Abstract
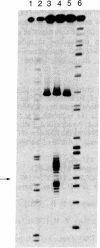
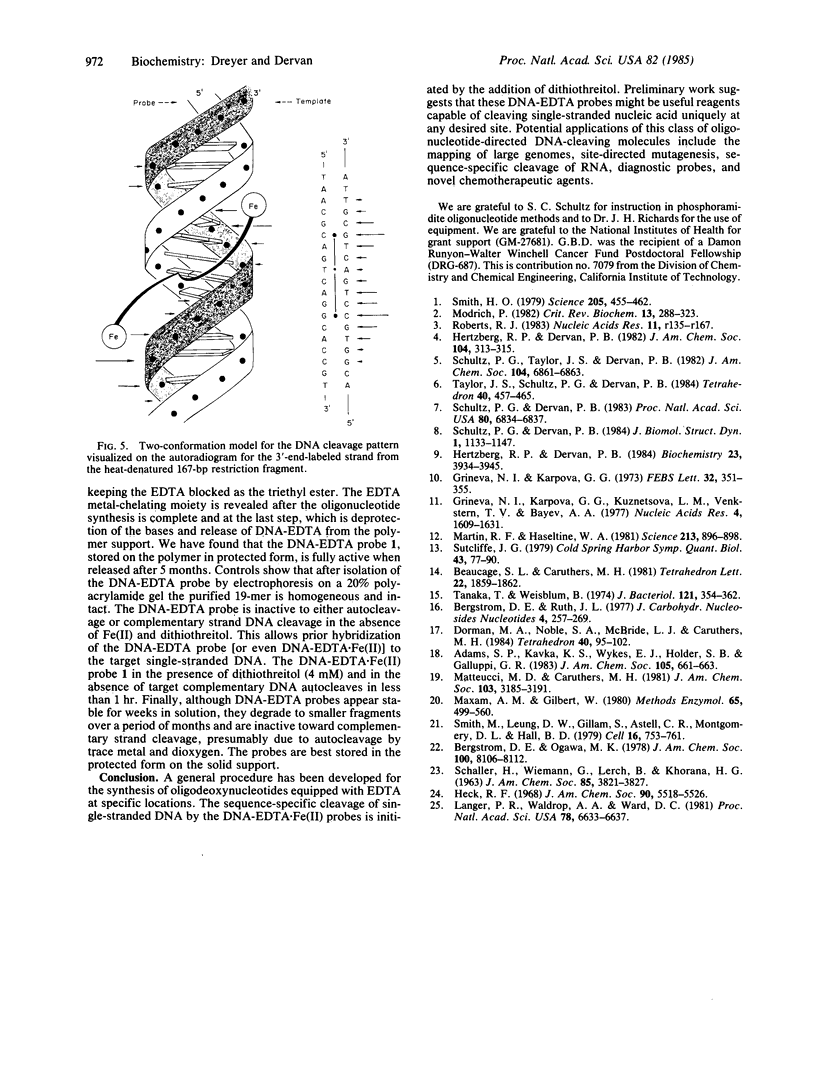
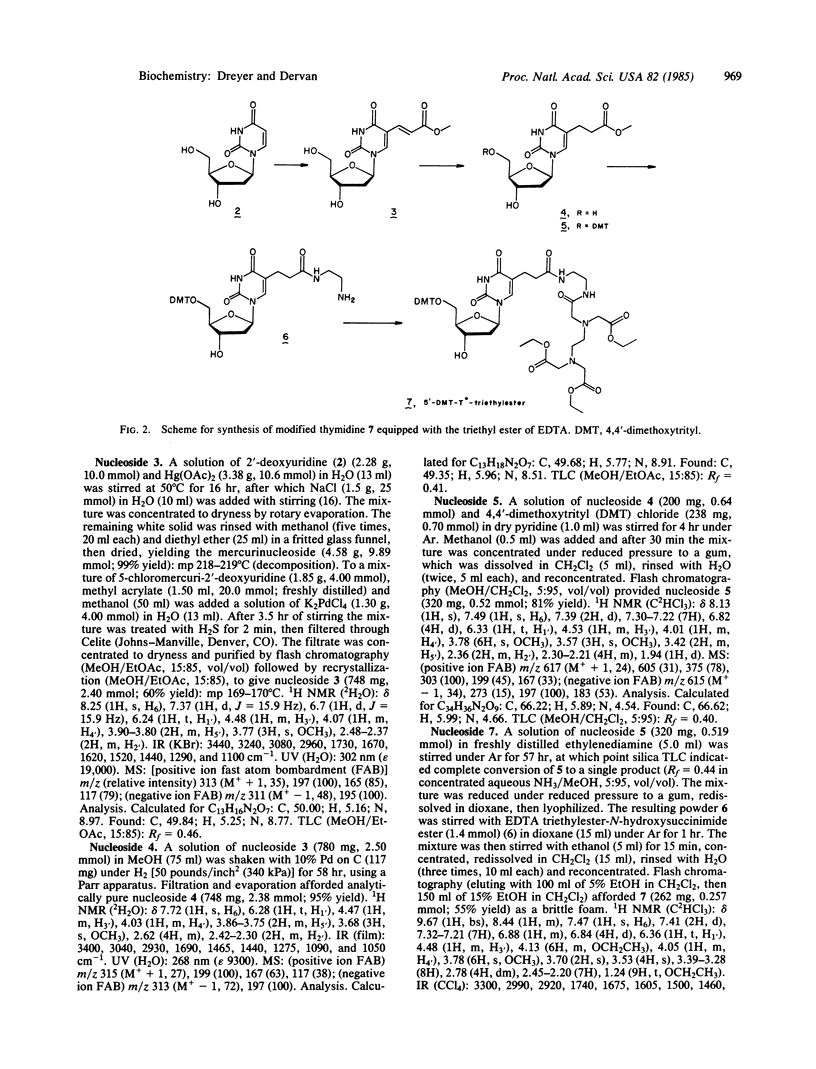
The synthesis of a DNA hybridization probe 19 nucleotides in length, equipped with the metal chelator EDTA at C-5 of thymidine in position 10 (indicated by T*) is described. DNA-EDTA 1 has the sequence 5'-T-A-A-C-G-C-A-G-T*-C-A-G-G-C-A-C-C-G-T-3', which is complementary to a 19-nucleotide sequence in the plasmid pBR322. In the presence of Fe(II), O2, and dithiothreitol, DNA-EDTA 1 affords specific cleavage (25 degrees C, pH 7.4, 60 min) at its complementary sequence in a heat-denatured 167-base-pair restriction fragment. Cleavage occurs over a range of 16 nucleotides at the site of hybridization of 1, presumably due to a diffusible reactive species. No other cleavage sites are observed in the 167-base-pair restriction fragment. The procedure used to synthesize DNA-EDTA probes is based on the incorporation of a thymidine modified at C-5 with the triethyl ester of EDTA. By using routine phosphoramidite procedures, thymidine-EDTA can be incorporated into oligodeoxynucleotides of any desired length and sequence. Because the efficiency of the DNA cleavage reaction is dependent on the addition of both Fe(II) and reducing agent (dithiothreitol), the initiation of the cleavage reaction can be controlled. These DNA-EDTA X Fe(II) probes should be useful for the sequence-specific cleavage of single-stranded DNA (and most likely RNA) under mild conditions.
Full text
PDF



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Grineva N. I., Karpova G. G. Complementarily addressed modification of rRNA with p-(chloroethylmethylamino)benzylidene hexanucleotides. FEBS Lett. 1973 Jun 1;32(2):351–355. doi: 10.1016/0014-5793(73)80871-3. [DOI] [PubMed] [Google Scholar]
- Grineva N. I., Karpova G. G., Kuznetsova L. M., Venkstern T. V., Bayev A. A. Complementary addressed modification of yeast tRNA Val 1 with alkylating derivative of d(pC-G)-A. The positions of the alkylated nucleotides and the course of the alkylation in the complex. Nucleic Acids Res. 1977;4(5):1609–1631. doi: 10.1093/nar/4.5.1609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hertzberg R. P., Dervan P. B. Cleavage of DNA with methidiumpropyl-EDTA-iron(II): reaction conditions and product analyses. Biochemistry. 1984 Aug 14;23(17):3934–3945. doi: 10.1021/bi00312a022. [DOI] [PubMed] [Google Scholar]
