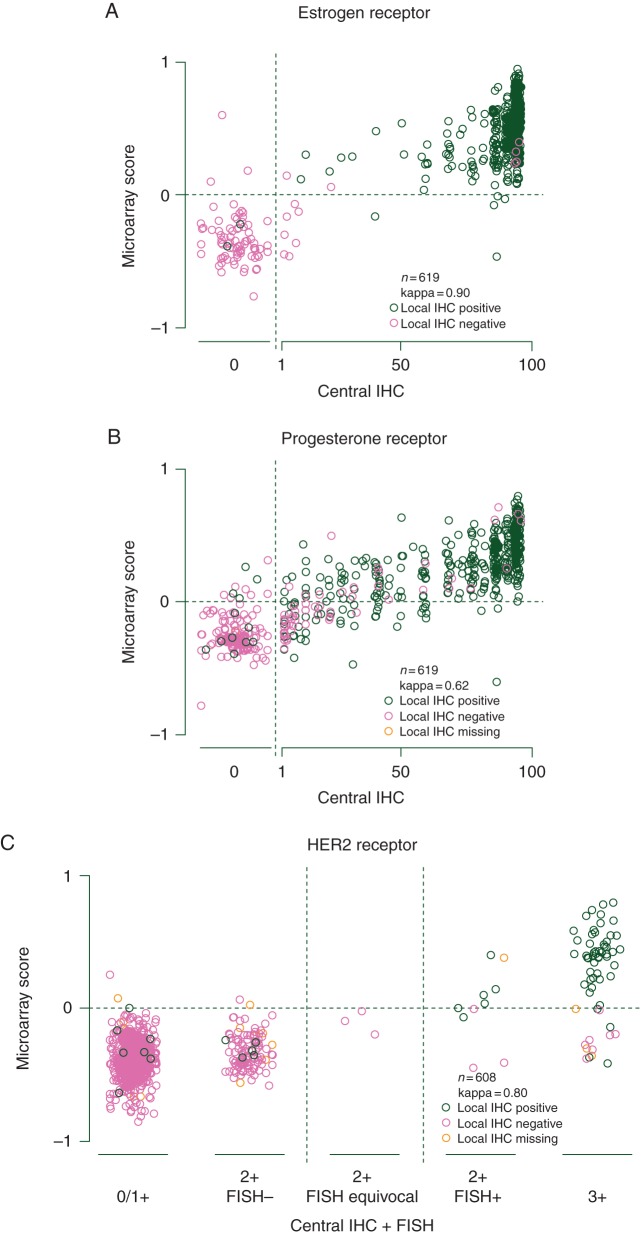
Figure 2.
Three-way comparison of gene expression scores on a continuous scale for TargetPrint versus central pathology assessment as integer percentage for ER (A) and PgR (B) and as five categories for HER2 (C). The corresponding local assessment is indicated by the color of the dots.
Figure 1 shows the local assessment of ER, PgR, and HER2 versus the percentage of immunoreactive cells registered by the central laboratory. Figure 2 shows the plots for the TargetPrint results versus central pathology assessment. All the plots illustrate the similarity between the TargetPrint and local laboratory results, as assessed by receptor status determined by central pathology. ER- and PgR-positive cases with a low percentage of immunoreactive cells had the most discordances by both TargetPrint and local assessment.
discussion
Locally and centrally assessed ER, PgR, and HER2 status in the first 800 (626 centrally assessed) MINDACT patient samples indicates a high level of quality for pathology in the local hospitals. Despite the high concordance, there are still concerns about the range of discordance and the false-negative rate of locally assessed ER (13 of 537, 2.4%) and PgR (52 of 490, 10.6%), and the false-positive rate (12 of 69, 17%) for HER2 status. These rates are similar to those observed in other clinical trials (e.g. HERA, ALTTO) and emphasize the need for a continuous effort to harmonize analytical performance and interpretative skills in the assessment of these important biological variables. Importantly, most centers participating in MINDACT had a good-quality pathology department able to comply with the complex study requirements [13]. This may in part explain the substantially high concordance rate between local and central assessments, which is usually lower for less well-performing local laboratories.
TargetPrint assessment of ER and HER2 (and to a lesser extent PgR) status gives results comparable with IHC/FISH and provides an objective and quantitative assessment of tumor-receptor status. Microarray readouts of ER, PgR, and HER2 by TargetPrint were previously shown to be strongly correlated with high-quality IHC/FISH assessment, with concordance rates of 93% (κ = 0.79) for ER, 83% (κ = 0.65) for PgR, and 96% for HER2 (κ = 0.88) [9]. The data indicate that TargetPrint can serve as a reliable alternative to local IHC/FISH.
