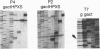Abstract
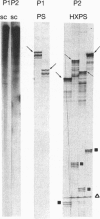
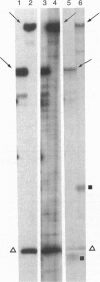
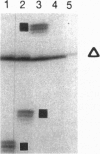
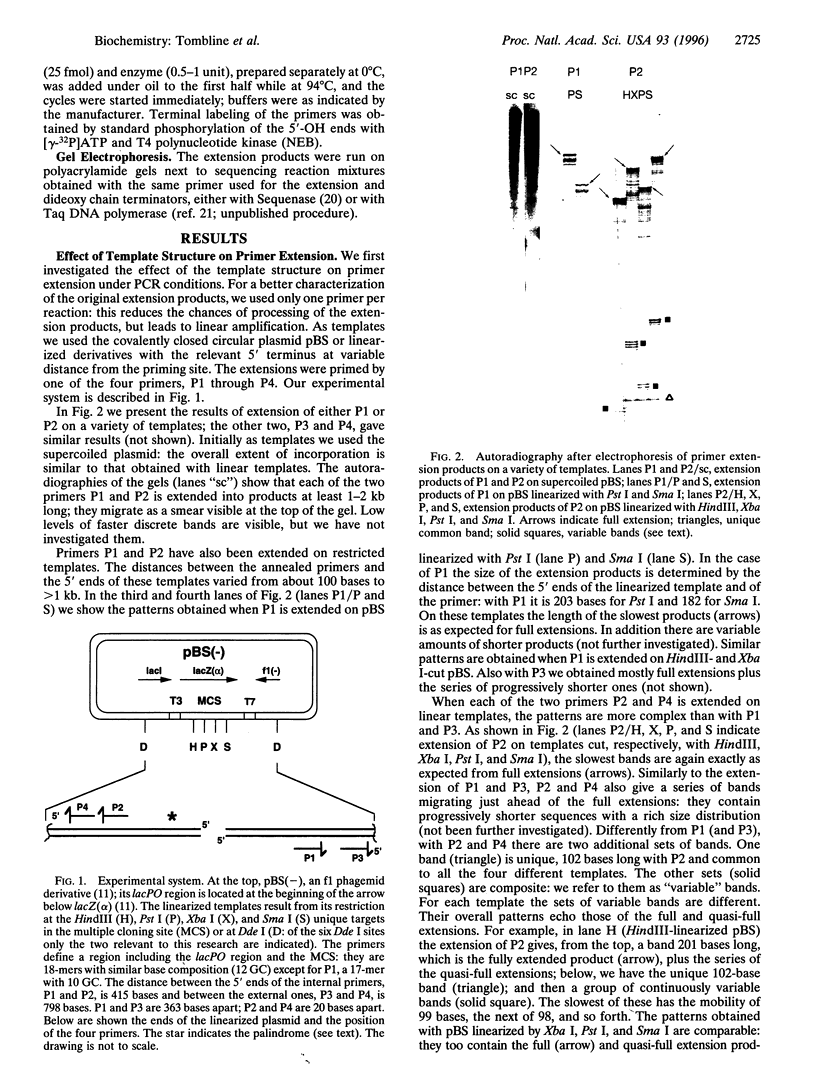
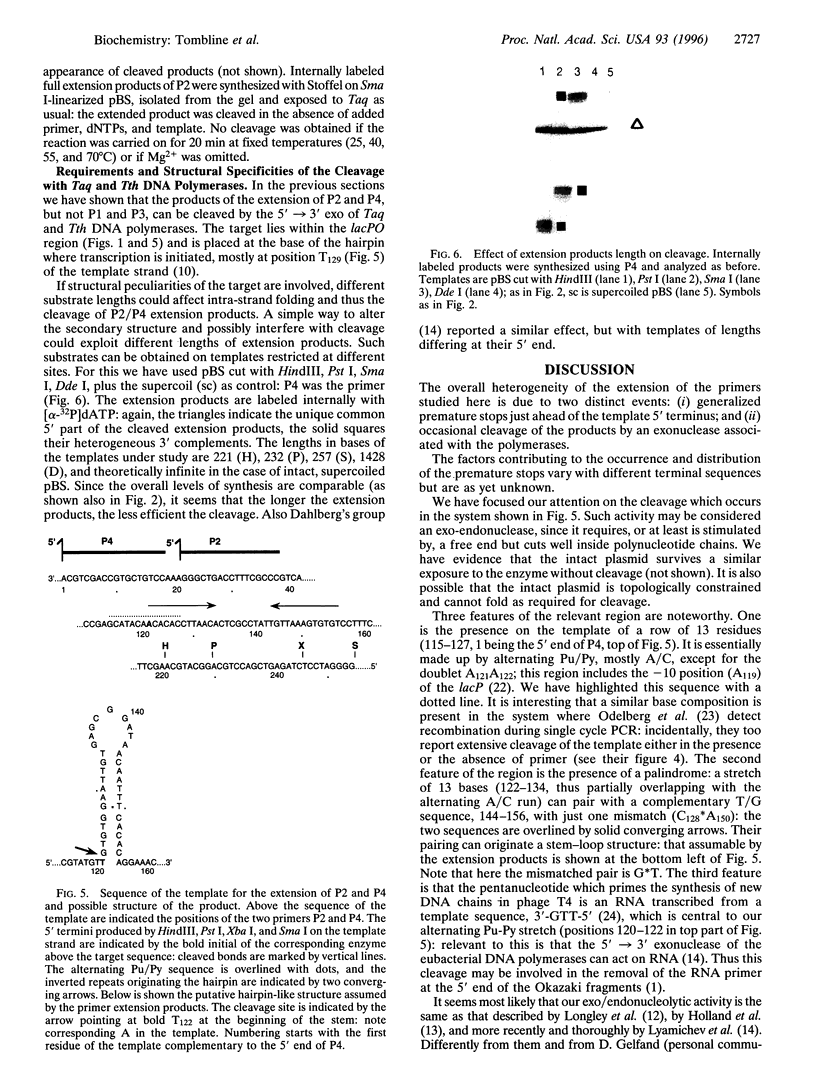
In PCR, DNA polymerases from thermophilic bacteria catalyze the extension of primers annealed to templates as well as the structure-specific cleavage of the products of primer extension. Here we show that cleavage by Thermus aquaticus and Thermus thermophilus DNA polymerases can be precise and substantial: it occurs at the base of the stem-loop structure assumed by the single strand products of primer extension using as template a common genetic element, the promoter-operator of the Escherichia coli lactose operon, and may involve up to 30% of the products. The cleavage is independent of primer, template, and triphosphates, is dependent on substrate length and temperature, requires free ends and Mg2+, and is absent in DNA polymerases lacking the 5'-->3' exonuclease, such as the Stoffel fragment and the T7 DNA polymerase. Heterogeneity of the extension products results also from premature detachment of the enzyme approaching the 5' end of the template.
Full text
PDF




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Barany F. The ligase chain reaction in a PCR world. PCR Methods Appl. 1991 Aug;1(1):5–16. doi: 10.1101/gr.1.1.5. [DOI] [PubMed] [Google Scholar]
- Bardwell A. J., Bardwell L., Tomkinson A. E., Friedberg E. C. Specific cleavage of model recombination and repair intermediates by the yeast Rad1-Rad10 DNA endonuclease. Science. 1994 Sep 30;265(5181):2082–2085. doi: 10.1126/science.8091230. [DOI] [PubMed] [Google Scholar]
- Barnes W. M. PCR amplification of up to 35-kb DNA with high fidelity and high yield from lambda bacteriophage templates. Proc Natl Acad Sci U S A. 1994 Mar 15;91(6):2216–2220. doi: 10.1073/pnas.91.6.2216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Birkenkamp K., Kemper B. In vitro processing of heteroduplex loops and mismatches by endonuclease VII. DNA Res. 1995;2(1):9–14. doi: 10.1093/dnares/2.1.9. [DOI] [PubMed] [Google Scholar]
- Cha T. A., Alberts B. M. Studies of the DNA helicase-RNA primase unit from bacteriophage T4. A trinucleotide sequence on the DNA template starts RNA primer synthesis. J Biol Chem. 1986 May 25;261(15):7001–7010. [PubMed] [Google Scholar]