- Langer P. R., Waldrop A. A., Ward D. C. Enzymatic synthesis of biotin-labeled polynucleotides: novel nucleic acid affinity probes. Proc Natl Acad Sci U S A. 1981 Nov;78(11):6633–6637. doi: 10.1073/pnas.78.11.6633. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin R. F., Haseltine W. A. Range of radiochemical damage to DNA with decay of iodine-125. Science. 1981 Aug 21;213(4510):896–898. doi: 10.1126/science.7256283. [DOI] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 1980;65(1):499–560. doi: 10.1016/s0076-6879(80)65059-9. [DOI] [PubMed] [Google Scholar]
- Modrich P. Studies on sequence recognition by type II restriction and modification enzymes. CRC Crit Rev Biochem. 1982;13(3):287–323. doi: 10.3109/10409238209114231. [DOI] [PubMed] [Google Scholar]
- Roberts R. J. Restriction and modification enzymes and their recognition sequences. Nucleic Acids Res. 1983 Jan 11;11(1):r135–r167. [PMC free article] [PubMed] [Google Scholar]
- Schultz P. G., Dervan P. B. Distamycin and penta-N-methylpyrrolecarboxamide binding sites on native DNA. A comparison of methidiumpropyl-EDTA-Fe(II) footprinting and DNA affinity cleaving. J Biomol Struct Dyn. 1984 Mar;1(5):1133–1147. doi: 10.1080/07391102.1984.10507508. [DOI] [PubMed] [Google Scholar]
- Schultz P. G., Dervan P. B. Sequence-specific double-strand cleavage of DNA by penta-N-methylpyrrolecarboxamide-EDTA X Fe(II). Proc Natl Acad Sci U S A. 1983 Nov;80(22):6834–6837. doi: 10.1073/pnas.80.22.6834. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith H. O. Nucleotide sequence specificity of restriction endonucleases. Science. 1979 Aug 3;205(4405):455–462. doi: 10.1126/science.377492. [DOI] [PubMed] [Google Scholar]
- Smith M., Leung D. W., Gillam S., Astell C. R., Montgomery D. L., Hall B. D. Sequence of the gene for iso-1-cytochrome c in Saccharomyces cerevisiae. Cell. 1979 Apr;16(4):753–761. doi: 10.1016/0092-8674(79)90091-6. [DOI] [PubMed] [Google Scholar]
- Sutcliffe J. G. Complete nucleotide sequence of the Escherichia coli plasmid pBR322. Cold Spring Harb Symp Quant Biol. 1979;43(Pt 1):77–90. doi: 10.1101/sqb.1979.043.01.013. [DOI] [PubMed] [Google Scholar]
- Tanaka T., Weisblum B. Construction of a colicin E1-R factor composite plasmid in vitro: means for amplification of deoxyribonucleic acid. J Bacteriol. 1975 Jan;121(1):354–362. doi: 10.1128/jb.121.1.354-362.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]