The positive and negative agreement (>95%) for ER in the current study indicate that TargetPrint is a very stable and reliable assay for this receptor [2]. For the small percentage of discordant samples, a prospective comparison of the two methods will ultimately establish whether mRNA readout for ER is the preferred technique, as suggested by a retrospective analysis [14]. For the majority of the discordant cases, the percentage of immunoreactive cells was quite low, potentially indicating remarkable intratumoral heterogeneity of ER expression. The evaluation of different tumor areas may well be the cause of discordant results in heterogeneous tumors.
PgR concordance was 85%, indicating discordance for ∼15% of cases. Compared with ER, the distribution of PgR IHC-positive and discordant cases is somewhat more homogeneously spread across different percentages of immunoreactive cells. Similar to ER, discordance was more likely for low percentages of immunoreactive cells. The concordance for mRNA and IHC assessment of PgR has been shown to be less than that for ER, but mRNA-derived receptor status is more strongly associated with clinical outcome, suggesting that mRNA may be a more reliable method for assessing receptor status [15, 16].
The positive agreement for TargetPrint with IHC/FISH for HER2 is comparable with other mRNA readouts [17, 18], indicating there are differences between the two methods. Intratumor heterogeneity of HER2 status may be one reason for the discordant results, but further research is warranted to determine the suggested appropriateness of mRNA readout for HER2 as alternate approach [19].
In summary, our work has two important implications: (i) the results of the MINDACT trial will not be affected by the fact that risk assessment by clinicopathological factors used local pathology and risk assessment by MammaPrint® was carried out centrally; (ii) TargetPrint can improve the reliability of hormone receptor and HER2 testing for those centers with a lower concordance rate with the reference laboratory, with the limitation of a positive agreement of 75% for HER2. TargetPrint consequently has important implications for treatment decisions in routine clinical practice and is a reliable alternative to local assessment for ER.
funding
This trial has funding grants from the European Commission Framework Programme VI (FP6-LSHC-CT-2004-503426), the Breast Cancer Research Foundation (BCRF 2007-2008), Novartis, F. Hoffman La Roche, Sanofi-Aventis, the National Cancer Institute (NCI), the EBCC-Breast Cancer Working Group (BCWG grant for the MINDACT biobank), the Jacqueline Seroussi Memorial Foundation (2006 JSMF award), Prix Mois du Cancer du Sein (2004 award), Susan G. Komen for the Cure (SG05-0922-02), Fondation Belge Contre le Cancer (SCIE 2005-27), Dutch Cancer Society (KWF), Association Le Cancer du Sein, Parlons-en!, Deutsche Krebshilfe, and the Grant Simpson Trust and Cancer Research UK. This trial was supported by the EORTC Charitable Trust. Whole genome analysis was provided in kind by Agendia.
disclosure
FdS, LS-S, JvdA, and AG are employees of Agendia. LvV is a founder of Agendia and has stock ownership. All other authors have declared no conflicts of interest.
Supplementary Material
acknowledgements
We are grateful to all women participating in this study, all the investigators, surgeons, pathologists, and research nurses, the National Coordinating Centers/BIG Groups (BOOG, CaCTUS, CEEOG, FNCLCC, GOIRC, IBCSG, SAKK, SOLTI, WSG), and World Courier.