- Gyllensten U. B., Erlich H. A. Generation of single-stranded DNA by the polymerase chain reaction and its application to direct sequencing of the HLA-DQA locus. Proc Natl Acad Sci U S A. 1988 Oct;85(20):7652–7656. doi: 10.1073/pnas.85.20.7652. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harrington J. J., Lieber M. R. The characterization of a mammalian DNA structure-specific endonuclease. EMBO J. 1994 Mar 1;13(5):1235–1246. doi: 10.1002/j.1460-2075.1994.tb06373.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holland P. M., Abramson R. D., Watson R., Gelfand D. H. Detection of specific polymerase chain reaction product by utilizing the 5'----3' exonuclease activity of Thermus aquaticus DNA polymerase. Proc Natl Acad Sci U S A. 1991 Aug 15;88(16):7276–7280. doi: 10.1073/pnas.88.16.7276. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jeffreys A. J., Wilson V., Neumann R., Keyte J. Amplification of human minisatellites by the polymerase chain reaction: towards DNA fingerprinting of single cells. Nucleic Acids Res. 1988 Dec 9;16(23):10953–10971. doi: 10.1093/nar/16.23.10953. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lawyer F. C., Stoffel S., Saiki R. K., Chang S. Y., Landre P. A., Abramson R. D., Gelfand D. H. High-level expression, purification, and enzymatic characterization of full-length Thermus aquaticus DNA polymerase and a truncated form deficient in 5' to 3' exonuclease activity. PCR Methods Appl. 1993 May;2(4):275–287. doi: 10.1101/gr.2.4.275. [DOI] [PubMed] [Google Scholar]
- Lewis S. M. P nucleotide insertions and the resolution of hairpin DNA structures in mammalian cells. Proc Natl Acad Sci U S A. 1994 Feb 15;91(4):1332–1336. doi: 10.1073/pnas.91.4.1332. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Longley M. J., Bennett S. E., Mosbaugh D. W. Characterization of the 5' to 3' exonuclease associated with Thermus aquaticus DNA polymerase. Nucleic Acids Res. 1990 Dec 25;18(24):7317–7322. doi: 10.1093/nar/18.24.7317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lyamichev V., Brow M. A., Dahlberg J. E. Structure-specific endonucleolytic cleavage of nucleic acids by eubacterial DNA polymerases. Science. 1993 May 7;260(5109):778–783. doi: 10.1126/science.7683443. [DOI] [PubMed] [Google Scholar]