references
- 1.Rutgers E, Piccart-Gebhart MJ, Bogaerts J, et al. The EORTC 10041/BIG 03-04 MINDACT trial is feasible: results of the pilot phase. Eur J Cancer. 2011;00:2742–2749. doi: 10.1016/j.ejca.2011.09.016. [DOI] [PubMed] [Google Scholar]
- 2.Hammond ME, Hayes DF, Dowsett M, et al. American Society of Clinical Oncology/College of American Pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. J Clin Oncol. 2010;28:2784–2795. doi: 10.1200/JCO.2009.25.6529. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Goldhirsch A, Wood WC, Coates AS, et al. Strategies for subtypes—dealing with the diversity of breast cancer: highlights of the St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2011. Ann Oncol. 2011;22:1736–1747. doi: 10.1093/annonc/mdr304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Aebi S, Davidson T, Gruber G, et al. Primary breast cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann Oncol. 2011;22(Suppl 6):vi12–vi24. doi: 10.1093/annonc/mdr371. [DOI] [PubMed] [Google Scholar]
- 5.National Comprehensive Cancer Network: NCCN Clinical Practice Guidelines in Oncology. 2013. Breast Cancer. Version 2.2013 http://www.nccn.org/professionals/physician_gls/pdf/breast.pdf. (23 December 2013, date last accessed)
- 6.Cardoso F, Saghatchian M, Thompson A, et al. Inconsistent criteria used in American Society of Clinical Oncology 2007 update of recommendations for the use of tumor markers in breast cancer. J Clin Oncol. 2008;26:2058–2059. doi: 10.1200/JCO.2007.15.6638. author reply 2060–2061. [DOI] [PubMed] [Google Scholar]
- 7.Bogaerts J, Cardoso F, Buyse M, et al. Gene signature evaluation as a prognostic tool: challenges in the design of the MINDACT trial. Nat Clin Pract Oncol. 2006;3:540–551. doi: 10.1038/ncponc0591. [DOI] [PubMed] [Google Scholar]
- 8.Rhodes A, Jasani B, Barnes DM, et al. Reliability of immunohistochemical demonstration of oestrogen receptors in routine practice: interlaboratory variance in the sensitivity of detection and evaluation of scoring systems. J Clin Pathol. 2000;53:125–130. doi: 10.1136/jcp.53.2.125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Roepman P, Horlings HM, Krijgsman O, et al. Microarray-based determination of estrogen receptor, progesterone receptor, and HER2 receptor status in breast cancer. Clin Cancer Res. 2009;15:7003–7011. doi: 10.1158/1078-0432.CCR-09-0449. [DOI] [PubMed] [Google Scholar]
- 10.Glas AM, Floore A, Delahaye LJ, et al. Converting a breast cancer microarray signature into a high-throughput diagnostic test. BMC Genomics. 2006;7:278. doi: 10.1186/1471-2164-7-278. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. US Food and Drug Administration: Guidance for industry and FDA staff: Statistical guidance on reporting results from studies evaluating diagnostic tests http://medical.cms.itri.org.tw/pdf/u14.pdf. (23 December 2013, date last accessed)
- 12.Cohen J. A coefficient of agreement for nominal scales. Educ Psychol Meas. 1960;20:37–46. [Google Scholar]
- 13.Cardoso F, Van't Veer L, Rutgers E, et al. Clinical application of the 70-gene profile: the MINDACT trial. J Clin Oncol. 2008;26:729–735. doi: 10.1200/JCO.2007.14.3222. [DOI] [PubMed] [Google Scholar]
- 14.Bordeaux JM, Cheng H, Welsh AW, et al. Quantitative in situ measurement of estrogen receptor mRNA predicts response to tamoxifen. PLoS One. 2012;7:e36559. doi: 10.1371/journal.pone.0036559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Nielsen TO, Parker JS, Leung S, et al. A comparison of PAM50 intrinsic subtyping with immunohistochemistry and clinical prognostic factors in tamoxifen-treated estrogen receptor-positive breast cancer. Clin Cancer Res. 2010;16:5222–5232. doi: 10.1158/1078-0432.CCR-10-1282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Ma XJ, Hilsenbeck SG, Wang W, et al. The HOXB13:IL17BR expression index is a prognostic factor in early-stage breast cancer. J Clin Oncol. 2006;24:4611–4619. doi: 10.1200/JCO.2006.06.6944. [DOI] [PubMed] [Google Scholar]
- 17.Baehner FL, Achacoso N, Maddala T, et al. Human epidermal growth factor receptor 2 assessment in a case-control study: comparison of fluorescence in situ hybridization and quantitative reverse transcription polymerase chain reaction performed by central laboratories. J Clin Oncol. 2010;28:4300–4306. doi: 10.1200/JCO.2009.24.8211. [DOI] [PubMed] [Google Scholar]
- 18.Dabbs DJ, Klein ME, Mohsin SK, et al. High false-negative rate of HER2 quantitative reverse transcription polymerase chain reaction of the Oncotype DX test: an independent quality assurance study. J Clin Oncol. 2011;29:4279–4285. doi: 10.1200/JCO.2011.34.7963. [DOI] [PubMed] [Google Scholar]
- 19.Bergqvist J, Ohd JF, Smeds J, et al. Quantitative real-time PCR analysis and microarray-based RNA expression of HER2 in relation to outcome. Ann Oncol. 2007;18:845–850. doi: 10.1093/annonc/mdm059. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
At the time of analysis, central pathology results were unavailable for 174 patients due to delayed submission of tumor samples. Among the 626 patients with central pathology results, 15 had incomplete data (7 for ER and PgR, 15 for HER2) and 3 equivocal HER2 IHC and FISH. For local pathological laboratory assessments, two patients had a missing PgR assessment and 28 had unknown HER2 status. TargetPrint readout was available for all 800 patients (Table 1).