- Malan T. P., McClure W. R. Dual promoter control of the Escherichia coli lactose operon. Cell. 1984 Nov;39(1):173–180. doi: 10.1016/0092-8674(84)90203-4. [DOI] [PubMed] [Google Scholar]
- Myers T. W., Gelfand D. H. Reverse transcription and DNA amplification by a Thermus thermophilus DNA polymerase. Biochemistry. 1991 Aug 6;30(31):7661–7666. doi: 10.1021/bi00245a001. [DOI] [PubMed] [Google Scholar]
- Odelberg S. J., Weiss R. B., Hata A., White R. Template-switching during DNA synthesis by Thermus aquaticus DNA polymerase I. Nucleic Acids Res. 1995 Jun 11;23(11):2049–2057. doi: 10.1093/nar/23.11.2049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olsen D. B., Eckstein F. Incomplete primer extension during in vitro DNA amplification catalyzed by Taq polymerase; exploitation for DNA sequencing. Nucleic Acids Res. 1989 Dec 11;17(23):9613–9620. doi: 10.1093/nar/17.23.9613. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robins P., Pappin D. J., Wood R. D., Lindahl T. Structural and functional homology between mammalian DNase IV and the 5'-nuclease domain of Escherichia coli DNA polymerase I. J Biol Chem. 1994 Nov 18;269(46):28535–28538. [PubMed] [Google Scholar]
- Saiki R. K., Gelfand D. H., Stoffel S., Scharf S. J., Higuchi R., Horn G. T., Mullis K. B., Erlich H. A. Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science. 1988 Jan 29;239(4839):487–491. doi: 10.1126/science.2448875. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Siegal G., Turchi J. J., Myers T. W., Bambara R. A. A 5' to 3' exonuclease functionally interacts with calf DNA polymerase epsilon. Proc Natl Acad Sci U S A. 1992 Oct 15;89(20):9377–9381. doi: 10.1073/pnas.89.20.9377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Welsh J., McClelland M. Fingerprinting genomes using PCR with arbitrary primers. Nucleic Acids Res. 1990 Dec 25;18(24):7213–7218. doi: 10.1093/nar/18.24.7213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williams J. G., Kubelik A. R., Livak K. J., Rafalski J. A., Tingey S. V. DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res. 1990 Nov 25;18(22):6531–6535. doi: 10.1093/nar/18.22.6531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zuker M., Stiegler P. Optimal computer folding of large RNA sequences using thermodynamics and auxiliary information. Nucleic Acids Res. 1981 Jan 10;9(1):133–148. doi: 10.1093/nar/9.1.133. [DOI] [PMC free article] [PubMed] [Google Scholar]